Structural and Immunoreactivity Properties of the SARS-CoV-2 Spike Protein upon the Development of an Inactivated Vaccine
Abstract
1. Introduction
2. Materials and Methods
2.1. Virus Growing and Inactivation
2.1.1. Virus and Cells
2.1.2. Infectious Activity Determination
2.1.3. Virus Inactivation
2.1.4. Virus Concentration and Purification
2.2. Electrophoretic Analysis and Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry (MALDI-TOF-MS) in-Gel Protein Identification
2.3. Transmission Electron Microscopy (TEM)
2.4. Enzyme-Linked Immunosorbent Assay (ELISA)
2.5. Statistical Analysis
3. Results
3.1. Validation of Inactivation
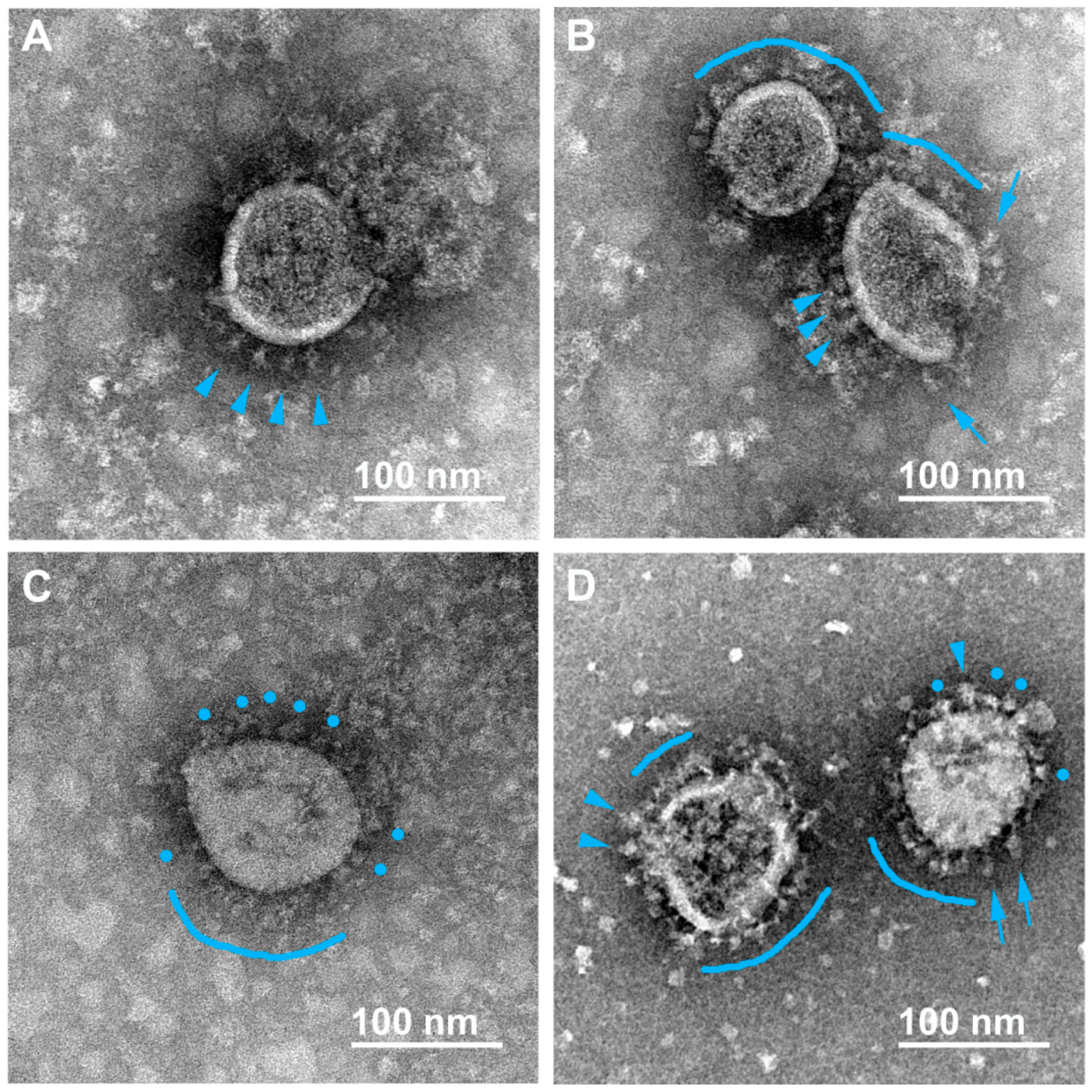
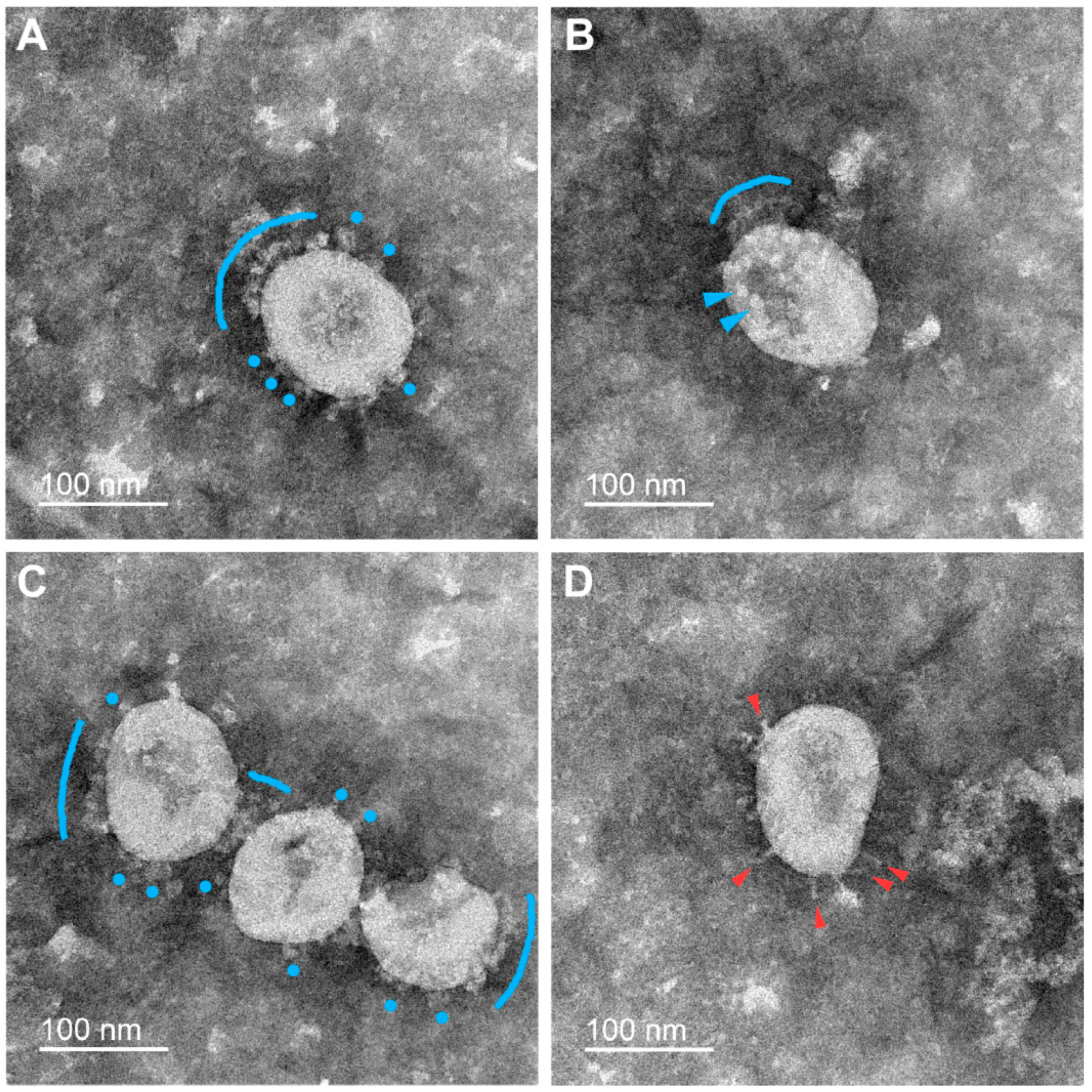
3.2. Various Inactivating Agents Affect the Virus Spikes’ Morphology Differently
