HIV-1 Latency in Monocytes/Macrophages
Abstract
:1. Introduction
2. General Mechanisms of Latency
2.1. Pre-integration Latency
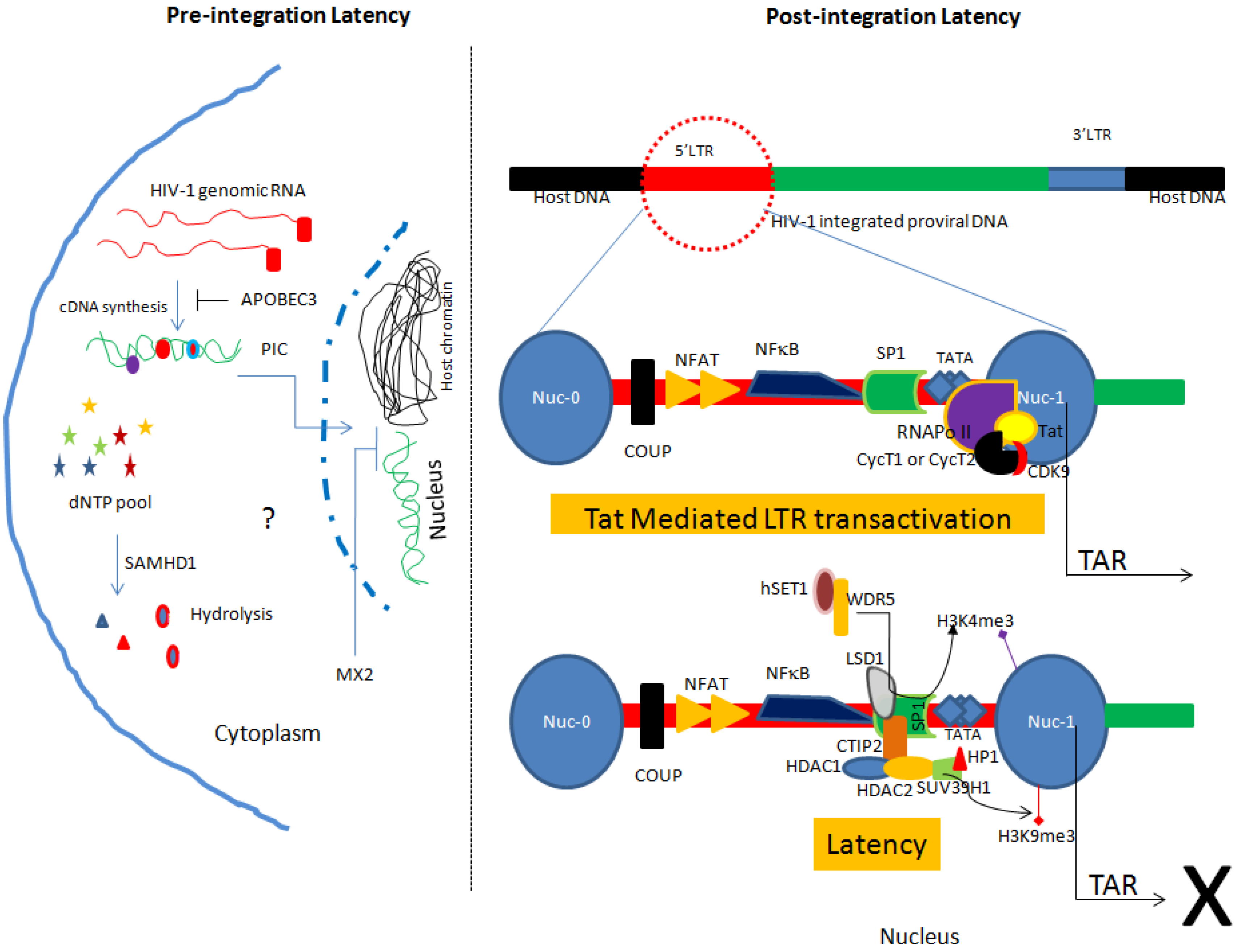
2.2. Post-integration Latency
2.2.1. Site of Integration and Chromatin Remodeling
2.2.2. Involvement of Host Transcription Factors and Viral Proteins in Latency
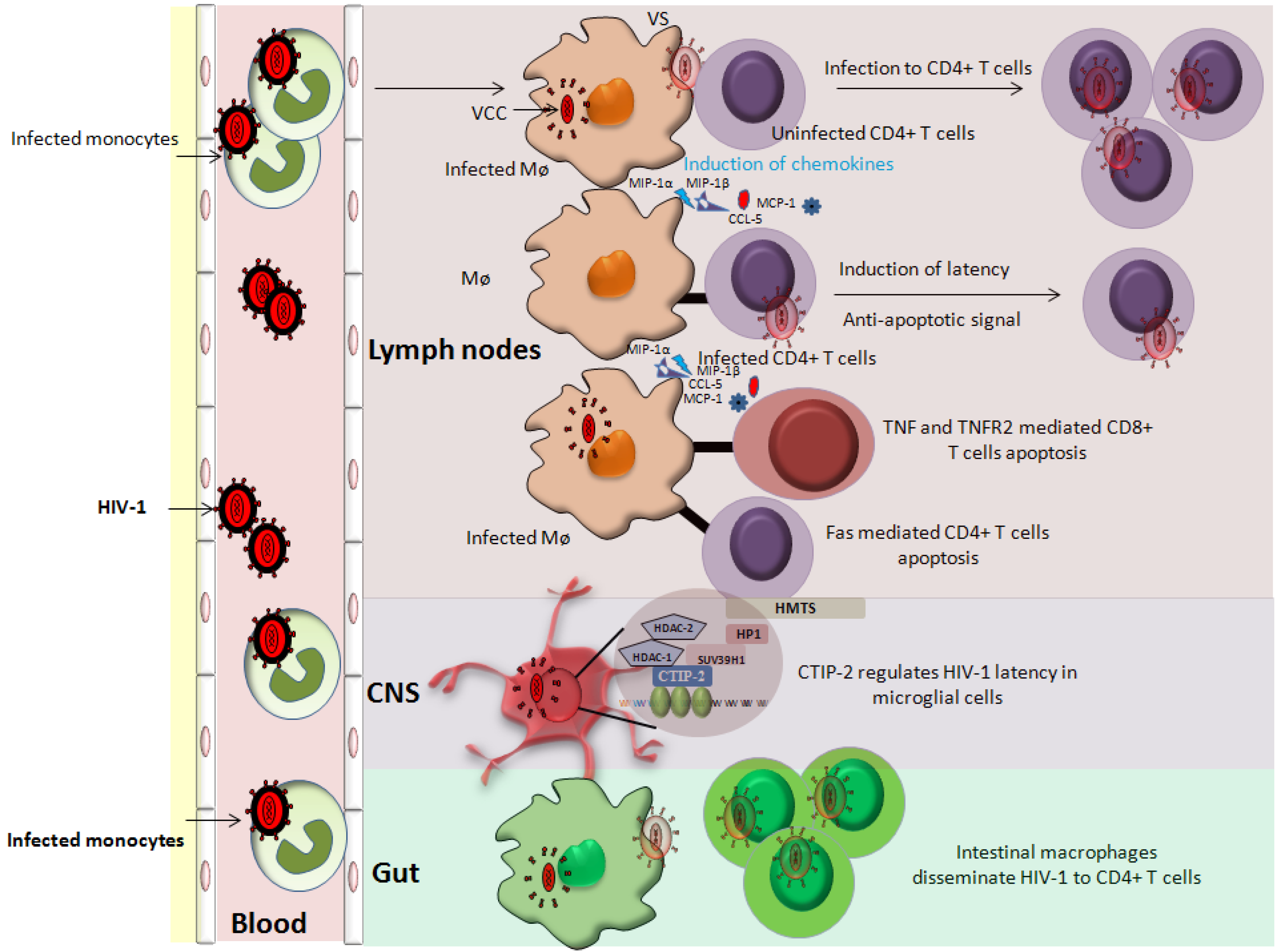
3. Latency in Monocyte/Macrophage Lineage Subsets
3.1. Latency in Monocytes
3.2. Latency in Macrophages
3.3. Latency in CNS Resident Macrophages
3.4. Latency in Gut-Associated Macrophages
3.5. Latency in Dendritic Cells
4. Role of Monocyte/Macrophage Lineage in Regulating Latency in CD4+ T Cells
5. Novel Therapeutic Strategies Against HIV-1 Latent Reservoirs
5.1. Integrase Inhibitors
5.2. “Flushing out” HIV-1 from the Latent Reservoirs
5.3. Apoptosis Inducing Agents
6. Conclusions
Acknowledgements
Author Contributions
Conflicts of Interest
References and Notes
- Cohen, M.S.; Chen, Y.Q.; McCauley, M.; Gamble, T.; Hosseinipour, M.C.; Kumarasamy, N.; Hakim, J.G.; Kumwenda, J.; Grinsztejn, B.; Pilotto, J.H.; et al. Prevention of HIV-1 infection with early antiretroviral therapy. N. Engl. J. Med. 2011, 365, 493–505. [Google Scholar] [CrossRef]
- Hammer, S.M.; Squires, K.E.; Hughes, M.D.; Grimes, J.M.; Demeter, L.M.; Currier, J.S.; Eron, J.J., Jr.; Feinberg, J.E.; Balfour, H.H., Jr.; Deyton, L.R.; et al. A controlled trial of two nucleoside analogues plus indinavir in persons with human immunodeficiency virus infection and CD4 cell counts of 200 per cubic millimeter or less. AIDS clinical trials group 320 study team. N. Engl. J. Med. 1997, 337, 725–733. [Google Scholar] [CrossRef]
- Davey, R.T., Jr.; Bhat, N.; Yoder, C.; Chun, T.W.; Metcalf, J.A.; Dewar, R.; Natarajan, V.; Lempicki, R.A.; Adelsberger, J.W.; Miller, K.D.; et al. HIV-1 and T cell dynamics after interruption of highly active antiretroviral therapy (HAART) in patients with a history of sustained viral suppression. Proc. Natl. Acad. Sci. USA 1999, 96, 15109–15114. [Google Scholar] [CrossRef]
- Varatharajan, L.; Thomas, S.A. The transport of anti-HIV drugs across blood-CNS interfaces: Summary of current knowledge and recommendations for further research. Antivir. Res. 2009, 82, A99–A109. [Google Scholar] [CrossRef]
- Van Lint, C.; Bouchat, S.; Marcello, A. HIV-1 transcription and latency: An update. Retrovirology 2013, 10. [Google Scholar] [CrossRef]
- Langer, R. Drug delivery. Drugs on target. Science 2001, 293, 58–59. [Google Scholar] [CrossRef]
- Wan, L.; Pooyan, S.; Hu, P.; Leibowitz, M.J.; Stein, S.; Sinko, P.J. Peritoneal macrophage uptake, pharmacokinetics and biodistribution of macrophage-targeted PEG-fMLF (N-formyl-methionyl-leucyl-phenylalanine) nanocarriers for improving HIV drug delivery. Pharm. Res. 2007, 24, 2110–2119. [Google Scholar] [CrossRef]
- Park, K. Nanotechnology: What it can do for drug delivery. J. Control. Release 2007, 120, 1–3. [Google Scholar] [CrossRef]
- Siliciano, R.F.; Greene, W.C. HIV latency. Cold Spring Harb. Perspect. Med. 2011, 1, a007096. [Google Scholar]
- Chun, T.W.; Engel, D.; Berrey, M.M.; Shea, T.; Corey, L.; Fauci, A.S. Early establishment of a pool of latently infected, resting CD4+ T cells during primary HIV-1 infection. Proc. Natl. Acad. Sci. USA 1998, 95, 8869–8873. [Google Scholar]
- Redel, L.; le Douce, V.; Cherrier, T.; Marban, C.; Janossy, A.; Aunis, D.; van Lint, C.; Rohr, O.; Schwartz, C. HIV-1 regulation of latency in the monocyte-macrophage lineage and in CD4+ T lymphocytes. J. Leukoc. Biol. 2010, 87, 575–588. [Google Scholar] [CrossRef]
- Finzi, D.; Hermankova, M.; Pierson, T.; Carruth, L.M.; Buck, C.; Chaisson, R.E.; Quinn, T.C.; Chadwick, K.; Margolick, J.; Brookmeyer, R.; et al. Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science 1997, 278, 1295–1300. [Google Scholar] [CrossRef]
- Eisele, E.; Siliciano, R.F. Redefining the viral reservoirs that prevent HIV-1 eradication. Immunity 2012, 37, 377–388. [Google Scholar] [CrossRef]
- Persaud, D.; Zhou, Y.; Siliciano, J.M.; Siliciano, R.F. Latency in human immunodeficiency virus type 1 infection: No easy answers. J. Virol. 2003, 77, 1659–1665. [Google Scholar] [CrossRef]
- Fu, M.; Gao, Y.; Zhou, Q.; Zhang, Q.; Peng, Y.; Tian, K.; Wang, J.; Zheng, X. Human cytomegalovirus latent infection alters the expression of cellular and viral miRNA. Gene 2014, 536, 272–278. [Google Scholar] [CrossRef]
- Tripathy, M.K.; Abbas, W.; Herbein, G. Epigenetic regulation of HIV-1 transcription. Epigenomics 2011, 3, 487–502. [Google Scholar] [CrossRef]
- Abbas, W.; Herbein, G. Plasma membrane signaling in HIV-1 infection. Biochim. Biophys. Acta 2014, 1838, 1132–1142. [Google Scholar] [CrossRef]
- Bukrinsky, M.I.; Sharova, N.; Dempsey, M.P.; Stanwick, T.L.; Bukrinskaya, A.G.; Haggerty, S.; Stevenson, M. Active nuclear import of human immunodeficiency virus type 1 preintegration complexes. Proc. Natl. Acad. Sci. USA 1992, 89, 6580–6584. [Google Scholar] [CrossRef]
