A Duplex Fluorescent Microsphere Immunoassay for Detection of Bluetongue and Epizootic Hemorrhagic Disease Virus Antibodies in Cattle Sera
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Viruses
2.2. Microsphere Conjugation
2.3. Cattle Sera
2.4. Fluorescent Microsphere Immunoassay (FMIA)
2.5. cELISA
2.6. Western Blots
3. Results
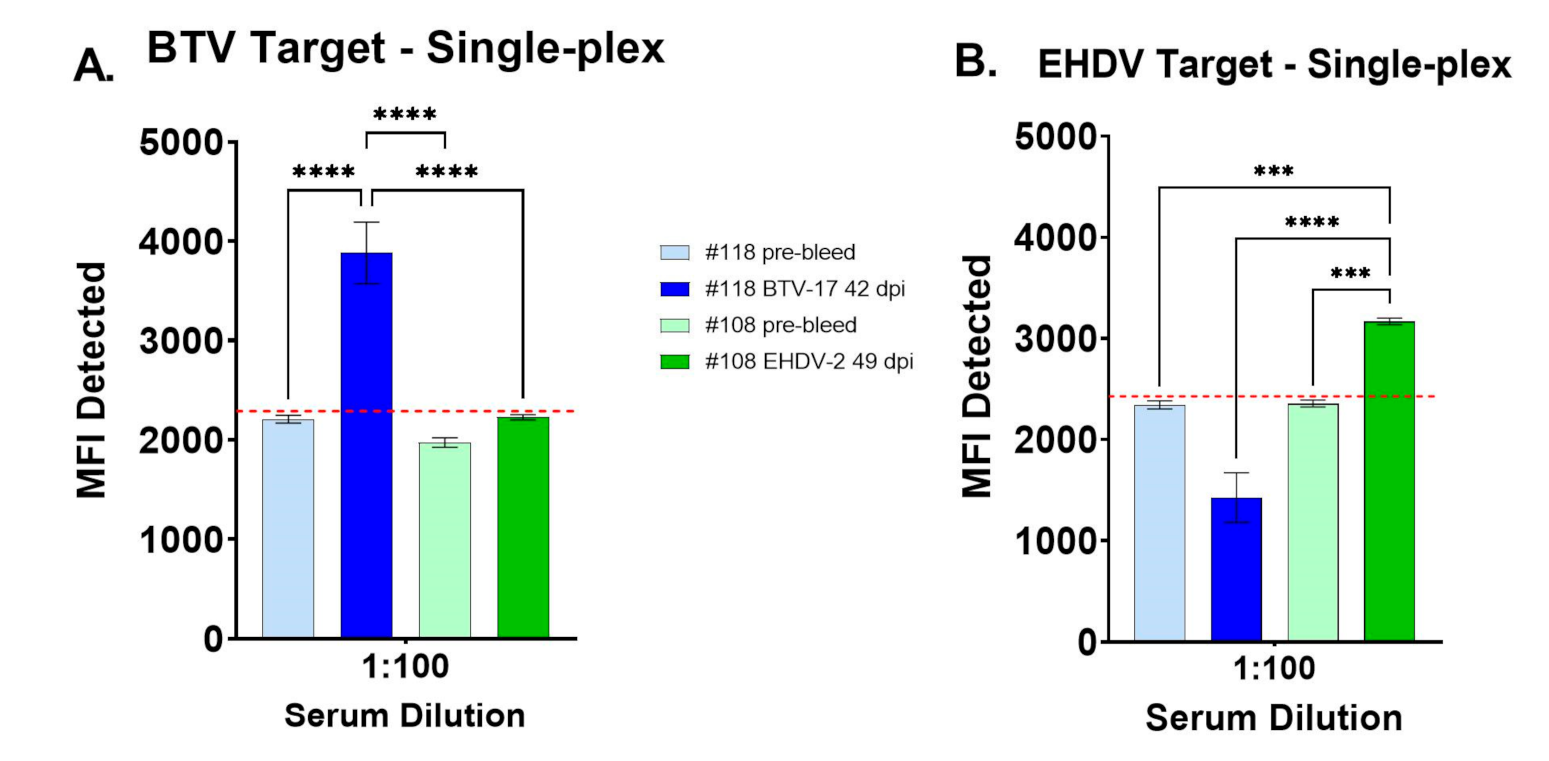
3.1. Evaluation of Single-Analyte FMIA with Control Cattle Sera
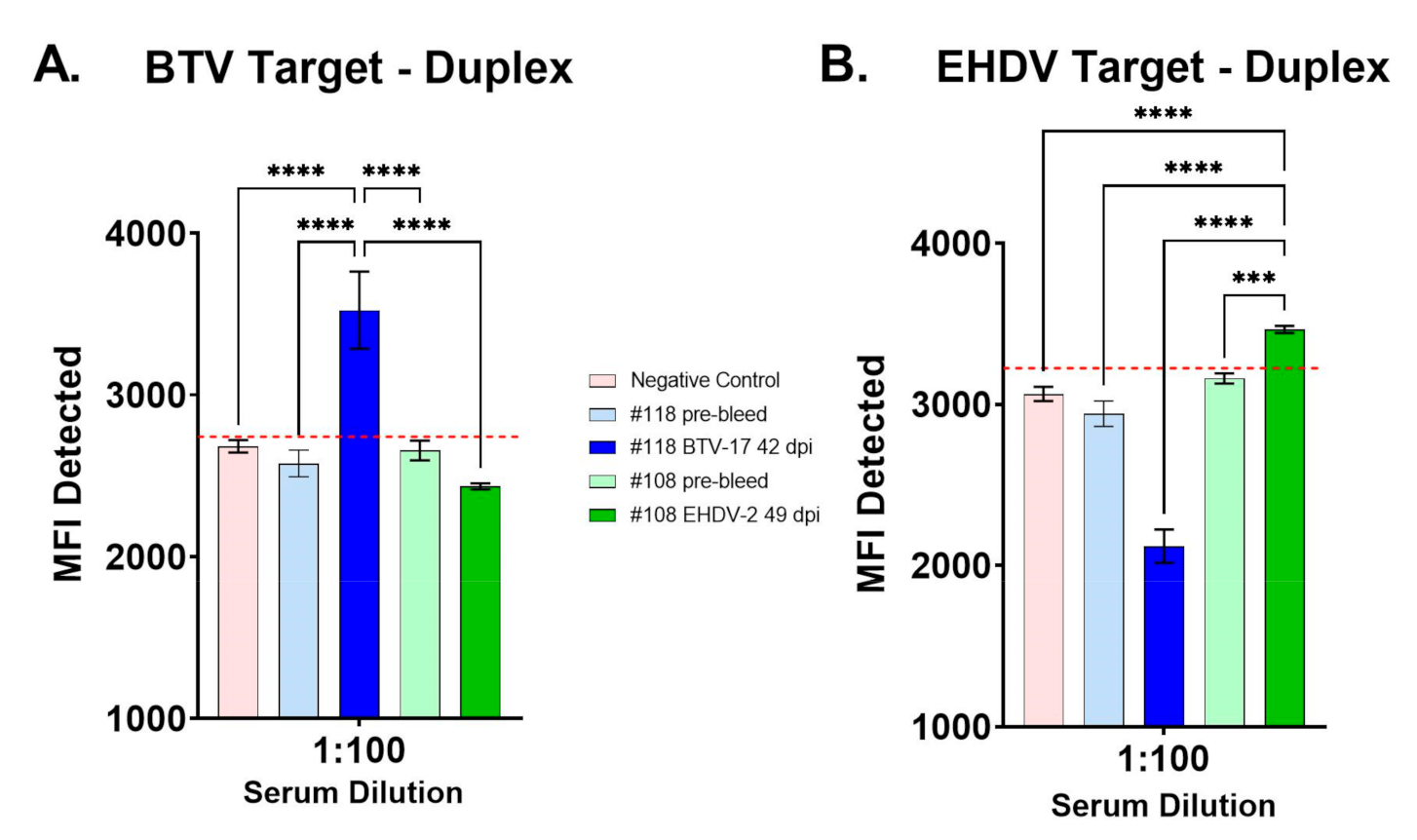
3.2. Evaluation of BTV/EHDV Duplex FMIA with Control Cattle Sera
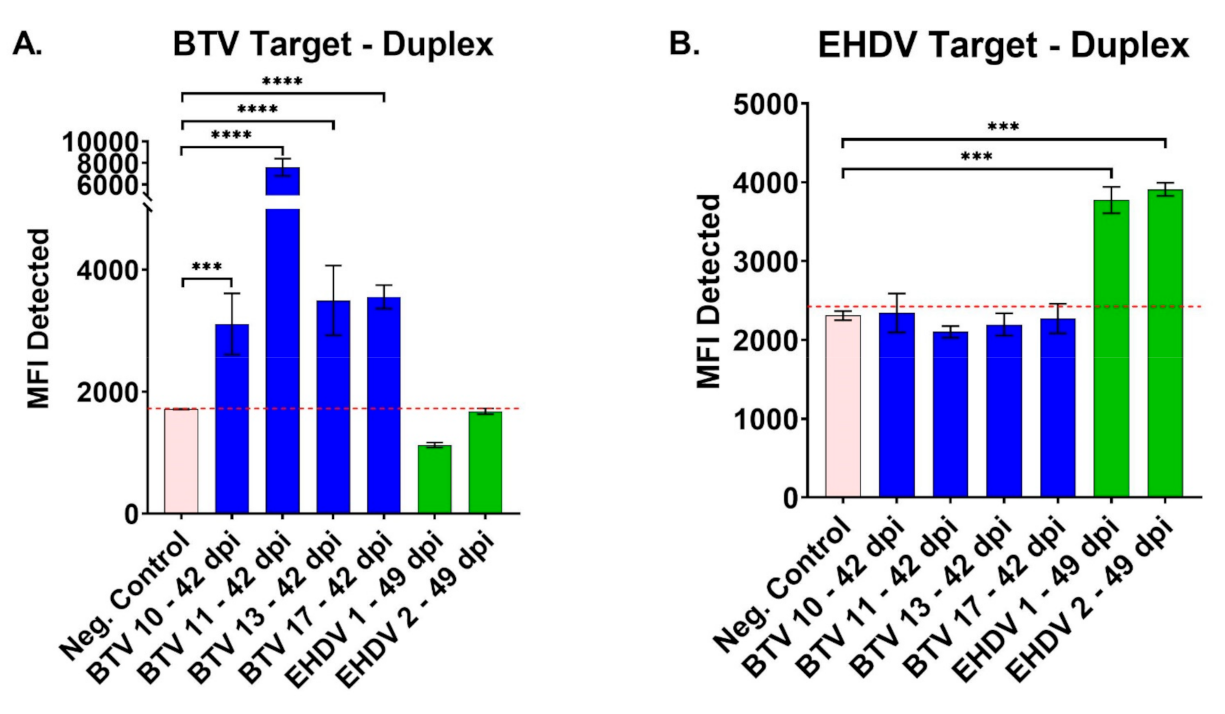
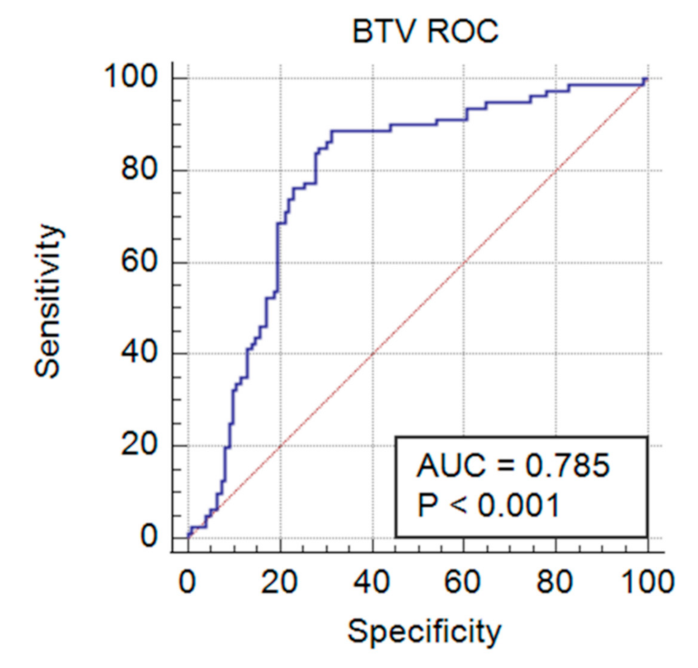
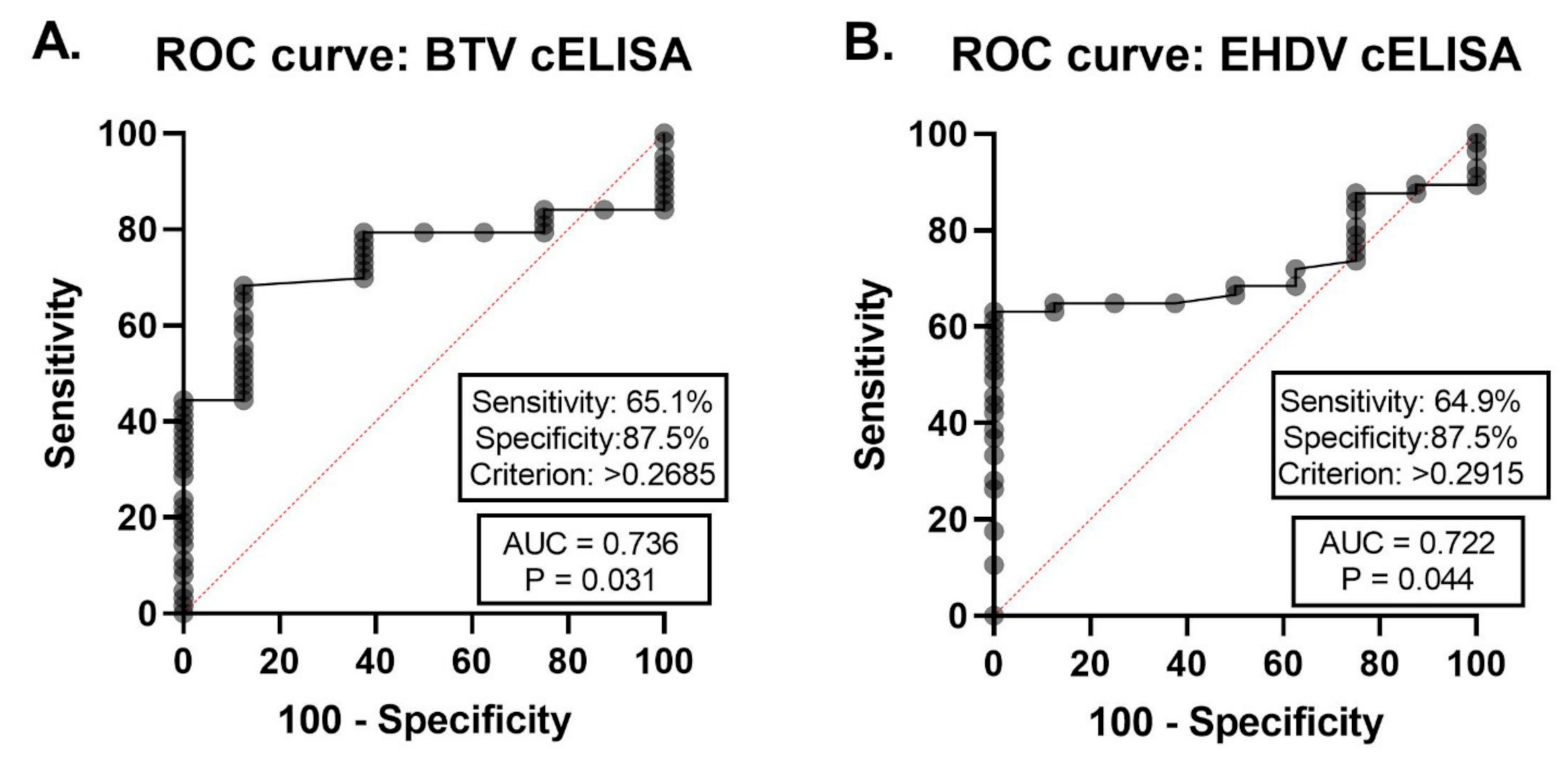
3.3. Validation of Duplex FMIA with Field-Collected Cattle Sera
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ruder, M.; Vigil, S.; Kienzle, C.; Stallknecht, D.E.; Corn, J.; Mertins, J. Update on SCWDS arthropod surveys, EHDV/BTV research and 2017 hemorrhagic disease activity. In Proceedings of the U.S. Animal Health Assoc., Kansas City, MO, USA, 18–24 October 2018; pp. 286–288. [Google Scholar]
- Johnson, D.J.; Ostlund, E.N.; Mertens, P.P.; Maan, S. Exotic bluetongue viruses identified from ruminants in the Southeastern U.S., from 1999–2006. In Proceedings of the United States Animal Health Association, Reno, NV, USA, 18–24 October 2007; pp. 209–210. [Google Scholar]
- MacLachlan, N.J.; Wilson, W.C.; Crossley, B.M.; Mayo, C.E.; Jasperson, D.C.; Breitmeyer, R.E.; Whiteford, A.M. Novel serotype of bluetongue virus, western North America. Emerg. Infect. Dis. 2013, 19, 665–666. [Google Scholar] [CrossRef] [PubMed]
- Allison, A.B.; Goekjian, V.H.; Potgieter, A.C.; Wilson, W.C.; Johnson, D.J.; Mertens, P.P.; Stallknecht, D.E. Detection of a novel reassortant epizootic hemorrhagic disease virus (EHDV) in the USA containing RNA segments derived from both exotic (EHDV-6) and endemic (EHDV-2) serotypes. J. Gen. Virol. 2010, 91, 430–439. [Google Scholar] [CrossRef] [PubMed]
- Elbers, A.R.; Backx, A.; Mintiens, K.; Gerbier, G.; Staubach, C.; Hendrickx, G.; van der Spek, A. Field observations during the bluetongue serotype 8 epidemic in 2006. II. Morbidity and mortality rate, case fatality and clinical recovery in sheep and cattle in the Netherlands. Prev. Vet. Med. 2008, 87, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Brenner, J.; Batten, C.; Yadin, H.; Bumbarov, V.; Friedgut, O.; Rotenberg, D.; Golender, N.; Oura, C.A. Clinical syndromes associated with the circulation of multiple serotypes of bluetongue virus in dairy cattle in Israel. Vet. Rec. 2011. [Google Scholar] [CrossRef]