3.2.1. Formaldehyde
3.2.2. β-Propiolactone (BPL)
3.2.3. UV Irradiation
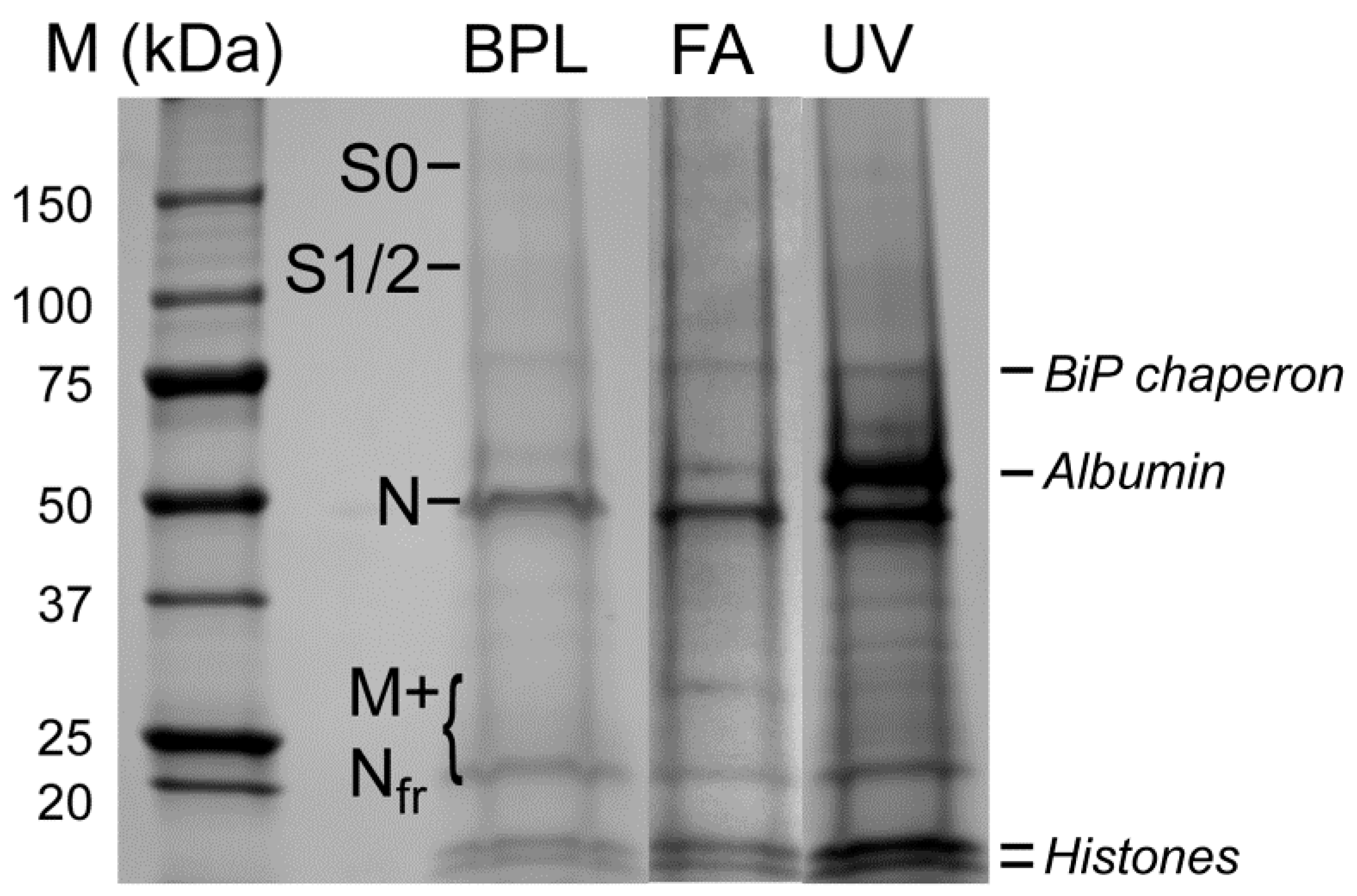
3.3. Electrophoretic and Mass Spectrometric Analysis of Inactivated Virus Preparations
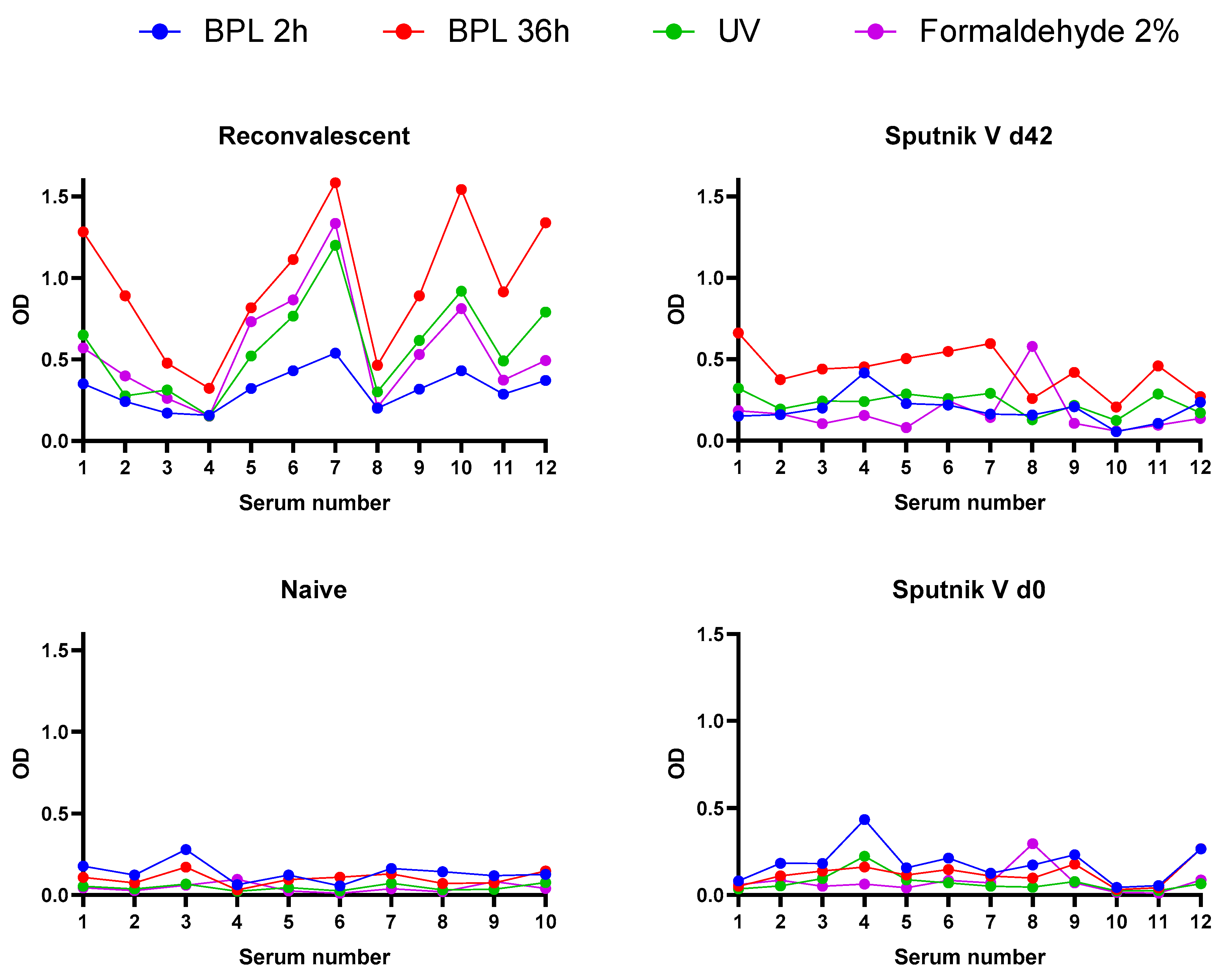
3.4. Immunoreactivity of Inactivated Virus Preparations
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Murdin, A.D.; Barreto, L.; Plotkin, S. Inactivated poliovirus vaccine: Past and present experience. Vaccine 1996, 14, 735–746. [Google Scholar] [CrossRef] [PubMed]
- Iversen, P.L.; Bavari, S. Inactivated COVID-19 vaccines to make a global impact. Lancet Infect. Dis. 2021, 21, 746–748. [Google Scholar] [CrossRef] [PubMed]
- Bridge Beijing. Available online: https://bridgebeijing.com (accessed on 1 October 2022).