- Marchand, C.; Johnson, A.A.; Semenova, E.; Pommier, Y. Mechanisms and inhibition of HIV integration. Drug Discov. Today Dis. Mech. 2006, 3, 253–260. [Google Scholar] [CrossRef]
- Han, Y.; Lassen, K.; Monie, D.; Sedaghat, A.R.; Shimoji, S.; Liu, X.; Pierson, T.C.; Margolick, J.B.; Siliciano, R.F.; Siliciano, J.D. Resting CD4+ T cells from human immunodeficiency virus type 1 (HIV-1)-infected individuals carry integrated HIV-1 genomes within actively transcribed host genes. J. Virol. 2004, 78, 6122–6133. [Google Scholar] [CrossRef]
- Koenig, S.; Gendelman, H.E.; Orenstein, J.M.; dal Canto, M.C.; Pezeshkpour, G.H.; Yungbluth, M.; Janotta, F.; Aksamit, A.; Martin, M.A.; Fauci, A.S. Detection of AIDS virus in macrophages in brain tissue from AIDS patients with encephalopathy. Science 1986, 233, 1089–1093. [Google Scholar]
- Koppensteiner, H.; Brack-Werner, R.; Schindler, M. Macrophages and their relevance in human immunodeficiency virus type 1 infection. Retrovirology 2012, 9. [Google Scholar] [CrossRef]
- Szabo, G.; Miller, C.L.; Kodys, K. Antigen presentation by the CD4 positive monocyte subset. J. Leukoc. Biol. 1990, 47, 111–120. [Google Scholar]
- Lee, B.; Sharron, M.; Montaner, L.J.; Weissman, D.; Doms, R.W. Quantification of CD4, CCR5, and CXCR4levels on lymphocyte subsets, dendritic cells, and differentially conditioned monocyte-derived macrophaes. Proc. Natl. Acad. Sci. USA 1999, 96, 5215–5220. [Google Scholar] [CrossRef]
- Le Douce, V.; Herbein, G.; Rohr, O.; Schwartz, C. Molecular mechanisms of HIV-1 persistence in the monocyte-macrophage lineage. Retrovirology 2010, 7. [Google Scholar] [CrossRef]
- Blankson, J.N.; Persaud, D.; Siliciano, R.F. The challenge of viral reservoirs in HIV-1 infection. Annu. Rev. Med. 2002, 53, 557–593. [Google Scholar] [CrossRef]
- Greene, W.C.; Peterlin, B.M. Charting HIV’s remarkable voyage through the cell: Basic science as a passport to future therapy. Nat. Med. 2002, 8, 673–680. [Google Scholar] [CrossRef]
- Zack, J.A.; Arrigo, S.J.; Weitsman, S.R.; Go, A.S.; Haislip, A.; Chen, I.S. HIV-1 entry into quiescent primary lymphocytes: Molecular analysis reveals a labile, latent viral structure. Cell 1990, 61, 213–222. [Google Scholar] [CrossRef]
- Bukrinsky, M.I.; Stanwick, T.L.; Dempsey, M.P.; Stevenson, M. Quiescent T lymphocytes as an inducible virus reservoir in HIV-1 infection. Science 1991, 254, 423–427. [Google Scholar]
- Chun, T.W.; Carruth, L.; Finzi, D.; Shen, X.; DiGiuseppe, J.A.; Taylor, H.; Hermankova, M.; Chadwick, K.; Margolick, J.; Quinn, T.C.; et al. Quantification of latent tissue reservoirs and total body viral load in HIV-1 infection. Nature 1997, 387, 183–188. [Google Scholar] [CrossRef]
- Pierson, T.C.; Zhou, Y.; Kieffer, T.L.; Ruff, C.T.; Buck, C.; Siliciano, R.F. Molecular characterization of preintegration latency in human immunodeficiency virus type 1 infection. J. Virol. 2002, 76, 8518–8531. [Google Scholar] [CrossRef]
- Kelly, J.; Beddall, M.H.; Yu, D.; Iyer, S.R.; Marsh, J.W.; Wu, Y. Human macrophages support persistent transcription from unintegrated HIV-1 DNA. Virology 2008, 372, 300–312. [Google Scholar] [CrossRef]
- Pang, S.; Koyanagi, Y.; Miles, S.; Wiley, C.; Vinters, H.V.; Chen, I.S. High levels of unintegrated HIV-1 DNA in brain tissue of AIDS dementia patients. Nature 1990, 343, 85–89. [Google Scholar] [CrossRef]
- Teo, I.; Veryard, C.; Barnes, H.; An, S.F.; Jones, M.; Lantos, P.L.; Luthert, P.; Shaunak, S. Circular forms of unintegrated human immunodeficiency virus type 1 DNA and high levels of viral protein expression: Association with dementia and multinucleated giant cells in the brains of patients with AIDS. J. Virol. 1997, 71, 2928–2933. [Google Scholar]
- Miyagi, E.; Schwartzkopff, F.; Plishka, R.; Buckler-White, A.; Clouse, K.A.; Strebel, K. APOBEC3G-independent reduction in virion infectivity during long-term HIV-1 replication in terminally differentiated macrophages. Virology 2008, 379, 266–274. [Google Scholar] [CrossRef]
- Lahouassa, H.; Daddacha, W.; Hofmann, H.; Ayinde, D.; Logue, E.C.; Dragin, L.; Bloch, N.; Maudet, C.; Bertrand, M.; Gramberg, T.; et al. SAMHD1 restricts the replication of human immunodeficiency virus type 1 by depleting the intracellular pool of deoxynucleoside triphosphates. Nat. Immunol. 2012, 13, 223–228. [Google Scholar]
- Goujon, C.; Moncorge, O.; Bauby, H.; Doyle, T.; Ward, C.C.; Schaller, T.; Hue, S.; Barclay, W.S.; Schulz, R.; Malim, M.H. Human MX2 is an interferon-induced post-entry inhibitor of HIV-1 infection. Nature 2013, 502, 559–562. [Google Scholar] [CrossRef]
- Liu, L.; Oliveira, N.M.; Cheney, K.M.; Pade, C.; Dreja, H.; Bergin, A.M.; Borgdorff, V.; Beach, D.H.; Bishop, C.L.; Dittmar, M.T.; et al. A whole genome screen for HIV restriction factors. Retrovirology 2011, 8. [Google Scholar] [CrossRef]
- Sheehy, A.M.; Gaddis, N.C.; Choi, J.D.; Malim, M.H. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature 2002, 418, 646–650. [Google Scholar] [CrossRef]
