Genetic Markers of the Host in Persons Living with HTLV-1, HIV and HCV Infections
Abstract
1. Introduction
2. HLA Risk for HAM/TSP
| HLA Allele | Japanese | Brazilians | Iranians | Spanish | Afro-Caribbean (Martinique) | Afro-Caribbean (London) | Jamaicans |
|---|---|---|---|---|---|---|---|
| A*02 | ++ | + | 0 | 0 | 0 | ++ | 0 |
| Cw*08 | ++ | 0 | 0 | 0 | |||
| A*24 | - | 0 | 0 | ||||
| B*07 | - | ± | 0 | - | 0 | 0 | |
| B*5401 | - | ᴓ | ᴓ | ᴓ | ᴓ | ᴓ | ᴓ |
| DRB1*0101 | ± | 0 | ± | - | 0 | 0 | |
| DRB1*11 | - | - | - | - | 0 | 0 |
3. Interferon Lambda 3 (IFN-λ3)
4. miRNA
| MiRNA | Regulation | miRNA Target | Function |
|---|---|---|---|
| miR-21 | Upregulated | PTEN | Antiapoptotic |
| miR-93 | Upregulated | p21 (WAF1/CIP1); MICB | Antiapoptotic |
| miR-132 | Downregulated | p300 | Immune evasion |
| miR-143-p3 | Upregulated | AChE; PKA; GRα | Increase of viral transcription |
| miR-146 a | Upregulated | Unknown | Pro-inflammatory |
| miR-149 | Downregulated | p300 | Proliferation |
| miR-155 | Upregulated | TP53INP1; Unknown | Proliferation |
| miR-873 | Downregulated | p300 | Proliferation |
5. Killer Cell Immunoglobulin-Like Receptors (KIRs)
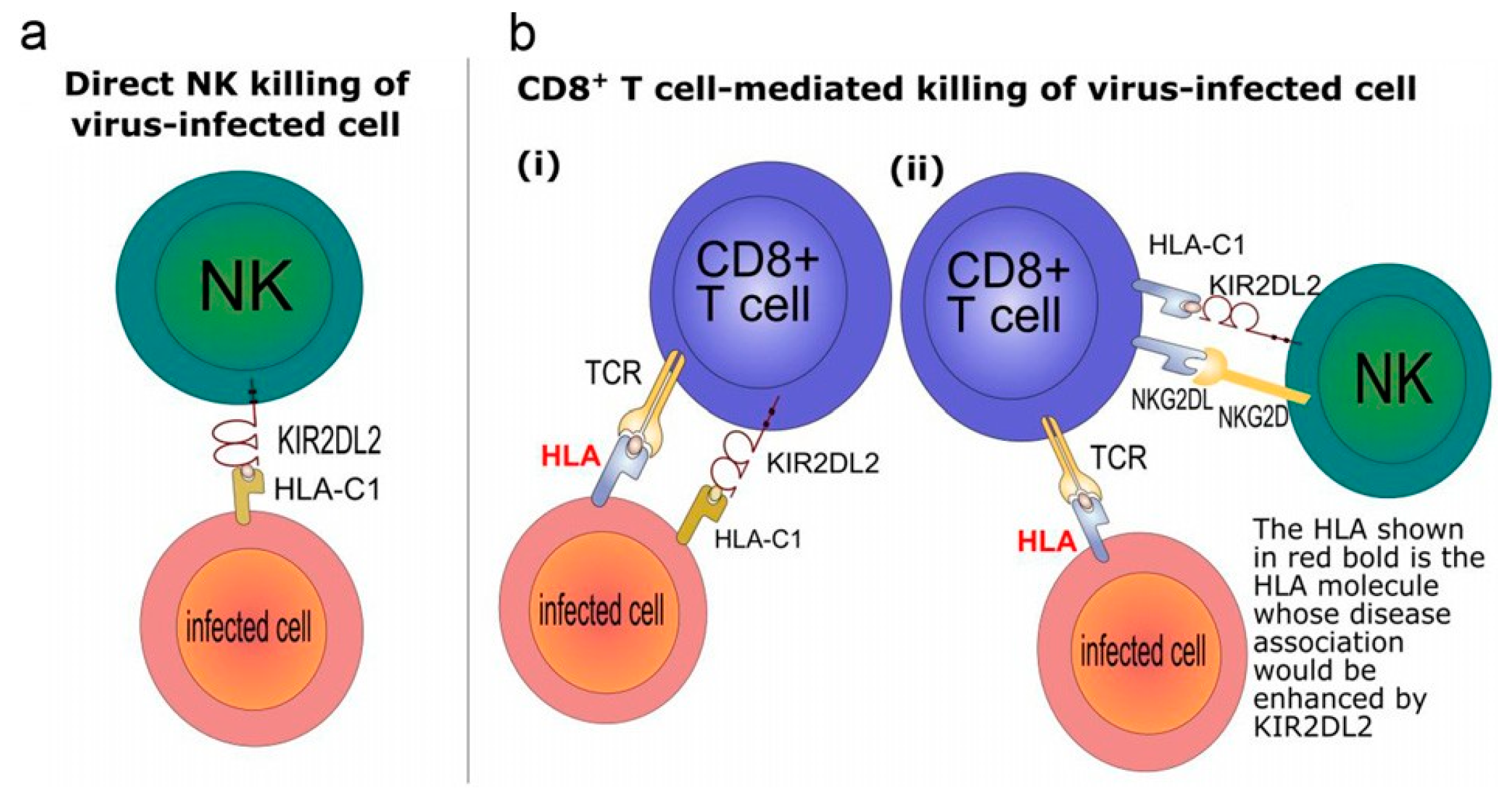
6. Genes and Susceptibility to HIV Infection
6.1. Genes that Influence the Dynamic Progression of AIDS
6.2. Genes Important to Anti-HIV Treatment
6.3. Co-Infection HIV–HTLV
7. HCV and Genetic Susceptibility
8. Concluding Remarks
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Cook, L.B.; Elemans, M.; Rowan, A.G.; Asquith, B. HTLV-1: Persistence and pathogenesis. Virology 2013, 435, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Jeffery, K.J.; Usuku, K.; Hall, S.E.; Matsumoto, W.; Taylor, G.P.; Procter, J.; Bunce, M.; Ogg, G.S.; Welsh, K.I.; Weber, J.N.; et al. HLA alleles determine human T-lymphotropic virus-I (HTLV-I) proviral load and the risk of HTLV-I-associated myelopathy. Proc. Natl. Acad. Sci. USA 1999, 96, 3848–3853. [Google Scholar] [CrossRef] [PubMed]
- Jeffery, K.J.; Siddiqui, A.A.; Bunce, M.; Lloyd, A.L.; Vine, A.M.; Witkover, A.D.; Izumo, S.; Usuku, K.; Welsh, K.I.; Osame, M.; et al. The influence of HLA class I alleles and heterozygosity on the outcome of human T cell lymphotropic virus type I infection. J.Immunol. 2000, 165, 7278–7284. [Google Scholar] [CrossRef] [PubMed]
- Bangham, C.R.; Hall, S.E.; Jeffery, K.J.; Vine, A.M.; Witkover, A.; Nowak, M.A.; Wodarz, D.; Usuku, K.; Osame, M.; et al. Genetic control and dynamics of the cellular immune response to the human T-cell leukaemia virus, HTLV-I. Philos. Trans. R. Soc. Lond. B Biol. Sci. 1999, 354, 691–700. [Google Scholar] [CrossRef] [PubMed]
- Borducchi, D.M.; Gerbase-DeLima, M.; Morgun, A.; Shulzhenko, N.; Pombo-de-Oliveira, M.S.; Kerbauy, J.; Rodrigues de Oliveira, J.S.; et al. Human leucocyte antigen and human T-cell lymphotropic virus type 1 associated diseases in Brazil. Br. J. Haematol. 2003, 123, 954–955. [Google Scholar] [CrossRef] [PubMed]
- Catalan-Soares, B.C.; Carneiro-Proietti, A.B.; Da Fonseca, F.G.; Correa-Oliveira, R.; Peralva-Lima, D.; Portela, R.; Ribas, J.G.; Goncalves, D.U.; Interdisciplinary, H.R.G.; Proietti, F.A.; et al. HLA class I alleles in HTLV-1-associated myelopathy and asymptomatic carriers from the Brazilian cohort GIPH. Med. Microbiol. Immunol. 2009, 198, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Deschamps, R.; Bera, O.; Belrose, G.; Lezin, A.; Bellance, R.; Signate, A.; Cabre, P.; Smadja, D.; Cesaire, R.; Olindo, S.; et al. Absence of consistent association between human leukocyte antigen-I and -II alleles and human T-lymphotropic virus type 1 (HTLV-1)-associated myelopathy/tropical spastic paraparesis risk in an HTLV-1 French Afro-Caribbean population. Int. J. Infect. Dis. 2010, 14, e986–e990. [Google Scholar] [CrossRef] [PubMed]
- Goedert, J.J.; Li, H.C.; Gao, X.J.; Chatterjee, N.; Sonoda, S.; Biggar, R.J.; Cranston, B.; Kim, N.; Carrington, M.; Morgan, O.; et al. Risk of human T-lymphotropic virus type I-associated diseases in Jamaica with common HLA types. Int. J. Cancer 2007, 121, 1092–1097. [Google Scholar] [CrossRef] [PubMed]
- Trevino, A.; Vicario, J.L.; Lopez, M.; Parra, P.; Benito, R.; Ortiz de Lejarazu, R.; Ramos, J.M.; Del Romero, J.; de Mendoza, C.; Soriano, V.; et al. Association between HLA alleles and HAM/TSP in individuals infected with HTLV-1. J. Neurol. 2013, 260, 2551–2555. [Google Scholar] [CrossRef] [PubMed]
- Sabouri, A.H.; Saito, M.; Usuku, K.; Bajestan, S.N.; Mahmoudi, M.; Forughipour, M.; Sabouri, Z.; Abbaspour, Z.; Goharjoo, M.E.; Khayami, E.; et al. Differences in viral and host genetic risk factors for development of human T-cell lymphotropic virus type 1 (HTLV-1)-associated myelopathy/tropical spastic paraparesis between Iranian and Japanese HTLV-1-infected individuals. J. Gen. Virol. 2005, 86 Pt 3, 773–781. [Google Scholar] [CrossRef] [PubMed]