- Our World in Data. Available online: https://ourworldindata.org (accessed on 1 October 2022).
- Mallapaty, S. China’s COVID vaccines have been crucial—Now immunity is waning. Nature 2021, 598, 398–399. [Google Scholar] [CrossRef] [PubMed]
- Jantarabenjakul, W.; Chantasrisawad, N.; Puthanakit, T.; Wacharapluesadee, S.; Hirankarn, N.; Ruenjaiman, V.; Paitoonpong, L.; Suwanpimolkul, G.; Torvorapanit, P.; Pradit, R.; et al. Short-Term Immune Response after Inactivated SARS-CoV-2 (CoronaVac®, Sinovac) And ChAdOx1 nCoV-19 (Vaxzevria®, Oxford-AstraZeneca) Vaccinations in Thai Health Care Workers. Asian Pac. J. Allergy Immunol. 2022, 40, 269–277. [Google Scholar] [CrossRef] [PubMed]
- Mok, C.K.P.; Cohen, C.A.; Cheng, S.M.S.; Chen, C.; Kwok, K.-O.; Yiu, K.; Chan, T.-O.; Bull, M.; Ling, K.C.; Dai, Z.; et al. Comparison of the Immunogenicity of BNT162b2 and CoronaVac COVID-19 Vaccines in Hong Kong: An Observational Cohort Study. Respirology 2022, 27, 301–310. [Google Scholar] [CrossRef]
- Liu, Y.; Zeng, Q.; Deng, C.; Li, M.; Li, L.; Liu, D.; Liu, M.; Ruan, X.; Mei, J.; Mo, R.; et al. Robust induction of B cell and T cell responses by a third dose of inactivated SARS-CoV-2 vaccine. Cell Discov. 2022, 8, 10. [Google Scholar] [CrossRef]
- Villela, D.A.M.; de Noronha, T.G.; Bastos, L.S.; Pacheco, A.G.; Cruz, O.G.; Carvalho, L.M.; Codeço, C.T.; da Costa Gomes, M.F.; Coelho, F.C.; Freitas, L.P.; et al. Effectiveness of Mass Vaccination in Brazil against Severe COVID-19 Cases. medRxiv 2021. [Google Scholar] [CrossRef]
- Ke, Z.; Oton, J.; Qu, K.; Cortese, M.; Zila, V.; McKeane, L.; Nakane, T.; Zivanov, J.; Neufeldt, C.J.; Cerikan, B.; et al. Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature 2020, 588, 498–502. [Google Scholar] [CrossRef]
- Liu, C.; Mendonça, L.; Yang, Y.; Gao, Y.; Shen, C.; Liu, J.; Ni, T.; Ju, B.; Liu, C.; Tang, X.; et al. The Architecture of Inactivated SARS-CoV-2 with Postfusion Spikes Revealed by Cryo-EM and Cryo-ET. Structure 2020, 28, 1218–1224.e4. [Google Scholar] [CrossRef]
- Yao, H.; Song, Y.; Chen, Y.; Wu, N.; Xu, J.; Sun, C.; Zhang, J.; Weng, T.; Zhang, Z.; Wu, Z.; et al. Molecular Architecture of the SARS-CoV-2 Virus. Cell 2020, 183, 730–738.e13. [Google Scholar] [CrossRef]
- Turoňová, B.; Sikora, M.; Schürmann, C.; Hagen, W.J.H.; Welsch, S.; Blanc, F.E.C.; von Bülow, S.; Gecht, M.; Bagola, K.; Hörner, C.; et al. In situ structural analysis of SARS-CoV-2 spike reveals flexibility mediated by three hinges. Science 2020, 370, 203–208. [Google Scholar] [CrossRef]
- Cai, Y.; Zhang, J.; Xiao, T.; Peng, H.; Sterling, S.M.; Walsh, R.M.; Rawson, S.; Rits-Volloch, S.; Chen, B. Distinct conformational states of SARS-CoV-2 spike protein. Science 2020, 369, 1586–1592. [Google Scholar] [CrossRef]
- Kordyukova, L.V.; Shanko, A.V. COVID-19: Myths and Reality. Biochemistry 2021, 86, 800–817. [Google Scholar] [CrossRef]