- Hrecka, K.; Hao, C.; Gierszewska, M.; Swanson, S.K.; Kesik-Brodacka, M.; Srivastava, S.; Florens, L.; Washburn, M.P.; Skowronski, J. Vpx relieves inhibition of HIV-1 infection of macrophages mediated by the SAMHD1 protein. Nature 2011, 474, 658–661. [Google Scholar] [CrossRef]
- Stopak, K.; de Noronha, C.; Yonemoto, W.; Greene, W.C. HIV-1 Vif blocks the antiviral activity of APOBEC3G by impairing both its translation and intracellular stability. Mol. Cell 2003, 12, 591–601. [Google Scholar] [CrossRef]
- Abbas, W.; Herbein, G. Molecular understanding of HIV-1 latency. Adv. Virol. 2012, 2012. [Google Scholar] [CrossRef]
- Demeret, C.; Vassetzky, Y.; Mechali, M. Chromatin remodelling and DNA replication: From nucleosomes to loop domains. Oncogene 2001, 20, 3086–3093. [Google Scholar] [CrossRef]
- Brady, T.; Kelly, B.; Male, F.; Roth, S.; Bailey, A.; Malani, N.; Gijsbers, R.; O’Doherty, U.; Bushman, F.D. Quantitation of HIV DNA integration: Effects of differential integration site distributions on Alu-PCR assays. J. Virol. Methods 2013, 189, 53–57. [Google Scholar] [CrossRef]
- Wu, X.; Li, Y.; Crise, B.; Burgess, S.M. Transcription start regions in the human genome are favored targets for MLV integration. Science 2003, 300, 1749–1751. [Google Scholar] [CrossRef]
- Taube, R.; Peterlin, M. Lost in transcription: Molecular mechanisms that control HIV latency. Viruses 2013, 5, 902–927. [Google Scholar] [CrossRef]
- Barr, S.D.; Ciuffi, A.; Leipzig, J.; Shinn, P.; Ecker, J.R.; Bushman, F.D. HIV integration site selection: Targeting in macrophages and the effects of different routes of viral entry. Mol. Ther. 2006, 14, 218–225. [Google Scholar] [CrossRef]
- Mack, K.D.; Jin, X.; Yu, S.; Wei, R.; Kapp, L.; Green, C.; Herndier, B.; Abbey, N.W.; Elbaggari, A.; Liu, Y.; et al. HIV insertions within and proximal to host cell genes are a common finding in tissues containing high levels of HIV DNA and macrophage-associated p24 antigen expression. J. Acquir. Immune Defic. Syndr. 2003, 33, 308–320. [Google Scholar] [CrossRef]
- Wellensiek, B.P.; Ramakrishnan, R.; Sundaravaradan, V.; Mehta, R.; Harris, D.T.; Ahmad, N. Differential HIV-1 integration targets more actively transcribed host genes in neonatal than adult blood mononuclear cells. Virology 2009, 385, 28–38. [Google Scholar] [CrossRef]
- Easley, R.; van Duyne, R.; Coley, W.; Guendel, I.; Dadgar, S.; Kehn-Hall, K.; Kashanchi, F. Chromatin dynamics associated with HIV-1 Tat-activated transcription. Biochim. Biophys. Acta 2010, 1799, 275–285. [Google Scholar] [CrossRef]
- Grewal, S.I.; Moazed, D. Heterochromatin and epigenetic control of gene expression. Science 2003, 301, 798–802. [Google Scholar] [CrossRef]
- Stewart, M.D.; Li, J.; Wong, J. Relationship between histone H3 lysine 9 methylation, transcription repression, and heterochromatin protein 1 recruitment. Mol. Cell. Biol. 2005, 25, 2525–2538. [Google Scholar] [CrossRef]
- Marban, C.; Suzanne, S.; Dequiedt, F.; de Walque, S.; Redel, L.; van Lint, C.; Aunis, D.; Rohr, O. Recruitment of chromatin-modifying enzymes by CTIP2 promotes HIV-1 transcriptional silencing. EMBO J. 2007, 26, 412–423. [Google Scholar] [CrossRef]
- Du Chene, I.; Basyuk, E.; Lin, Y.L.; Triboulet, R.; Knezevich, A.; Chable-Bessia, C.; Mettling, C.; Baillat, V.; Reynes, J.; Corbeau, P.; et al. SUV39H1 and HPL1gamma are responsible for chromatin-mediated HIV-1 transcriptional silencing and post-integration latency. EMBO J. 2007, 26, 424–435. [Google Scholar] [CrossRef]
- Alexaki, A.; Liu, Y.; Wigdahl, B. Cellular reservoirs of HIV-1 and their role in viral persistence. Curr. HIV Res. 2008, 6, 388–400. [Google Scholar] [CrossRef]
- Bouchat, S.; Gatot, J.S.; Kabeya, K.; Cardona, C.; Colin, L.; Herbein, G.; de Wit, S.; Clumeck, N.; Lambotte, O.; Rouzioux, C.; et al. Histone methyltransferase inhibitors induce HIV-1 recovery in resting CD4+ T cells from HIV-1-infected HAART-treated patients. AIDS 2012, 26, 1473–1482. [Google Scholar] [CrossRef]
- Verdin, E.; Paras, P., Jr.; van Lint, C. Chromatin disruption in the promoter of human immunodeficiency virus type 1 during transcriptional activation. EMBO J. 1993, 12, 3249–3259. [Google Scholar]
- Jiang, G.; Espeseth, A.; Hazuda, D.J.; Margolis, D.M. C-Myc and Sp1 contribute to proviral latency by recruiting histone deacetylase 1 to the human immunodeficiency virus type 1 promoter. J. Virol. 2007, 81, 10914–10923. [Google Scholar] [CrossRef]
- Coull, J.J.; Romerio, F.; Sun, J.M.; Volker, J.L.; Galvin, K.M.; Davie, J.R.; Shi, Y.; Hansen, U.; Margolis, D.M. The human factors YY1 and LSF repress the human immunodeficiency virus type 1 long terminal repeat via recruitment of histone deacetylase 1. J. Virol. 2000, 74, 6790–6799. [Google Scholar] [CrossRef]
- He, G.; Margolis, D.M. Counterregulation of chromatin deacetylation and histone deacetylase occupancy at the integrated promoter of human immunodeficiency virus type 1 (HIV-1) by the HIV-1 repressor YY1 and HIV-1 activator Tat. Mol. Cell. Biol. 2002, 22, 2965–2973. [Google Scholar] [CrossRef]
