Enteric Viral Co-Infections: Pathogenesis and Perspective
Abstract
1. Introduction
2. Virus–Virus Co-Infection Prevalence
2.1. Co-Infection in Acute Gastroenteritis (AGE)
2.2. Co-Infection in Asymptomatic Individuals
2.3. Co-Infection Effects on Vaccines and Treatments
3. Cellular and Molecular Consequences of Co-Infection
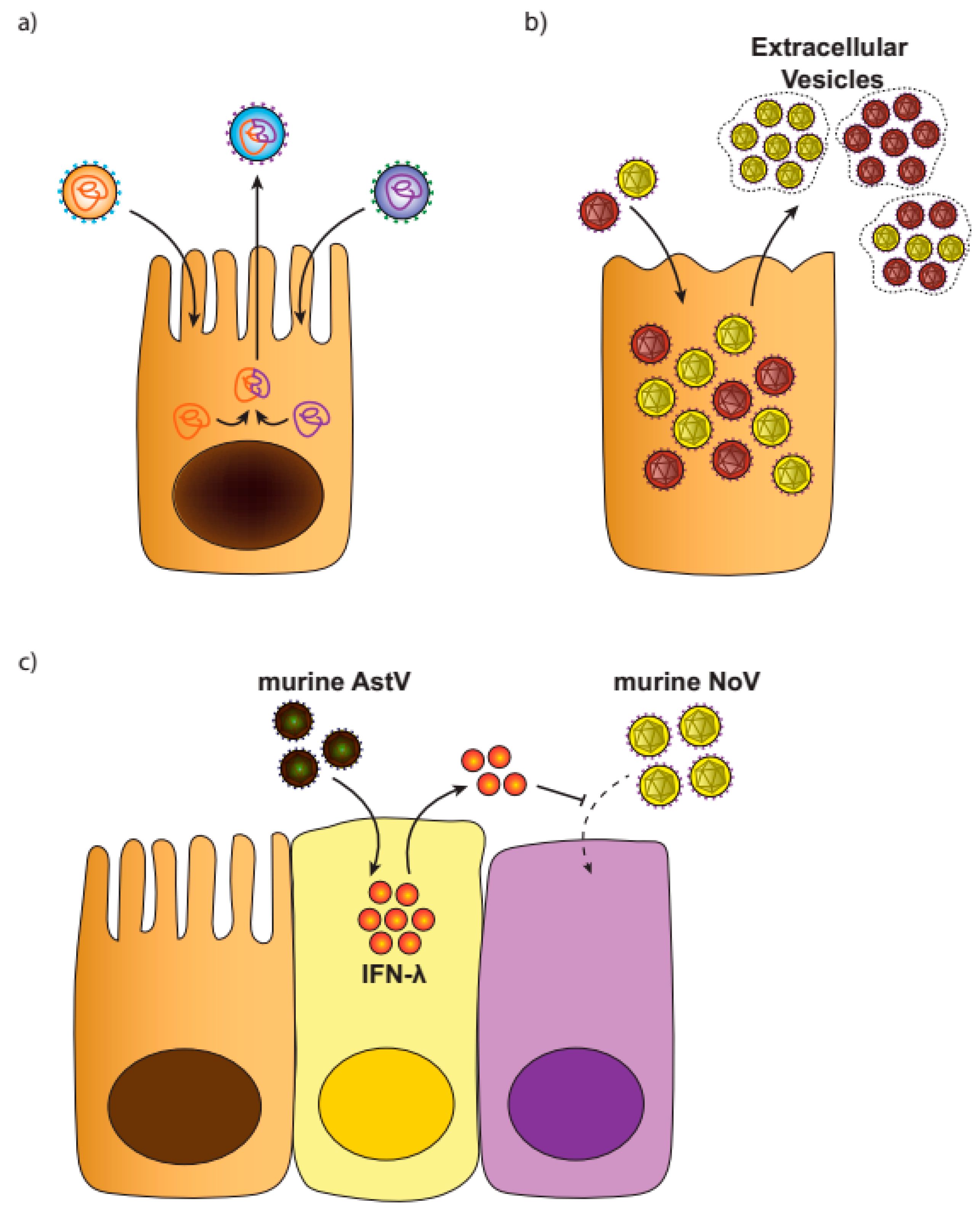
3.1. Viral Recombination or Reassortment
3.2. Virus Aggregates
3.3. Viral Interference
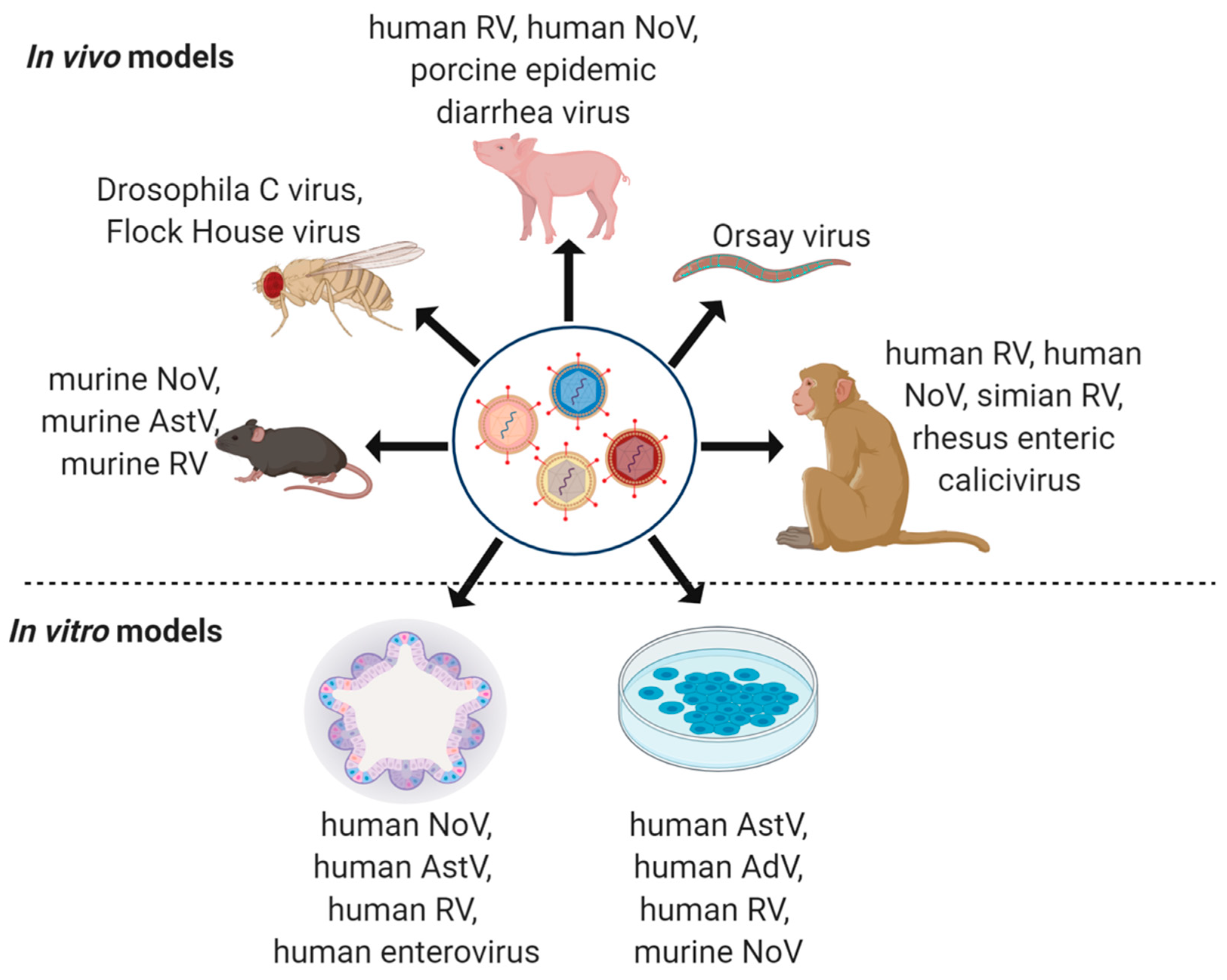
4. Model Systems to Study Virus Co-Infections
4.1. Non-Human Primates
4.2. Pig Models
4.3. Mouse Models
4.4. Drosophila melanogaster
4.5. Caenorhabditis elegans
4.6. Immortalized In Vitro Cell Lines
4.7. Intestinal Organoids and Enteroids
5. Conclusions and Future Directions
Author Contributions
Funding
Conflicts of Interest
References
- Parashar, U.D.; Hummelman, E.G.; Bresee, J.S.; Miller, M.A.; Glass, R.I. Global illness and deaths caused by rotavirus disease in children. Emerg. Infect. Dis. 2003, 9, 565–572. [Google Scholar] [CrossRef] [PubMed]
- Collaborators, G.B.D. Estimates of global, regional, and national morbidity, mortality, and aetiologies of diarrhoeal diseases: A systematic analysis for the global burden of disease study 2015. Lancet Infect. Dis. 2017, 17, 909–948. [Google Scholar] [CrossRef]
- Mills, J.A.P.; Liu, J.; Rogawski, E.T.; Kabir, F.; Lertsethtakarn, P.; Siguas, M.; Investigators, M.-E.N. Use of quantitative molecular diagnostic methods to assess the aetiology, burden, and clinical characteristics of diarrhoea in children in low-resource settings: A reanalysis of the MAL-ED cohort study. Lancet Glob. Health 2018, 6, e1309–e1318. [Google Scholar] [CrossRef]
- Graves, N.S. Acute gastroenteritis. Prim. Care 2013, 40, 727–741. [Google Scholar] [CrossRef] [PubMed]
- Eckardt, A.J.; Baumgart, D.C. Viral gastroenteritis in adults. Recent Pat. Antiinfect. Drug Discov. 2011, 6, 54–63. [Google Scholar] [CrossRef]
- Burnett, E.; Jonesteller, C.L.; Tate, J.E.; Yen, C.; Parashar, U.D. Global impact of rotavirus vaccination on childhood hospitalizations and mortality from diarrhea. J. Infect. Dis. 2017, 215, 1666–1672. [Google Scholar] [CrossRef]
- Vu, D.L.; Bosch, A.; Pinto, R.M.; Guix, S. Epidemiology of classic and novel human astrovirus: Gastroenteritis and beyond. Viruses 2017, 9, 33. [Google Scholar] [CrossRef]
- Aktas, O.; Aydin, H.; Timurkan, M.O. A molecular study on the prevalence and co-infections of rotavirus, norovirus, astrovirus and adenovirus in children with gastroenteritis. Minerva Pediatr. 2019, 71, 431–437. [Google Scholar] [CrossRef]
- Ahmed, S.M.; Hall, A.J.; Robinson, A.E.; Verhoef, L.; Premkumar, P.; Parashar, U.D.; Lopman, B.A. Global prevalence of norovirus in cases of gastroenteritis: A systematic review and meta-analysis. Lancet Infect. Dis. 2014, 14, 725–730. [Google Scholar] [CrossRef]
- Xing, W.; Liao, Q.; Viboud, C.; Zhang, J.; Sun, J.; Wu, J.T.; Yu, H. Hand, foot, and mouth disease in China, 2008–2012: An epidemiological study. Lancet Infect. Dis. 2014, 14, 308–318. [Google Scholar] [CrossRef]
- Jevsnik, M.; Steyer, A.; Zrim, T.; Pokorn, M.; Mrvic, T.; Grosek, S.; Petrovec, M. Detection of human coronaviruses in simultaneously collected stool samples and nasopharyngeal swabs from hospitalized children with acute gastroenteritis. Virol. J. 2013, 10, 46. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, T.; Perlot, T.; Rehman, A.; Trichereau, J.; Ishiguro, H.; Paolino, M.; Penninger, J.M. ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature 2012, 487, 477–481. [Google Scholar] [CrossRef] [PubMed]
- Burgueno, J.F.; Reich, A.; Hazime, H.; Quintero, M.A.; Fernandez, I.; Fritsch, J.; Abreu, M.T. Expression of SARS-CoV-2 entry molecules ACE2 and TMPRSS2 in the gut of patients with IBD. Inflamm. Bowel Dis. 2020, 26, 797–808. [Google Scholar] [CrossRef] [PubMed]
- Holtz, L.R.; Cao, S.; Zhao, G.; Bauer, I.K.; Denno, D.M.; Klein, E.J.; Wang, D. Geographic variation in the eukaryotic virome of human diarrhea. Virology 2014, 468–470, 556–564. [Google Scholar] [CrossRef] [PubMed]
- Foca, A.; Liberto, M.C.; Quirino, A.; Marascio, N.; Zicca, E.; Pavia, G. Gut inflammation and immunity: What is the role of the human gut virome? Mediat. Inflamm. 2015, 2015, 326032. [Google Scholar] [CrossRef] [PubMed]
- Neil, J.A.; Cadwell, K. The Intestinal virome and immunity. J. Immunol. 2018, 201, 1615–1624. [Google Scholar] [CrossRef]
- Reyes, A.; Haynes, M.; Hanson, N.; Angly, F.E.; Heath, A.C.; Rohwer, F.; Gordon, J.I. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 2010, 466, 334–338. [Google Scholar] [CrossRef]
- Roux, S.; Hawley, A.K.; Beltran, M.T.; Scofield, M.; Schwientek, P.; Stepanauskas, R.; Woyke, T.; Hallam, S.J.; Sullivan, M.B. Ecology and evolution of viruses infecting uncultivated SUP05 bacteria as revealed by single-cell- and meta-genomics. eLife 2014, 3, e03125. [Google Scholar] [CrossRef]
- Labonte, J.M.; Swan, B.K.; Poulos, B.; Luo, H.; Koren, S.; Hallam, S.J.; Sullivan, M.B.; Woyke, T.; Wommack, E.C.; Stepanauskas, R. Single-cell genomics-based analysis of virus-host interactions in marine surface bacterioplankton. ISME J. 2015, 9, 2386–2399. [Google Scholar] [CrossRef]
- Munoz, S.L.D. Viral co-infection is shaped by host ecology and virus-virus interactions across diverse microbial taxa and environments. Virus Evol. 2017, 3. [Google Scholar] [CrossRef]
- Dzunkova, M.; Low, S.J.; Daly, J.N.; Deng, L.; Rinke, C.; Hugenholtz, P. Defining the human gut host-phage network through single-cell viral tagging. Nat. Microbiol. 2019, 4, 2192–2203. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Zhao, C.; Zhang, H.; Mattei, L.; Mix, S.S.; Bittinger, K.; Kessler, L.R.; Wu, G.D.; Baldassano, R.N.; DeRusso, P.; et al. The stepwise assembly of the neonatal virome is modulated by breastfeeding. Nature 2020, 581, 470–474. [Google Scholar] [CrossRef] [PubMed]
- Maqsood, R.; Rodgers, R.; Rodriguez, C.; Handley, S.A.; Ndao, I.M.; Tarr, P.I.; Holtz, L.R. Discordant transmission of bacteria and viruses from mothers to babies at birth. Microbiome 2019, 7, 156. [Google Scholar] [CrossRef] [PubMed]
- Lim, E.S.; Zhou, Y.; Zhao, G.; Bauer, I.K.; Droit, L.; Ndao, I.M.; Holtz, L.R. Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat. Med. 2015, 21, 1228–1234. [Google Scholar] [CrossRef]
