Heat Inactivation of Different Types of SARS-CoV-2 Samples: What Protocols for Biosafety, Molecular Detection and Serological Diagnostics?
Abstract
1. Introduction
2. Materials and Methods
2.1. Virus Strain and Titration
2.2. Samples Used for Heat Inactivation
2.3. Heat Inactivation of SARS-CoV-2 Samples
2.4. Integrity of SARS-CoV-2 RNA before and after Heat Inactivation
2.5. Impact of 56 °C-30 min Heating on Results of Serological Assays
2.5.1. Detection of SARS-CoV-2 IgG by ELISA
2.5.2. Detection of SARS-CoV-2 Neutralizing Antibodies
3. Results
3.1. Heat Inactivation of SARS-CoV-2 Samples
3.1.1. Heat Inactivation of SARS-CoV-2 Infected Cell Supernatant
3.1.2. Heat Inactivation of SARS-CoV-2 Spiked Nasopharyngeal Samples
3.1.3. Heat Inactivation of SARS-CoV-2 Spiked Blood Donor Sera
3.2. Impact of 56 °C-30 min Protocol on the Results of Serological Assays
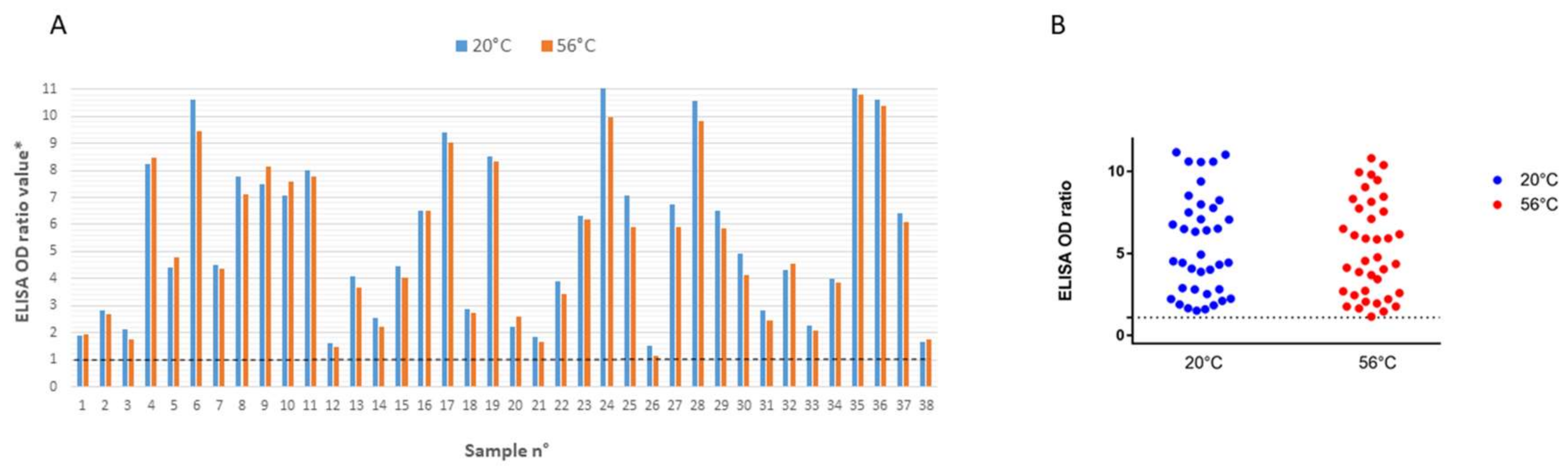
3.2.1. ELISA
3.2.2. Virus Neutralization Test
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Otter, J.A.; Donskey, C.; Yezli, S.; Douthwaite, S.; Goldenberg, S.D.; Weber, D.J. Transmission of SARS and MERS coronaviruses and influenza virus in healthcare settings: The possible role of dry surface contamination. J. Hosp. Infect. 2016, 92, 235–250. [Google Scholar] [CrossRef] [PubMed]
- Xiao, S.; Li, Y.; Wong, T.; Hui, D.S.C. Role of fomites in SARS transmission during the largest hospital outbreak in Hong Kong. PLoS ONE 2017, 12, e0181558. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.W.X.; Tan, Y.K.; Chia, P.Y.; Lee, T.H.; Ng, O.T.; Wong, M.S.Y.; Marimuthu, K. Air, Surface Environmental, and Personal Protective Equipment Contamination by Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) From a Symptomatic Patient. JAMA 2020. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Laboratory Biosafety Guidance Related to Coronavirus Disease 2019 ( COVID-19): Interim Guidance; WHO: Geneva, Switzerland, 12 February; Available online: https://apps.who.int/iris/handle/10665/331138 (accessed on 12 February 2020).
