Retrospective Analysis of the SARS-CoV-2 Infection Profile in COVID-19 Positive Patients in Vitoria da Conquista, Northeast Brazil
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics and Patients
2.2. Clinical-Epidemiological Data Analysis
2.3. Gene Expression Analysis
2.4. Viral Sequencing
2.4.1. cDNA Synthesis and Whole Genome Sequencing
2.4.2. Generation of Consensus Sequences
2.4.3. Data Quality Control and Global Data Set Collection
2.4.4. Phylogenetic Analysis
2.4.5. Lineage Classification
3. Results
3.1. Clinical-Epidemiological Data Analysis
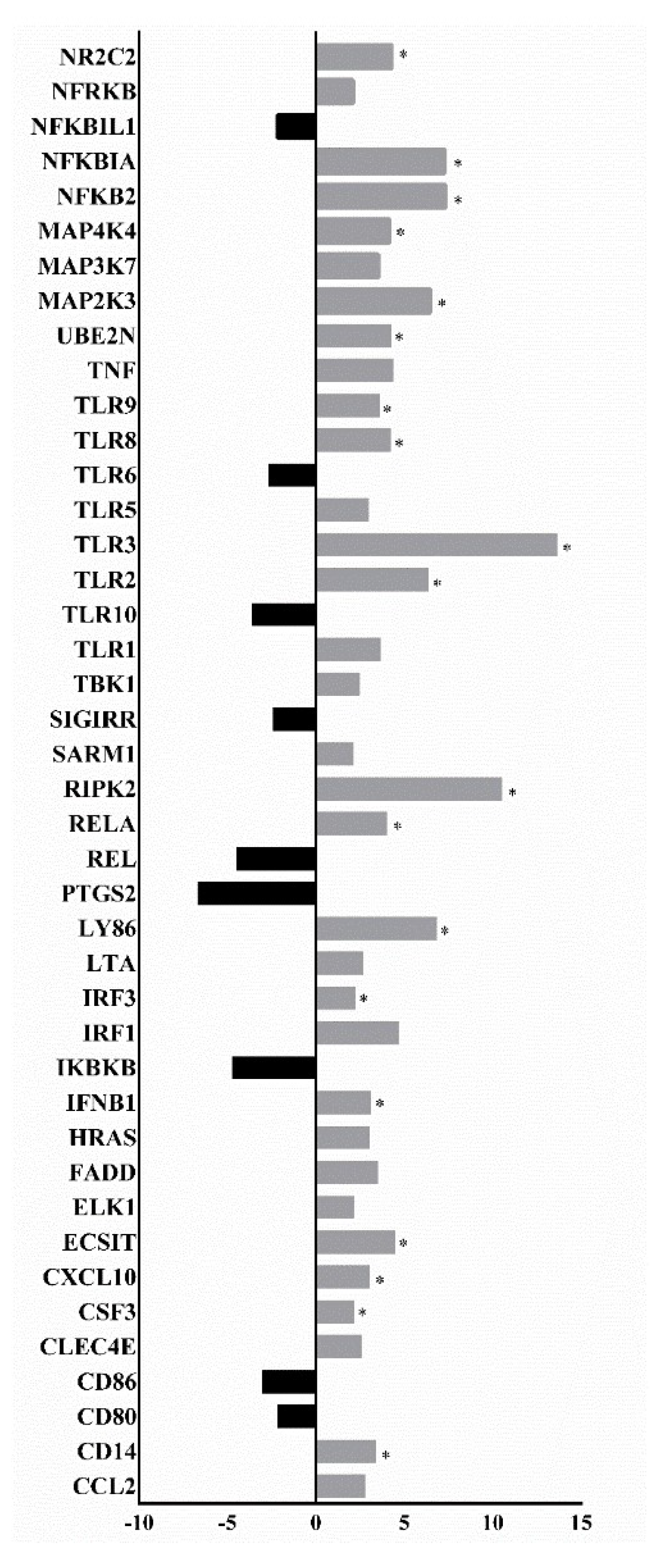
3.2. Analysis of the Expression Profile of Inflammation Pathway Activation Markers
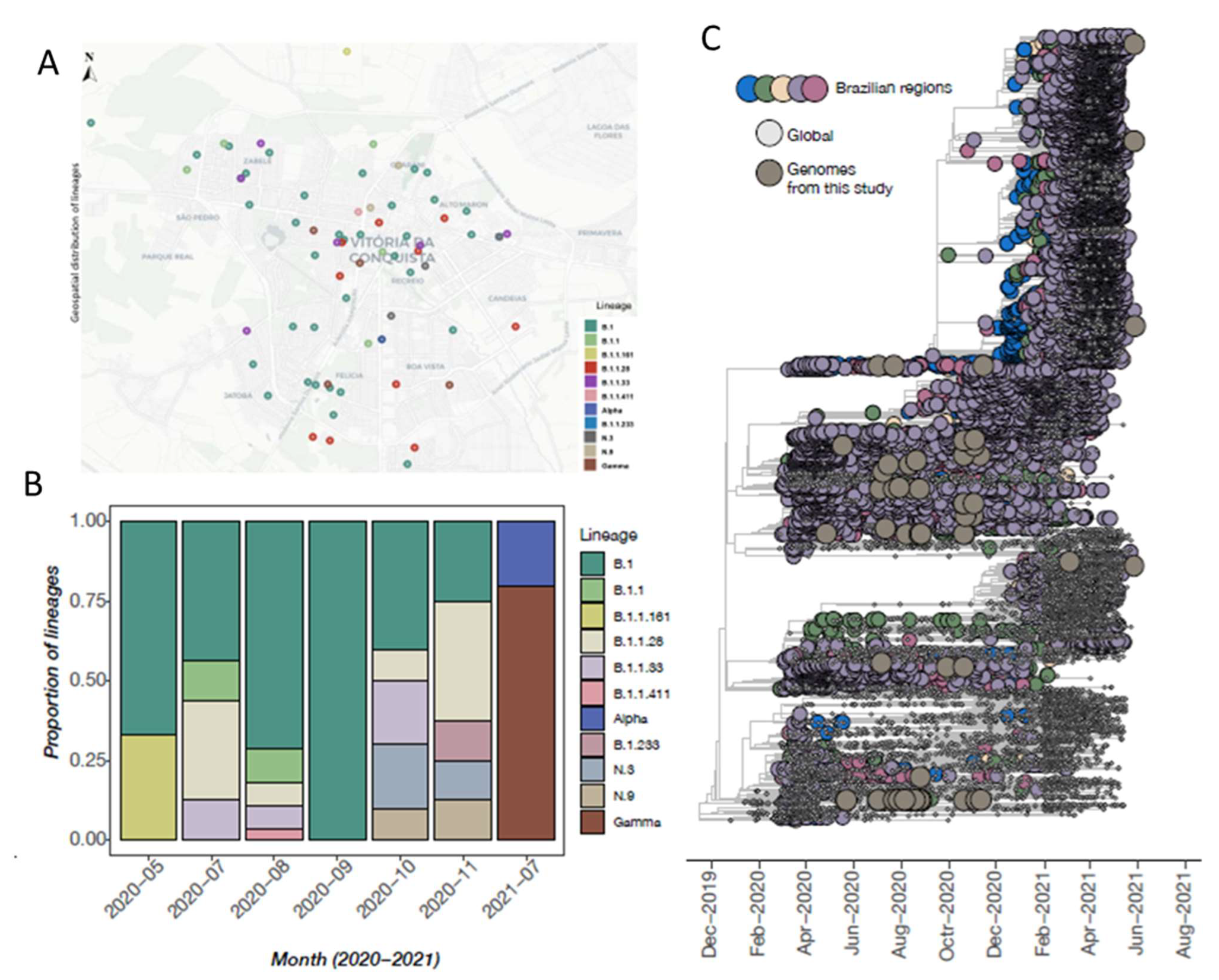
3.3. SARS-CoV-2 Genomic Data
3.4. Phylogenetic Inference and Lineage Diversity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Organização Pan-Americana de Saúde. Histórico da Pandemia de COVID-19. Available online: https://www.paho.org/pt/covid19/historico-da-pandemia-covid-19 (accessed on 27 November 2021).