- Dal Pozzo, F.; Saegerman, C.; Thiry, E. Bovine infection with bluetongue virus with special emphasis on European serotype 8. Vet. J. 2009, 182, 142–151. [Google Scholar] [CrossRef]
- Darpel, K.E.; Batten, C.A.; Veronesi, E.; Shaw, A.E.; Anthony, S.; Bachanek-Bankowska, K.; Kgosana, L.; bin-Tarif, A.; Carpenter, S.; Muller-Doblies, U.U.; et al. Clinical signs and pathology shown by British sheep and cattle infected with bluetongue virus serotype 8 derived from the 2006 outbreak in northern Europe. Vet. Rec. 2007, 161, 253–261. [Google Scholar] [CrossRef]
- Muller, U.; Kemmerling, K.; Straet, D.; Janowitz, U.; Sauerwein, H. Effects of bluetongue virus infection on sperm quality in bulls: A preliminary report. Vet. J. 2010, 186, 402–403. [Google Scholar] [CrossRef]
- De Clercq, K.; De Leeuw, I.; Verheyden, B.; Vandemeulebroucke, E.; Vanbinst, T.; Herr, C.; Meroc, E.; Bertels, G.; Steurbaut, N.; Miry, C.; et al. Transplacental infection and apparently immunotolerance induced by a wild-type bluetongue virus serotype 8 natural infection. Transbound. Emerg. Dis. 2008, 55, 352–359. [Google Scholar] [CrossRef]
- Zanella, G.; Durand, B.; Sellal, E.; Breard, E.; Sailleau, C.; Zientara, S.; Batten, C.A.; Mathevet, P.; Audeval, C. Bluetongue virus serotype 8: Abortion and transplacental transmission in cattle in the Burgundy region, France, 2008–2009. Theriogenology 2011. [Google Scholar] [CrossRef]
- Vercauteren, G.; Miry, C.; Vandenbussche, F.; Ducatelle, R.; Van der Heyden, S.; Vandemeulebroucke, E.; De Leeuw, I.; Deprez, P.; Chiers, K.; De Clercq, K. Bluetongue virus serotype 8-associated congenital hydranencephaly in calves. Transbound. Emerg. Dis. 2008, 55, 293–298. [Google Scholar] [CrossRef]
- Santman-Berends, I.M.; van Wuijckhuise, L.; Vellema, P.; van Rijn, P.A. Vertical transmission of bluetongue virus serotype 8 virus in Dutch dairy herds in 2007. Vet. Microbiol. 2010, 141, 31–35. [Google Scholar] [CrossRef]