- Rafatpanah, H.; Pravica, V.; Faridhosseini, R.; Tabatabaei, A.; Ollier, W.; Poulton, K.; Thomson, W.; Hutchinson, I.; et al. Association between HLA-DRB1*01 and HLA-Cw*08 and outcome following HTLV-I infection. Iran. J. Immunol. 2007, 4, 94–100. [Google Scholar] [PubMed]
- Taghaddosi, M.; Rezaee, S.A.; Rafatpanah, H.; Rajaei, T.; Farid Hosseini, R.; Narges, V. Association between HLA Class I Alleles and Proviral Load in HTLV-I Associated Myelopathy/Tropical Spastic Paraperesis (HAM/TSP) Patients in Iranian Population. Iran. J. Basic Med. Sci. 2013, 16, 264–267. [Google Scholar] [PubMed]
- Imanishi, T.; Akaza, T.; Kimura, A.; Tokunaga, K.; Gojobori, T. Allele and haplotype frequencies for HLA and complement loci in various ethnic groups. In HLA. 1; Tsuji, K., Aizawa, M., Sasazuki, T., Eds.; Oxford University Press: New York, NY, USA, 1992; p. 1065. [Google Scholar]
- Macnamara, A.; Rowan, A.; Hilburn, S.; Kadolsky, U.; Fujiwara, H.; Suemori, K.; Yasukawa, M.; Taylor, G.; Bangham, C.R.; Asquith, B.; et al. HLA class I binding of HBZ determines outcome in HTLV-1 infection. PLoS Pathog. 2010, 6, e1001117. [Google Scholar] [CrossRef] [PubMed]
- MacNamara, A.; Kadolsky, U.; Bangham, C.R.; Asquith, B. T-cell epitope prediction: Rescaling can mask biological variation between MHC molecules. PLoS Comput. Biol. 2009, 5, e1000327. [Google Scholar] [CrossRef] [PubMed]
- Suppiah, V.; Moldovan, M.; Ahlenstiel, G.; Berg, T.; Weltman, M.; Abate, M.L.; Bassendine, M.; Spengler, U.; Dore, G.J.; Powell, E.; et al. IL28B is associated with response to chronic hepatitis C interferon-alpha and ribavirin therapy. Nat. Genet. 2009, 41, 1100–1104. [Google Scholar] [CrossRef] [PubMed]
- Sanabani, S.S.; Nukui, Y.; Pereira, J.; da Costa, A.C.; de Oliveira, A.C.; Pessoa, R.; Leal, F.E.; Segurado, A.C.; Kallas, E.G.; Sabino, E.C.; et al. Lack of evidence to support the association of a single IL28B genotype SNP rs12979860 with the HTLV-1 clinical outcomes and proviral load. BMC Infect. Dis. 2012, 12, 374. [Google Scholar] [CrossRef] [PubMed]
- Trevino, A.; Lopez, M.; Vispo, E.; Aguilera, A.; Ramos, J.M.; Benito, R.; Roc, L.; Eiros, J.M.; de Mendoza, C.; Soriano, V.; et al. Development of tropical spastic paraparesis in human T-lymphotropic virus type 1 carriers is influenced by interleukin 28B gene polymorphisms. Clin. Infect. Dis. 2012, 55, e1–e4. [Google Scholar] [CrossRef] [PubMed]
- Assone, T.; de Souza, F.V.; Gaester, K.O.; Fonseca, L.A.; Luiz Odo, C.; Malta, F.; Pinho, J.R.; Goncalves Fde, T.; Duarte, A.J.; de Oliveira, A.C.; et al. IL28B gene polymorphism SNP rs8099917 genotype GG is associated with HTLV-1-associated myelopathy/tropical spastic paraparesis (HAM/TSP) in HTLV-1 carriers. PLoS Negl. Trop. Dis. 2014, 8, e3199. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.P.; Qi, Y.; Goedert, J.J.; Hussain, S.K.; Kirk, G.D.; Hoots, W.K.; Buchbinder, S.; Carrington, M.; Thio, C.L.; et al. IL28B polymorphism does not determine outcomes of hepatitis B virus or HIV infection. J. Infect. Dis. 2010, 202, 1749–1753. [Google Scholar] [CrossRef] [PubMed]
- Machmach, K.; Abad-Molina, C.; Romero-Sanchez, M.C.; Abad, M.A.; Ferrando-Martinez, S.; Genebat, M.; Pulido, I.; Viciana, P.; Gonzalez-Escribano, M.F.; Leal, M.; et al. IL28B single-nucleotide polymorphism rs12979860 is associated with spontaneous HIV control in white subjects. J. Infect. Diseases. 2013, 207, 651–655. [Google Scholar] [CrossRef] [PubMed]
- Ge, D.; Fellay, J.; Thompson, A.J.; Simon, J.S.; Shianna, K.V.; Urban, T.J.; Heinzen, E.L.; Qiu, P.; Bertelsen, A.H.; Muir, A.J.; et al. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature 2009, 461, 399–401. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, D.F.; Souza, F.V.; Fonseca, L.A.; Duarte, A.J.; Casseb, J. Influence of human T-cell lymphotropic virus type 1 (HTLV-1) Infection on laboratory parameters of patients with chronic hepatitis C virus. Rev. Inst. Med. Trop. Sao Paulo 2009, 51, 325–329. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Brites, C.; Sampalo, J.; Oliveira, A. HIV/human T-cell lymphotropic virus coinfection revisited: Impact on AIDS progression. AIDS Rev. 2009, 11, 8–16. [Google Scholar] [PubMed]
- Montanheiro, P.A.; Penalva de Oliveira, A.C.; Smid, J.; Fukumori, L.M.; Olah, I.; Da, S.D.A.J.; Casseb, J.; et al. The elevated interferon gamma production is an important immunological marker in HAM/TSP pathogenesis. Scand. J. Immunol. 2009, 70, 403–407. [Google Scholar] [CrossRef] [PubMed]
- Balagopal, A.; Thomas, D.L.; Thio, C.L. IL28B and the control of hepatitis C virus infection. Gastroenterology 2010, 139, 1865–1876. [Google Scholar] [CrossRef] [PubMed]