- World Health Organization. WHO Coronavirus (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 2 August 2022).
- Worldometer. Total Coronavirus Cases in Brazil. Available online: https://www.worldometers.info/coronavirus/country/brazil/ (accessed on 2 August 2022).
- Magno, L. Desafios e propostas para ampliação da testagem e diagnóstico para COVID-19 no Brasil. Ciência Saúde Coletiva 2020, 25, 3355–3364. [Google Scholar] [CrossRef] [PubMed]
- Secretaria de Saúde do Estado da Bahia. Boletim Epidemiológico COVID-19 nº 861/2022. Available online: http://www.saude.ba.gov.br/wp-content/uploads/2022/08/BOLETIM_ELETRONICO_BAHIAN_861___08022022.pdf (accessed on 2 August 2022).
- Prefeitura Municipal de Vitória da Conquista. Boletim Epidemiológico. Available online: https://www.pmvc.ba.gov.br/coronavirus/ (accessed on 1 August 2022).
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Jafarzadeh, A.; Chauhan, P.; Saha, B.; Jafarzadeh, S.; Nemati, M. Contribution of monocytes and macrophages to the local tissue inflammation and cytokine storm in COVID-19: Lessons from SARS and MERS, and potential therapeutic interventions. Life Sci. 2020, 257, 118102. [Google Scholar] [CrossRef]
- Naqvi, A.A.T.; Fatima, K.; Mohammad, T.; Fatima, U.; Singh, I.K.; Singh, A.; Atif, S.M.; Hariprasad, G.; Hasan, G.M.; Hassan, I. Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: Structural genomics approach. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165878. [Google Scholar] [CrossRef] [PubMed]
- Hue, S.; Beldi-Ferchiou, A.; Bendib, I.; Surenaud, M.; Fourati, S.; Frapard, T.; Rivoal, S.; Razazi, K.; Carteaux, G.; Delfau-Larue, M.-H.; et al. Uncontrolled Innate and Impaired Adaptive Immune Responses in Patients with COVID-19 Acute Respiratory Distress Syndrome. Am J Respir Crit Care Med. 2020, 202, 1509–1519. [Google Scholar] [CrossRef] [PubMed]
- Gustine, J.N.; Jones, D. Immunopathology of Hyperinflammation in COVID-19. Am. J. Pathol. 2021, 191, 4–17. [Google Scholar] [CrossRef]
- Raskin, S. Genetics of COVID-19. J. Pediatr. 2021, 97, 378–386. [Google Scholar] [CrossRef]
- Candido, D.S.; Claro, I.M.; De Jesus, J.G.; Souza, W.M.; Moreira, F.R.R.; Dellicour, S.; Mellan, T.A.; Du Plessis, L.; Pereira, R.H.M.; Sales, F.C.S.; et al. Evolution and epidemic spread of SARS-CoV-2 in Brazil. Science 2020, 369, 1255–1260. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. CDC 2019-Novel Coronavirus (2019-nCoV) Real-Time RT-PCR Diagnostic Panel; Centers for Disease Control and Prevention: Atlanta, GA, USA, 2019. [Google Scholar]
- World Health Organization. Clinical Management of COVID-19: Interim Guidance; World Health Organization: Geneva, Switzerland, 2020; WHO reference number: WHO/2019-nCoV/clinical/2020.5. [Google Scholar]
- Quick, J. nCoV-2019 Sequencing Protocol; Protocol: Berkeley, CA, USA, 2020. [Google Scholar] [CrossRef]
- Vilsker, M.; Moosa, Y.; Nooij, S.; Fonseca, V.; Ghysens, Y.; Dumon, K.; Pauwels, R.; Alcantara, L.C.; Vanden Eynden, E.; Vandamme, A.-M.; et al. Genome Detective: An automated system for virus identification from high-throughput sequencing data. Bioinformatics 2019, 35, 871–873. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A.; Lam, T.T.; Max Carvalho, L.; Pybus, O.G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2016, 2, vew007. [Google Scholar] [CrossRef] [PubMed]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [PubMed]
- Sagulenko, P.; Puller, V.; Neher, R.A. TreeTime: Maximum-likelihood phylodynamic analysis. Virus Evol. 2018, 4, vex042. [Google Scholar] [CrossRef]
- Rambaut, A.; Holmes, E.C.; O’Toole, Á.; Hill, V.; McCrone, J.T.; Ruis, C.; du Plessis, L.; Pybus, O.G. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat Microbiol. 2020, 5, 1403–1407. [Google Scholar] [CrossRef]
- World Health Organization. A Year without Precedent: WHO’s COVID-19 Response. Available online: https://www.who.int/news-room/spotlight/a-year-without-precedent-who-s-covid-19-response (accessed on 27 November 2021).