- Pijnacker, R.; Pelt, W.V.; Vennema, H.; Kortbeek, L.M.; Notermans, D.W.; Franz, E.; Gras, L.M. Clinical relevance of enteropathogen co-infections in preschool children-a population-based repeated cross-sectional study. Clin. Microbiol. Infect. 2019, 25, 1013–1037. [Google Scholar] [CrossRef]
- Kapusinszky, B.; Minor, P.; Delwart, E. Nearly constant shedding of diverse enteric viruses by two healthy infants. J. Clin. Microbiol. 2012, 50, 3427–3434. [Google Scholar] [CrossRef]
- Tandukar, S.; Sherchand, J.B.; Karki, S.; Malla, B.; Shrestha, R.G.; Bhandari, D.; Haramoto, E. Co-infection by waterborne enteric viruses in children with gastroenteritis in nepal. Healthcare 2019, 7, 9. [Google Scholar] [CrossRef]
- Mengelle, C.; Mansuy, J.M.; Prere, M.F.; Grouteau, E.; Claudet, I.; Kamar, N.; Izopet, J. Simultaneous detection of gastrointestinal pathogens with a multiplex Luminex-based molecular assay in stool samples from diarrhoeic patients. Clin. Microbiol. Infect. 2013, 19, E458–E465. [Google Scholar] [CrossRef]
- Eibach, D.; Krumkamp, R.; Hahn, A.; Sarpong, N.; Sarkodie, Y.A.; Leva, A.; Tannich, E. Application of a multiplex PCR assay for the detection of gastrointestinal pathogens in a rural African setting. BMC Infect. Dis. 2016, 16, 150. [Google Scholar] [CrossRef]
- Mokomane, M.; Tate, J.E.; Steenhoff, A.P.; Esona, M.D.; Bowen, M.D.; Lechiile, K.; Goldfarb, D.M. Evaluation of the influence of gastrointestinal coinfections on rotavirus vaccine effectiveness in botswana. Pediatr. Infect. Dis. J. 2018, 37, e58–e62. [Google Scholar] [CrossRef]
- Mills, J.A.P.; Babji, S.; Bodhidatta, L.; Gratz, J.; Haque, R.; Havt, A.; Investigators, M.-E.N. Pathogen-specific burdens of community diarrhoea in developing countries: A multisite birth cohort study (MAL-ED). Lancet Glob. Health. 2015, 3, e564–e575. [Google Scholar] [CrossRef]
- Clark, A.; Black, R.; Tate, J.; Roose, A.; Kotloff, K.; Lam, D.; Steele, D. Estimating global, regional and national rotavirus deaths in children aged <5 years: Current approaches, new analyses and proposed improvements. PLoS ONE 2017, 12, e0183392. [Google Scholar] [CrossRef]
- Deen, J.; Lopez, A.L.; Kanungo, S.; Wang, X.Y.; Anh, D.D.; Tapia, M.; Grais, R.F. Improving rotavirus vaccine coverage: Can newer-generation and locally produced vaccines help? Hum. Vaccin. Immunother. 2018, 14, 495–499. [Google Scholar] [CrossRef] [PubMed]
- Pindyck, T.; Tate, J.E.; Parashar, U.D. A decade of experience with rotavirus vaccination in the United States -vaccine uptake, effectiveness, and impact. Expert Rev. Vaccines 2018, 17, 593–606. [Google Scholar] [CrossRef] [PubMed]
- Ramig, R.F. Pathogenesis of intestinal and systemic rotavirus infection. J. Virol. 2004, 78, 10213–10220. [Google Scholar] [CrossRef] [PubMed]
- McAtee, C.L.; Webman, R.; Gilman, R.H.; Mejia, C.; Bern, C.; Apaza, S.; Torrico, F. Burden of norovirus and rotavirus in children after rotavirus vaccine introduction, Cochabamba, Bolivia. Am. J. Trop Med. Hyg. 2016, 94, 212–217. [Google Scholar] [CrossRef]
- Hassan, E.; Baldridge, M.T. Norovirus encounters in the gut: Multifaceted interactions and disease outcomes. Mucosal Immunol. 2019, 12, 1259–1267. [Google Scholar] [CrossRef]
- Graaf, M.d.; Beek, J.V.; Koopmans, M.P. Human norovirus transmission and evolution in a changing world. Nat. Rev. Microbiol. 2016, 14, 421–433. [Google Scholar] [CrossRef]
- Zhang, S.X.; Zhou, Y.M.; Xu, W.; Tian, L.G.; Chen, J.X.; Chen, S.H.; Zhou, X.N. Impact of co-infections with enteric pathogens on children suffering from acute diarrhea in southwest China. Infect. Dis. Poverty 2016, 5, 64. [Google Scholar] [CrossRef]
- Green, K.Y.; Kaufman, S.S.; Nagata, B.M.; Chaimongkol, N.; Kim, D.Y.; Levenson, E.A.; Sosnovtsev, S.V. Human norovirus targets enteroendocrine epithelial cells in the small intestine. Nat. Commun. 2020, 11, 2759. [Google Scholar] [CrossRef]
- Karandikar, U.C.; Crawford, S.E.; Ajami, N.J.; Murakami, K.; Kou, B.; Ettayebi, K.; Estes, M.K. Detection of human norovirus in intestinal biopsies from immunocompromised transplant patients. J. Gen. Virol. 2016, 97, 2291–2300. [Google Scholar] [CrossRef] [PubMed]
- Dreps, S.B.; Bucardo, F.; Vinje, J. Sapovirus: An important cause of acute gastroenteritis in children. Lancet Child. Adolesc. Health 2019, 3, 758–759. [Google Scholar] [CrossRef]
- Varela, M.F.; Rivadulla, E.; Lema, A.; Romalde, J.L. Human sapovirus among outpatients with acute gastroenteritis in spain: A one-year study. Viruses 2019, 11, 144. [Google Scholar] [CrossRef] [PubMed]
- Roels, G.L.; Cramer, J.P.; Mendelman, P.M.; Sherwood, J.; Clemens, R.; Aerssens, A.; Damme, P.V. Safety and immunogenicity of different formulations of norovirus vaccine candidate in healthy adults: A randomized, controlled, double-blind clinical trial. J. Infect. Dis. 2018, 217, 597–607. [Google Scholar] [CrossRef] [PubMed]
- Hallowell, B.D.; Parashar, U.D.; Hall, A.J. Epidemiologic challenges in norovirus vaccine development. Hum. Vaccin Immunother. 2019, 15, 1279–1283. [Google Scholar] [CrossRef] [PubMed]
- Koopmans, M.P.; Bijen, M.H.; Monroe, S.S.; Vinje, J. Age-stratified seroprevalence of neutralizing antibodies to astrovirus types 1 to 7 in humans in The Netherlands. Clin. Diagn. Lab. Immunol. 1998, 5, 33–37. [Google Scholar] [CrossRef]
- Kriston, S.; Willcocks, M.M.; Carter, M.J.; Cubitt, W.D. Seroprevalence of astrovirus types 1 and 6 in London, determined using recombinant virus antigen. Epidemiol. Infect. 1996, 117, 159–164. [Google Scholar] [CrossRef]
- Wunderli, W.; Meerbach, A.; Gungor, T.; Berger, C.; Greiner, O.; Caduff, R.; Tapparel, C. Astrovirus infection in hospitalized infants with severe combined immunodeficiency after allogeneic hematopoietic stem cell transplantation. PLoS ONE 2011, 6, e27483. [Google Scholar] [CrossRef]
- Quan, P.L.; Wagner, T.A.; Briese, T.; Torgerson, T.R.; Hornig, M.; Tashmukhamedova, A.; Lipkin, W.I. Astrovirus encephalitis in boy with X-linked agammaglobulinemia. Emerg. Infect. Dis. 2010, 16, 918–925. [Google Scholar] [CrossRef]
- Brown, J.R.; Morfopoulou, S.; Hubb, J.; Emmett, W.A.; Ip, W.; Shah, D.; Breuer, J. Astrovirus VA1/HMO-C: An increasingly recognized neurotropic pathogen in immunocompromised patients. Clin. Infect. Dis. 2015, 60, 881–888. [Google Scholar] [CrossRef]
- Cordey, S.; Vu, D.L.; Schibler, M.; L’Huillier, A.G.; Brito, F.; Docquier, M.; Kaiser, L. Astrovirus MLB2, a new gastroenteric virus associated with meningitis and disseminated infection. Emerg. Infect. Dis. 2016, 22, 846–853. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Kuroda, M.; Kasai, M.; Matsui, H.; Fukuyama, T.; Katano, H.; Taya, K.T. Acute encephalopathy in an immunocompromised boy with astrovirus-MLB1 infection detected by next generation sequencing. J. Clin. Virol. 2016, 78, 66–70. [Google Scholar] [CrossRef] [PubMed]