- Santman-Berends, I.M.; Hage, J.J.; van Rijn, P.A.; Stegeman, J.A.; van Schaik, G. Bluetongue virus serotype 8 (BTV-8) infection reduces fertility of Dutch dairy cattle and is vertically transmitted to offspring. Theriogenology 2010, 74, 1377–1384. [Google Scholar] [CrossRef]
- Saegerman, C.; Bolkaerts, B.; Baricalla, C.; Raes, M.; Wiggers, L.; de Leeuw, I.; Vandenbussche, F.; Zimmer, J.Y.; Haubruge, E.; Cassart, D.; et al. The impact of naturally-occurring, trans-placental bluetongue virus serotype-8 infection on reproductive performance in sheep. Vet. J. 2011, 187, 72–80. [Google Scholar] [CrossRef]
- Menzies, F.D.; McCullough, S.J.; McKeown, I.M.; Forster, J.L.; Jess, S.; Batten, C.; Murchie, A.K.; Gloster, J.; Fallows, J.G.; Pelgrim, W.; et al. Evidence for transplacental and contact transmission of bluetongue virus in cattle. Vet. Rec. 2008, 163, 203–209. [Google Scholar] [CrossRef]
- Backx, A.; Heutink, R.; van Rooij, E.; van Rijn, P. Transplacental and oral transmission of wild-type bluetongue virus serotype 8 in cattle after experimental infection. Vet. Microbiol. 2009, 138, 235–243. [Google Scholar] [CrossRef]
- Allen, S.E.; Rothenburger, J.L.; Jardine, C.M.; Ambagala, A.; Hooper-McGrevy, K.; Colucci, N.; Furukawa-Stoffer, T.; Vigil, S.; Ruder, M.; Nemeth, N.M. Epizootic Hemorrhagic Disease in White-Tailed Deer, Canada. Emerg. Infect. Dis. 2019, 25, 832–834. [Google Scholar] [CrossRef]
- MacLachlan, N.J.; Osburn, B.I. Impact of bluetongue virus infection on the international movement and trade of ruminants. J. Am. Vet. Med. Assoc. 2006, 228, 1346–1349. [Google Scholar] [CrossRef]
- Hoar, B.R.; Carpenter, T.E.; Singer, R.S.; Gardner, I.A. Regional risk of exporting cattle seropositive for bluetongue virus from the United States. Am. J. Vet. Res. 2003, 64, 520–529. [Google Scholar] [CrossRef]
- Roberts, D.H.; Lucas, M.H.; Bell, R.A. Animal and animal product importation and the assessment of risk from bluetongue and other ruminant orbiviruses. Br. Vet. J. 1993, 149, 87–99. [Google Scholar] [CrossRef]
- McVey, D.S.; Drolet, B.S.; Ruder, M.G.; Wilson, W.C.; Nayduch, D.; Pfannenstiel, R.; Cohnstaedt, L.W.; MacLachlan, N.J.; Gay, C.G. Orbiviruses: A North American perspective. Vector Borne Zoonotic Dis. 2015, 15, 335–338. [Google Scholar] [CrossRef]
- Jimenez-Cabello, L.; Utrilla-Trigo, S.; Calvo-Pinilla, E.; Moreno, S.; Nogales, A.; Ortego, J.; Marin-Lopez, A. Viral vector vaccines against bluetongue virus. Microorganisms 2020, 9, 42. [Google Scholar] [CrossRef] [PubMed]