- Chan-Yeung, M.; Xu, R.H. SARS: Epidemiology. Respirology 2003, 8 (Suppl. S1), S9–S14. [Google Scholar] [CrossRef]
- Chams, N.; Chams, S.; Badran, R.; Shams, A.; Araji, A.; Raad, M.; Mukhopadhyay, S.; Stroberg, E.; Duval, E.J.; Barton, L.M.; et al. COVID-19: A Multidisciplinary Review. Front Public Health 2020, 8, 383. [Google Scholar] [CrossRef]
- Hu, B.; Huang, S.; Yin, L. The cytokine storm and COVID-19. J. Med. Virol. 2021, 93, 250–256. [Google Scholar] [CrossRef]
- Fang, X.; Li, S.; Yu, H.; Wang, P.; Zhang, Y.; Chen, Z.; Li, Y.; Cheng, L.; Li, W.; Jia, H.; et al. Epidemiological, comorbidity factors with severity and prognosis of COVID-19: A systematic review and meta-analysis. Aging 2020, 12, 12493–12503. [Google Scholar] [CrossRef]
- Kragholm, K.; Andersen, M.P.; A Gerds, T.; Butt, J.H.; Østergaard, L.; Polcwiartek, C.; Phelps, M.; Andersson, C.; Gislason, G.H.; Torp-Pedersen, C.; et al. Association Between Male Sex and Outcomes of Coronavirus Disease 2019 (COVID-19)-A Danish Nationwide, Register-based Study. Clin. Infect. Dis. 2021, 73, e4025–e4030. [Google Scholar] [CrossRef]
- Petrilli, C.M.; Jones, S.A.; Yang, J.; Rajagopalan, H.; O’Donnell, L.; Chernyak, Y.; Tobin, K.A.; Cerfolio, R.J.; Francois, F.; Horwitz, L.I. Factors associated with hospital admission and critical illness among 5279 people with coronavirus disease 2019 in New York City: Prospective cohort study. BMJ 2020, 369, m1966. [Google Scholar] [CrossRef] [PubMed]
- Global Health 5050. The Sex, Gender and COVID-19 Project. Available online: https://globalhealth5050.org/the-sex-gender-and-covid-19-project/the-data-tracker/?explore=country&country=Brazil#search (accessed on 11 February 2022).