- Kolawole, A.O.; Mirabelli, C.; Hill, D.R.; Svoboda, S.A.; Janowski, A.B.; Passalacqua, K.D.; Wobus, C.E. Astrovirus replication in human intestinal enteroids reveals multi-cellular tropism and an intricate host innate immune landscape. PLoS Pathog. 2019, 15, e1008057. [Google Scholar] [CrossRef] [PubMed]
- Mehndiratta, M.M.; Mehndiratta, P.; Pande, R. Poliomyelitis: Historical facts, epidemiology, and current challenges in eradication. Neurohospitalist 2014, 4, 223–229. [Google Scholar] [CrossRef]
- Lancaster, K.Z.; Pfeiffer, J.K. Limited trafficking of a neurotropic virus through inefficient retrograde axonal transport and the type I interferon response. PLoS Pathog. 2010, 6, e1000791. [Google Scholar] [CrossRef] [PubMed]
- Mwengee, W.; Okeibunor, J.; Poy, A.; Shaba, K.; Kinuani, L.M.; Minkoulou, E.; Mkanda, P. Polio eradication initiative: Contribution to improved communicable diseases surveillance in WHO African region. Vaccine 2016, 34, 5170–5174. [Google Scholar] [CrossRef]
- Salk, J.E. Considerations in the preparation and use of poliomyelitis virus vaccine. J. Am. Med. Assoc. 1955, 158, 1239–1248. [Google Scholar] [CrossRef]
- Krugman, S.; Warren, J.; Eiger, M.S.; Berman, P.H.; Michaels, R.M.; Sabin, A.B. Immunization with live attenuated poliovirus vaccine. Am. J. Dis. Child. 1961, 101, 23–29. [Google Scholar] [CrossRef]
- Shin, W.J.; Hara, D.; Gbormittah, F.; Chang, H.; Chang, B.S.; Jung, J.U. Development of thermostable lyophilized sabin inactivated poliovirus vaccine. mBio 2018, 9. [Google Scholar] [CrossRef]
- Platt, L.R.; Estivariz, C.F.; Sutter, R.W. Vaccine-associated paralytic poliomyelitis: A review of the epidemiology and estimation of the global burden. J. Infect. Dis. 2014, 210, S380–S389. [Google Scholar] [CrossRef]
- Jorba, J.; Diop, O.M.; Iber, J.; Henderson, E.; Zhao, K.; Quddus, A.; Burns, C.C. Update on vaccine-derived poliovirus outbreaks-worldwide, January 2018–June 2019. MMWR Morb. Mortal. Wkly. Rep. 2019, 68, 1024–1028. [Google Scholar] [CrossRef] [PubMed]
- Koh, W.M.; Bogich, T.; Siegel, K.; Jin, J.; Chong, E.Y.; Tan, C.Y.; Cook, A.R. The epidemiology of hand, foot and mouth disease in Asia: A systematic review and analysis. Pediatr. Infect. Dis. J. 2016, 35, e285–e300. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Wu, S.; Xiong, Y.; Li, T.; Wen, Z.; Yan, M.; Wu, J. Co-circulation and genomic recombination of coxsackievirus A16 and enterovirus 71 during a large outbreak of hand, foot, and mouth disease in Central China. PLoS ONE 2014, 9, e96051. [Google Scholar] [CrossRef] [PubMed]
- Han, J.F.; Zhang, Y.; Hou, P.Q.; Zhu, S.Y.; Wu, X.Y.; Zhao, H.; Qin, C.F. Human enterovirus co-infection in severe HFMD patients in China. J. Clin. Virol. 2014, 61, 621–622. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Du, Z.; Feng, M.; Che, Y.; Li, Q. A severe case of co-infection with Enterovirus 71 and vaccine-derived Poliovirus type II. J. Clin. Virol. 2015, 72, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Patil, P.R.; Chitambar, S.D.; Gopalkrishna, V. Molecular surveillance of non-polio enterovirus infections in patients with acute gastroenteritis in Western India: 2004–2009. J. Med. Virol. 2015, 87, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Chansaenroj, J.; Tuanthap, S.; Thanusuwannasak, T.; Duang, A.I.; Klinfueng, S.; Thaneskongtong, N.; Poovorawan, Y. Human enteroviruses associated with and without diarrhea in Thailand between 2010 and 2016. PLoS ONE 2017, 12, e0182078. [Google Scholar] [CrossRef]
- Kumthip, K.; Khamrin, P.; Ushijima, H.; Maneekarn, N. Multiple enterovirus genotypes circulating in children hospitalized with acute gastroenteritis in Thailand. Infect. Genet. Evol. 2017, 55, 324–331. [Google Scholar] [CrossRef]
- Lynch, J.P.; Fishbein, M.; Echavarria, M. Adenovirus. Semin. Respir. Crit. Care Med. 2011, 32, 494–511. [Google Scholar] [CrossRef]
- Qiu, F.Z.; Shen, X.X.; Li, G.X.; Zhao, L.; Chen, C.; Duan, S.X.; Ma, X.J. Adenovirus associated with acute diarrhea: A case-control study. BMC Infect. Dis. 2018, 18, 450. [Google Scholar] [CrossRef]
- Kumthip, K.; Khamrin, P.; Ushijima, H.; Maneekarn, N. Enteric and non-enteric adenoviruses associated with acute gastroenteritis in pediatric patients in Thailand, 2011 to 2017. PLoS ONE 2019, 14, e0220263. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.D.P.; Manna, B.; Sarkar, M.C. Molecular characterization of enteric adenovirus genotypes 40 and 41 identified in children with acute gastroenteritis in Kolkata, India during 2013–2014. J. Med. Virol. 2017, 89, 606–614. [Google Scholar] [CrossRef] [PubMed]
- Uhnoo, I.; Wadell, G.; Svensson, L.; Johansson, M.E. Importance of enteric adenoviruses 40 and 41 in acute gastroenteritis in infants and young children. J. Clin. Microbiol. 1984, 20, 365–372. [Google Scholar] [CrossRef] [PubMed]
- Filho, E.P.; Faria, N.R.d.C.; Fialho, A.M.; Assis, R.S.d.; Almeida, M.M.S.; Rocha, M.; Leite, J.P.G. Adenoviruses associated with acute gastroenteritis in hospitalized and community children up to 5 years old in Rio de Janeiro and Salvador, Brazil. J. Med. Microbiol. 2007, 56, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Kosulin, K. Intestinal HAdV infection: Tissue specificity, persistence, and implications for antiviral therapy. Viruses 2019, 11, 804. [Google Scholar] [CrossRef] [PubMed]
- Lion, T. Adenovirus infections in immunocompetent and immunocompromised patients. Clin. Microbiol. Rev. 2014, 27, 441–462. [Google Scholar] [CrossRef] [PubMed]
- Echavarria, M. Adenoviruses in immunocompromised hosts. Clin. Microbiol. Rev. 2008, 21, 704–715. [Google Scholar] [CrossRef]
- Zhang, S.Y.; Luo, Y.P.; Huang, D.D.; Fan, H.; Lu, Q.B.; Wo, Y.; Liu, W. Fatal pneumonia cases caused by human adenovirus 55 in immunocompetent adults. Infect. Dis. 2016, 48, 40–47. [Google Scholar] [CrossRef]
- Thomas, P.D.; Pollok, R.C.; Gazzard, B.G. Enteric viral infections as a cause of diarrhoea in the acquired immunodeficiency syndrome. HIV Med. 1999, 1, 19–24. [Google Scholar] [CrossRef]
- Roy, S.; Calcedo, R.; Jaszek, A.M.; Keough, M.; Peng, H.; Wilson, J.M. Adenoviruses in lymphocytes of the human gastro-intestinal tract. PLoS ONE 2011, 6, e24859. [Google Scholar] [CrossRef]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H. First case of 2019 novel coronavirus in the United States. N. Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef] [PubMed]