- Roy, P. Highly efficient vaccines for Bluetongue virus and a related Orbivirus based on reverse genetics. Curr. Opin. Virol. 2020, 44, 35–41. [Google Scholar] [CrossRef] [PubMed]
- van Rijn, P.A. Prospects of next-generation vaccines for bluetongue. Front. Vet. Sci. 2019, 6, 407. [Google Scholar] [CrossRef] [PubMed]
- Sunwoo, S.Y.; Noronha, L.E.; Morozov, I.; Trujillo, J.D.; Kim, I.J.; Schirtzinger, E.E.; Faburay, B.; Drolet, B.S.; Urbaniak, K.; McVey, D.S.; et al. Evaluation of A baculovirus-expressed VP2 subunit vaccine for the protection of white-tailed deer (Odocoileus virginianus) from epizootic hemorrhagic disease. Vaccines 2020, 8, 59. [Google Scholar] [CrossRef]
- Spooner, T.; Jones, A.E.; Fearnley, J.; Savani, R.; Turner, J.; Baylis, M. Bayesian optimisation of restriction zones for bluetongue control. Sci. Rep. 2020, 10, 15139. [Google Scholar] [CrossRef]
- Cappai, S.; Loi, F.; Coccollone, A.; Contu, M.; Capece, P.; Fiori, M.; Canu, S.; Foxi, C.; Rolesu, S. Retrospective analysis of bluetongue farm risk profile definition, based on biology, farm management practices and climatic data. Prev. Vet. Med. 2018, 155, 75–85. [Google Scholar] [CrossRef]
- Pfannenstiel, R.S.; Mullens, B.A.; Ruder, M.G.; Zurek, L.; Cohnstaedt, L.W.; Nayduch, D. Management of North American Culicoides biting midges: Current knowledge and research needs. Vector Borne Zoonotic Dis. 2015, 15, 374–384. [Google Scholar] [CrossRef]
- Maclachlan, N.J.; Mayo, C.E. Potential strategies for control of bluetongue, a globally emerging, Culicoides-transmitted viral disease of ruminant livestock and wildlife. Antivir. Res. 2013, 99, 79–90. [Google Scholar] [CrossRef]
- McVey, D.S.; MacLachlan, N.J. Vaccines for prevention of bluetongue and epizootic hemorrhagic disease in livestock: A North American perspective. Vector Borne Zoonotic Dis. 2015, 15, 385–396. [Google Scholar] [CrossRef]
- Savini, G.; Maclachlan, N.J.; Sanchez-Vizcaino, J.M.; Zientara, S. Vaccines against bluetongue in Europe. Comp. Immunol. Microbiol. Infect. Dis. 2008, 31, 101–120. [Google Scholar] [CrossRef]
- MacLachlan, N.J. The pathogenesis and immunology of bluetongue virus infection of ruminants. Comp. Immunol. Microbiol. Infect. Dis. 1994, 17, 197–206. [Google Scholar] [CrossRef]