- Tyagi, M.; Karn, J. CBF-1 promotes transcriptional silencing during the establishment of HIV-1 latency. EMBO J. 2007, 26, 4985–4995. [Google Scholar] [CrossRef]
- Desplats, P.; Dumaop, W.; Smith, D.; Adame, A.; Everall, I.; Letendre, S.; Ellis, R.; Cherner, M.; Grant, I.; Masliah, E. Molecular and pathologic insights from latent HIV-1 infection in the human brain. Neurology 2013, 80, 1415–1423. [Google Scholar] [CrossRef]
- Le Douce, V.; Colin, L.; Redel, L.; Cherrier, T.; Herbein, G.; Aunis, D.; Rohr, O.; van Lint, C.; Schwartz, C. LSD1 cooperates with CTIP2 to promote HIV-1 transcriptional silencing. Nucleic Acids Res. 2012, 40, 1904–1915. [Google Scholar] [CrossRef]
- Le Douce, V.; Janossy, A.; Hallay, H.; Ali, S.; Riclet, R.; Rohr, O.; Schwartz, C. Achieving a cure for HIV infection: Do we have reasons to be optimistic? J. Antimicrob. Chemother. 2012, 67, 1063–1074. [Google Scholar] [CrossRef]
- Sakane, N.; Kwon, H.S.; Pagans, S.; Kaehlcke, K.; Mizusawa, Y.; Kamada, M.; Lassen, K.G.; Chan, J.; Greene, W.C.; Schnoelzer, M.; et al. Activation of HIV transcription by the viral Tat protein requires a demethylation step mediated by lysine-specific demethylase 1 (LSD1/KDM1). PLoS Pathog. 2011, 7, e1002184. [Google Scholar] [CrossRef]
- Cherrier, T.; le Douce, V.; Eilebrecht, S.; Riclet, R.; Marban, C.; Dequiedt, F.; Goumon, Y.; Paillart, J.C.; Mericskay, M.; Parlakian, A.; et al. CTIP2 is a negative regulator of P-TEFb. Proc. Natl. Acad. Sci. USA 2013, 110, 12655–12660. [Google Scholar] [CrossRef]
- Le Douce, V.; Cherrier, T.; Riclet, R.; Rohr, O.; Schwartz, C. The many lives of CTIP2: From aids to cancer and cardiac hypertrophy. J. Cell. Physiol. 2014, 229, 533–537. [Google Scholar] [CrossRef]
- Henderson, L.J.; Narasipura, S.D.; Adarichev, V.; Kashanchi, F.; Al-Harthi, L. Identification of novel T cell factor 4 (TCF-4) binding sites on the HIV long terminal repeat which associate with TCF-4, beta-catenin, and SMAR1 to repress HIV transcription. J. Virol. 2012, 86, 9495–9503. [Google Scholar] [CrossRef]
- Kumar, A.; Zloza, A.; Moon, R.T.; Watts, J.; Tenorio, A.R.; Al-Harthi, L. Active beta-catenin signaling is an inhibitory pathway for human immunodeficiency virus replication in peripheral blood mononuclear cells. J. Virol. 2008, 82, 2813–2820. [Google Scholar] [CrossRef]
- Karn, J. The molecular biology of HIV latency: Breaking and restoring the Tat-dependent transcriptional circuit. Curr. Opin. HIV AIDS 2011, 6, 4–11. [Google Scholar] [CrossRef]
- Price, D.H. P-TEFb, a cyclin-dependent kinase controlling elongation by RNA polymerase II. Mol. Cell. Biol. 2000, 20, 2629–2634. [Google Scholar] [CrossRef]
- Breuer, D.; Kotelkin, A.; Ammosova, T.; Kumari, N.; Ivanov, A.; Ilatovskiy, A.V.; Beullens, M.; Roane, P.R.; Bollen, M.; Petukhov, M.G.; et al. CDK2 regulates HIV-1 transcription by phosphorylation of CDK9 on serine 90. Retrovirology 2012, 9. [Google Scholar] [CrossRef]
- Ammosova, T.; Berro, R.; Kashanchi, F.; Nekhai, S. RNA interference directed to CDK2 inhibits HIV-1 transcription. Virology 2005, 341, 171–178. [Google Scholar] [CrossRef]
- Van Duyne, R.; Guendel, I.; Narayanan, A.; Gregg, E.; Shafagati, N.; Tyagi, M.; Easley, R.; Klase, Z.; Nekhai, S.; Kehn-Hall, K.; et al. Varying modulation of HIV-1 LTR activity by BAF complexes. J. Mol. Biol. 2011, 411, 581–596. [Google Scholar] [CrossRef]
- Dong, C.; Kwas, C.; Wu, L. Transcriptional restriction of human immunodeficiency virus type 1 gene expression in undifferentiated primary monocytes. J. Virol. 2009, 83, 3518–3527. [Google Scholar] [CrossRef]
- Carlson, K.A.; Leisman, G.; Limoges, J.; Pohlman, G.D.; Horiba, M.; Buescher, J.; Gendelman, H.E.; Ikezu, T. Molecular characterization of a putative antiretroviral transcriptional factor, OTK18. J. Immunol. 2004, 172, 381–391. [Google Scholar]
- Narayanan, A.; Kehn-Hall, K.; Bailey, C.; Kashanchi, F. Analysis of the roles of HIV-derived miRNAs. Expert Opin. Biol. Ther. 2011, 11, 17–29. [Google Scholar] [CrossRef]
- Haasnoot, J.; Berkhout, B. RNAi and cellular miRNAs in infections by mammalian viruses. Methods Mol. Biol. 2011, 721, 23–41. [Google Scholar] [CrossRef]
- Triboulet, R.; Mari, B.; Lin, Y.L.; Chable-Bessia, C.; Bennasser, Y.; Lebrigand, K.; Cardinaud, B.; Maurin, T.; Barbry, P.; Baillat, V.; et al. Suppression of miRNA-silencing pathway by HIV-1 during virus replication. Science 2007, 315, 1579–1582. [Google Scholar] [CrossRef]
- Wang, X.; Ye, L.; Hou, W.; Zhou, Y.; Wang, Y.J.; Metzger, D.S.; Ho, W.Z. Cellular miRNA expression correlates with susceptibility of monocytes/macrophages to HIV-1 infection. Blood 2009, 113, 671–674. [Google Scholar] [CrossRef]
- McElrath, M.J.; Steinman, R.M.; Cohn, Z.A. Latent HIV-1 infection in enriched populations of blood monocytes and T cells from seropositive patients. J. Clin. Invest. 1991, 87, 27–30. [Google Scholar] [CrossRef]