- Plavec, Z.; Domanska, A.; Liu, X.; Laine, P.; Paulin, L.; Varjosalo, M.; Auvinen, P.; Wolf, S.G.; Anastasina, M.; Butcher, S.J. SARS-CoV-2 Production, Purification Methods and UV Inactivation for Proteomics and Structural Studies. Viruses 2022, 14, 1989. [Google Scholar] [CrossRef]
- Darnell, M.E.R.; Subbarao, K.; Feinstone, S.M.; Taylor, D.R. Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV. J. Virol. Methods 2004, 121, 85–91. [Google Scholar] [CrossRef]
- Herrera-Rodriguez, J.; Signorazzi, A.; Holtrop, M.; de Vries-Idema, J.; Huckriede, A. Inactivated or damaged? Comparing the effect of inactivation methods on influenza virions to optimize vaccine production. Vaccine 2019, 37, 1630–1637. [Google Scholar] [CrossRef]
- Sabbaghi, A.; Miri, S.M.; Keshavarz, M.; Zargar, M.; Ghaemi, A. Inactivation methods for whole influenza vaccine production. Rev. Med. Virol. 2019, 29, e2074. [Google Scholar] [CrossRef]
- Elveborg, S.; Monteil, V.; Mirazimi, A. Methods of Inactivation of Highly Pathogenic Viruses for Molecular, Serology or Vaccine Development Purposes. Pathogens 2022, 11, 271. [Google Scholar] [CrossRef]
- Xia, S.; Duan, K.; Zhang, Y.; Zhao, D.; Zhang, H.; Xie, Z.; Li, X.; Peng, C.; Zhang, Y.; Zhang, W.; et al. Effect of an Inactivated Vaccine against SARS-CoV-2 on Safety and Immunogenicity Outcomes: Interim Analysis of 2 Randomized Clinical Trials. JAMA 2020, 324, 951–960. [Google Scholar] [CrossRef]
- Kandeil, A.; Mostafa, A.; Hegazy, R.R.; El-Shesheny, R.; El Taweel, A.; Gomaa, M.R.; Shehata, M.; Elbaset, M.A.; Kayed, A.E.; Mahmoud, S.H.; et al. Immunogenicity and Safety of an Inactivated SARS-CoV-2 Vaccine: Preclinical Studies. Vaccines 2021, 9, 214. [Google Scholar] [CrossRef]
- Gao, Q.; Bao, L.; Mao, H.; Wang, L.; Xu, K.; Yang, M.; Li, Y.; Zhu, L.; Wang, N.; Lv, Z.; et al. Development of an inactivated vaccine candidate for SARS-CoV-2. Science 2020, 369, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Wei, Y.; Liang, H.; Ji, W.; Chang, Z.; Xie, S.; Wang, Y.; Li, W.; Liu, Y.; Wu, H.; et al. Comparison of Physical and Biochemical Characterizations of SARS-CoV-2 Inactivated by Different Treatments. Viruses 2022, 14, 1938. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Xie, Z.; Long, R.; Fan, S.; Li, H.; He, Z.; Xu, K.; Liao, Y.; Wang, L.; Zhang, Y.; et al. Immunological evaluation of an inactivated SARS-CoV-2 vaccine in rhesus macaques. Mol. Ther. Methods Clin. Dev. 2021, 23, 108–118. [Google Scholar] [CrossRef] [PubMed]
- Jeyanathan, M.; Afkhami, S.; Smaill, F.; Miller, M.S.; Lichty, B.D.; Xing, Z. Immunological considerations for COVID-19 vaccine strategies. Nat. Rev. Immunol. 2020, 20, 615–632. [Google Scholar] [CrossRef]
- Wan, Y.; Shang, J.; Sun, S.; Tai, W.; Chen, J.; Geng, Q.; He, L.; Chen, Y.; Wu, J.; Shi, Z.; et al. Molecular Mechanism for Antibody-Dependent Enhancement of Coronavirus Entry. J. Virol. 2020, 94, e02015–e02019. [Google Scholar] [CrossRef]