- Merrill, M.M.; Boughton, R.K.; Lollis, L.O.; Sayler, K.A.; Wisely, S.M. Epidemiology of Bluetongue Virus and Epizootic Hemorrhagic Disease Virus in Beef Cattle on a Ranch in South-Central Florida. Vector Borne Zoonotic Dis. 2019, 19, 752–757. [Google Scholar] [CrossRef]
- Maclachlan, N.J.; Zientara, S.; Savini, G.; Daniels, P.W. Epizootic haemorrhagic disease. Rev. Sci. Tech. 2015, 34, 341–351. [Google Scholar] [CrossRef]
- MacLachlan, N.J. Bluetongue: Pathogenesis and duration of viraemia. Vet. Ital. 2004, 40, 462–467. [Google Scholar]
- Mejri, S.; Dhaou, S.B.; Jemli, M.; Breard, E.; Sailleau, C.; Sghaier, S.; Zouari, M.; Lorusso, A.; Savini, G.; Zientara, S.; et al. Epizootic haemorrhagic disease virus circulation in Tunisia. Vet. Ital. 2018, 54, 87–90. [Google Scholar] [CrossRef]
- Cetre-Sossah, C.; Roger, M.; Sailleau, C.; Rieau, L.; Zientara, S.; Breard, E.; Viarouge, C.; Beral, M.; Esnault, O.; Cardinale, E. Epizootic haemorrhagic disease virus in Reunion Island: Evidence for the circulation of a new serotype and associated risk factors. Vet. Microbiol. 2014, 170, 383–390. [Google Scholar] [CrossRef]
- Temizel, E.M.; Yesilbag, K.; Batten, C.; Senturk, S.; Maan, N.S.; Mertens, P.P.C.; Batmaz, H. Epizootic hemorrhagic disease in cattle, Western Turkey. Emerg. Infect. Dis. 2009, 15, 317–319. [Google Scholar] [CrossRef]
- Yadin, H.; Brenner, J.; Bumbrov, V.; Oved, Z.; Stram, Y.; Klement, E.; Perl, S.; Anthony, S.; Maan, S.; Batten, C.; et al. Epizootic haemorrhagic disease virus type 7 infection in cattle in Israel. Vet. Rec. 2008, 162, 53–56. [Google Scholar] [CrossRef]
- Eschbaumer, M.; Wernike, K.; Batten, C.A.; Savini, G.; Edwards, L.; Di Gennaro, A.; Teodori, L.; Oura, C.A.; Beer, M.; Hoffmann, B. Epizootic hemorrhagic disease virus serotype 7 in European cattle and sheep: Diagnostic considerations and effect of previous BTV exposure. Vet. Microbiol. 2012, 159, 298–306. [Google Scholar] [CrossRef]
- Stott, J.L.; Osburn, B.I. Immune response to bluetongue virus infection. Curr. Top. Microbiol. Immunol. 1990, 162, 163–178. [Google Scholar] [CrossRef]
- Chand, K.; Biswas, S.K.; Sing, B.; De, A.; Mondal, B. A sandwich ELISA for the detection of bluetongue virus in cell culture using antiserum against the recombinant VP7 protein. Vet. Ital. 2009, 45, 443–448. [Google Scholar] [PubMed]