- Mikovits, J.A.; Lohrey, N.C.; Schulof, R.; Courtless, J.; Ruscetti, F.W. Activation of infectious virus from latent human immunodeficiency virus infection of monocytes in vivo. J. Clin. Invest. 1992, 90, 1486–1491. [Google Scholar] [CrossRef]
- Lambotte, O.; Taoufik, Y.; de Goer, M.G.; Wallon, C.; Goujard, C.; Delfraissy, J.F. Detection of infectious HIV in circulating monocytes from patients on prolonged highly active antiretroviral therapy. J. Acquir. Immune Defic. Syndr. 2000, 23, 114–119. [Google Scholar] [CrossRef]
- Zhu, T. HIV-1 in peripheral blood monocytes: An underrated viral source. J. Antimicrob. Chemother. 2002, 50, 309–311. [Google Scholar] [CrossRef]
- Zhu, T.; Muthui, D.; Holte, S.; Nickle, D.; Feng, F.; Brodie, S.; Hwangbo, Y.; Mullins, J.I.; Corey, L. Evidence for human immunodeficiency virus type 1 replication in vivo in CD14+ monocytes and its potential role as a source of virus in patients on highly active antiretroviral therapy. J. Virol. 2002, 76, 707–716. [Google Scholar] [CrossRef]
- Coleman, C.M.; Wu, L. HIV interactions with monocytes and dendritic cells: Viral latency and reservoirs. Retrovirology 2009, 6. [Google Scholar] [CrossRef]
- Perno, C.F.; Svicher, V.; Ceccherini-Silberstein, F. Novel drug resistance mutations in HIV: Recognition and clinical relevance. AIDS Rev. 2006, 8, 179–190. [Google Scholar]
- Khan, K.A.; Abbas, W.; Varin, A.; Kumar, A.; di Martino, V.; Dichamp, I.; Herbein, G. HIV-1 Nef interacts with HCV core, recruits TRAF2, TRAF5 and TRAF6, and stimulates HIV-1 replication in macrophages. J. Innate Immun. 2013, 5, 639–656. [Google Scholar] [CrossRef]
- Brown, A.; Zhang, H.; Lopez, P.; Pardo, C.A.; Gartner, S. In vitro modeling of the HIV-macrophage reservoir. J. Leukoc. Biol. 2006, 80, 1127–1135. [Google Scholar] [CrossRef]
- Gillim-Ross, L.; Cara, A.; Klotman, M.E. HIV-1 extrachromosomal 2-LTR circular DNA is long-lived in human macrophages. Viral. Immunol. 2005, 18, 190–196. [Google Scholar] [CrossRef]
- Caselli, E.; Galvan, M.; Cassai, E.; Caruso, A.; Sighinolfi, L.; di Luca, D. Human herpesvirus 8 enhances human immunodeficiency virus replication in acutely infected cells and induces reactivation in latently infected cells. Blood 2005, 106, 2790–2797. [Google Scholar] [CrossRef]
- Cassol, E.; Cassetta, L.; Rizzi, C.; Alfano, M.; Poli, G. M1 and M2a polarization of human monocyte-derived macrophages inhibits HIV-1 replication by distinct mechanisms. J. Immunol. 2009, 182, 6237–6246. [Google Scholar] [CrossRef]
- Herbein, G.; Varin, A. The macrophage in HIV-1 infection: From activation to deactivation? Retrovirology 2010, 7. [Google Scholar] [CrossRef]
- Williams, K.C.; Hickey, W.F. Central nervous system damage, monocytes and macrophages, and neurological disorders in AIDS. Annu. Rev. Neurosci. 2002, 25, 537–562. [Google Scholar] [CrossRef]
- He, J.; Chen, Y.; Farzan, M.; Choe, H.; Ohagen, A.; Gartner, S.; Busciglio, J.; Yang, X.; Hofmann, W.; Newman, W.; et al. CCR3 and CCR5 are co-receptors for HIV-1 infection of microglia. Nature 1997, 385, 645–649. [Google Scholar] [CrossRef]
- Cosenza, M.A.; Zhao, M.L.; Si, Q.; Lee, S.C. Human brain parenchymal microglia express CD14 and CD45 and are productively infected by HIV-1 in HIV-1 encephalitis. Brain Pathol. 2002, 12, 442–455. [Google Scholar] [CrossRef]
- Garden, G.A. Microglia in human immunodeficiency virus-associated neurodegeneration. Glia 2002, 40, 240–251. [Google Scholar] [CrossRef]
- Chakrabarti, L.; Hurtrel, M.; Maire, M.A.; Vazeux, R.; Dormont, D.; Montagnier, L.; Hurtrel, B. Early viral replication in the brain of SIV-infected rhesus monkeys. Am. J. Pathol. 1991, 139, 1273–1280. [Google Scholar]
- Davis, L.E.; Hjelle, B.L.; Miller, V.E.; Palmer, D.L.; Llewellyn, A.L.; Merlin, T.L.; Young, S.A.; Mills, R.G.; Wachsman, W.; Wiley, C.A. Early viral brain invasion in iatrogenic human immunodeficiency virus infection. Neurology 1992, 42, 1736–1739. [Google Scholar] [CrossRef]
- Smith, P.D.; Meng, G.; Salazar-Gonzalez, J.F.; Shaw, G.M. Macrophage HIV-1 infection and the gastrointestinal tract reservoir. J. Leukoc. Biol. 2003, 74, 642–649. [Google Scholar] [CrossRef]
- Shen, R.; Meng, G.; Ochsenbauer, C.; Clapham, P.R.; Grams, J.; Novak, L.; Kappes, J.C.; Smythies, L.E.; Smith, P.D. Stromal down-regulation of macrophage CD4/CCR5 expression and NF-kappaB activation mediates HIV-1 non-permissiveness in intestinal macrophages. PLoS Pathog. 2011, 7, e1002060. [Google Scholar] [CrossRef]
- Blauvelt, A.; Asada, H.; Saville, M.W.; Klaus-Kovtun, V.; Altman, D.J.; Yarchoan, R.; Katz, S.I. Productive infection of dendritic cells by HIV-1 and their ability to capture virus are mediated through separate pathways. J. Clin. Invest. 1997, 100, 2043–2053. [Google Scholar] [CrossRef]