- Xiao, F.; Tang, M.; Zheng, X.; Liu, Y.; Li, X.; Shan, H. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology 2020, 158, 1831–1833. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Li, X.; Zhu, B.; Liang, H.; Fang, C.; Gong, Y.; Gong, S. Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral shedding. Nat. Med. 2020, 26, 502–505. [Google Scholar] [CrossRef] [PubMed]
- Munster, V.J.; Feldmann, F.; Williamson, B.N.; Doremalen, N.V.; Perez, L.P.; Schulz, J.; Wit, E.d. Respiratory disease in rhesus macaques inoculated with SARS-CoV-2. Nature 2020. [Google Scholar] [CrossRef]
- Kim, Y.I.; Kim, S.G.; Kim, S.M.; Kim, E.H.; Park, S.J.; Yu, K.M.; Choi, Y.K. Infection and rapid transmission of SARS-CoV-2 in Ferrets. Cell Host Microbe 2020, 27, 704–709. [Google Scholar] [CrossRef]
- Lu, X.; Zhang, L.; Du, H.; Zhang, J.; Li, Y.Y.; Qu, J.; Chinese pediatric novel coronavirus study, T. SARS-CoV-2 infection in children. N. Engl. J. Med. 2020, 382, 1663–1665. [Google Scholar] [CrossRef]
- Nowak, M.D.; Sordillo, E.M.; Gitman, M.R.; Paniz Mondolfi, A.E. Co-infection in SARS-CoV-2 infected patients: Where are influenza virus and rhinovirus/enterovirus? J. Med. Virol. 2020. [Google Scholar] [CrossRef]
- Zang, R.; Castro, M.F.G.; McCune, B.T.; Zeng, Q.; Rothlauf, P.W.; Sonnek, N.M.; Ding, S. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 2020, 5. [Google Scholar] [CrossRef]
- Xiao, F.; Sun, J.; Xu, Y.; Li, F.; Huang, X.; Li, H.; Zhao, J. Infectious SARS-CoV-2 in feces of patient with severe COVID-19. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef]
- Ruggeri, F.M.; Delogu, R.; Petouchoff, T.; Tcheremenskaia, O.; Petris, S.D.; Fiore, L. RotaNet-Italy study group molecular characterization of rotavirus strains from children with diarrhea in Italy, 2007–2009. J. Med. Virol. 2011, 83, 1657–1668. [Google Scholar] [CrossRef]
- Eiden, J.; Losonsky, G.A.; Johnson, J.; Yolken, R.H. Rotavirus RNA variation during chronic infection of immunocompromised children. Pediatr. Infect. Dis. 1985, 4, 632–637. [Google Scholar] [CrossRef] [PubMed]
- Saito, M.; Apaza, S.G.; Espetia, S.; Velasquez, D.; Cabrera, L.; Loli, S. Multiple norovirus infections in a birth cohort in a Peruvian Periurban community. Clin. Infect. Dis. 2014, 58, 483–491. [Google Scholar] [CrossRef] [PubMed]
- Gruber, J.F.; Bowman, N.M.; Dreps, S.B.; Reyes, Y.; Belson, C.; Michaels, K.C.; Bucardo, F. Risk factors for norovirus gastroenteritis among nicaraguan children. Am. J. Trop. Med. Hyg. 2017, 97, 937–943. [Google Scholar] [CrossRef] [PubMed]
- Menon, V.K.; George, S.; Sarkar, R.; Giri, S.; Samuel, P.; Vivek, R.; Kang, G. Norovirus gastroenteritis in a birth cohort in Southern India. PLoS ONE 2016, 11, e0157007. [Google Scholar] [CrossRef] [PubMed]
- Rao, D.C.; Babu, M.A.; Raghavendra, A.; Dhananjaya, D.; Kumar, S.; Maiya, P.P. Non-polio enteroviruses and their association with acute diarrhea in children in India. Infect. Genet. Evol. 2013, 17, 153–161. [Google Scholar] [CrossRef]
- Wu, D.; Changwen, K.; Wei, L.; Monagin, C.; Jin, Y.; Cong, M.; Jun, S. A large outbreak of hand, foot, and mouth disease caused by EV71 and CAV16 in Guangdong, China, 2009. Arch. Virol. 2011, 156, 945–953. [Google Scholar] [CrossRef]
- Rasanen, S.; Lappalainen, S.; Kaikkonen, S.; Hamalainen, M.; Salminen, M.; Vesikari, T. Mixed viral infections causing acute gastroenteritis in children in a waterborne outbreak. Epidemiol. Infect. 2010, 138, 1227–1234. [Google Scholar] [CrossRef]
- Chitambar, S.; Gopalkrishna, V.; Chhabra, P.; Patil, P.; Verma, H.; Lahon, A.; Pawar, S. Diversity in the enteric viruses detected in outbreaks of gastroenteritis from Mumbai, Western India. Int. J. Environ. Res. Public Health 2012, 9, 895–915. [Google Scholar] [CrossRef]
- Roman, E.; Wilhelmi, I.; Colomina, J.; Villar, J.; Cilleruelo, M.L.; Nebreda, V.; Fauquier, A.S. Acute viral gastroenteritis: Proportion and clinical relevance of multiple infections in Spanish children. J. Med. Microbiol. 2001, 52, 435–440. [Google Scholar] [CrossRef]
- Bon, F.; Fascia, P.; Dauvergne, M.; Tenenbaum, D.; Planson, H.; Petion, A.M.; Kohli, E. Prevalence of group a rotavirus, human calicivirus, astrovirus, and adenovirus type 40 and 41 infections among children with acute gastroenteritis in Dijon, France. J. Clin. Microbiol. 1999, 37, 3055–3058. [Google Scholar] [CrossRef]
- Leblanc, D.; Inglis, G.D.; Boras, V.F.; Brassard, J.; Houde, A. The prevalence of enteric RNA viruses in stools from diarrheic and non-diarrheic people in southwestern Alberta, Canada. Arch. Virol. 2017, 162, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Vu, D.L.; Sabria, A.; Aregall, N.; Michl, K.; Garrido, V.R.; Goterris, L.; Guix, S. Novel human astroviruses: Prevalence and association with common enteric viruses in undiagnosed gastroenteritis cases in Spain. Viruses 2019, 11, 585. [Google Scholar] [CrossRef]
- Wayman, S.D.; Fahle, G.; Palmore, T.; Green, K.Y.; Prevots, D.R. Norovirus, astrovirus, and sapovirus among immunocompromised patients at a tertiary care research hospital. Diagn. Microbiol. Infect. Dis. 2018, 92, 143–146. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.R.; Shah, D.; Breuer, J. Viral gastrointestinal infections and norovirus genotypes in a paediatric UK hospital, 2014–2015. J. Clin. Virol. 2016, 84, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Green, K.Y. Norovirus infection in immunocompromised hosts. Clin. Microbiol. Infect. 2014, 20, 717–723. [Google Scholar] [CrossRef] [PubMed]
- Beek, J.V.; Graaf, M.d.; Smits, S.; Schapendonk, C.M.E.; Verjans, G.; Vennema, H.; Koopmans, M. Whole-genome next-generation sequencing to study within-host evolution of norovirus (NoV) among immunocompromised patients with chronic NoV infection. J. Infect. Dis. 2017, 216, 1513–1524. [Google Scholar] [CrossRef] [PubMed]
- Munir, N.; Liu, P.; Gastanaduy, P.; Montes, J.; Shane, A.; Moe, C. Norovirus infection in immunocompromised children and children with hospital-acquired acute gastroenteritis. J. Med. Virol. 2014, 86, 1203–1209. [Google Scholar] [CrossRef]
- Chong, P.P.; Atmar, R.L. Norovirus in health care and implications for the immunocompromised host. Curr. Opin. Infect. Dis. 2019, 32, 348–355. [Google Scholar] [CrossRef]