- Drolet, B.S.; van Rijn, P.; Howerth, E.W.; Beer, M.; Mertens, P.P. A review of knowledge gaps and tools for orbivirus research. Vector Borne Zoonotic Dis. 2015, 15, 339–347. [Google Scholar] [CrossRef] [PubMed]
- Breard, E.; Viarouge, C.; Donnet, F.; Sailleau, C.; Rossi, S.; Pourquier, P.; Vitour, D.; Comtet, L.; Zientara, S. Evaluation of a commercial ELISA for detection of epizootic haemorrhagic disease antibodies in domestic and wild ruminant sera. Transbound. Emerg. Dis. 2020, 67, 2475–2481. [Google Scholar] [CrossRef] [PubMed]
- Afshar, A. Bluetongue: Laboratory diagnosis. Comp. Immunol. Microbiol. Infect. Dis. 1994, 17, 221–242. [Google Scholar] [CrossRef]
- Patton, J.F.; Work, T.M.; Jessup, D.A.; Hietala, S.K.; Oliver, M.N.; Maclachlan, N.J. Serologic detection of bluetongue virus infection of black-tailed deer: Comparison of serum neutralization, agar gel immunodiffusion, and competitive ELISA assays. J. Wildl. Dis. 1994, 30, 99–102. [Google Scholar] [CrossRef][Green Version]
- Ward, M.P.; Gardner, I.A.; Flanagan, M. Evaluation of an agar gel immunodiffusion test to detect infection of cattle with bluetongue viruses in Queensland, Australia. Vet. Microbiol. 1995, 45, 27–34. [Google Scholar] [CrossRef]
- OIE. Bluetongue (Infection with Bluetongue Virus), 8th ed.; World Organisation for Animal Health: Paris, France, 2018; Volume 3. [Google Scholar]
- Stevens, G.; McCluskey, B.; King, A.; O’Hearn, E.; Mayr, G. Review of the 2012 epizootic hemorrhagic disease outbreak in domestic ruminants in the United States. PLoS ONE 2015, 10, e0133359. [Google Scholar] [CrossRef]
- Mandrekar, J.N. Receiver operating characteristic curve in diagnostic test assessment. J. Thorac. Oncol. 2010, 5, 1315–1316. [Google Scholar] [CrossRef]
- Afshar, A.; Thomas, F.C.; Wright, P.F.; Shapiro, J.L.; Anderson, J. Comparison of competitive ELISA, indirect ELISA and standard AGID tests for detecting blue-tongue virus antibodies in cattle and sheep. Vet. Rec. 1989, 124, 136–141. [Google Scholar] [CrossRef]
- Ji, C.; Wei, Y.; Wang, J.; Zeng, Y.; Pan, H.; Liang, G.; Ma, J.; Gong, L.; Zhang, W.; Zhang, G.; et al. Development of a Dual Fluorescent Microsphere Immunological Assay for Detection of Pseudorabies Virus gE and gB IgG Antibodies. Viruses 2020, 12, 912. [Google Scholar] [CrossRef]