- Sagripanti, J.-L.; Hülseweh, B.; Grote, G.; Voß, L.; Böhling, K.; Marschall, H.-J. Microbial Inactivation for Safe and Rapid Diagnostics of Infectious Samples. Appl. Environ. Microbiol. 2011, 77, 7289–7295. [Google Scholar] [CrossRef] [PubMed]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I.; et al. Aerosol and Surface Stability of SARS-CoV-2 as Compared with SARS-CoV-1. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Reed, L.J.; Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Huang, Y.-J.S.; Hsu, W.-W.; Higgs, S.; Vanlandingham, D.L. Temperature Tolerance and Inactivation of Chikungunya Virus. Vector Borne Zoonotic Dis. 2015, 15, 674–677. [Google Scholar] [CrossRef] [PubMed]
- Nurtop, E.; Villarroel, P.M.S.; Pastorino, B.; Ninove, L.; Drexler, J.F.; Roca, Y.; Gake, B.; Dubot-Peres, A.; Grard, G.; Peyrefitte, C.; et al. Combination of ELISA screening and seroneutralisation tests to expedite Zika virus seroprevalence studies. Virol. J. 2018, 15, 192. [Google Scholar] [CrossRef] [PubMed]
- Isnard, C.; Pastorino, B. Absence of Effect of 56°C-30 min Treatment on SARS-CoV-2 ELISA Using Pre-Pandemic Serum Samples; Unité des Virus Émergents (UVE: Aix-Marseille Univ-IRD 190-Inserm 1207): Marseille, France, 2020. [Google Scholar]
- Saknimit, M.; Inatsuki, I.; Sugiyama, Y.; Yagami, K. Virucidal efficacy of physico-chemical treatments against coronaviruses and parvoviruses of laboratory animals. Jikken Dobutsu 1988, 37, 341–345. [Google Scholar] [PubMed]
- Cartwright, S.F.; Harris, H.M.; Blandford, T.B.; Fincham, I.; Gitter, M. A cytopathic virus causing a transmissible gastroenteritis in swine. I. Isolation and properties. J. Comp. Pathol. 1965, 75, 387–396. [Google Scholar] [CrossRef] [PubMed]
- Pastorino, B.; Touret, F.; Gilles, M.; Luciani, L.; de Lamballerie, X.; Charrel, R.N. Evaluation of Chemical Protocols for Inactivating SARS-CoV-2 Infectious Samples. Viruses 2020, 12, 624. [Google Scholar] [CrossRef] [PubMed]
- Batéjat, C.; Grassin, Q.; Manuguerra, J.-C.; Leclercq, I. Heat inactivation of the Severe Acute Respiratory Syndrome Coronavirus 2. bioRxiv 2020. [Google Scholar] [CrossRef]
- Pastorino, B.; Bessaud, M.; Grandadam, M.; Murri, S.; Tolou, H.J.; Peyrefitte, C.N. Development of a TaqMan® RT-PCR assay without RNA extraction step for the detection and quantification of African Chikungunya viruses. J. Virol. Methods 2005, 124, 65–71. [Google Scholar] [CrossRef] [PubMed]
- Bruce, E.A.; Huang, M.L.; Perchetti, G.A.; Tighe, S.; Laaguiby, P.; Hoffman, J.J.; Gerrard, D.L.; Nalla, A.K.; Wei, Y.; Greninger, A.L.; et al. Direct RT-qPCR detection of SARS-CoV-2 RNA from patient nasopharyngeal swabs without an RNA extraction step. bioRxiv 2020. [Google Scholar] [CrossRef]
- Pan, Y.; Long, L.; Zhang, D.; Yan, T.; Cui, S.; Yang, P.; Wang, Q.; Ren, S. Potential false-negative nucleic acid testing results for Severe Acute Respiratory Syndrome Coronavirus 2 from thermal inactivation of samples with low viral loads. Clin. Chem. 2020. [Google Scholar] [CrossRef] [PubMed]
- General Procedures for Inactivation of Potentially Infectious Samples with Ebola Virus and Other Highly Pathogenic Viral Agents. 2014. Available online: https://www.paho.org/hq/dmdocuments/2014/2014-cha-procedures-inactivation-ebola.pdf (accessed on 26 June 2020).