- Jacobson, S.; Krichavsky, M.; Flerlage, N.; Levin, M. Immunopathogenesis of HTLV-I associated neurologic disease: Massive latent HTLV-I infection in bone marrow of HAM/TSP patients. Leukemia 1997, 11 (Suppl. 3), 73–75. [Google Scholar] [PubMed]
- Ferri, C.; Monti, M.; La Civita, L.; Careccia, G.; Mazzaro, C.; Longombardo, G.; Lombardini, F.; Greco, F.; Pasero, G.; Bombardieri, S.; et al. Hepatitis C virus infection in non-Hodgkin’s B-cell lymphoma complicating mixed cryoglobulinaemia. Eur. J. Clin. Investig. 1994, 24, 781–784. [Google Scholar] [CrossRef]
- Koziel, M.J.; Dudley, D.; Wong, J.T.; Dienstag, J.; Houghton, M.; Ralston, R.; Walker, B.D. Intrahepatic cytotoxic T lymphocytes specific for hepatitis C virus in persons with chronic hepatitis. J.Immunol. 1992, 149, 3339–3344. [Google Scholar] [PubMed]
- Bellon, M.; Lepelletier, Y.; Hermine, O.; Nicot, C. Deregulation of microRNA involved in hematopoiesis and the immune response in HTLV-I adult T-cell leukemia. Blood 2009, 113, 4914–4917. [Google Scholar] [CrossRef] [PubMed]
- Ruggero, K.; Corradin, A.; Zanovello, P.; Amadori, A.; Bronte, V.; Ciminale, V.; D'Agostino, D.M. Role of microRNAs in HTLV-1 infection and transformation. Mol. Aspects Med. 2010, 31, 367–382. [Google Scholar] [CrossRef] [PubMed]
- Colonna, M.; Borsellino, G.; Falco, M.; Ferrara, G.B.; Strominger, J.L. HLA-C is the inhibitory ligand that determines dominant resistance to lysis by NK1- and NK2-specific natural killer cells. Proc. Natl. Acad. Sci. USA 1993, 90, 12000–12004. [Google Scholar] [CrossRef] [PubMed]
- Wagtmann, N.; Rajagopalan, S.; Winter, C.C.; Peruzzi, M.; Long, E.O. Killer cell inhibitory receptors specific for HLA-C and HLA-B identified by direct binding and by functional transfer. Immunity 1995, 3, 801–809. [Google Scholar] [CrossRef]
- Numasaki, M.; Tagawa, M.; Iwata, F.; Suzuki, T.; Nakamura, A.; Okada, M.; Iwakura, Y.; Aiba, S.; Yamaya, M. IL-28 elicits antitumor responses against murine fibrosarcoma. J. Immunol. 2007, 178, 5086–5098. [Google Scholar] [CrossRef] [PubMed]
- Sato, A.; Ohtsuki, M.; Hata, M.; Kobayashi, E.; Murakami, T. Antitumor activity of IFN-lambda in murine tumor models. J. Immunol. 2006, 176, 7686–7694. [Google Scholar] [CrossRef] [PubMed]
- Khakoo, S.I.; Thio, C.L.; Martin, M.P.; Brooks, C.R.; Gao, X.; Astemborski, J.; Cheng, J.; Goedert, J.J.; Vlahov, D.; Hilgartner, M.; et al. HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science 2004, 305, 872–874. [Google Scholar] [CrossRef] [PubMed]
- Knapp, S.; Warshow, U.; Hegazy, D.; Brackenbury, L.; Guha, I.N.; Fowell, A.; Little, A.M.; Alexander, G.J.; Rosenberg, W.M.; Cramp, M.E.; et al. Consistent beneficial effects of killer cell immunoglobulin-like receptor 2DL3 and group 1 human leukocyte antigen-C following exposure to hepatitis C virus. Hepatology 2010, 51, 1168–1175. [Google Scholar] [CrossRef] [PubMed]
- Vidal-Castineira, J.R.; Lopez-Vazquez, A.; Martinez-Borra, J.; Martinez-Camblor, P.; Prieto, J.; Lopez-Rodriguez, R.; Sanz-Cameno, P.; de la Vega, J.; Rodrigo, L.; Perez-Lopez, R.; et al. Diversity of killer cell immunoglobulin-like receptor (KIR) genotypes and KIR2DL2/3 variants in HCV treatment outcome. PLoS ONE 2014, 9, e99426. [Google Scholar]
- Ahlenstiel, G.; Martin, M.P.; Gao, X.; Carrington, M.; Rehermann, B. Distinct KIR/HLA compound genotypes affect the kinetics of human antiviral natural killer cell responses. J. Clin. Investig. 2008, 118, 1017–1026. [Google Scholar] [CrossRef] [PubMed]
- Suppiah, V.; Gaudieri, S.; Armstrong, N.J.; O'Connor, K.S.; Berg, T.; Weltman, M.; Abate, M.L.; Spengler, U.; Bassendine, M.; Dore, G.J.; et al. IL28B, HLA-C, and KIR variants additively predict response to therapy in chronic hepatitis C virus infection in a European Cohort: A cross-sectional study. PLoS Med. 2011, 8, e1001092. [Google Scholar] [CrossRef] [PubMed]
- Kamihira, S.; Usui, T.; Ichikawa, T.; Uno, N.; Morinaga, Y.; Mori, S.; Nagai, K.; Sasaki, D.; Hasegawa, H.; Yanagihara, K.; et al. Paradoxical expression of IL-28B mRNA in peripheral blood in human T-cell leukemia virus type-1 mono-infection and co-infection with hepatitis C virus. Virol. J. 2012, 9, 40. [Google Scholar] [CrossRef] [PubMed]
- Seich Al Basatena, N.K.; Macnamara, A.; Vine, A.M.; Thio, C.L.; Astemborski, J.; Usuku, K.; Osame, M.; Kirk, G.D.; Donfield, S.M.; Goedert, J.J.; et al. KIR2DL2 enhances protective and detrimental HLA class I-mediated immunity in chronic viral infection. PLoS Pathog. 2011, 7, e1002270. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Paxton, W.A.; Choe, S.; Ceradini, D.; Martin, S.R.; Horuk, R.; MacDonald, M.E.; Stuhlmann, H.; Koup, R.A.; Landau, N.R. Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell 1996, 86, 367–377. [Google Scholar] [CrossRef]