- Liste, M.B.; Natera, I.; Suarez, J.A.; Pujol, F.H.; Liprandi, F.; Ludert, J.E. Enteric virus infections and diarrhea in healthy and human immunodeficiency virus-infected children. J. Clin. Microbiol. 2000, 38, 2873–2877. [Google Scholar] [CrossRef]
- Portes, S.A.R.; Costa, F.A.C.; Rocha, M.S.; Fumian, T.M.; Maranhao, A.G.; Assis, R.M.d.; Volotao, E.M. Enteric viruses in HIV-1 seropositive and HIV-1 seronegative children with diarrheal diseases in Brazil. PLoS ONE 2017, 12, e0183196. [Google Scholar] [CrossRef]
- Lewandowska, D.W.; Capaul, R.; Prader, S.; Zagordi, O.; Geissberger, F.D.; Kugler, M.; Huber, M. Persistent mammalian orthoreovirus, coxsackievirus and adenovirus co-infection in a child with a primary immunodeficiency detected by metagenomic sequencing: A case report. BMC Infect. Dis. 2018, 18, 33. [Google Scholar] [CrossRef] [PubMed]
- Santos, H.C.; Borges, F.P.; Arantes, A.M.; Souza, M. Detection of human adenovirus (species-C, -D and -F) in an allogeneic stem cell transplantation recipient: A case report. Rev. Bras. Hematol. Hemoter. 2017, 39, 60–62. [Google Scholar] [CrossRef][Green Version]
- Bodhidatta, L.; McDaniel, P.; Sornsakrin, S.; Srijan, A.; Serichantalergs, O.; Mason, C.J. Case-control study of diarrheal disease etiology in a remote rural area in Western Thailand. Am. J. Trop Med. Hyg. 2010, 83, 1106–1109. [Google Scholar] [CrossRef] [PubMed]
- O’Ryan, M.L.; Lucero, Y.; Prado, V.; Santolaya, M.E.; Rabello, M.; Solis, Y.; Mamani, N. Symptomatic and asymptomatic rotavirus and norovirus infections during infancy in a Chilean birth cohort. Pediatr. Infect. Dis. J. 2009, 28, 879–884. [Google Scholar] [CrossRef] [PubMed]
- Okitsu, S.; Khamrin, P.; Takanashi, S.; Thongprachum, A.; Hoque, S.A.; Takeuchi, H.; Ushijima, H. Molecular detection of enteric viruses in the stool samples of children without diarrhea in Bangladesh. Infect. Genet. Evol. 2020, 77, 104055. [Google Scholar] [CrossRef] [PubMed]
- Ayukekbong, J.; Lindh, M.; Nenonen, N.; Tah, F.; Akenji, T.N.; Bergstrom, T. Enteric viruses in healthy children in Cameroon: Viral load and genotyping of norovirus strains. J. Med. Virol. 2011, 83, 2135–2142. [Google Scholar] [CrossRef]
- Akihara, S.; Phan, T.G.; Nguyen, T.A.; Hansman, G.; Okitsu, S.; Ushijima, H. Existence of multiple outbreaks of viral gastroenteritis among infants in a day care center in Japan. Arch. Virol. 2005, 150, 2061–2075. [Google Scholar] [CrossRef]
- Nathanson, N.; Kew, O.M. From emergence to eradication: The epidemiology of poliomyelitis deconstructed. Am. J. Epidemiol. 2010, 172, 1213–1229. [Google Scholar] [CrossRef]
- Ramani, S.; Harris, V. Going viral! Unraveling the impact of nonpolio enteroviruses on oral vaccine responses. J. Infect. Dis. 2019, 219, 1173–1175. [Google Scholar] [CrossRef]
- Parker, E.P.; Kampmann, B.; Kang, G.; Grassly, N.C. Influence of enteric infections on response to oral poliovirus vaccine: A systematic review and meta-analysis. J. Infect. Dis. 2014, 210, 853–864. [Google Scholar] [CrossRef]
- Praharaj, I.; Parker, E.P.K.; Giri, S.; Allen, D.J.; Silas, S.; Revathi, R.; Kang, G. Influence of nonpolio enteroviruses and the bacterial gut microbiota on oral poliovirus vaccine response: A study from South India. J. Infect. Dis. 2019, 219, 1178–1186. [Google Scholar] [CrossRef] [PubMed]
- Taniuchi, M.; Mills, J.A.P.; Begum, S.; Uddin, M.J.; Sobuz, S.U.; Liu, J.; Houpt, E.R. Impact of enterovirus and other enteric pathogens on oral polio and rotavirus vaccine performance in Bangladeshi infants. Vaccine 2016, 34, 3068–3075. [Google Scholar] [CrossRef] [PubMed]
- Parker, E.P.K.; Praharaj, I.; Zekavati, A.; Lazarus, R.P.; Giri, S.; Operario, D.J.; Grassly, N.C. Influence of the intestinal microbiota on the immunogenicity of oral rotavirus vaccine given to infants in south India. Vaccine 2018, 36, 264–272. [Google Scholar] [CrossRef] [PubMed]
- Patel, M.; Steele, A.D.; Parashar, U.D. Influence of oral polio vaccines on performance of the monovalent and pentavalent rotavirus vaccines. Vaccine 2012, A30–A35. [Google Scholar] [CrossRef] [PubMed]
- Groome, M.J.; Page, N.; Cortese, M.M.; Moyes, J.; Zar, H.J.; Kapongo, C.N.; Madhi, S.A. Effectiveness of monovalent human rotavirus vaccine against admission to hospital for acute rotavirus diarrhoea in South African children: A case-control study. Lancet Infect. Dis. 2014, 14, 1096–1104. [Google Scholar] [CrossRef]
- Praharaj, I.; Mills, J.A.P.; Taneja, S.; Antony, K.; Yuhas, K.; Flores, J.; Kang, G. Diarrheal etiology and impact of coinfections on rotavirus vaccine efficacy estimates in a clinical trial of a monovalent human-bovine (116E) oral rotavirus vaccine, Rotavac, India. Clin. Infect. Dis. 2019, 69, 243–250. [Google Scholar] [CrossRef]
- Brouwer, L.; Benschop, K.S.M.; Nguyen, D.; Kamau, E.; Pajkrt, D.; Simmonds, P.; Wolthers, K.C. Recombination analysis of non-poliovirus members of the enterovirus C species; restriction of recombination events to members of the same 3DPol cluster. Viruses 2020, 12, 706. [Google Scholar] [CrossRef]
- Muslin, C.; Kain, A.M.; Bessaud, M.; Blondel, B.; Delpeyroux, F. Recombination in enteroviruses, a multi-step modular evolutionary process. Viruses 2019, 11, 859. [Google Scholar] [CrossRef]
- Bessaud, M.; Joffret, M.L.; Blondel, B.; Delpeyroux, F. Exchanges of genomic domains between poliovirus and other cocirculating species C enteroviruses reveal a high degree of plasticity. Sci. Rep. 2016, 6, 38831. [Google Scholar] [CrossRef]
- Wohlgemuth, N.; Honce, R.; Cherry, S.S. Astrovirus evolution and emergence. Infect. Genet. Evol. 2019, 69, 30–37. [Google Scholar] [CrossRef]
- Begall, L.F.L.; Mauroy, A.; Thiry, E. Norovirus recombinants: Recurrent in the field, recalcitrant in the lab-a scoping review of recombination and recombinant types of noroviruses. J. Gen. Virol. 2018, 99, 970–988. [Google Scholar] [CrossRef] [PubMed]
- McDonald, S.M.; Nelson, M.I.; Turner, P.E.; Patton, J.T. Reassortment in segmented RNA viruses: Mechanisms and outcomes. Nat. Rev. Microbiol. 2016, 14, 448–460. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Liu, W.J.; Xu, W.; Jin, T.; Zhao, Y.; Song, J.; Gao, G.F. A bat-derived putative cross-family recombinant coronavirus with a reovirus gene. PLoS Pathog. 2016, 12, e1005883. [Google Scholar] [CrossRef] [PubMed]
- Santiana, M.; Ghosh, S.; Ho, B.A.; Rajasekaran, V.; Du, W.L.; Mutsafi, Y.; Bonnet, N.A. Vesicle-cloaked virus clusters are optimal units for inter-organismal viral transmission. Cell Host Microbe 2018, 24, 208–220. [Google Scholar] [CrossRef]
