Ross River Virus Provokes Differentially Expressed MicroRNA and RNA Interference Responses in Aedes aegypti Mosquitoes
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Mosquito Infections with Ross River Virus
2.3. Mosquito Dissection and RNA Extraction
2.4. Library Preparations and Sequencing
2.5. Small RNA Analysis
2.6. RNAi Activity Analysis
2.7. The miRNA Target Identification
2.8. Gene Ontology
2.9. RT-qPCR Analysis
3. Results and Discussion
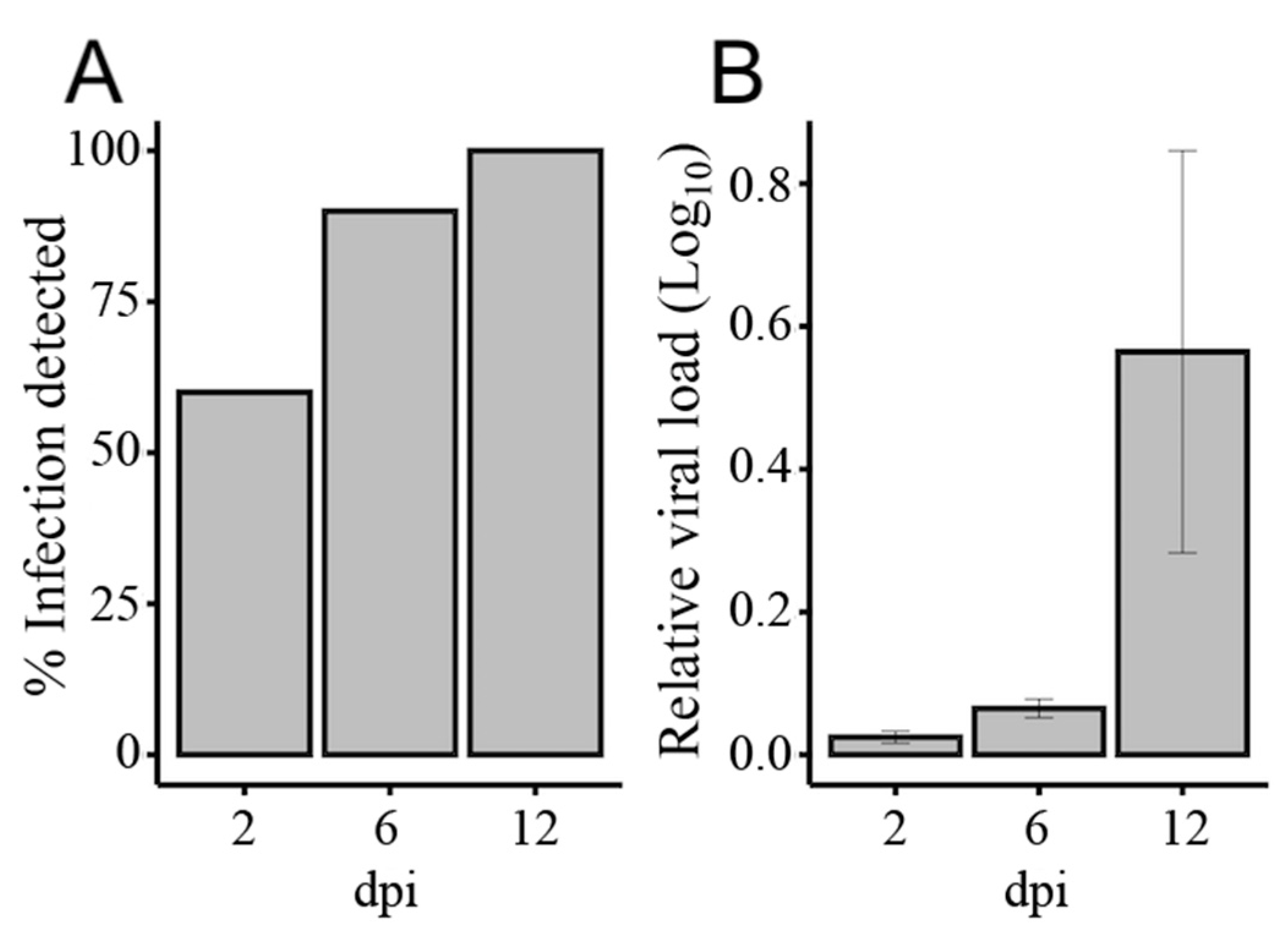
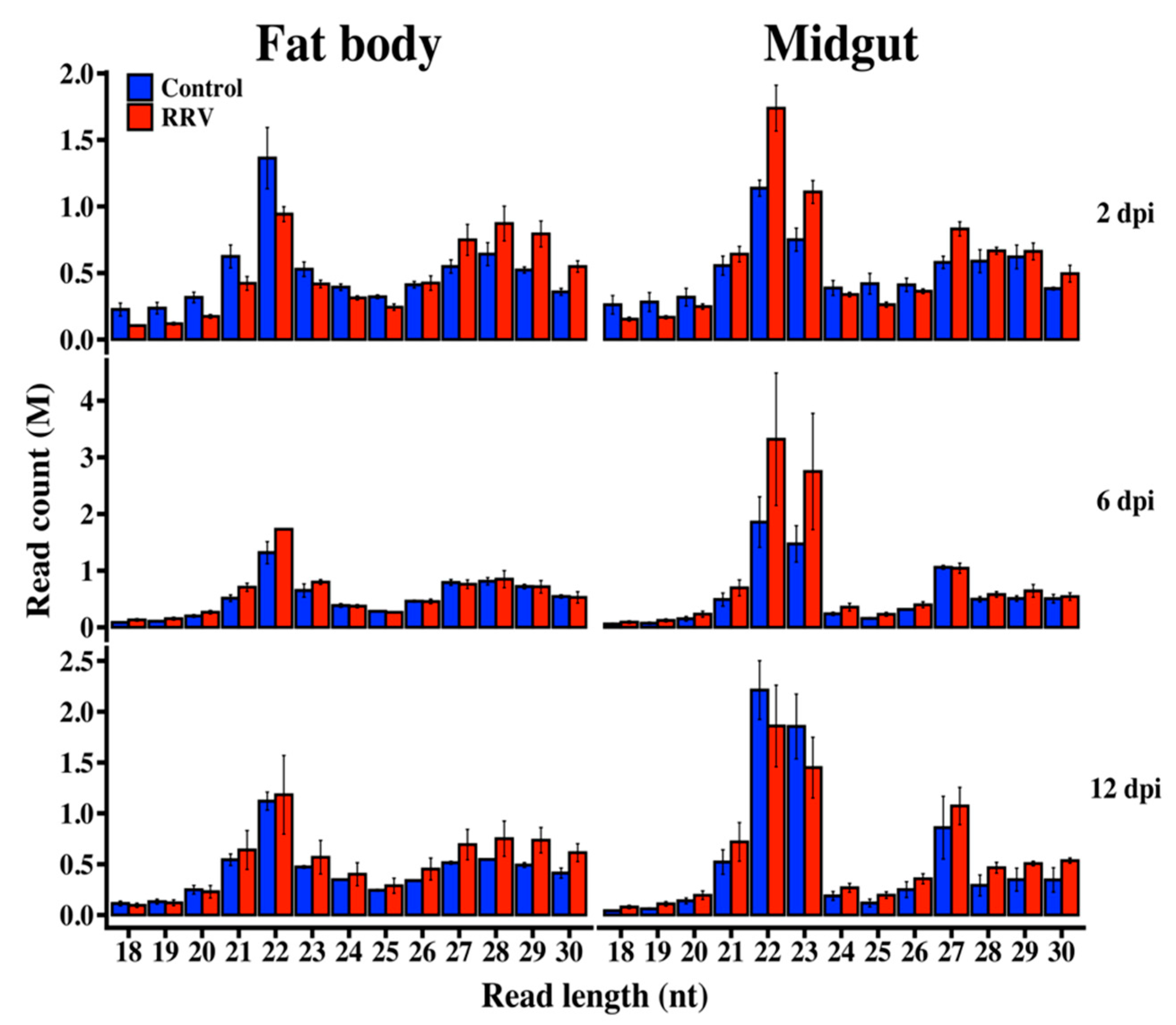
3.1. Illumina Sequencing of Small RNAs
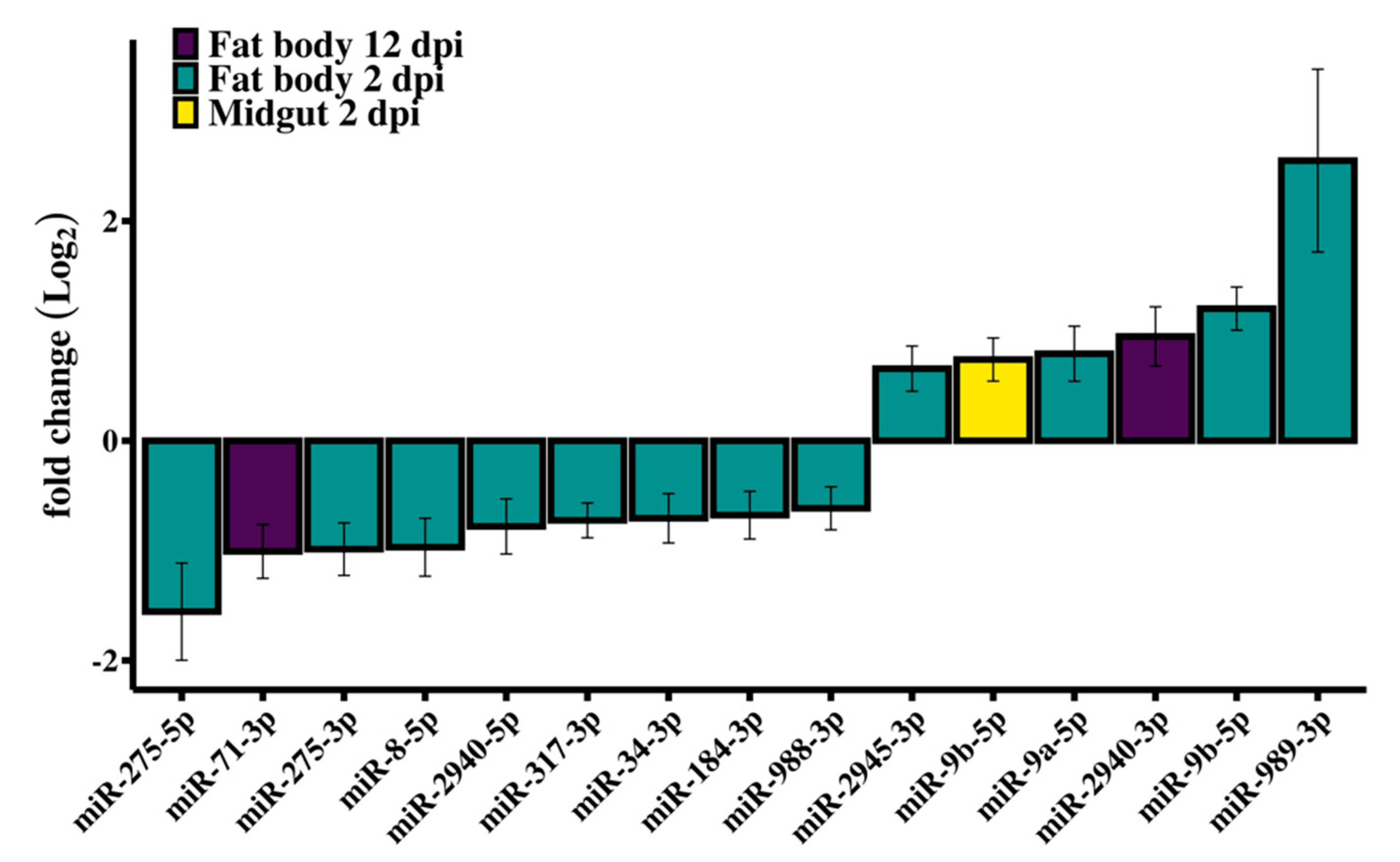
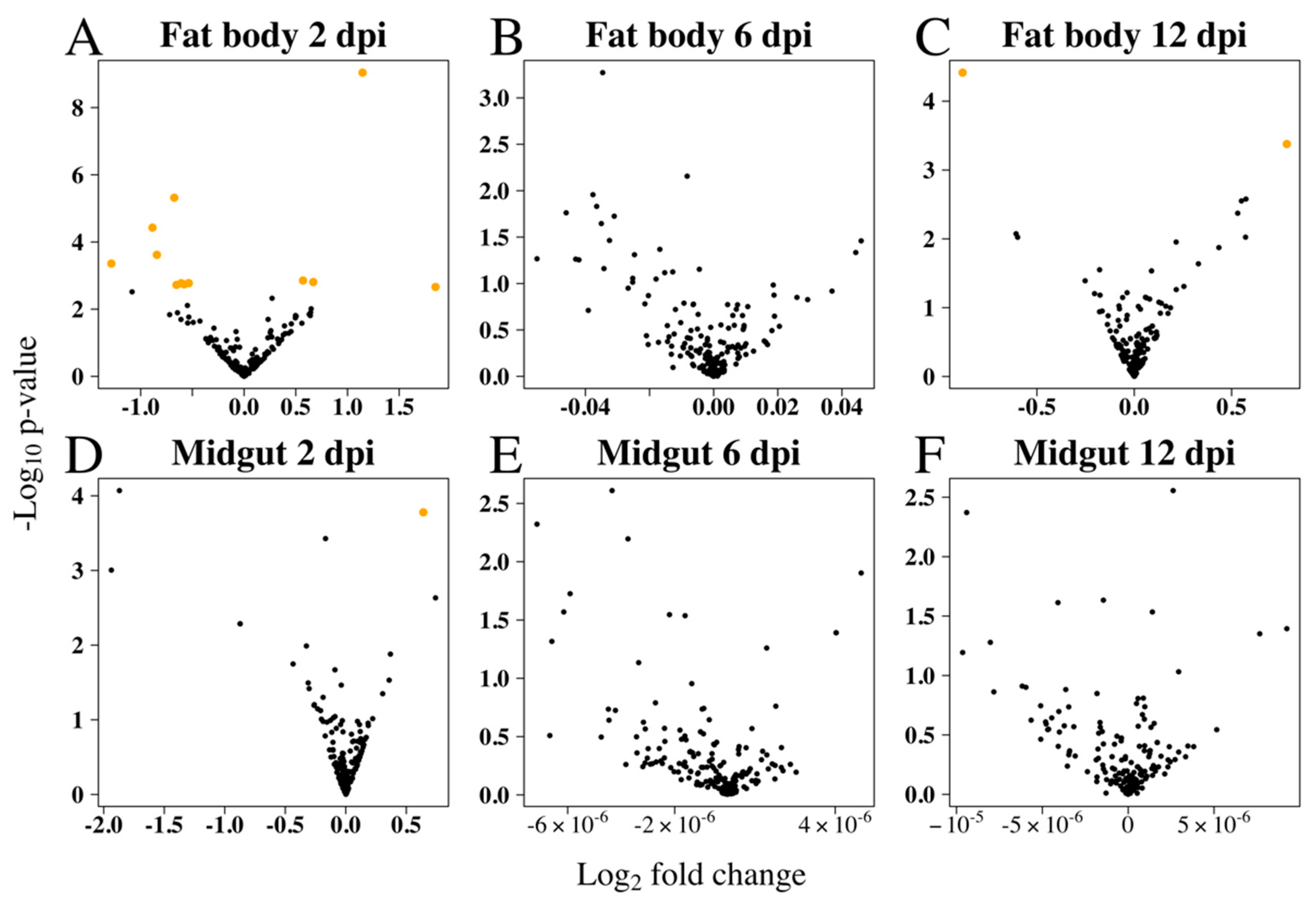
3.2. Differential Expression of Ae. aegypti miRNAs in Response to RRV Infection