- Dean, M.; Carrington, M.; Winkler, C.; Huttley, G.A.; Smith, M.W.; Allikmets, R.; Goedert, J.J.; Buchbinder, S.P.; Vittinghoff, E.; Gomperts, E.; et al. Genetic restriction of HIV-1 infection and progression to AIDS by a deletion allele of the CKR5 structural gene. Hemophilia Growth and Development Study, Multicenter AIDS Cohort Study, Multicenter Hemophilia Cohort Study, San Francisco City Cohort, ALIVE Study. Science 1996, 273, 1856–1862. [Google Scholar] [PubMed]
- Samson, M.; Libert, F.; Doranz, B.J.; Rucker, J.; Liesnard, C.; Farber, C.M.; Saragosti, S.; Lapoumeroulie, C.; Cognaux, J.; Forceille, C.; et al. Resistance to HIV-1 infection in caucasian individuals bearing mutant alleles of the CCR-5 chemokine receptor gene. Nature 1996, 382, 722–725. [Google Scholar] [CrossRef] [PubMed]
- Schinkel, J.; Langendam, M.W.; Coutinho, R.A.; Krol, A.; Brouwer, M.; Schuitemaker, H. No evidence for an effect of the CCR5 delta32/+ and CCR2b 64I/+ mutations on human immunodeficiency virus (HIV)-1 disease progression among HIV-1-infected injecting drug users. J. Infect. Dis. 1999, 179, 825–831. [Google Scholar] [CrossRef] [PubMed]
- Stewart, G.J.; Ashton, L.J.; Biti, R.A.; Ffrench, R.A.; Bennetts, B.H.; Newcombe, N.R.; Benson, E.M.; Carr, A.; Cooper, D.A.; Kaldor, J.M. Increased frequency of CCR-5 delta 32 heterozygotes among long-term non-progressors with HIV-1 infection. The Australian Long-Term Non-Progressor Study Group. AIDS 1997, 11, 1833–1838. [Google Scholar] [CrossRef] [PubMed]
- Michael, N.L.; Louie, L.G.; Sheppard, H.W. CCR5-delta 32 gene deletion in HIV-1 infected patients. Lancet 1997, 350, 741–742. [Google Scholar] [CrossRef]
- Martinson, J.J.; Chapman, N.H.; Rees, D.C.; Liu, Y.T.; Clegg, J.B. Global distribution of the CCR5 gene 32-basepair deletion. Nat. Genet. 1997, 16, 100–103. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Li, X.; Tang, J.; Jin, T.; Liao, Q.; Hu, G. Association between chemotactic chemokine ligand 5 -403G/A polymorphism and risk of human immunodeficiency virus-1 infection: A meta-analysis. OncoTargets Ther. 2015, 8, 727–734. [Google Scholar]
- Gong, X.; Liu, Y.; Liu, F.L.; Jin, L.; Wang, H.; Zheng, Y.T. A SDF1 genetic variant confers resistance to HIV-1 infection in intravenous drug users in China. Infect. Genet. Evol. 2015, 34, 137–142. [Google Scholar] [CrossRef] [PubMed]
- An, P.; Goedert, J.J.; Donfield, S.; Buchbinder, S.; Kirk, G.D.; Detels, R.; Winkler, C.A. Regulatory variation in HIV-1 dependency factor ZNRD1 associates with host resistance to HIV-1 acquisition. J. Infect. Dis. 2014, 210, 1539–1548. [Google Scholar] [CrossRef] [PubMed]
- Fahrbach, K.M.; Barry, S.M.; Ayehunie, S.; Lamore, S.; Klausner, M.; Hope, T.J. Activated CD34-derived Langerhans cells mediate transinfection with human immunodeficiency virus. J. Virol. 2007, 81, 6858–6868. [Google Scholar] [CrossRef] [PubMed]
- de Witte, L.; Nabatov, A.; Pion, M.; Fluitsma, D.; de Jong, M.A.; de Gruijl, T.; Piguet, V.; van Kooyk, Y.; Geijtenbeek, T.B. Langerin is a natural barrier to HIV-1 transmission by Langerhans cells. Nat. Med. 2007, 13, 367–371. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, O. Langerhans cells lap up HIV-1. Nat. Med. 2007, 13, 245–246. [Google Scholar] [CrossRef] [PubMed]
- Cole, A.M.; Ganz, T. Human antimicrobial peptides: Analysis and application. BioTechniques 2000, 29, 822–826, 828,830–831. [Google Scholar] [PubMed]
- Jenssen, H.; Hamill, P.; Hancock, R.E. Peptide antimicrobial agents. Clin. Microbiol. Rev. 2006, 19, 491–511. [Google Scholar] [CrossRef] [PubMed]
- Klotman, M.E.; Chang, T.L. Defensins in innate antiviral immunity. Nat. Rev. Immunol. 2006, 6, 447–456. [Google Scholar] [CrossRef] [PubMed]
- Furci, L.; Sironi, F.; Tolazzi, M.; Vassena, L.; Lusso, P. Alpha-defensins block the early steps of HIV-1 infection: Interference with the binding of gp120 to CD4. Blood 2007, 109, 2928–2935. [Google Scholar] [PubMed]
- Chang, T.L.; Vargas, J., Jr.; DelPortillo, A.; Klotman, M.E. Dual role of alpha-defensin-1 in anti-HIV-1 innate immunity. J. Clin. Investig. 2005, 115, 765–773. [Google Scholar] [CrossRef] [PubMed]
- Weinberg, A.; Quinones-Mateu, M.E.; Lederman, M.M. Role of human beta-defensins in HIV infection. Adv. Dent. Res. 2006, 19, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Schutte, B.C.; Mitros, J.P.; Bartlett, J.A.; Walters, J.D.; Jia, H.P.; Welsh, M.J.; Casavant, T.L.; McCray, P.B. Discovery of five conserved beta-defensin gene clusters using a computational search strategy. Proc. Natl. Acad. Sci. USA 2002, 99, 2129–2133. [Google Scholar] [CrossRef] [PubMed]