- Aguilera, E.R.; Erickson, A.K.; Jesudhasan, P.R.; Robinson, C.M.; Pfeiffer, J.K. Plaques formed by mutagenized viral populations have elevated co-infection frequencies. mBio 2017, 8. [Google Scholar] [CrossRef]
- Aguilera, E.R.; Pfeiffer, J.K. Strength in numbers: Mechanisms of viral co-infection. Virus Res. 2019, 265, 43–46. [Google Scholar] [CrossRef]
- Moshe, A.; Gorovits, R. Virus-induced aggregates in infected cells. Viruses 2012, 4, 2218–2232. [Google Scholar] [CrossRef]
- Moreno, I.A.; Sanjuan, R. Collective infection of cells by viral aggregates promotes early viral proliferation and reveals a cellular-level allee effect. Curr. Biol. 2018, 28, 3212–3219.e3214. [Google Scholar] [CrossRef]
- Erickson, A.K.; Jesudhasan, P.R.; Mayer, M.J.; Narbad, A.; Winter, S.E.; Pfeiffer, J.K. Bacteria facilitate enteric virus co-infection of mammalian cells and promote genetic recombination. Cell Host Microbe 2018, 23, 77–88. [Google Scholar] [CrossRef]
- Dianzani, F. Viral interference and interferon. Ric. Clin. Lab. 1975, 5, 196–213. [Google Scholar] [CrossRef]
- Ingle, H.; Lee, S.; Ai, T.; Orvedahl, A.; Rodgers, R.; Zhao, G.; Baldridge, M.T. Viral complementation of immunodeficiency confers protection against enteric pathogens via interferon-lambda. Nat. Microbiol. 2019, 4, 1120–1128. [Google Scholar] [CrossRef] [PubMed]
- Sestak, K. Non-human primate models of enteric viral infections. Viruses 2018, 10, 544. [Google Scholar] [CrossRef] [PubMed]
- Chege, G.K.; Steele, A.D.; Hart, C.A.; Snodgrass, D.R.; Omolo, E.O.; Mwenda, J.M. Experimental infection of non-human primates with a human rotavirus isolate. Vaccine 2005, 23, 1522–1528. [Google Scholar] [CrossRef] [PubMed]
- Bentes, G.A.; Guimaraes, J.R.; Volotao, E.M.; Fialho, A.M.; Hooper, C.; Ganime, A.C.; Pinto, M.A. Cynomolgus monkeys (Macaca fascicularis) as an experimental infection model for human group a rotavirus. Viruses 2018, 10, 355. [Google Scholar] [CrossRef]
- Rockx, B.H.; Bogers, W.M.; Heeney, J.L.; Amerongen, G.V.; Koopmans, M.P. Experimental norovirus infections in non-human primates. J. Med. Virol. 2005, 75, 313–320. [Google Scholar] [CrossRef] [PubMed]
- Farkas, T.; Yang, K.; Pendu, J.L.; Baines, J.D.; Cardin, R.D. The coxsackievirus and adenovirus receptor, a required host factor for recovirus infection, is a putative enteric calicivirus receptor. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Sestak, K.; McNeal, M.M.; Choi, A.; Cole, M.J.; Ramesh, G.; Alvarez, X.; Ward, R.L. Defining T-cell-mediated immune responses in rotavirus-infected juvenile rhesus macaques. J. Virol. 2004, 78, 10258–10264. [Google Scholar] [CrossRef]
- McNeal, M.M.; Sestak, K.; Choi, A.H.; Basu, M.; Cole, M.J.; Aye, P.P.; Ward, R.L. Development of a rotavirus-shedding model in rhesus macaques, using a homologous wild-type rotavirus of a new P genotype. J. Virol. 2005, 79, 944–954. [Google Scholar] [CrossRef]
- Hoshino, Y.; Honma, S.; Jones, R.W.; Santos, N.; Nakagomi, O.; Nakagomi, T.; Thouless, M.E. A rotavirus strain isolated from pig-tailed macaque (Macaca nemestrina) with diarrhea bears a P6[1]: G8 specificity. Virology 2006, 345, 1–12. [Google Scholar] [CrossRef]
- D’Arc, M.; Furtado, C.; Siqueira, J.D.; Seuanez, H.N.; Ayouba, A.; Peeters, M.; Soares, M.A. Assessment of the gorilla gut virome in association with natural simian immunodeficiency virus infection. Retrovirology 2018, 15, 19. [Google Scholar] [CrossRef]
- Handley, S.A.; Thackray, L.B.; Zhao, G.; Presti, R.; Miller, A.D.; Droit, L.; Virgin, H.W. Pathogenic simian immunodeficiency virus infection is associated with expansion of the enteric virome. Cell 2012, 151, 253–266. [Google Scholar] [CrossRef] [PubMed]
- Handley, S.A.; Desai, C.; Zhao, G.; Droit, L.; Monaco, C.L.; Schroeder, A.C.; Virgin, H.W. SIV infection-mediated changes in gastrointestinal bacterial microbiome and virome are associated with immunodeficiency and prevented by vaccination. Cell Host Microbe 2016, 19, 323–335. [Google Scholar] [CrossRef] [PubMed]
- Saif, L.J.; Ward, L.A.; Yuan, L.; Rosen, B.I.; To, T.L. The gnotobiotic piglet as a model for studies of disease pathogenesis and immunity to human rotaviruses. Arch. Virol. Suppl. 1996, 12, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Ramesh, A.; Mao, J.; Lei, S.; Twitchell, E.; Shiraz, A.; Jiang, X.; Yuan, A.L. Parenterally administered P24-VP8* nanoparticle vaccine conferred strong protection against rotavirus diarrhea and virus shedding in gnotobiotic pigs. Vaccines 2019, 7, 177. [Google Scholar] [CrossRef]
- Yuan, L.; Ward, L.A.; Rosen, B.I.; To, T.L.; Saif, L.J. Systematic and intestinal antibody-secreting cell responses and correlates of protective immunity to human rotavirus in a gnotobiotic pig model of disease. J. Virol. 1996, 70, 3075–3083. [Google Scholar] [CrossRef]
- Troeger, H.; Loddenkemper, C.; Schneider, T.; Schreier, E.; Epple, H.J.; Zeitz, M.; Schulzke, J.D. Structural and functional changes of the duodenum in human norovirus infection. Gut 2009, 58, 1070–1077. [Google Scholar] [CrossRef]
- Sugieda, M.; Nakajima, S. Viruses detected in the caecum contents of healthy pigs representing a new genetic cluster in genogroup II of the genus “Norwalk-like viruses”. Virus Res. 2002, 87, 165–172. [Google Scholar] [CrossRef]
- Poel, W.H.v.D.; Vinje, J.; Heide, R.v.D.; Herrera, M.I.; Vivo, A.; Koopmans, M.P. Norwalk-like calicivirus genes in farm animals. Emerg. Infect. Dis. 2000, 6, 36–41. [Google Scholar] [CrossRef]
- Wang, Q.H.; Han, M.G.; Cheetham, S.; Souza, M.; Funk, J.A.; Saif, L.J. Porcine noroviruses related to human noroviruses. Emerg. Infect. Dis. 2005, 11, 1874–1881. [Google Scholar] [CrossRef]
- Koonpaew, S.; Teeravechyan, S.; Frantz, P.N.; Chailangkarn, T.; Jongkaewwattana, A. PEDV and PDCoV pathogenesis: The interplay between host innate immune responses and porcine enteric coronaviruses. Front. Vet. Sci. 2019, 6, 34. [Google Scholar] [CrossRef]
- Zhao, Z.P.; Yang, Z.; Lin, W.D.; Wang, W.Y.; Yang, J.; Jin, W.J.; Qin, A.J. The rate of co-infection for piglet diarrhea viruses in China and the genetic characterization of porcine epidemic diarrhea virus and porcine kobuvirus. Acta Virol. 2016, 60, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Williams, S.H.; Che, X.; Garcia, J.A.; Klena, J.D.; Lee, B.; Muller, D.; Lipkin, W.I. Viral diversity of house mice in New York City. mBio 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Cortez, V.; Sharp, B.; Yao, J.; Livingston, B.; Vogel, P.; Cherry, S.S. Characterizing a murine model for astrovirus using viral isolates from persistently infected immunocompromised mice. J. Virol. 2019, 93. [Google Scholar] [CrossRef] [PubMed]