- Bienvenu, L.A.; Noonan, J.; Wang, X.; Peter, K. Higher mortality of COVID-19 in males: Sex differences in immune response and cardiovascular comorbidities. Cardiovasc. Res. 2020, 116, 2197–2206. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Cai, Y.; Zhang, K.; Zhou, L.; Zhang, Y.; Zhang, X.; Li, Q.; Li, W.; Yang, S.; Zhao, X.; et al. Association of hypertension and antihypertensive treatment with COVID-19 mortality: A retrospective observational study. Eur. Heart J. 2020, 41, 2058–2066. [Google Scholar] [CrossRef]
- Bloomgarden, Z.T. Diabetes e COVID-19. J Diabetes 2020, 12, 347–348. [Google Scholar] [CrossRef] [PubMed]
- Ruan, Q.; Yang, K.; Wang, W.; Jiang, L.; Song, J. Clinical predictors of mortality due to COVID-19 based on an analysis of data of 150 patients from Wuhan, China. Intensive Care Med. 2020, 46, 846–848. [Google Scholar] [CrossRef]
- Ejaz, H.; Alsrhani, A.; Zafar, A.; Javed, H.; Junaid, K.; Abdalla, A.E.; Abosalif, K.O.; Ahmed, Z.; Younas, S. COVID-19 and comorbidities: Deleterious impact on infected patients. J Infect Public Health 2020, 13, 1833–1839. [Google Scholar] [CrossRef]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef]
- Salvatore, P.P.; Dawson, P.; Wadhwa, A.; Rabold, E.M.; Buono, S.; A Dietrich, E.; E Reses, H.; Vuong, J.; Pawloski, L.; Dasu, T.; et al. Epidemiological Correlates of Polymerase Chain Reaction Cycle Threshold Values in the Detection of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Clin. Infect Dis. 2021, 72, e761–e767. [Google Scholar] [CrossRef]
- Choudhuri, J.; Carter, J.; Nelson, R.; Skalina, K.; Osterbur-Badhey, M.; Johnston, A.; Goldstein, D.; Paroder, M.; Szymanski, J. SARS-CoV-2 PCR cycle threshold at hospital admission associated with patient mortality. PLoS ONE. 2020, 15, e0244777. [Google Scholar] [CrossRef]
- Ibrahim, F.; Natasha, A.; Saharman, Y.R.; Yasmon, A.; Fithriyah, F.; Karuniawati, A.; Ganiesa, S.; Sudarmono, P. Consideration of the Cycle Threshold Values from Real-Time RT-PCR SARS-CoV-2 Interpretation for the Clinicians: Analysis of 339 Positive Cases from a Referral Laboratory in Jakarta, Indonesia. Acta Med. Indones 2021, 53, 13–17. [Google Scholar]
- Waudby-West, R.; Parcell, B.J.; Palmer, I.; Bell, S.; Chalmers, J.D.; Siddiqui, M.K. The association between SARS-CoV-2 RT-PCR cycle threshold and mortality in a community cohort. Eur. Respir. J. 2021, 58, 2100360. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Fan, J.; Yu, F.; Feng, B.; Lou, B.; Zou, Q.; Xie, G.; Lin, S.; Wang, R.; Yang, X.; et al. Viral load dynamics and disease severity in patients infected with SARS-CoV-2 in Zhejiang province, China, January-March 2020: Retrospective cohort study. BMJ 2020, 369, m1443. [Google Scholar] [CrossRef] [PubMed]
- Jose, R.J.; Manuel, A. COVID-19 cytokine storm: The interplay between inflammation and coagulation. Lancet Respir. Med. 2020, 8, e46–e47. [Google Scholar] [CrossRef]