- Cole, A.M.; Hong, T.; Boo, L.M.; Nguyen, T.; Zhao, C.; Bristol, G.; Zack, J.A.; Waring, A.J.; Yang, O.O.; Lehrer, R.I. Retrocyclin: A primate peptide that protects cells from infection by T- and M-tropic strains of HIV-1. Proc. Natl. Acad. Sci. USA 2002, 99, 1813–1818. [Google Scholar] [CrossRef] [PubMed]
- Gallo, S.A.; Wang, W.; Rawat, S.S.; Jung, G.; Waring, A.J.; Cole, A.M.; Lu, H.; Yan, X.; Daly, N.L.; Craik, D.J.; et al. Theta-defensins prevent HIV-1 Env-mediated fusion by binding gp41 and blocking 6-helix bundle formation. J. Biol. Chem. 2006, 281, 18787–18792. [Google Scholar] [CrossRef] [PubMed]
- Cole, A.M.; Lehrer, R.I. Minidefensins: Antimicrobial peptides with activity against HIV-1. Curr. Pharm. Des. 2003, 9, 1463–1473. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, S.L.; de Sa, N.B.; Campos, D.P.; Coelho, A.B.; Guimaraes, M.L.; Leite, T.C.; Veloso, V.G.; Morgado, M.G. Association of the HLA-B*52 allele with non-progression to AIDS in Brazilian HIV-1-infected individuals. Genes Immun. 2014, 15, 256–262. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; O'Brien, T.R.; Welzel, T.M.; Marti, D.; Qi, Y.; Goedert, J.J.; Phair, J.; Pfeiffer, R.; Carrington, M. HLA-B alleles associate consistently with HIV heterosexual transmission, viral load, and progression to AIDS, but not susceptibility to infection. AIDS 2010, 24, 1835–1840. [Google Scholar] [CrossRef] [PubMed]
- Klein, M.R.; Keet, I.P.; D'Amaro, J.; Bende, R.J.; Hekman, A.; Mesman, B.; Koot, M.; de Waal, L.P.; Coutinho, R.A.; Miedema, F. Associations between HLA frequencies and pathogenic features of human immunodeficiency virus type 1 infection in seroconverters from the Amsterdam cohort of homosexual men. J. Infect. Dis. 1994, 169, 1244–1249. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Nelson, G.W.; Karacki, P.; Martin, M.P.; Phair, J.; Kaslow, R.; Goedert, J.J.; Buchbinder, S.; Hoots, K.; Vlahov, D.; et al. Effect of a single amino acid change in MHC class I molecules on the rate of progression to AIDS. N. Engl. J. Med. 2001, 344, 1668–1675. [Google Scholar] [CrossRef] [PubMed]
- Migueles, S.A.; Sabbaghian, M.S.; Shupert, W.L.; Bettinotti, M.P.; Marincola, F.M.; Martino, L.; Hallahan, C.W.; Selig, S.M.; Schwartz, D.; Sullivan, J.; et al. HLA B*5701 is highly associated with restriction of virus replication in a subgroup of HIV-infected long term nonprogressors. Proc. Natl. Acad. Sci. USA 2000, 97, 2709–2714. [Google Scholar] [CrossRef] [PubMed]
- Kaslow, R.A.; Carrington, M.; Apple, R.; Park, L.; Munoz, A.; Saah, A.J.; Goedert, J.J.; Winkler, C.; O'Brien, S.J.; Rinaldo, C.; et al. Influence of combinations of human major histocompatibility complex genes on the course of HIV-1 infection. Nat. Med. 1996, 2, 405–411. [Google Scholar] [CrossRef] [PubMed]
- Hendel, H.; Caillat-Zucman, S.; Lebuanec, H.; Carrington, M.; O'Brien, S.; Andrieu, J.M.; Schachter, F.; Zagury, D.; Rappaport, J.; Winkler, C.; et al. New class I and II HLA alleles strongly associated with opposite patterns of progression to AIDS. J. Immunol. 1999, 162, 6942–6946. [Google Scholar] [PubMed]
- Altfeld, M.; Addo, M.M.; Rosenberg, E.S.; Hecht, F.M.; Lee, P.K.; Vogel, M.; Yu, X.G.; Draenert, R.; Johnston, M.N.; Strick, D.; et al. Influence of HLA-B57 on clinical presentation and viral control during acute HIV-1 infection. AIDS 2003, 17, 2581–2591. [Google Scholar] [CrossRef] [PubMed]
- Catano, G.; Kulkarni, H.; He, W.; Marconi, V.C.; Agan, B.K.; Landrum, M.; Anderson, S.; Delmar, J.; Telles, V.; Song, L.; et al. HIV-1 disease-influencing effects associated with ZNRD1, HCP5 and HLA-C alleles are attributable mainly to either HLA-A10 or HLA-B*57 alleles. PLoS ONE 2008, 3, e3636. [Google Scholar] [CrossRef] [PubMed]
- Antoni, G.; Guergnon, J.; Meaudre, C.; Samri, A.; Boufassa, F.; Goujard, C.; Lambotte, O.; Autran, B.; Rouzioux, C.; Costagliola, D.; et al. MHC-driven HIV-1 control on the long run is not systematically determined at early times post-HIV-1 infection. AIDS 2013, 27, 1707–1716. [Google Scholar] [CrossRef] [PubMed]
- Moroni, M.; Ghezzi, S.; Baroli, P.; Heltai, S.; De Battista, D.; Pensieroso, S.; Cavarelli, M.; Dispinseri, S.; Vanni, I.; Pastori, C.; et al. Spontaneous control of HIV-1 viremia in a subject with protective HLA-B plus HLA-C alleles and HLA-C associated single nucleotide polymorphisms. J. Transl. Med. 2014, 12, 335. [Google Scholar] [CrossRef] [PubMed]
- Ballana, E.; Ruiz-de Andres, A.; Mothe, B.; Ramirez de Arellano, E.; Aguilar, F.; Badia, R.; Grau, E.; Clotet, B.; del Val, M.; Brander, C.; et al. Differential prevalence of the HLA-C—35 CC genotype among viremic long term non-progressor and elite controller HIV+ individuals. Immunobiology 2012, 217, 889–894. [Google Scholar] [CrossRef] [PubMed]