- Reed, L.J.; Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of Structural Proteins during the Assembly of the Head of Bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Kordyukova, L.V.; Serebryakova, M.V.; Polyakov, V.Y.; Ovchinnikova, T.V.; Smirnova, Y.A.; Fedorova, N.V.; Baratova, L.A. Influenza A virus M1 protein structure probed by in situ limited proteolysis with bromelain. Protein Pept. Lett. 2008, 15, 922–930. [Google Scholar] [CrossRef]
- Barchuk, A.; Shirokov, D.; Sergeeva, M.; Tursun zade, R.; Dudkina, O.; Tychkova, V.; Barabanova, L.; Skougarevskiy, D.; Danilenko, D. Evaluation of the performance of SARS-CoV-2 antibody assays for a longitudinal population based study of COVID-19 spread in St. Petersburg, Russia. J. Med. Virol. 2021, 93, 5846–5852. [Google Scholar] [CrossRef]
- Neuman, B.W.; Kiss, G.; Kunding, A.H.; Bhella, D.; Baksh, M.F.; Connelly, S.; Droese, B.; Klaus, J.P.; Makino, S.; Sawicki, S.G.; et al. A structural analysis of M protein in coronavirus assembly and morphology. J. Struct. Biol. 2011, 174, 11–22. [Google Scholar] [CrossRef]
- Thura, M.; Sng, J.X.E.; Ang, K.H.; Li, J.; Gupta, A.; Hong, J.M.; Hong, C.W.; Zeng, Q. Targeting intra-viral conserved nucleocapsid (N) proteins as novel vaccines against SARS-CoVs. Biosci. Rep. 2021, 41, BSR20211491. [Google Scholar] [CrossRef]
- Bar-On, Y.M.; Flamholz, A.; Phillips, R.; Milo, R. SARS-CoV-2 (COVID-19) by the numbers. eLife 2020, 9, e57309. [Google Scholar] [CrossRef]
- Watanabe, Y.; Allen, J.D.; Wrapp, D.; McLellan, J.S.; Crispin, M. Site-specific glycan analysis of the SARS-CoV-2 spike. Science 2020, 369, 330–333. [Google Scholar] [CrossRef]
- Voß, D.; Pfefferle, S.; Drosten, C.; Stevermann, L.; Traggiai, E.; Lanzavecchia, A.; Becker, S. Studies on membrane topology, N-glycosylation and functionality of SARS-CoV membrane protein. Virol. J. 2009, 6, 79. [Google Scholar] [CrossRef]
- Liu, D.; Si, B.; Li, C.; Mi, Z.; An, X.; Qin, C.; Liu, W.; Tong, Y. Prokaryotic expression and purification of HA1 and HA2 polypeptides for serological analysis of the 2009 pandemic H1N1 influenza virus. J. Virol. Methods 2011, 172, 16–21. [Google Scholar] [CrossRef]
- Ragan, I.K.; Hartson, L.M.; Dutt, T.S.; Obregon-Henao, A.; Maison, R.M.; Gordy, P.; Fox, A.; Karger, B.R.; Cross, S.T.; Kapuscinski, M.L.; et al. A Whole Virion Vaccine for COVID-19 Produced via a Novel Inactivation Method and Preliminary Demonstration of Efficacy in an Animal Challenge Model. Vaccines 2021, 9, 340. [Google Scholar] [CrossRef]
- Goodrich, R.P.; Edrich, R.A.; Li, J.; Seghatchian, J. The Mirasol PRT system for pathogen reduction of platelets and plasma: An overview of current status and future trends. Transfus. Apher. Sci. 2006, 35, 5–17. [Google Scholar] [CrossRef]
- Patterson, E.I.; Prince, T.; Anderson, E.R.; Casas-Sanchez, A.; Smith, S.L.; Cansado-Utrilla, C.; Solomon, T.; Griffiths, M.J.; Acosta-Serrano, Á.; Turtle, L.; et al. Methods of Inactivation of SARS-CoV-2 for Downstream Biological Assays. J. Infect. Dis. 2020, 222, 1462–1467. [Google Scholar] [CrossRef]
- Uittenbogaard, J.P.; Zomer, B.; Hoogerhout, P.; Metz, B. Reactions of β-Propiolactone with Nucleobase Analogues, Nucleosides, and Peptides. J. Biol. Chem. 2011, 286, 36198–36214. [Google Scholar] [CrossRef]