- Bastard, P.; Rosen, L.B.; Zhang, Q.; Michailidis, E.; Hoffmann, H.-H.; Zhang, Y.; Dorgham, K.; Philippot, Q.; Rosain, J.; Béziat, V.; et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science 2020, 370, eabd4585. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Bastard, P.; Liu, Z.; Le Pen, J.; Moncada-Velez, M.; Chen, J.; Ogishi, M.; Sabli, I.K.D.; Hodeib, S.; Korol, C.; et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 2020, 370, eabd4570. [Google Scholar] [CrossRef] [PubMed]
- Karki, R.; Sharma, B.R.; Tuladhar, S.; Williams, E.P.; Zalduondo, L.; Samir, P.; Zheng, M.; Sundaram, B.; Banoth, B.; Malireddi, R.K.S.; et al. Synergism of TNF-a and IFN-g Triggers Inflammatory Cell Death, Tissue Damage, and Mortality in SARS-CoV-2 Infection and Cytokine Shock Syndromes. Cell 2021, 184, 149–168. [Google Scholar] [CrossRef]
- Catanzaro, M.; Fagiani, F.; Racchi, M.; Corsini, E.; Govoni, S.; Lanni, C. Immune response in COVID-19: Addressing a pharmacological challenge by targeting pathways triggered by SARS-CoV-2. Signal Transduct Target Ther. 2020, 5, 84. [Google Scholar] [CrossRef]
- Chen, D.; Khan, N.; Close, B.J.; Goel, R.K.; Blum, B.; Tavares, A.H.; Kenney, D.; Conway, H.L.; Ewoldt, J.K.; Chitalia, V.C.; et al. SARS-CoV-2 Disrupts Proximal Elements in the JAK-STAT Pathway. J Virol. 2021, 95, e0086221. [Google Scholar] [CrossRef]
- Sun, X.; Wang, T.; Cai, D.; Hu, Z.; Chen, J.; Liao, H.; Zhi, L.; Wei, H.; Zhang, Z.; Qiu, Y.; et al. Cytokine storm intervention in the early stages of COVID-19 pneumonia. Cytokine Growth Factor Rev. 2020, 53, 38–42. [Google Scholar] [CrossRef]
- Nikolich-Zugich, J.; Knox, K.S.; Rios, C.T.; Natt, B.; Bhattacharya, D.; Fain, M.J. SARS-CoV-2 and COVID-19 in older adults: What we may expect regarding pathogenesis, immune responses, and outcomes. Geroscience 2020, 42, 505–514. [Google Scholar] [CrossRef]
- Zheng, M.; Karki, R.; Williams, E.P.; Yang, D.; Fitzpatrick, E.; Vogel, P.; Jonsson, C.B.; Kanneganti, T.-D. TLR2 senses the SARS-CoV-2 envelope protein to produce inflammatory cytokines. Nat. Immunol. 2021, 22, 829–838. [Google Scholar] [CrossRef] [PubMed]
- Brasil. Ministério da Saúde. Secretaria de Ciência, Tecnologia, Inovação e Insumos Estratégicos em Saúde. In Departamento de Ciência e Tecnologia. Informe Semanal n° 37 de Evidências sobre Variantes de Atenção de SARS-CoV-2 [Recurso eletrônico]/Ministério da Saúde. Secretaria de Ciência, Tecnologia, Inovação e Insumos Estratégicos em Saúde; Departamento de Ciência e Tecnolog–a-Brasília: Ministério da Saúde, Brasília, 2021; p. 112. Available online: https://www.gov.br/saude/pt-br/coronavirus/publicacoes-tecnicas/informes-de-variantes/informe_s37.pdf/view (accessed on 27 November 2021).
- Strowig, T.; Henao-Mejia, J.; Elinav, E.; Flavell, R. Inflammasomes in health and disease. Nature 2012, 481, 278–286. [Google Scholar] [CrossRef] [PubMed]
- Khandia, R.; Singhal, S.; Alqahtani, T.; Kamal, M.A.; El-Shall, N.A.; Nainu, F.; Desingu, P.A.; Dhama, K. Emergence of SARS-CoV-2 Omicron (B.1.1.529) variant, salient features, high global health concerns and strategies to counter it amid ongoing COVID-19 pandemic. Environ. Res. 2022, 209, 112816. [Google Scholar] [CrossRef] [PubMed]