- Julg, B.; Moodley, E.S.; Qi, Y.; Ramduth, D.; Reddy, S.; Mncube, Z.; Gao, X.; Goulder, P.J.; Detels, R.; Ndung'u, T.; Walker, B.D.; Carrington, M. Possession of HLA class II DRB1*1303 associates with reduced viral loads in chronic HIV-1 clade C and B infection. J. Infect. Dis. 2011, 203, 803–809. [Google Scholar] [CrossRef] [PubMed]
- Ranasinghe, S.; Cutler, S.; Davis, I.; Lu, R.; Soghoian, D.Z.; Qi, Y.; Sidney, J.; Kranias, G.; Flanders, M.D.; Lindqvist, M.; et al. Association of HLA-DRB1-restricted CD4(+) T cell responses with HIV immune control. Nat. Med. 2013, 19, 930–933. [Google Scholar] [CrossRef] [PubMed]
- Ramana, G.V.; Vasanthi, A.; Khaja, M.; Su, B.; Govindaiah, V.; Jin, L.; Singh, L.; Chakraborty, R. Distribution of HIV-1 resistance-conferring polymorphic alleles SDF-1-3′A, CCR2-64I and CCR5-Delta32 in diverse populations of Andhra Pradesh, South India. J. Genet. 2001, 80, 137–140. [Google Scholar] [CrossRef] [PubMed]
- Voevodin, A.; Samilchuk, E.; Dashti, S. Frequencies of SDF-1 chemokine, CCR-5, and CCR-2 chemokine receptor gene alleles conferring resistance to human immunodeficiency virus type 1 and AIDS in Kuwaitis. J. Med. Virol. 1999, 58, 54–58. [Google Scholar] [CrossRef]
- Chatterjee, A.; Rathore, A.; Vidyant, S.; Kakkar, K.; Dhole, T.N. Chemokines and chemokine receptors in susceptibility to HIV-1 infection and progression to AIDS. Dis. Markers 2012, 32, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.W.; Carrington, M.; Winkler, C.; Lomb, D.; Dean, M.; Huttley, G.; O'Brien, S.J. CCR2 chemokine receptor and AIDS progression. Nat. Med. 1997, 3, 1052–1053. [Google Scholar] [CrossRef] [PubMed]
- Lama, J.; Planelles, V. Host factors influencing susceptibility to HIV infection and AIDS progression. Retrovirology 2007, 4, 52. [Google Scholar] [CrossRef] [PubMed]
- Faure, S.; Meyer, L.; Costagliola, D.; Vaneensberghe, C.; Genin, E.; Autran, B.; Delfraissy, J.F.; McDermott, D.H.; Murphy, P.M.; Debre, P.; et al. Rapid progression to AIDS in HIV+ individuals with a structural variant of the chemokine receptor CX3CR1. Science 2000, 287, 2274–2277. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.K.; Hughes, M.D.; Chen, J.; Spector, S.A. Genetic polymorphisms in CX3CR1 predict HIV-1 disease progression in children independently of CD4+ lymphocyte count and HIV-1 RNA load. J. Infect. Dis. 2005, 191, 1971–1980. [Google Scholar] [CrossRef] [PubMed]
- Vidal, F.; Vilades, C.; Domingo, P.; Broch, M.; Pedrol, E.; Dalmau, D.; Knobel, H.; Peraire, J.; Gutierrez, C.; Sambeat, M.A.; et al. Spanish HIV-1-infected long-term nonprogressors of more than 15 years have an increased frequency of the CX3CR1 249I variant allele. J. Acquir. Immune Defic. Syndr. 2005, 40, 527–531. [Google Scholar] [CrossRef] [PubMed]
- Menten, P.; Wuyts, A.; van Damme, J. Macrophage inflammatory protein-1. Cytokine Growth Factor Rev. 2002, 13, 455–481. [Google Scholar] [CrossRef]
- Irving, S.G.; Zipfel, P.F.; Balke, J.; McBride, O.W.; Morton, C.C.; Burd, P.R.; Siebenlist, U.; Kelly, K. Two inflammatory mediator cytokine genes are closely linked and variably amplified on chromosome 17q. Nucleic Acids Res. 1990, 18, 3261–3270. [Google Scholar] [CrossRef] [PubMed]
- Nasi, M.; Riva, A.; Borghi, V.; D'Amico, R.; Del Giovane, C.; Casoli, C.; Galli, M.; Vicenzi, E.; Gibellini, L.; De Biasi, S.; et al. Novel genetic association of TNF-alpha-238 and PDCD1–7209 polymorphisms with long-term non-progressive HIV-1 infection. Int. J. Infect. Dis. 2013, 17, e845–e850. [Google Scholar] [CrossRef] [PubMed]
- Hetherington, S.; McGuirk, S.; Powell, G.; Cutrell, A.; Naderer, O.; Spreen, B.; Lafon, S.; Pearce, G.; Steel, H. Hypersensitivity reactions during therapy with the nucleoside reverse transcriptase inhibitor abacavir. Clin. Ther. 2001, 23, 1603–1614. [Google Scholar] [CrossRef]
- Mallal, S.; Nolan, D.; Witt, C.; Masel, G.; Martin, A.M.; Moore, C.; Sayer, D.; Castley, A.; Mamotte, C.; Maxwell, D.; et al. Association between presence of HLA-B*5701, HLA-DR7, and HLA-DQ3 and hypersensitivity to HIV-1 reverse-transcriptase inhibitor abacavir. Lancet 2002, 359, 727–732. [Google Scholar] [CrossRef]
- Hetherington, S.; Hughes, A.R.; Mosteller, M.; Shortino, D.; Baker, K.L.; Spreen, W.; Lai, E.; Davies, K.; Handley, A.; Dow, D.J.; et al. Genetic variations in HLA-B region and hypersensitivity reactions to abacavir. Lancet 2002, 359, 1121–1122. [Google Scholar] [CrossRef]
- Martin, A.M.; Nolan, D.; Gaudieri, S.; Almeida, C.A.; Nolan, R.; James, I.; Carvalho, F.; Phillips, E.; Christiansen, F.T.; Purcell, A.W.; et al. Predisposition to abacavir hypersensitivity conferred by HLA-B*5701 and a haplotypic Hsp70-Hom variant. Proc. Natl. Acad. Sci. USA 2004, 101, 4180–4185. [Google Scholar] [CrossRef] [PubMed]