- Lindahl, J.F.; Ragan, I.K.; Rowland, R.R.; Wainaina, M.; Mbotha, D.; Wilson, W. A multiplex fluorescence microsphere immunoassay for increased understanding of Rift Valley fever immune responses in ruminants in Kenya. J. Virol. Methods 2019, 269, 70–76. [Google Scholar] [CrossRef]
- Ragan, I.K.; Davis, A.S.; McVey, D.S.; Richt, J.A.; Rowland, R.R.; Wilson, W.C. Evaluation of Fluorescence Microsphere Immunoassay for Detection of Antibodies to Rift Valley Fever Virus Nucleocapsid Protein and Glycoproteins. J. Clin. Microbiol. 2018, 56. [Google Scholar] [CrossRef]
- Langenhorst, R.J.; Lawson, S.; Kittawornrat, A.; Zimmerman, J.J.; Sun, Z.; Li, Y.; Christopher-Hennings, J.; Nelson, E.A.; Fang, Y. Development of a fluorescent microsphere immunoassay for detection of antibodies against porcine reproductive and respiratory syndrome virus using oral fluid samples as an alternative to serum-based assays. Clin. Vaccine Immunol. 2012, 19, 180–189. [Google Scholar] [CrossRef]
- Gimenez-Lirola, L.G.; Zhang, J.; Carrillo-Avila, J.A.; Chen, Q.; Magtoto, R.; Poonsuk, K.; Baum, D.H.; Pineyro, P.; Zimmerman, J. Reactivity of Porcine Epidemic Diarrhea Virus Structural Proteins to Antibodies against Porcine Enteric Coronaviruses: Diagnostic Implications. J. Clin. Microbiol. 2017, 55, 1426–1436. [Google Scholar] [CrossRef]
- Hossain, M.M.; Wilson, W.C.; Faburay, B.; Richt, J.; McVey, D.S.; Rowland, R.R. Multiplex Detection of IgG and IgM to Rift Valley Fever Virus Nucleoprotein, Nonstructural Proteins, and Glycoprotein in Ovine and Bovine. Vector Borne Zoonotic Dis. 2016, 16, 550–557. [Google Scholar] [CrossRef]
- Pinette, M.M.; Rodriguez-Lecompte, J.C.; Pasick, J.; Ojkic, D.; Leith, M.; Suderman, M.; Berhane, Y. Development of a duplex Fluorescent Microsphere Immunoassay (FMIA) for the detection of antibody responses to influenza A and newcastle disease viruses. J. Immunol. Methods 2014, 405, 167–177. [Google Scholar] [CrossRef]
- Feichtner, F.; Schachner, A.; Berger, E.; Hess, M. Fiber-based fluorescent microsphere immunoassay (FMIA) as a novel multiplex serodiagnostic tool for simultaneous detection and differentiation of all clinically relevant fowl adenovirus (FAdV) serotypes. J. Immunol. Methods 2018, 458, 33–43. [Google Scholar] [CrossRef]
- Trevethan, R. Sensitivity, Specificity, and Predictive Values: Foundations, Pliabilities, and Pitfalls in Research and Practice. Front. Public Health 2017, 5, 307. [Google Scholar] [CrossRef]