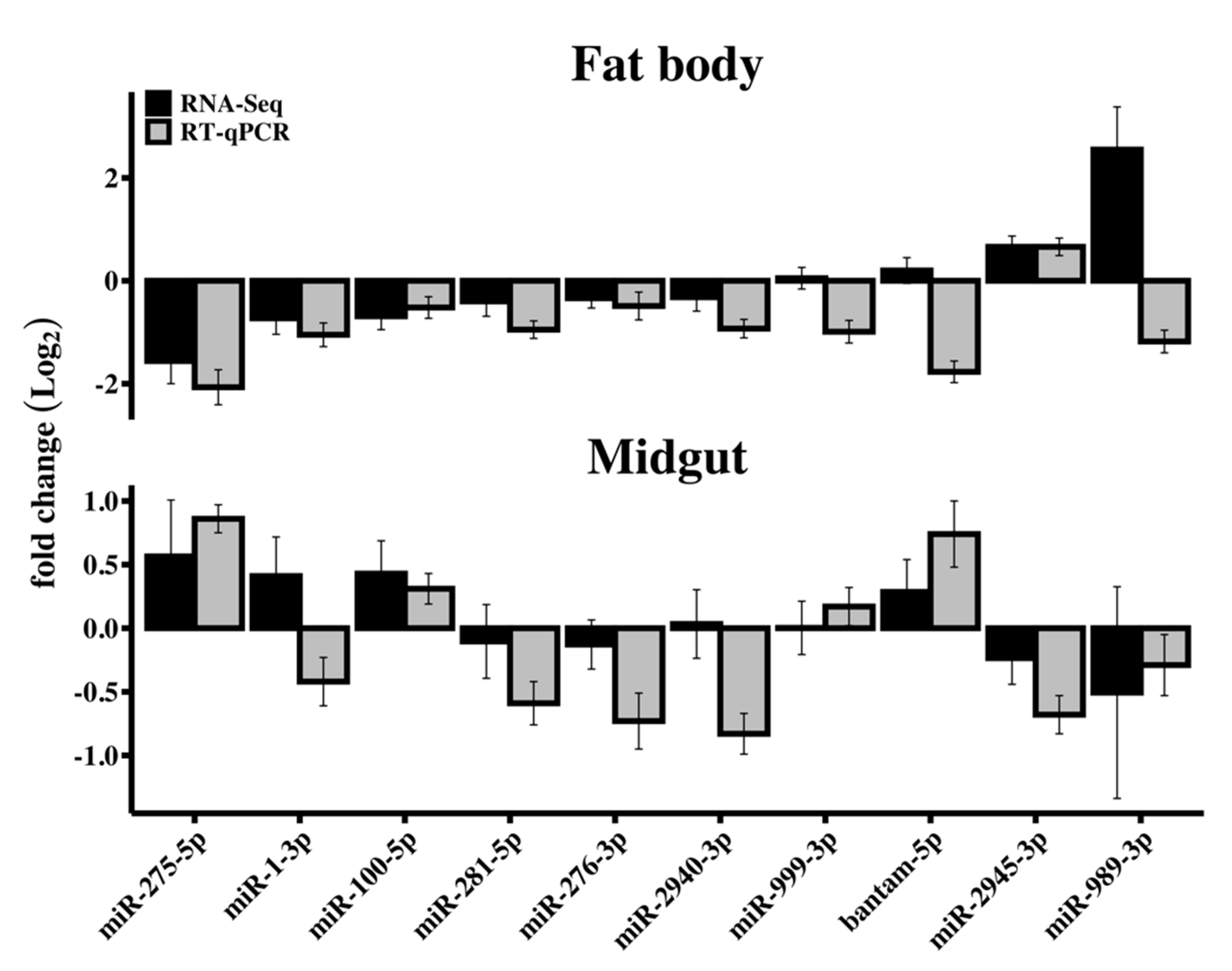
3.3. RT-qPCR Validation of Differentially Expressed miRNAs
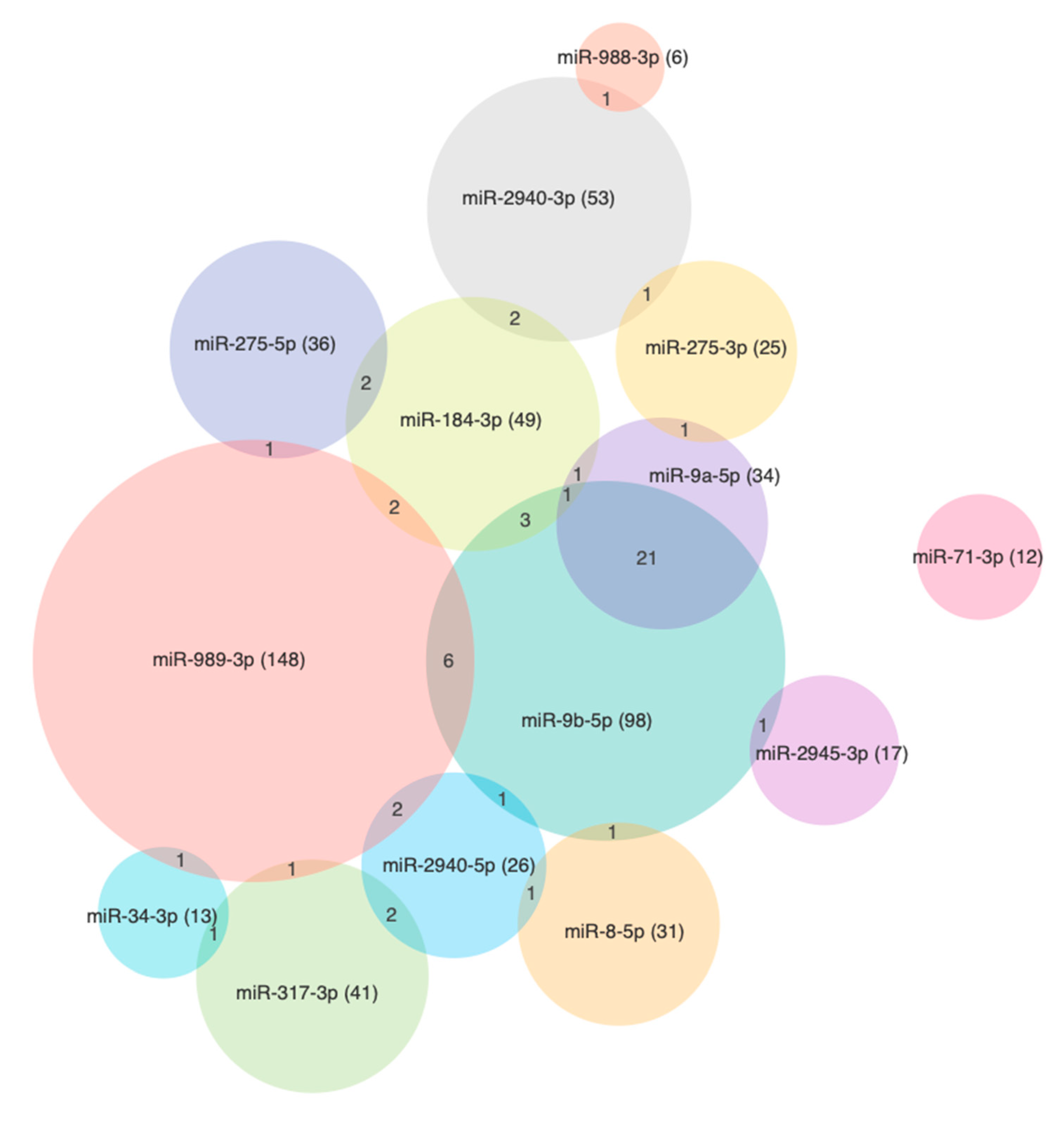
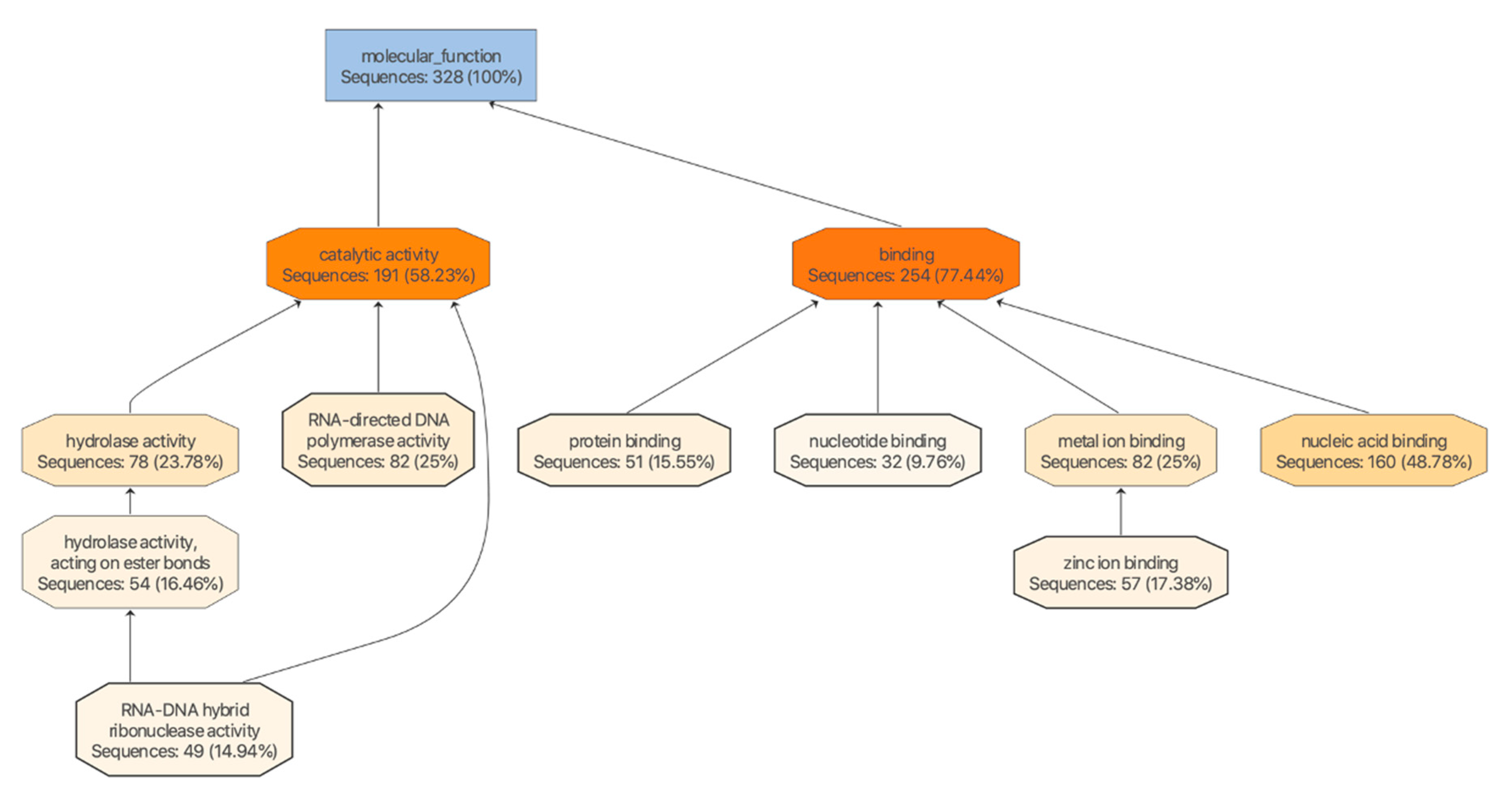
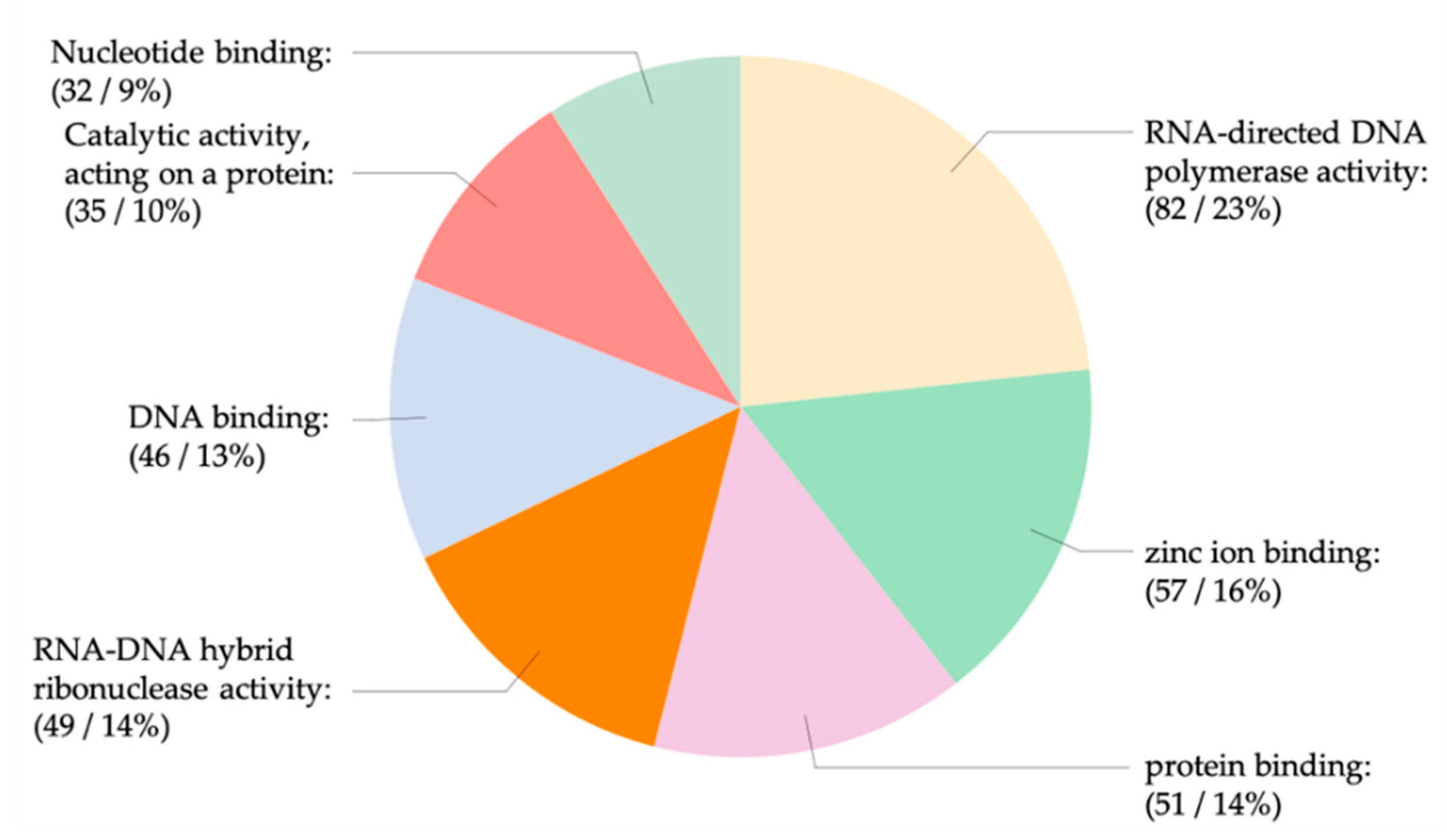
3.4. The miRNA Target Analysis
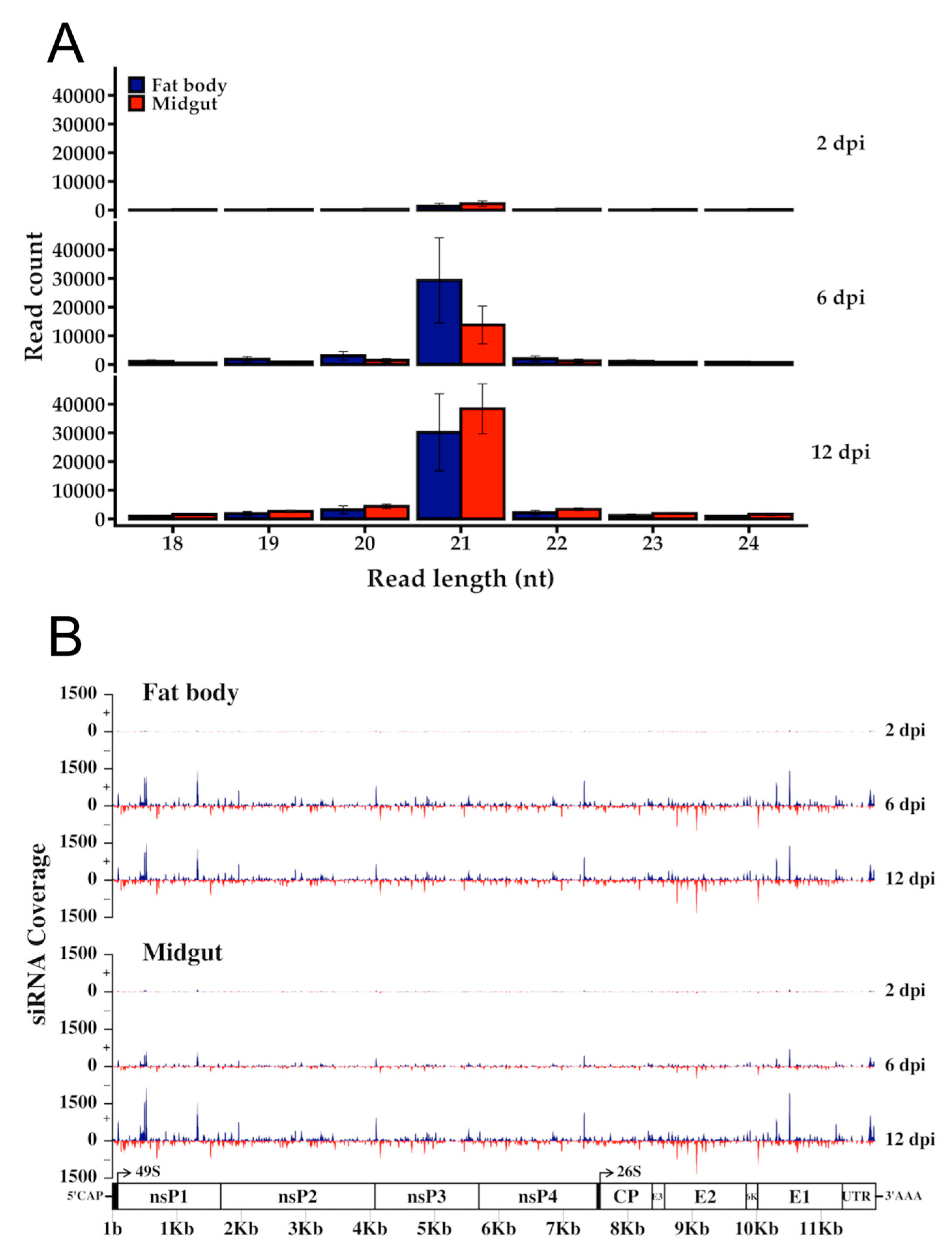
3.5. RRV is a Target of the Ae. aegypti RNAi Response