- Resende, P.C.; Delatorre, E.; Gräf, T.; Mir, D.; Motta, F.C.; Appolinario, L.R.; da Paixão, A.C.D.; Mendonça, A.C.D.F.; Ogrzewalska, M.; Caetano, B.; et al. Evolutionary Dynamics and Dissemination Pattern of the SARS-CoV-2 Lineage B.1.1.33 During the Early Pandemic Phase in Brazil. Front. Microbiol. 2021, 11, 615280. [Google Scholar] [CrossRef]
- Naveca, F.; Nascimento, V.; Souza, V.; Corado, A.; Nascimento, F.; Silva, G.; Costa, Á.; Duarte, D.; Pessoa, K.; Gonçalves, L.; et al. Relação Filogenética de Sequências SARS-CoV-2 do Amazonas com Variantes Emergentes Brasileiras que Abrigam Mutações E484K e N501Y na Proteína Spike. In Nota Técnica 2021/01–Rede Genômica FIOCRUZ/Ministério da Saúde; Instituto Oswaldo Cruz (IOC): Manaus, Brazil, 2021. [Google Scholar]
- Nonaka, C.K.V.; Franco, M.M.; Gräf, T.; Barcia, C.A.d.L.; de Ávila, R.N.; de Sousa, K.A.F.; Neiva, L.M.C.; Fosenca, V.; Mendes, A.V.A.; de Aguiar, R.S.; et al. Genomic Evidence of SARS-CoV-2 Reinfection Involving E484K Spike Mutation, Brazil. Emerg. Infect. Dis. 2021, 27, 1522–1524. [Google Scholar] [CrossRef]
- Wolf, J.M.; Kipper, D.; Borges, G.R.; Streck, A.F.; Lunge, V.R. Temporal spread and evolution of SARS-CoV-2 in the second pandemic wave in Brazil. J. Med. Virol. 2021, 94, 926–936. [Google Scholar] [CrossRef] [PubMed]
- Dejnirattisai, W.; Zhou, D.; Supasa, P.; Liu, C.; Mentzer, A.J.; Ginn, H.M.; Zhao, Y.; Duyvesteyn, H.M.; Tuekprakhon, A.; Nutalai, R.; et al. Antibody evasion by the P.1 strain of SARS-CoV-2. Cell 2021, 184, 2939–2954.e9. [Google Scholar] [CrossRef]
| Symptoms | Comorbidities |
|---|---|
| Fever Low fever or chills Cough Odynophagia | Cardiovascular disease Hypertension Diabetes Liver disease |
| Coryza Nasal congestion Mild breathing difficulty Progressive breathing difficulty Ageusia or anosmia Urinary deficit Syncope SpO2 saturation lower than 94% Mental confusion Signs of cyanosis Drowsiness Fast or slow breathing Myalgia/arthralgia Diarrhea Emesis Headache Irritability/confusion Adynamia Sputum production Conjunctival congestion Dysphagia Lymphadenopathy Nasal flaring Dehydration Inappetence | Chronic neurological or neuromuscular disease Immunosuppression Human Immunodeficiency Virus (HIV) Kidney disease Chronic lung disease Neoplasia (solid or hematological tumor) Obesity Asthma |
| Variables | n | % |
|---|---|---|
| Sex | ||
| Male | 292 | 37.3 |
| Female | 491 | 62.7 |
| Age (years) | ||
| ≤19 | 50 | 6.4 |
| 20–59 | 490 | 62.6 |
| ≥60 | 243 | 31.0 |
| Health professional | ||
| Yes | 194 | 24.8 |
| No | 589 | 75.2 |
| Disease severity | ||
| Asymptomatic | 36 | 4.6 |
| Mildly symptomatic | 131 | 16.7 |
| Moderately/severely symptomatic | 616 | 78.7 |
| Interval between appearance of symptoms and collection of sample | ||
| ≤8 days | 621 | 83.1 |
| >9 days | 126 | 16.9 |
| Number of comorbidities | ||
| 0 | 483 | 61.7 |
| 1 | 205 | 26.2 |
| ≥2 | 95 | 12.1 |
| Ct a | ||
| <22 | 212 | 27.1 |
| ≥23 | 571 | 72.9 |
| Variables | Disease Severity a n = 783 | Death n = 35 | OR (Crude) | OR (Adjusted) b | ||||
|---|---|---|---|---|---|---|---|---|
| OR | CI 95% | p-Value | OR | CI 95% | p-Value | |||
| n (%) | ||||||||
| Age (years) | ||||||||
| ≤19 | 50 (6.39) | 1 (2.85) | ||||||