- Patterson, S.; Rae, A.; Hockey, N.; Gilmour, J.; Gotch, F. Plasmacytoid dendritic cells are highly susceptible to human immunodeficiency virus type 1 infection and release infectious virus. J. Virol. 2001, 75, 6710–6713. [Google Scholar] [CrossRef]
- Kawamura, T.; Gulden, F.O.; Sugaya, M.; McNamara, D.T.; Borris, D.L.; Lederman, M.M.; Orenstein, J.M.; Zimmerman, P.A.; Blauvelt, A. R5 HIV productively infects langerhans cells, and infection levels are regulated by compound CCR5 polymorphisms. Proc. Natl. Acad. Sci. USA 2003, 100, 8401–8406. [Google Scholar] [CrossRef]
- Haase, A.T.; Henry, K.; Zupancic, M.; Sedgewick, G.; Faust, R.A.; Melroe, H.; Cavert, W.; Gebhard, K.; Staskus, K.; Zhang, Z.Q.; et al. Quantitative image analysis of HIV-1 infection in lymphoid tissue. Science 1996, 274, 985–989. [Google Scholar] [CrossRef]
- Otero, M.; Nunnari, G.; Leto, D.; Sullivan, J.; Wang, F.X.; Frank, I.; Xu, Y.; Patel, C.; Dornadula, G.; Kulkosky, J.; et al. Peripheral blood dendritic cells are not a major reservoir for HIV type 1 in infected individuals on virally suppressive HAART. AIDS Res. Hum. Retroviruses 2003, 19, 1097–1103. [Google Scholar] [CrossRef]
- Evans, V.A.; Kumar, N.; Filali, A.; Procopio, F.A.; Yegorov, O.; Goulet, J.P.; Saleh, S.; Haddad, E.K.; da Fonseca Pereira, C.; Ellenberg, P.C.; et al. Myeloid dendritic cells induce HIV-1 latency in non-proliferating CD4+ T cells. PLoS Pathog. 2013, 9, e1003799. [Google Scholar] [CrossRef]
- Welsch, S.; Groot, F.; Krausslich, H.G.; Keppler, O.T.; Sattentau, Q.J. Architecture and regulation of the HIV-1 assembly and holding compartment in macrophages. J. Virol. 2013, 85, 7922–7927. [Google Scholar]
- Gaudin, R.; Berre, S.; de Alencar, B.C.; Decalf, J.; Schindler, M.; Gobert, F.X.; Jouve, M.; Benaroch, P. Dynamics of HIV-containing compartments in macrophages reveal sequestration of virions and transient surface connections. PLoS One 2013, 8, e69450. [Google Scholar] [CrossRef]
- Tan, J.; Sattentau, Q.J. The HIV-1-containing macrophage compartment: A perfect cellular niche? Trends Microbiol. 2013, 21, 405–412. [Google Scholar] [CrossRef]
- Carr, J.M.; Hocking, H.; Li, P.; Burrell, C.J. Rapid and efficient cell-to-cell transmission of human immunodeficiency virus infection from monocyte-derived macrophages to peripheral blood lymphocytes. Virology 1999, 265, 319–329. [Google Scholar] [CrossRef]
- Sharova, N.; Swingler, C.; Sharkey, M.; Stevenson, M. Macrophages archive HIV-1 virions for dissemination in trans. EMBO J. 2005, 24, 2481–2489. [Google Scholar] [CrossRef]
- Groot, F.; Welsch, S.; Sattentau, Q.J. Efficient HIV-1 transmission from macrophages to T cells across transient virological synapses. Blood 2008, 111, 4660–4663. [Google Scholar] [CrossRef]
- Berre, S.; Gaudin, R.; de Alencar, B.C.; Desdouits, M.; Chabaud, M.; Naffakh, N.; Rabaza-Gairi, M.; Gobert, F.X.; Jouve, M.; Benaroch, P. CD36-specific antibodies block release of HIV-1 from infected primary macrophages and its transmission to T cells. J. Exp. Med. 2013, 210, 2523–2538. [Google Scholar] [CrossRef]
- Herbein, G.; Gras, G.; Khan, K.A.; Abbas, W. Macrophage signaling in HIV-1 infection. Retrovirology 2010, 7. [Google Scholar] [CrossRef]
- Badley, A.D.; Dockrell, D.; Simpson, M.; Schut, R.; Lynch, D.H.; Leibson, P.; Paya, C.V. Macrophage-dependent apoptosis of CD4+ T lymphocytes from HIV-infected individuals is mediated by FASL and tumor necrosis factor. J. Exp. Med. 1997, 185, 55–64. [Google Scholar] [CrossRef]
- Herbein, G.; Mahlknecht, U.; Batliwalla, F.; Gregersen, P.; Pappas, T.; Butler, J.; O’Brien, W.A.; Verdin, E. Apoptosis of CD8+ T cells is mediated by macrophages through interaction of HIV gp120 with chemokine receptor cxcr4. Nature 1998, 395, 189–194. [Google Scholar] [CrossRef]
- Mahlknecht, U.; Deng, C.; Lu, M.C.; Greenough, T.C.; Sullivan, J.L.; O’Brien, W.A.; Herbein, G. Resistance to apoptosis in HIV-infected CD4+ T lymphocytes is mediated by macrophages: Role for Nef and immune activation in viral persistence. J. Immunol. 2000, 165, 6437–6446. [Google Scholar]
- Devadas, K.; Hardegen, N.J.; Wahl, L.M.; Hewlett, I.K.; Clouse, K.A.; Yamada, K.M.; Dhawan, S. Mechanisms for macrophage-mediated HIV-1 induction. J. Immunol. 2004, 173, 6735–6744. [Google Scholar]
- Hazuda, D.J.; Felock, P.; Witmer, M.; Wolfe, A.; Stillmock, K.; Grobler, J.A.; Espeseth, A.; Gabryelski, L.; Schleif, W.; Blau, C.; et al. Inhibitors of strand transfer that prevent integration and inhibit HIV-1 replication in cells. Science 2000, 287, 646–650. [Google Scholar] [CrossRef]
- Malet, I.; Calvez, V.; Marcelin, A.G. The future of integrase inhibitors of HIV-1. Curr. Opin. Virol. 2012, 2, 580–587. [Google Scholar] [CrossRef]
- Bera, S.; Pandey, K.K.; Vora, A.C.; Grandgenett, D.P. Molecular interactions between HIV-1 integrase and the two viral DNA ends within the synaptic complex that mediates concerted integration. J. Mol. Biol. 2009, 389, 183–198. [Google Scholar] [CrossRef]