- Hughes, A.R.; Mosteller, M.; Bansal, A.T.; Davies, K.; Haneline, S.A.; Lai, E.H.; Nangle, K.; Scott, T.; Spreen, W.R.; Warren, L.L.; et al. Association of genetic variations in HLA-B region with hypersensitivity to abacavir in some, but not all, populations. Pharmacogenomics 2004, 5, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Phillips, E.J.; Wong, G.A.; Kaul, R.; Shahabi, K.; Nolan, D.A.; Knowles, S.R.; Martin, A.M.; Mallal, S.A.; Shear, N.H. Clinical and immunogenetic correlates of abacavir hypersensitivity. AIDS 2005, 19, 979–981. [Google Scholar] [CrossRef] [PubMed]
- Rauch, A.; Nolan, D.; Martin, A.; McKinnon, E.; Almeida, C.; Mallal, S. Prospective genetic screening decreases the incidence of abacavir hypersensitivity reactions in the Western Australian HIV cohort study. Clin. Infect. Dis. 2006, 43, 99–102. [Google Scholar] [CrossRef] [PubMed]
- Mallal, S.; Phillips, E.; Carosi, G.; Molina, J.M.; Workman, C.; Tomazic, J.; Jagel-Guedes, E.; Rugina, S.; Kozyrev, O.; Cid, J.F.; et al. HLA-B*5701 screening for hypersensitivity to abacavir. N. Engl. J. Med. 2008, 358, 568–579. [Google Scholar] [CrossRef] [PubMed]
- Gilliam, B.L.; Riedel, D.J.; Redfield, R.R. Clinical use of CCR5 inhibitors in HIV and beyond. J. Transl. Med. 2011, 9 (Suppl. 1), S9. [Google Scholar] [CrossRef] [PubMed]
- Yukl, S.A.; Boritz, E.; Busch, M.; Bentsen, C.; Chun, T.W.; Douek, D.; Eisele, E.; Haase, A.; Ho, Y.C.; Hutter, G.; et al. Challenges in detecting HIV persistence during potentially curative interventions: A study of the Berlin patient. PLoS Pathog. 2013, 9, e1003347. [Google Scholar] [CrossRef] [PubMed]
- Zack, J.A.; Cann, A.J.; Lugo, J.P.; Chen, I.S. HIV-1 production from infected peripheral blood T cells after HTLV-I induced mitogenic stimulation. Science 1988, 240, 1026–1029. [Google Scholar] [CrossRef] [PubMed]
- Bartholomew, C.; Blattner, W.; Cleghorn, F. Progression to AIDS in homosexual men co-infected with HIV and HTLV-I in Trinidad. Lancet 1987, 2, 1469. [Google Scholar] [CrossRef]
- Sobesky, M.; Couppie, P.; Pradinaud, R.; Godard, M.C.; Alvarez, F.; Benoit, B.; Carme, B.; Lebeux, P.; et al. Coinfection with HIV and HTLV-I infection and survival in AIDS stage. French Guiana Study. GECVIG (Clinical HIV Study Group in Guiana). Presse Med. 2000, 29, 413–416. [Google Scholar] [PubMed]
- Bovolenta, C.; Pilotti, E.; Mauri, M.; Turci, M.; Ciancianaini, P.; Fisicaro, P.; Bertazzoni, U.; Poli, G.; Casoli, C. Human T-cell leukemia virus type 2 induces survival and proliferation of CD34(+) TF-1 cells through activation of STAT1 and STAT5 by secretion of interferon-gamma and granulocyte macrophage-colony-stimulating factor. Blood 2002, 99, 224–231. [Google Scholar] [CrossRef] [PubMed]
- Casoli, C.; Vicenzi, E.; Cimarelli, A.; Magnani, G.; Ciancianaini, P.; Cattaneo, E.; Dall'Aglio, P.; Poli, G.; Bertazzoni, U. HTLV-II down-regulates HIV-1 replication in IL-2-stimulated primary PBMC of coinfected individuals through expression of MIP-1alpha. Blood 2000, 95, 2760–2769. [Google Scholar] [PubMed]
- Balistrieri, G.; Barrios, C.; Castillo, L.; Umunakwe, T.C.; Giam, C.Z.; Zhi, H.; Beilke, M.A.; et al. Induction of CC-chemokines with antiviral function in macrophages by the human T lymphotropic virus type 2 transactivating protein, Tax2. Viral Immunol. 2013, 26, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Lewis, M.J.; Gautier, V.W.; Wang, X.P.; Kaplan, M.H.; Hall, W.W. Spontaneous production of C-C chemokines by individuals infected with human T lymphotropic virus type II (HTLV-II) alone and HTLV-II/HIV-1 coinfected individuals. J. Immunol. 2000, 165, 4127–4132. [Google Scholar] [CrossRef] [PubMed]
- Dragic, T.; Litwin, V.; Allaway, G.P.; Martin, S.R.; Huang, Y.; Nagashima, K.A.; Cayanan, C.; Maddon, P.J.; Koup, R.A.; Moore, J.P.; et al. HIV-1 entry into CD4+ cells is mediated by the chemokine receptor CC-CKR-5. Nature 1996, 381, 667–673. [Google Scholar] [CrossRef] [PubMed]
- Connor, R.I.; Sheridan, K.E.; Ceradini, D.; Choe, S.; Landau, N.R. Change in coreceptor use correlates with disease progression in HIV-1--infected individuals. J. Exp. Med. 1997, 185, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Ullum, H.; Cozzi Lepri, A.; Victor, J.; Aladdin, H.; Phillips, A.N.; Gerstoft, J.; Skinhoj, P.; Pedersen, B.K. Production of beta-chemokines in human immunodeficiency virus (HIV) infection: Evidence that high levels of macrophage inflammatory protein-1beta are associated with a decreased risk of HIV disease progression. J. Infect. Dis. 1998, 177, 331–336. [Google Scholar] [CrossRef] [PubMed]