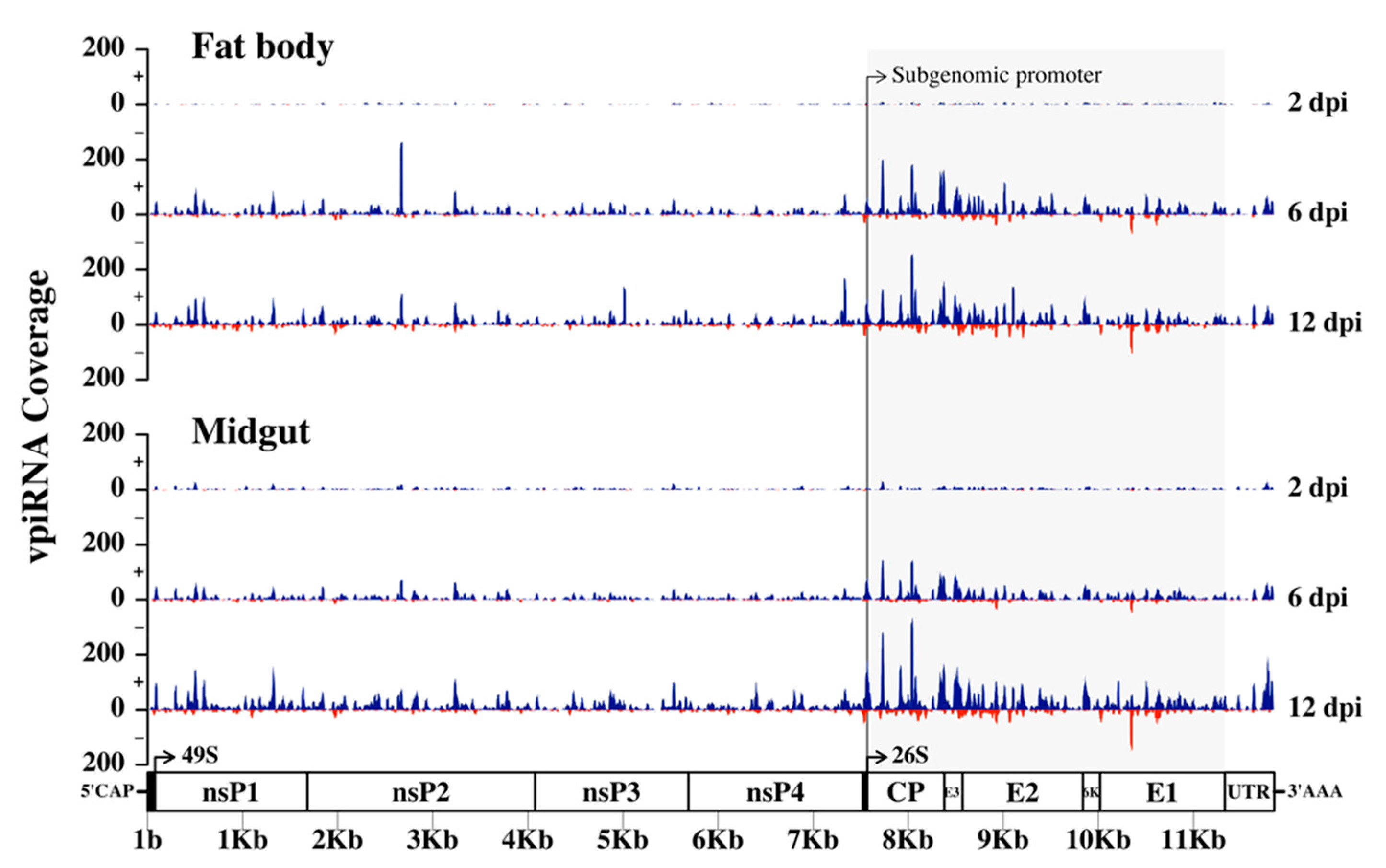
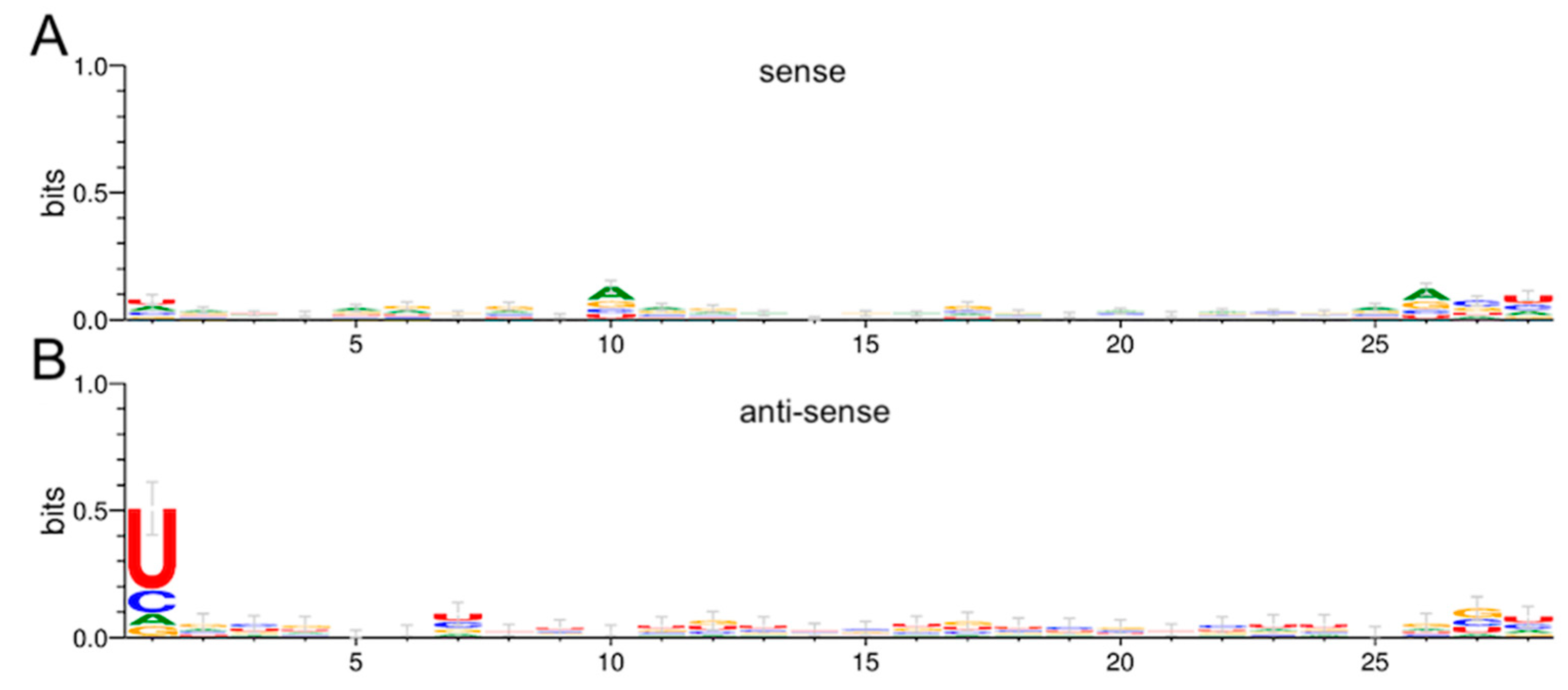
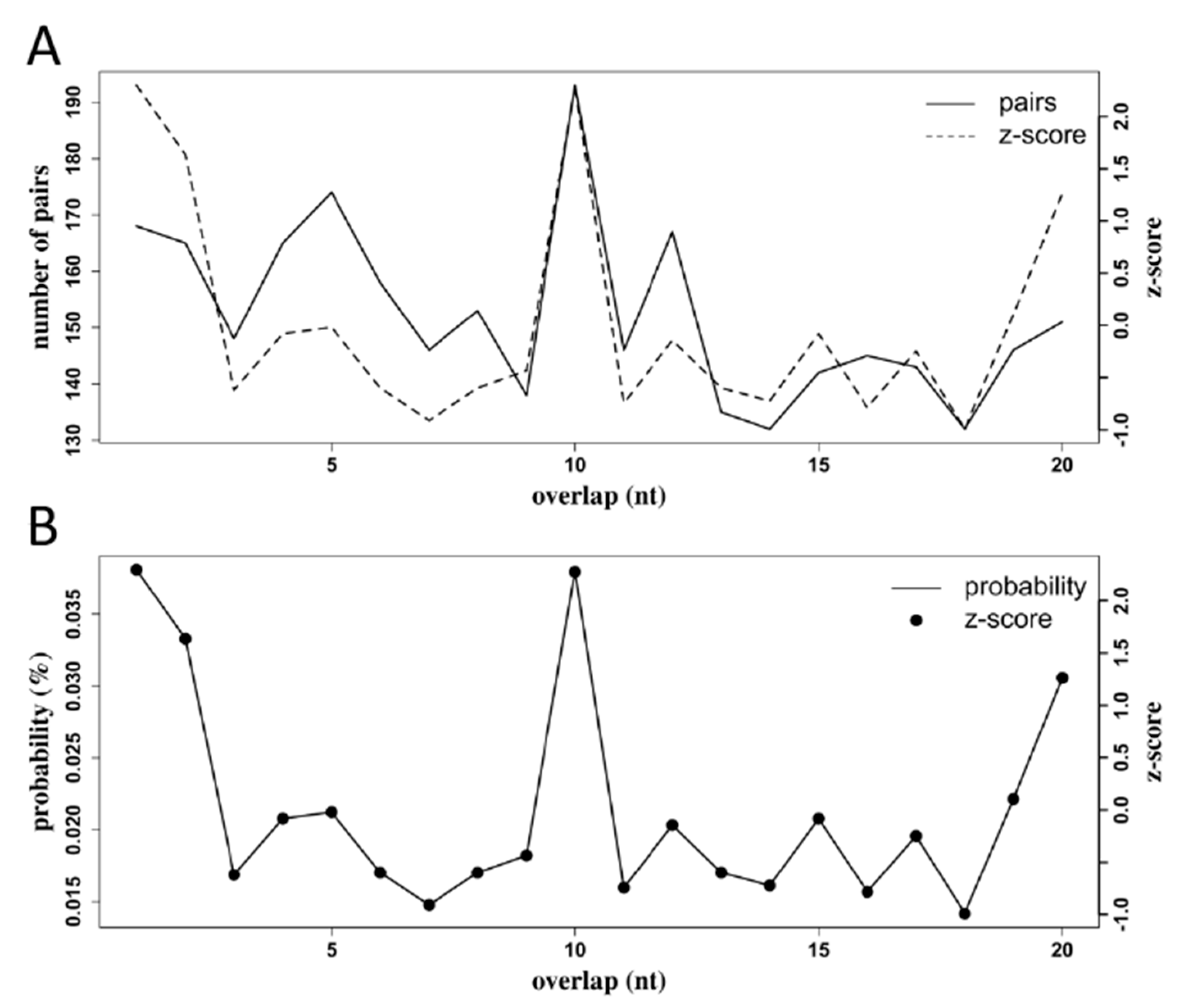
3.6. Production of RRV-Derived vpiRNAs
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Contigiani, M.S.; Diaz, L.A. In Arthropod Borne Diseases; Springer: Berlin/Heidelberg, Germany, 2017; pp. 115–135. [Google Scholar]