- Scopelliti, F.; Pollicita, M.; Ceccherini-Silberstein, F.; di Santo, F.; Surdo, M.; Aquaro, S.; Perno, C.F. Comparative antiviral activity of integrase inhibitors in human monocyte-derived macrophages and lymphocytes. Antivir. Res. 2011, 92, 255–261. [Google Scholar] [CrossRef]
- Marsden, M.D.; Avancena, P.; Kitchen, C.M.; Hubbard, T.; Zack, J.A. Single mutations in HIV integrase confer high-level resistance to raltegravir in primary human macrophages. Antimicrob. Agents Chemother. 2011, 55, 3696–3702. [Google Scholar] [CrossRef]
- Christ, F.; Voet, A.; Marchand, A.; Nicolet, S.; Desimmie, B.A.; Marchand, D.; Bardiot, D.; van der Veken, N.J.; Van Remoortel, B.; Strelkov, S.V.; et al. Rational design of small-molecule inhibitors of the LEDGF/p75-integrase interaction and HIV replication. Nat. Chem. Biol. 2010, 6, 442–448. [Google Scholar] [CrossRef]
- Christ, F.; Shaw, S.; Demeulemeester, J.; Desimmie, B.A.; Marchand, A.; Butler, S.; Smets, W.; Chaltin, P.; Westby, M.; Debyser, Z.; et al. Small-molecule inhibitors of the LEDGF/p75 binding site of integrase block HIV replication and modulate integrase multimerization. Antimicrob. Agents Chemother. 2012, 56, 4365–4374. [Google Scholar] [CrossRef]
- Christ, F.; Debyser, Z. The LEDGF/p75 integrase interaction, a novel target for anti-HIV therapy. Virology 2013, 435, 102–109. [Google Scholar] [CrossRef]
- Badley, A.D.; Sainski, A.; Wightman, F.; Lewin, S.R. Altering cell death pathways as an approach to cure HIV infection. Cell Death Dis. 2013, 4, e718. [Google Scholar] [CrossRef]
- Matalon, S.; Rasmussen, T.A.; Dinarello, C.A. Histone deacetylase inhibitors for purging HIV-1 from the latent reservoir. Mol. Med. 2011, 17, 466–472. [Google Scholar]
- Archin, N.M.; Liberty, A.L.; Kashuba, A.D.; Choudhary, S.K.; Kuruc, J.D.; Crooks, A.M.; Parker, D.C.; Anderson, E.M.; Kearney, M.F.; Strain, M.C.; et al. Administration of vorinostat disrupts HIV-1 latency in patients on antiretroviral therapy. Nature 2012, 487, 482–485. [Google Scholar] [CrossRef]
- Elliott, J.; Soloman, A.; Whightman, F.; Smith, M.; Palmer, S.; Prince, M.; Watson, J.; Hoy, J.; McMahon, J.; Lewin, S.R. The safety and effect of multiple doses of vorinostat on HIV transcription in HIV+ patients receiving cART. In Proceedings of the 20th Conference on Retroviruses and Opportunistic Infections, Atlanta, GA, USA, 3–6 March 2013. Abstract 50LB.
- Shehu-Xhilaga, M.; Rhodes, D.; Wightman, F.; Liu, H.B.; Solomon, A.; Saleh, S.; Dear, A.E.; Cameron, P.U.; Lewin, S.R. The novel histone deacetylase inhibitors metacept-1 and metacept-3 potently increase HIV-1 transcription in latently infected cells. AIDS 2009, 23, 2047–2050. [Google Scholar] [CrossRef]
- Warrilow, D.; Gardner, J.; Darnell, G.A.; Suhrbier, A.; Harrich, D. HIV type 1 inhibition by protein kinase C modulatory compounds. AIDS Res. Hum. Retroviruses 2006, 22, 854–864. [Google Scholar] [CrossRef]
- Korin, Y.D.; Zack, J.A. Nonproductive human immunodeficiency virus type 1 infection in nucleoside-treated G0 lymphocytes. J. Virol. 1999, 73, 6526–6532. [Google Scholar]
- Gulakowski, R.J.; McMahon, J.B.; Buckheit, R.W., Jr.; Gustafson, K.R.; Boyd, M.R. Antireplicative and anticytopathic activities of prostratin, a non-tumor-promoting phorbol ester, against human immunodeficiency virus (HIV). Antivir. Res. 1997, 33, 87–97. [Google Scholar] [CrossRef]
- Colin, L.; Van Lint, C. Molecular control of HIV-1 postintegration latency: Implications for the development of new therapeutic strategies. Retrovirology 2009, 6. [Google Scholar] [CrossRef]
- Greiner, D.; Bonaldi, T.; Eskeland, R.; Roemer, E.; Imhof, A. Identification of a specific inhibitor of the histone methyltransferase SU(VAR)3–9. Nat. Chem. Biol. 2005, 1, 143–145. [Google Scholar] [CrossRef]
- Miranda, T.B.; Cortez, C.C.; Yoo, C.B.; Liang, G.; Abe, M.; Kelly, T.K.; Marquez, V.E.; Jones, P.A. Dznep is a global histone methylation inhibitor that reactivates developmental genes not silenced by DNA methylation. Mol. Cancer Ther. 2009, 8, 1579–1588. [Google Scholar] [CrossRef]
- Fernandez, G.; Zaikos, T.D.; Khan, S.Z.; Jacobi, A.M.; Behlke, M.A.; Zeichner, S.L. Targeting Iκb proteins for HIV latency activation: The role of individual ikappab and NF-κB proteins. J. Virol. 2009, 87, 3966–3978. [Google Scholar]
- Kumar, A.; Abbas, W.; Herbein, G. TNF and TNF receptor superfamily members in HIV infection: New cellular targets for therapy? Mediat. Inflamm. 2013, 2013. [Google Scholar] [CrossRef]
- Chun, T.W.; Engel, D.; Mizell, S.B.; Hallahan, C.W.; Fischette, M.; Park, S.; Davey, R.T., Jr.; Dybul, M.; Kovacs, J.A.; Metcalf, J.A.; et al. Effect of interleukin-2 on the pool of latently infected, resting CD4+ T cells in HIV-1-infected patients receiving highly active anti-retroviral therapy. Nat. Med. 1999, 5, 651–655. [Google Scholar] [CrossRef]
- Scripture-Adams, D.D.; Brooks, D.G.; Korin, Y.D.; Zack, J.A. Interleukin-7 induces expression of latent human immunodeficiency virus type 1 with minimal effects on T-cell phenotype. J. Virol. 2002, 76, 13077–13082. [Google Scholar] [CrossRef]