- Taube, S.; Kolawole, A.O.; Hohne, M.; Wilkinson, J.E.; Handley, S.A.; Perry, J.W.; Wobus, C.E. A mouse model for human norovirus. mBio 2013, 4. [Google Scholar] [CrossRef]
- Karst, S.M.; Wobus, C.E.; Lay, M.; Davidson, J.; Virgin, H.W. STAT1-dependent innate immunity to a Norwalk-like virus. Science 2003, 299, 1575–1578. [Google Scholar] [CrossRef]
- Baldridge, M.T.; Nice, T.J.; McCune, B.T.; Yokoyama, C.C.; Kambal, A.; Wheadon, M.; Virgin, H.W. Commensal microbes and interferon-lambda determine persistence of enteric murine norovirus infection. Science 2015, 347, 266–269. [Google Scholar] [CrossRef]
- Orchard, R.C.; Wilen, C.B.; Doench, J.G.; Baldridge, M.T.; McCune, B.T.; Lee, Y.C.; Virgin, H.W. Discovery of a proteinaceous cellular receptor for a norovirus. Science 2016, 353, 933–936. [Google Scholar] [CrossRef]
- Nice, T.J.; Baldridge, M.T.; McCune, B.T.; Norman, J.M.; Lazear, H.M.; Artyomov, M.; Virgin, H.W. Interferon-lambda cures persistent murine norovirus infection in the absence of adaptive immunity. Science 2015, 347, 269–273. [Google Scholar] [CrossRef]
- Wilen, C.B.; Lee, S.; Hsieh, L.L.; Orchard, R.C.; Desai, C.; Hykes, B.L.J.; Virgin, H.W. Tropism for tuft cells determines immune promotion of norovirus pathogenesis. Science 2018, 360, 204–208. [Google Scholar] [CrossRef]
- Roth, A.N.; Helm, E.W.; Mirabelli, C.; Kirsche, E.; Smith, J.C.; Eurell, L.B.; Karst, S.M. Norovirus infection causes acute self-resolving diarrhea in wild-type neonatal mice. Nat. Commun. 2020, 11, 2968. [Google Scholar] [CrossRef]
- Broome, R.L.; Vo, P.T.; Ward, R.L.; Clark, H.F.; Greenberg, H.B. Murine rotavirus genes encoding outer capsid proteins VP4 and VP7 are not major determinants of host range restriction and virulence. J. Virol. 1993, 67, 2448–2455. [Google Scholar] [CrossRef] [PubMed]
- Knipping, K.; McNeal, M.M.; Crienen, A.; Amerongen, G.V.; Garssen, J.; Land, B.V. A gastrointestinal rotavirus infection mouse model for immune modulation studies. Virol. J. 2011, 8, 109. [Google Scholar] [CrossRef] [PubMed]
- Cortez, V.; Boyd, D.F.; Crawford, J.C.; Sharp, B.; Livingston, B.; Rowe, H.M.; Cherry, S.S. Astrovirus infects actively secreting goblet cells and alters the gut mucus barrier. Nat. Commun. 2020, 11, 2097. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, C.C.; Loh, J.; Zhao, G.; Stappenbeck, T.S.; Wang, D.; Huang, H.V.; Thackray, L.B. Adaptive immunity restricts replication of novel murine astroviruses. J. Virol. 2012, 86, 12262–12270. [Google Scholar] [CrossRef]
- Lee, S.; Wilen, C.B.; Orvedahl, A.; McCune, B.T.; Kim, K.W.; Orchard, R.C.; Virgin, H.W. Norovirus cell tropism is determined by combinatorial action of a viral non-structural protein and host cytokine. Cell Host Microbe 2017, 22, 449–459. [Google Scholar] [CrossRef]
- Nice, T.J.; Strong, D.W.; McCune, B.T.; Pohl, C.S.; Virgin, H.W. A single-amino-acid change in murine norovirus NS1/2 is sufficient for colonic tropism and persistence. J. Virol. 2013, 87, 327–334. [Google Scholar] [CrossRef]
- Zhu, S.; Regev, D.; Watanabe, M.; Hickman, D.; Moussatche, N.; Jesus, D.M.; Karst, S.M. Identification of immune and viral correlates of norovirus protective immunity through comparative study of intra-cluster norovirus strains. PLoS Pathog. 2013, 9, e1003592. [Google Scholar] [CrossRef]
- Tomov, V.T.; Palko, O.; Lau, C.W.; Pattekar, A.; Sun, Y.; Tacheva, R.; Wherry, E.J. Differentiation and protective capacity of virus-specific CD8(+) T cells suggest murine norovirus persistence in an immune-privileged enteric niche. Immunity 2017, 47, 723–738. [Google Scholar] [CrossRef]
- Kounatidis, I.; Ligoxygakis, P. Drosophila as a model system to unravel the layers of innate immunity to infection. Open Biol. 2012, 2, 120075. [Google Scholar] [CrossRef]
- Lemaitre, B.; Hoffmann, J. The host defense of drosophila melanogaster. Annu. Rev. Immunol. 2007, 25, 697–743. [Google Scholar] [CrossRef]
- Segrist, E.; Cherry, S. Using diverse model systems to define intestinal epithelial defenses to enteric viral infections. Cell Host Microbe 2020, 27, 329–344. [Google Scholar] [CrossRef] [PubMed]
- Sansone, C.L.; Cohen, J.; Yasunaga, A.; Xu, J.; Osborn, G.; Subramanian, H.; Cherry, S. Microbiota-dependent priming of antiviral intestinal immunity in drosophila. Cell Host Microbe 2015, 18, 571–581. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Hopkins, K.; Sabin, L.; Yasunaga, A.; Subramanian, H.; Lamborn, I.; Cherry, S. ERK signaling couples nutrient status to antiviral defense in the insect gut. Proc. Natl. Acad. Sci. USA 2013, 110, 15025–15030. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Felix, M.A.; Wang, D. Natural viruses of caenorhabditis nematodes. Annu. Rev. Genet. 2019, 53, 313–326. [Google Scholar] [CrossRef]
- Martin, A.; Rex, E.A.; Ishidate, T.; Lin, R.; Gammon, D.B. Infection of Caenorhabditis elegans with vesicular stomatitis virus via microinjection. Bio. Protoc. 2017, 7. [Google Scholar] [CrossRef]
- Hewes, S.A.; Wilson, R.L.; Estes, M.K.; Shroyer, N.F.; Blutt, S.E.; Allen, K.J.G. In vitro models of the small intestine: Engineering challenges and engineering solutions. Tissue Eng. Part B Rev. 2020. [Google Scholar] [CrossRef]
- Pearce, S.C.; Coia, H.G.; Karl, J.P.; Feliciano, I.G.P.; Zachos, N.C.; Racicot, K. Intestinal in vitro and ex vivo models to study host-microbiome interactions and acute stressors. Front. Physiol. 2018, 9, 1584. [Google Scholar] [CrossRef]
- Le, M.L.V.; Servin, A.L. Pathogenesis of human enterovirulent bacteria: Lessons from cultured, fully differentiated human colon cancer cell lines. Microbiol. Mol. Biol. Rev. 2013, 77, 380–439. [Google Scholar] [CrossRef]
- Oka, T.; Stoltzfus, G.T.; Zhu, C.; Jung, K.; Wang, Q.; Saif, L.J. Attempts to grow human noroviruses, a sapovirus, and a bovine norovirus in vitro. PLoS ONE 2018, 13, e0178157. [Google Scholar] [CrossRef]
- Straub, T.M.; Bartholomew, R.A.; Valdez, C.O.; Valentine, N.B.; Dohnalkova, A.; Ozanich, R.M.; Call, D.R. Human norovirus infection of caco-2 cells grown as a three-dimensional tissue structure. J. Water Health 2011, 9, 225–240. [Google Scholar] [CrossRef] [PubMed]
- Svensson, L.; Finlay, B.B.; Bass, D.; Bonsdorff, C.H.V.; Greenberg, H.B. Symmetric infection of rotavirus on polarized human intestinal epithelial (Caco-2) cells. J. Virol. 1991, 65, 4190–4197. [Google Scholar] [CrossRef] [PubMed]