- Cocchi, F.; DeVico, A.L.; Yarchoan, R.; Redfield, R.; Cleghorn, F.; Blattner, W.A.; Garzino-Demo, A.; Colombini-Hatch, S.; Margolis, D.; Gallo, R.C. Higher macrophage inflammatory protein (MIP)-1alpha and MIP-1beta levels from CD8+ T cells are associated with asymptomatic HIV-1 infection. Proc. Natl. Acad. Sci. USA 2000, 97, 13812–13817. [Google Scholar] [CrossRef] [PubMed]
- Ferbas, J.; Giorgi, J.V.; Amini, S.; Grovit-Ferbas, K.; Wiley, D.J.; Detels, R.; Plaeger, S. Antigen-specific production of RANTES, macrophage inflammatory protein (MIP)-1alpha, and MIP-1beta in vitro is a correlate of reduced human immunodeficiency virus burden in vivo. J. Infect. Dis. 2000, 182, 1247–1250. [Google Scholar] [CrossRef] [PubMed]
- Magnani, G.; Elia, G.; Casoli, C.; Calzetti, C.; Degli Antoni, A.; Fiaccadori, F. HTLV-II does not adversely affect the natural history of HIV-1 infection in intravenous drug users. Infection 1995, 23, 63. [Google Scholar] [CrossRef] [PubMed]
- Cimarelli, A.; Duclos, C.A.; Gessain, A.; Cattaneo, E.; Casoli, C.; Biglione, M.; Mauclere, P.; Bertazzoni, U. Quantification of HTLV-II proviral copies by competitive polymerase chain reaction in peripheral blood mononuclear cells of Italian injecting drug users, central Africans, and Amerindians. J. Acquir. Immune Defic. Syndr. Hum. Retrovirol. 1995, 10, 198–204. [Google Scholar] [CrossRef] [PubMed]
- Beilke, M.A.; Theall, K.P.; O'Brien, M.; Clayton, J.L.; Benjamin, S.M.; Winsor, E.L.; Kissinger, P.J. Clinical outcomes and disease progression among patients coinfected with HIV and human T lymphotropic virus types 1 and 2. Clin. Infect. Dis. 2004, 39, 256–263. [Google Scholar] [CrossRef] [PubMed]
- Bassani, S.; Lopez, M.; Toro, C.; Jimenez, V.; Sempere, J.M.; Soriano, V.; Benito, J.M. Influence of human T cell lymphotropic virus type 2 coinfection on virological and immunological parameters in HIV type 1-infected patients. Clin. Infect. Dis. 2007, 44, 105–110. [Google Scholar] [CrossRef] [PubMed]
- National Institutes of Health. Consensus Development Conference Statement: Management of hepatitis C 2002 (June 10–12, 2002). Gastroenterology 2002, 123, 2082–2099. [Google Scholar]
- Mohd-Hanafiah, K.; Groeger, J.; Flaxman, A.D.; Wiersma, S.T. Global epidemiology of hepatitis C virus infection: New estimates of age-specific antibody to HCV seroprevalence. Hepatology 2013, 57, 1333–1342. [Google Scholar] [CrossRef] [PubMed]
- Shebl, F.M.; Pfeiffer, R.M.; Buckett, D.; Muchmore, B.; Chen, S.; Dotrang, M.; Prokunina-Olsson, L.; Edlin, B.R.; O'Brien, T.R. IL28B rs12979860 genotype and spontaneous clearance of hepatitis C virus in a multi-ethnic cohort of injection drug users: Evidence for a supra-additive association. J. Infect. Dis. 2011, 204, 1843–1847. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.L.; Thio, C.L.; Martin, M.P.; Qi, Y.; Ge, D.; O'Huigin, C.; Kidd, J.; Kidd, K.; Khakoo, S.I.; Alexander, G.; et al. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature 2009, 461, 798–801. [Google Scholar] [CrossRef] [PubMed]
- Aka, P.V.; Kuniholm, M.H.; Pfeiffer, R.M.; Wang, A.S.; Tang, W.; Chen, S.; Astemborski, J.; Plankey, M.; Villacres, M.C.; Peters, M.G.; et al. Association of the IFNL4-DeltaG Allele With Impaired Spontaneous Clearance of Hepatitis C Virus. J. Infect. Dis. 2014, 209, 350–354. [Google Scholar] [CrossRef] [PubMed]
- De Re, V.; Caggiari, L.; De Zorzi, M.; Repetto, O.; Zignego, A.L.; Izzo, F.; Tornesello, M.L.; Buonaguro, F.M.; Mangia, A.; Sansonno, D.; et al. Genetic diversity of the KIR/HLA system and susceptibility to hepatitis C virus-related diseases. PLoS ONE 2015, 10, e0117420. [Google Scholar]
- de Vasconcelos, J.M.; de Jesus Maues Pereira Moia, L.; Amaral Ido, S.; Miranda, E.C.; Cicalisetakeshita, L.Y.; de Oliveira, L.F.; de Araujo Melo Mendes, L.; Sastre, D.; Tamegao-Lopes, B.P.; de Aquino Pedroza, L.S.; et al. Association of killer cell immunoglobulin-like receptor polymorphisms with chronic hepatitis C and responses to therapy in Brazil. Genet. Mol. Biol. 2013, 36, 22–27. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Assone, T.; Paiva, A.; Fonseca, L.A.M.; Casseb, J. Genetic Markers of the Host in Persons Living with HTLV-1, HIV and HCV Infections. Viruses 2016, 8, 38. https://doi.org/10.3390/v8020038
Assone T, Paiva A, Fonseca LAM, Casseb J. Genetic Markers of the Host in Persons Living with HTLV-1, HIV and HCV Infections. Viruses. 2016; 8(2):38. https://doi.org/10.3390/v8020038
Chicago/Turabian StyleAssone, Tatiane, Arthur Paiva, Luiz Augusto M. Fonseca, and Jorge Casseb. 2016. "Genetic Markers of the Host in Persons Living with HTLV-1, HIV and HCV Infections" Viruses 8, no. 2: 38. https://doi.org/10.3390/v8020038
APA StyleAssone, T., Paiva, A., Fonseca, L. A. M., & Casseb, J. (2016). Genetic Markers of the Host in Persons Living with HTLV-1, HIV and HCV Infections. Viruses, 8(2), 38. https://doi.org/10.3390/v8020038