- Oguariri, R.M.; Brann, T.W.; Imamichi, T. Hydroxyurea and interleukin-6 synergistically reactivate HIV-1 replication in a latently infected promonocytic cell line via Sp1/Sp3 transcription factors. J. Biol. Chem. 2007, 282, 3594–3604. [Google Scholar] [CrossRef]
- Zhang, H. Reversal of HIV-1 latency with anti-miRNA inhibitors. Int. J. Biochem. Cell Biol. 2009, 41, 451–454. [Google Scholar] [CrossRef]
- Shan, L.; Deng, K.; Shroff, N.S.; Durand, C.M.; Rabi, S.A.; Yang, H.C.; Zhang, H.; Margolick, J.B.; Blankson, J.N.; Siliciano, R.F. Stimulation of HIV-1-specific cytolyticT lymphocytes facilitates elimination of latent viral reservoir after virus reactivation. Immunity 2012, 36, 491–501. [Google Scholar] [CrossRef]
- Kim, D.; Cheng, G.Z.; Lindsley, C.W.; Yang, H.; Cheng, J.Q. Targeting the phosphatidylinositol-3 kinase/akt pathway for the treatment of cancer. Curr. Opin. Investig. Drugs 2005, 6, 1250–1258. [Google Scholar]
- Mita, A.C.; Mita, M.M.; Nawrocki, S.T.; Giles, F.J. Survivin: Key regulator of mitosis and apoptosis and novel target for cancer therapeutics. Clin. Cancer Res. 2008, 14, 5000–5005. [Google Scholar] [CrossRef]
- Azmi, A.S.; Wang, Z.; Philip, P.A.; Mohammad, R.M.; Sarkar, F.H. Emerging BCL-2 inhibitors for the treatment of cancer. Expert Opin. Emerg. Drugs 2011, 16, 59–70. [Google Scholar] [CrossRef]
- Tanaka, Y.; Kameoka, M.; Ota, K.; Itaya, A.; Ikuta, K.; Yoshihara, K. Establishment of persistentinfection with HIV-1 abrogates the caspase-3-dependent apoptotic signaling pathway in U937 cells. Exp. Cell Res. 1999, 247, 514–524. [Google Scholar] [CrossRef]
- Larrosa, P.N.F.; Croci, D.O.; Riva, D.A.; Bibini, M.; Luzzi, R.; Saracco, M.; Mersich, S.E.; Rabinovich, G.A.; Martinez Peralta, L. Apoptosis resistance in HIV-1 persistently-infected cells is independent of active viral replication and involves modulation of the apoptotic mitochondrial pathway. Retrovirology 2008, 5. [Google Scholar] [CrossRef]
- Hamilton, J.A. Colony-stimulating factors in inflammation and autoimmunity. Nat. Rev. Immunol. 2008, 8, 533–544. [Google Scholar] [CrossRef]
- Swingler, S.; Mann, A.M.; Zhou, J.; Swingler, C.; Stevenson, M. Apoptotic killing of HIV-1-infected macrophages is subverted by the viral envelope glycoprotein. PLoS Pathog. 2007, 3, 1281–1290. [Google Scholar]
- Busca, A.; Saxena, M.; Kumar, A. Critical role for antiapoptoticBCL-XL and MCL-1 in human macrophage survival and cellular IAP1/2 (CIAP1/2) in resistance to HIV-Vpr-induced apoptosis. J. Biol. Chem. 2012, 287, 15118–15133. [Google Scholar] [CrossRef]
- Abbas, W.; Khan, K.A.; Kumar, A.; Tripathy, M.K.; Dichamp, I.; Keita, M.; Mahlknecht, U.; Rohr, O.; Herbein, G. Blockade of BFA-mediated apoptosis in macrophages by the HIV-1 Nef protein. Cell Death Dis. 2014, 5, e1080. [Google Scholar] [CrossRef]
- Saxena, M.; Busca, A.; Pandey, S.; Kryworuchko, M.; Kumar, A. Cpg protects human monocytic cells against HIV-Vpr-induced apoptosis by cellular inhibitor of apoptosis-2 through the calcium-activated JNK pathway in a TLR9-independent manner. J. Immunol. 2011, 187, 5865–5878. [Google Scholar] [CrossRef]
- Herbein, G.; Varin, A.; Larbi, A.; Fortin, C.; Mahlknecht, U.; Fulop, T.; Aggarwal, B.B. Nef and TNF alpha are coplayers that favor HIV-1 replication in monocytic cells and primary macrophages. Curr. HIV Res. 2008, 6, 117–129. [Google Scholar] [CrossRef]
- Olivares, I.; Ballester, A.; Lombardia, L.; Dominguez, O.; Lopez-Galindez, C. Human immunodeficiency virus type 1 chronic infection is associated with different gene expression in MT-4, H9 and U937 cell lines. Virus Res. 2009, 139, 22–31. [Google Scholar] [CrossRef]
- Giri, M.S.; Nebozyhn, M.; Raymond, A.; Gekonge, B.; Hancock, A.; Creer, S.; Nicols, C.; Yousef, M.; Foulkes, A.S.; Mounzer, K.; et al. Circulating monocytes in HIV-1-infected viremic subjects exhibit an antiapoptosis gene signature and virus- and host-mediated apoptosis resistance. J. Immunol. 2009, 182, 4459–4470. [Google Scholar] [CrossRef]
- Hanauske-Abel, H.M.; Saxena, D.; Palumbo, P.E.; Hanauske, A.R.; Luchessi, A.D.; Cambiaghi, T.D.; Hoque, M.; Spino, M.; Gandolfi, D.D.; Heller, D.S.; et al. Drug-induced reactivation of apoptosis abrogates HIV-1 infection. PLoS One 2013, 8, e74414. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kumar, A.; Abbas, W.; Herbein, G. HIV-1 Latency in Monocytes/Macrophages. Viruses 2014, 6, 1837-1860. https://doi.org/10.3390/v6041837
Kumar A, Abbas W, Herbein G. HIV-1 Latency in Monocytes/Macrophages. Viruses. 2014; 6(4):1837-1860. https://doi.org/10.3390/v6041837
Chicago/Turabian StyleKumar, Amit, Wasim Abbas, and Georges Herbein. 2014. "HIV-1 Latency in Monocytes/Macrophages" Viruses 6, no. 4: 1837-1860. https://doi.org/10.3390/v6041837
APA StyleKumar, A., Abbas, W., & Herbein, G. (2014). HIV-1 Latency in Monocytes/Macrophages. Viruses, 6(4), 1837-1860. https://doi.org/10.3390/v6041837