- Martins, T.B. Development of internal controls for the Luminex instrument as part of a multiplex seven-analyte viral respiratory antibody profile. Clin. Diagn. Lab. Immunol. 2002, 9, 41–45. [Google Scholar] [CrossRef]
- IDEXX. ELISA Technical Guide; IDEXX Laboratories, Inc.: Westbrook, ME, USA, 2019. [Google Scholar]

| Criterion | Sensitivity | 95% CI | Specificity | 95% CI | +LR 1 | −LR 2 |
|---|---|---|---|---|---|---|
| >−0.1061 | 90.00 | 81.2–95.6 | 55.74 | 46.5–64.7 | 2.03 | 0.18 |
| >−0.1013 | 88.75 | 79.7–94.7 | 55.74 | 46.5–64.7 | 2.01 | 0.20 |
| >−0.0167 * | 88.75 | 79.7–94.7 | 68.85 | 59.8–76.9 | 2.85 | 0.16 |
| >−0.0003 | 86.25 | 76.7–92.9 | 68.85 | 59.8–76.9 | 2.77 | 0.20 |
| >−0.0004 | 86.25 | 76.7–92.9 | 69.67 | 60.7–77.7 | 2.84 | 0.20 |
| Criterion | Sensitivity | 95% CI | Specificity | 95% CI | +LR 1 | −LR 2 |
|---|---|---|---|---|---|---|
| >0.0401 | 76.27 | 67.6–83.6 | 69.05 | 58.0–78.7 | 2.46 | 0.34 |
| >0.0437 | 75.42 | 66.6–82.9 | 69.05 | 58.0–78.7 | 2.44 | 0.36 |
| >0.0623 * | 75.42 | 66.6–82.9 | 71.43 | 60.5–80.8 | 2.64 | 0.34 |
| >0.089 | 72.88 | 63.9–80.7 | 71.43 | 60.5–80.8 | 2.55 | 0.38 |
| >0.1019 | 72.88 | 63.9–80.7 | 72.62 | 61.8–81.8 | 2.66 | 0.37 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Drolet, B.S.; Reister-Hendricks, L.M. A Duplex Fluorescent Microsphere Immunoassay for Detection of Bluetongue and Epizootic Hemorrhagic Disease Virus Antibodies in Cattle Sera. Viruses 2021, 13, 682. https://doi.org/10.3390/v13040682
Drolet BS, Reister-Hendricks LM. A Duplex Fluorescent Microsphere Immunoassay for Detection of Bluetongue and Epizootic Hemorrhagic Disease Virus Antibodies in Cattle Sera. Viruses. 2021; 13(4):682. https://doi.org/10.3390/v13040682
Chicago/Turabian StyleDrolet, Barbara S., and Lindsey M. Reister-Hendricks. 2021. "A Duplex Fluorescent Microsphere Immunoassay for Detection of Bluetongue and Epizootic Hemorrhagic Disease Virus Antibodies in Cattle Sera" Viruses 13, no. 4: 682. https://doi.org/10.3390/v13040682
APA StyleDrolet, B. S., & Reister-Hendricks, L. M. (2021). A Duplex Fluorescent Microsphere Immunoassay for Detection of Bluetongue and Epizootic Hemorrhagic Disease Virus Antibodies in Cattle Sera. Viruses, 13(4), 682. https://doi.org/10.3390/v13040682