- Chaibi, C.; Laffitte, J.C.; Sandre, C.; Esclatine, A.; Servin, A.L.; Quero, A.M.; Legendre, M.G. Rotavirus induces apoptosis in fully differentiated human intestinal Caco-2 cells. Virology 2005, 332, 480–490. [Google Scholar] [CrossRef] [PubMed]
- Vu, D.L.; Bosch, A.; Pinto, R.M.; Ribes, E.; Guix, S. Human astrovirus MLB replication in vitro: Persistence in extraintestinal cell lines. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Meliopoulos, V.A.; Marvin, S.A.; Freiden, P.; Moser, L.A.; Nighot, P.; Ali, R.; Cherry, S.S. Oral administration of astrovirus capsid protein is sufficient to induce acute diarrhea in vivo. mBio 2016, 7. [Google Scholar] [CrossRef]
- Wang, H.; Moon, S.; Wang, Y.; Jiang, B. Multiple virus infection alters rotavirus replication and expression of cytokines and Toll-like receptors in intestinal epithelial cells. Virus Res. 2012, 167, 48–55. [Google Scholar] [CrossRef]
- Altamimi, M.; Abdelhay, O.; Rastall, R.A. Effect of oligosaccharides on the adhesion of gut bacteria to human HT-29 cells. Anaerobe 2016, 39, 136–142. [Google Scholar] [CrossRef]
- Raja, S.B.; Murali, M.R.; Devaraj, H.; Devaraj, S.N. Differential expression of gastric MUC5AC in colonic epithelial cells: TFF3-wired IL1 beta/Akt crosstalk-induced mucosal immune response against shigella dysenteriae infection. J. Cell Sci. 2012, 125, 703–713. [Google Scholar] [CrossRef]
- Rajan, A.; Persson, B.D.; Frangsmyr, L.; Olofsson, A.; Sandblad, L.; Heino, J.; Arnberg, N. Enteric species F human adenoviruses use laminin-binding integrins as co-receptors for infection of Ht-29 Cells. Sci. Rep. 2018, 8, 10019. [Google Scholar] [CrossRef]
- Chattopadhyay, S.; Basak, T.; Nayak, M.K.; Bhardwaj, G.; Mukherjee, A.; Bhowmick, R.; Sarkar, M.C. Identification of cellular calcium binding protein calmodulin as a regulator of rotavirus A infection during comparative proteomic study. PLoS ONE 2013, 8, e56655. [Google Scholar] [CrossRef]
- Chanda, S.; Nandi, S.; Sarkar, M.C. Rotavirus-induced miR-142-5p elicits proviral milieu by targeting non-canonical transforming growth factor beta signalling and apoptosis in cells. Cell Microbiol. 2016, 18, 733–747. [Google Scholar] [CrossRef]
- Sato, T.; Vries, R.G.; Snippert, H.J.; Wetering, M.V.d.; Barker, N.; Stange, D.E.; Clevers, H. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262–265. [Google Scholar] [CrossRef]
- Sato, T.; Stange, D.E.; Ferrante, M.; Vries, R.G.; Van, J.H.; Brink, S.V.D.; Clevers, H. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 2011, 141, 1762–1772. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Clevers, H. Growing self-organizing mini-guts from a single intestinal stem cell: Mechanism and applications. Science 2013, 340, 1190–1194. [Google Scholar] [CrossRef] [PubMed]
- Zachos, N.C.; Kovbasnjuk, O.; Abel, J.F.; In, J.; Blutt, S.E.; Jonge, H.R.d.; Donowitz, M. Human enteroids/colonoids and intestinal organoids functionally recapitulate normal intestinal physiology and pathophysiology. J. Biol. Chem. 2016, 291, 3759–3766. [Google Scholar] [CrossRef] [PubMed]
- Saxena, K.; Blutt, S.E.; Ettayebi, K.; Zeng, X.L.; Broughman, J.R.; Crawford, S.E.; Estes, M.K. Human intestinal enteroids: A new model to study human rotavirus infection, host restriction, and pathophysiology. J. Virol. 2015, 90, 43–56. [Google Scholar] [CrossRef] [PubMed]
- Saxena, K.; Simon, L.M.; Zeng, X.L.; Blutt, S.E.; Crawford, S.E.; Sastri, N.P.; Estes, M.K. A paradox of transcriptional and functional innate interferon responses of human intestinal enteroids to enteric virus infection. Proc. Natl. Acad. Sci. USA 2017, 114, E570–E579. [Google Scholar] [CrossRef]
- Ettayebi, K.; Crawford, S.E.; Murakami, K.; Broughman, J.R.; Karandikar, U.; Tenge, V.R.; Estes, M.K. Replication of human noroviruses in stem cell-derived human enteroids. Science 2016, 353, 1387–1393. [Google Scholar] [CrossRef]
- Drummond, C.G.; Bolock, A.M.; Ma, C.; Luke, C.J.; Good, M.; Coyne, C.B. Enteroviruses infect human enteroids and induce antiviral signaling in a cell lineage-specific manner. Proc. Natl. Acad. Sci. USA 2017, 114, 1672–1677. [Google Scholar] [CrossRef]
- Good, C.; Wells, A.I.; Coyne, C.B. Type III interferon signaling restricts enterovirus 71 infection of goblet cells. Sci. Adv. 2019, 5, 4255. [Google Scholar] [CrossRef]
| Classification | Genome | Major Agents | Disease | Prevalence | Reference |
|---|---|---|---|---|---|
| Class I Adenoviridae | dsDNA 30–40 kb | Adenovirus (AdV) serotypes 40, 41 (species F) | Gastroenteritis | 15% in children and 1.5 to 5.4% in adults | [5] |
| Class III Reoviridae | dsRNA 16–27 kb | Rotavirus (RV) | Gastroenteritis | In children <5: pre-vaccine 40% post-vaccine 19% | [5,6] |
| Class IV Astroviridae | ssRNA (+) 6.8–7.9 kb | Astrovirus (AstV) | Nosocomial or epidemic diarrhea | ~5% in children with AGE, 3–9% in hospitalized patients with diarrhea | [5,7] |
| Class IV Caliciviridae | ssRNA (+) 7.4–8.3 kb | Norovirus (NoV), Sapovirus (SaV) | Epidemic diarrhea | 18% in all AGE cases, as well as 24% in community | [5,8,9] |
| Class IV Picornaviridae | ssRNA (+) 7.1–8.9 kb | Enterovirus species A and B (Enteroviruses and Coxsackieviruses), species C (Polioviruses) | Gastroenteritis, Hand, foot, and mouth disease (HFMD), Poliomyelitis | ~5% in children with AGE; ~4% in children with HFMD | [10] |
| Class IV Coronaviridae | ssRNA (+) 27–30 kb | Severe acute respiratory syndrome (SARS-CoV) and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) | Acute diarrhea, severe acute respiratory syndrome, and coronavirus disease 2019 (COVID-19) | Ongoing global pandemic—to be determined | [11,12,13] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Makimaa, H.; Ingle, H.; Baldridge, M.T. Enteric Viral Co-Infections: Pathogenesis and Perspective. Viruses 2020, 12, 904. https://doi.org/10.3390/v12080904
Makimaa H, Ingle H, Baldridge MT. Enteric Viral Co-Infections: Pathogenesis and Perspective. Viruses. 2020; 12(8):904. https://doi.org/10.3390/v12080904
Chicago/Turabian StyleMakimaa, Heyde, Harshad Ingle, and Megan T. Baldridge. 2020. "Enteric Viral Co-Infections: Pathogenesis and Perspective" Viruses 12, no. 8: 904. https://doi.org/10.3390/v12080904
APA StyleMakimaa, H., Ingle, H., & Baldridge, M. T. (2020). Enteric Viral Co-Infections: Pathogenesis and Perspective. Viruses, 12(8), 904. https://doi.org/10.3390/v12080904