- She, Y.; Cheng, K.; Farnsworth, A.; Li, X.; Cyr, T.D. Surface modifications of influenza proteins upon virus inactivation by β-propiolactone. Proteomics 2013, 13, 3537–3547. [Google Scholar] [CrossRef]
- Scheres, S.H.W. RELION: Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 2012, 180, 519–530. [Google Scholar] [CrossRef] [PubMed]

| № | Inactivation | Inactivating Agent | Inactivation Conditions | Temperature | |
|---|---|---|---|---|---|
| Chemical | Physical | ||||
| 1 |  | BPL, 1:1000 | 2 h | +4–8 °C | |
| 2 |  | BPL, 1:2000 | 36 h | +4–8 °C | |
| 3 |  | formaldehyde, 2% | 1 h | +37 °C | |
| 4 |  | formaldehyde, 4% | 1 h | +37 °C | |
| 5 |  | UV + Riboflavin | 0.027–0.474 J/cm2 | RT | |
| 6 |  | UV | 0.027–0.474 J/cm2 | RT | |
| Spike Morphology/Inactivated Sample | Number of Virions | Number of Spikes * | Shape of the Spike, Number (%) | ||
|---|---|---|---|---|---|
| Flail-Like | Needle-Like | Unidentified | |||
| Formaldehyde (2%) | 44 | 405 | 381 (94) | 15 (4) | 9 (2) |
| UV (1 min) | 52 | 595 | 541 (91) | 31 (5) | 23 (4) |
| BPL (1:2000; 36 h) | 47 | 346 | 246 (71) | 78 (23) | 22 (6) |
| BPL (1:1000; 2 h) | 51 | 229 | 135 (59) | 75 (33) | 19 (8) |
| Inactivated Sample/Sera * | Area under Curve (AUC) ** | |||
|---|---|---|---|---|
| Formaldehyde (2%) | BPL (1:2000, 36 h) | BPL (1:1000, 2 h) | UV (3 min) | |
| R | 6.204 | 10.33 | 3.464 | 6.282 |
| Sputnik V d42 | 1.897 | 4.735 | 2.117 | 2.523 |
| Naïve | 0.41 | 0.889 | 1.225 | 0.4055 |
| Sputnik V d0 | 0.863 | 1.292 | 1.972 | 0.803 |
| R/Naïve | 15.13 | 11.62 | 2.83 | 15.49 |
| d42/d0 | 2.20 | 3.66 | 1.07 | 3.14 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kordyukova, L.V.; Moiseenko, A.V.; Serebryakova, M.V.; Shuklina, M.A.; Sergeeva, M.V.; Lioznov, D.A.; Shanko, A.V. Structural and Immunoreactivity Properties of the SARS-CoV-2 Spike Protein upon the Development of an Inactivated Vaccine. Viruses 2023, 15, 480. https://doi.org/10.3390/v15020480
Kordyukova LV, Moiseenko AV, Serebryakova MV, Shuklina MA, Sergeeva MV, Lioznov DA, Shanko AV. Structural and Immunoreactivity Properties of the SARS-CoV-2 Spike Protein upon the Development of an Inactivated Vaccine. Viruses. 2023; 15(2):480. https://doi.org/10.3390/v15020480
Chicago/Turabian StyleKordyukova, Larisa V., Andrey V. Moiseenko, Marina V. Serebryakova, Marina A. Shuklina, Maria V. Sergeeva, Dmitry A. Lioznov, and Andrei V. Shanko. 2023. "Structural and Immunoreactivity Properties of the SARS-CoV-2 Spike Protein upon the Development of an Inactivated Vaccine" Viruses 15, no. 2: 480. https://doi.org/10.3390/v15020480
APA StyleKordyukova, L. V., Moiseenko, A. V., Serebryakova, M. V., Shuklina, M. A., Sergeeva, M. V., Lioznov, D. A., & Shanko, A. V. (2023). Structural and Immunoreactivity Properties of the SARS-CoV-2 Spike Protein upon the Development of an Inactivated Vaccine. Viruses, 15(2), 480. https://doi.org/10.3390/v15020480