- Hu, X.; An, T.; Situ, B.; Hu, Y.; Ou, Z.; Li, Q.; He, X.; Zhang, Y.; Tian, P.; Sun, D.; et al. Heat inactivation of serum interferes with the immunoanalysis of antibodies to SARS-CoV-2. medRxiv 2020. [Google Scholar] [CrossRef]
- Meyer, D.; Petrov, A.; Becher, P. Inactivation of Classical Swine Fever Virus in Porcine Serum Samples Intended for Antibody Detection. Pathogens 2019, 8, 286. [Google Scholar] [CrossRef] [PubMed]
- Kenyon, R.H.; Peters, C.J. Actions of complement on Junin virus. Rev. Infect. Dis. 1989, 11 (Suppl. 4), S771–S776. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Freed, D.C.; Tang, A.; Rustandi, R.R.; Troutman, M.C.; Espeseth, A.S.; Zhang, N.; An, Z.; McVoy, M.; Zhu, H.; et al. Complement enhances in vitro neutralizing potency of antibodies to human cytomegalovirus glycoprotein B (gB) and immune sera induced by gB/MF59 vaccination. NPJ Vaccines 2017, 2, 36. [Google Scholar] [CrossRef] [PubMed]

| Type of Sample | Heating Protocol | Viral Titer (TCID50/mL) a | Log10 Reduction Factor (LRF) | Number of RNA Copies | Ct Var b | |||
|---|---|---|---|---|---|---|---|---|
| Before Heat Inactivation | After Heat Inactivation | Before Heat Inactivation | After Heat Inactivation | |||||
| No BSA | 3g/L BSA | |||||||
| SARS-CoV-2 infected cell supernatant c | 56 °C, 30 min | 3.3 ± 2.3 × 106 | 8.5 ± 7 | ND (0/2) e | 5 < LRF < 6 | 8.01 × 106 | 5.16 × 106 | <0.7 |
| 60 °C, 60 min | 3.3 ± 2.3 × 106 | ND (0/2) | 5 ± 2.8 | 5 < LRF < 6 | 8.01 × 106 | 4.54 × 106 | <0.8 | |
| 92 °C, 15 min | 3.3 ± 2.3 × 106 | ND (0/2) | ND (0/2) | LRF > 6 | 8.01 × 106 | 1.6 × 105 | >5 | |
| SARS-CoV-2 spiked nasopharyngeal sampled | 56 °C, 30 min | 3.5 ± 2.3 × 105 | ND (0/6) | LRF > 5 | 7.5 × 105 | 2.1 × 105 | <1.5 | |
| 60 °C, 60 min | 3.5 ± 2.3 × 105 | ND (0/6) | LRF > 5 | 7.5 × 105 | 1.5 × 105 | <2 | ||
| SARS-CoV-2 spiked blood donor sera d | 56 °C, 30 min | 3.5 ± 2.3 × 105 | ND (0/6) | LRF > 5 | 7.5 × 105 | 3.5 × 105 | <1 | |
| 60 °C, 60 min | 3.5 ± 2.3 × 105 | ND (0/6) | LRF > 5 | 7.5 × 105 | 1.5 × 105 | <2 | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pastorino, B.; Touret, F.; Gilles, M.; de Lamballerie, X.; Charrel, R.N. Heat Inactivation of Different Types of SARS-CoV-2 Samples: What Protocols for Biosafety, Molecular Detection and Serological Diagnostics? Viruses 2020, 12, 735. https://doi.org/10.3390/v12070735
Pastorino B, Touret F, Gilles M, de Lamballerie X, Charrel RN. Heat Inactivation of Different Types of SARS-CoV-2 Samples: What Protocols for Biosafety, Molecular Detection and Serological Diagnostics? Viruses. 2020; 12(7):735. https://doi.org/10.3390/v12070735
Chicago/Turabian StylePastorino, Boris, Franck Touret, Magali Gilles, Xavier de Lamballerie, and Remi N. Charrel. 2020. "Heat Inactivation of Different Types of SARS-CoV-2 Samples: What Protocols for Biosafety, Molecular Detection and Serological Diagnostics?" Viruses 12, no. 7: 735. https://doi.org/10.3390/v12070735
APA StylePastorino, B., Touret, F., Gilles, M., de Lamballerie, X., & Charrel, R. N. (2020). Heat Inactivation of Different Types of SARS-CoV-2 Samples: What Protocols for Biosafety, Molecular Detection and Serological Diagnostics? Viruses, 12(7), 735. https://doi.org/10.3390/v12070735