| 20–59 | 490 (62.58) | 7 (20.0) | 0.74 | 0.09–5.89 | 0.751 | 0.47 | 0.06–4.03 | 0.493 |
| ≥60 | 243 (31.03) | 27 (77.15) | 6.12 | 0.81–46.17 | 0.079 | 3.18 | 0.40–25.18 | 0.274 |
| Sex | ||||||||
| Male | 292 (37.29) | 19 (54.3) | 0.48 | 0.24–0.95 | 0.037 * | 0.53 | 0.26–1.07 | 0.078 |
| Female | 491 (62.71) | 16 (45.7) | ||||||
| Number of comorbidities | ||||||||
| 0 | 483 (61.7) | 11 (31.4) | ||||||
| 1 | 205 (26.2) | 15 (42.9) | 3.39 | 1.53–7.51 | 0.003 * | 2.33 | 1.00–5.41 | 0.049 * |
| ≥2 | 95 (12.1) | 9 (25.7) | 4.49 | 1.81–11.16 | 0.001 * | 2.88 | 1.10–7.56 | 0.032 * |
| Cardiovascular disease | ||||||||
| Yes | 645 (82.38) | 16 (45.7) | 4.32 | 2.16–8.63 | <0.001 * | |||
| No | 138 (17.62) | 19 (54.3) | ||||||
| Diabetes | ||||||||
| Yes | 678 (86.59) | 11 (31.4) | 3.19 | 1.51–6.72 | 0.002 * | |||
| No | 105 (13.41) | 24 (68.6) | ||||||
| Obesity | ||||||||
| Yes | 713 (91.06) | 2 (5.7) | 0.61 | 0.14–2.58 | 0.498 | |||
| No | 70 (8.94) | 33 (94.3) | ||||||
| COPDc | ||||||||
| Yes | 740 (94.51) | 32 (91.4) | 11.59 | 2.77–48.46 | 0.001 * | |||
| No | 43 (5.49) | 3 (8.6) | ||||||
| Ctd | ||||||||
| <22 | 212 (27.1) | 21 (60.0) | 3.41 | 1.72–6.77 | <0.001 * | 3.36 | 1.64–6.90 | 0.001 * |
| ≥23 | 571 (72.9) | 14 (40.0) | ||||||
| Interval between symptoms and sample collection | ||||||||
| ≤8 days | 621 (83.1) | 30 (85.7) | ||||||
| >9 days | 126 (16.9) | 5 (14.3) | 0.81 | 0.31–2.14 | 0.677 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dantas, A.C.S.; Oliveira, H.B.M.; Gomes, C.P.; Alves, D.L.; Infante, P.D.B.; Caitité, R.d.J.A.; Fritsch, H.M.; Cucco, M.S.; Silva, L.S.C.; Oliveira, C.N.T.; et al. Retrospective Analysis of the SARS-CoV-2 Infection Profile in COVID-19 Positive Patients in Vitoria da Conquista, Northeast Brazil. Viruses 2022, 14, 2424. https://doi.org/10.3390/v14112424
Dantas ACS, Oliveira HBM, Gomes CP, Alves DL, Infante PDB, Caitité RdJA, Fritsch HM, Cucco MS, Silva LSC, Oliveira CNT, et al. Retrospective Analysis of the SARS-CoV-2 Infection Profile in COVID-19 Positive Patients in Vitoria da Conquista, Northeast Brazil. Viruses. 2022; 14(11):2424. https://doi.org/10.3390/v14112424
Chicago/Turabian StyleDantas, Anna Carolina S., Hellen B. M. Oliveira, Camila P. Gomes, Daniele L. Alves, Priscilla D. B. Infante, Rosimara de J. A. Caitité, Hegger M. Fritsch, Marina S. Cucco, Lucas S. C. Silva, Caline N. T. Oliveira, and et al. 2022. "Retrospective Analysis of the SARS-CoV-2 Infection Profile in COVID-19 Positive Patients in Vitoria da Conquista, Northeast Brazil" Viruses 14, no. 11: 2424. https://doi.org/10.3390/v14112424
APA StyleDantas, A. C. S., Oliveira, H. B. M., Gomes, C. P., Alves, D. L., Infante, P. D. B., Caitité, R. d. J. A., Fritsch, H. M., Cucco, M. S., Silva, L. S. C., Oliveira, C. N. T., Bittencourt, R. d. S., Amorim, A. T., Nascimento, A. L. P., Marinho, F. A. G. C., de Medeiros, D. S., de Oliveira, M. G. G., Mistro, S., de Melo, F. F., Pereira, T. T. S., ... Marques, L. M. (2022). Retrospective Analysis of the SARS-CoV-2 Infection Profile in COVID-19 Positive Patients in Vitoria da Conquista, Northeast Brazil. Viruses, 14(11), 2424. https://doi.org/10.3390/v14112424