- Weaver, S.C.; Barrett, A.D.T. Transmission cycles, host range, evolution, and emergence of arboviral disease. Nat. Rev. Genet. 2004, 2, 789–801. [Google Scholar] [CrossRef] [PubMed]
- Koolhof, I.S.; Carver, S. Epidemic host community contribution to mosquito-borne disease transmission: Ross River virus. Epidemiol. Infect. 2016, 145, 656–666. [Google Scholar] [CrossRef] [PubMed]
- Russell, R.C. Ross River virus: Ecology and distribution. Annu. Rev. Entomol. 2002, 47, 1–31. [Google Scholar] [CrossRef]
- Lau, C.L.; Aubry, M.; Nhan, D.M.T.-X.; Teissier, A.; Paulous, S.; Desprès, P.; de Lamballerie, X.; Pastorino, B.; Cao-Lormeau, V.-M.; Weinstein, P. New evidence for endemic circulation of Ross River virus in the Pacific Islands and the potential for emergence. Int. J. Infect. Dis. 2017, 57, 73–76. [Google Scholar] [CrossRef] [PubMed]
- Togami, E.; Gyawali, N.; Ong, O.; Kama, M.; Cao-Lormeau, V.-M.; Aubry, M.; Ko, A.I.; Nilles, E.J.; Collins-Emerson, J.M.; Devine, G.J.; et al. First evidence of concurrent enzootic and endemic transmission of Ross River virus in the absence of marsupial reservoirs in Fiji. Int. J. Infect. Dis. 2020, 96, 94–96. [Google Scholar] [CrossRef]
- NNDSS. National Notifiable Diseases Surveillance Fortnightly Summary Notes—4 August to 17 August 2018. Available online: https://www1.health.gov.au/internet/main/publishing.nsf/Content/cdnareport-fn16-18.htm (accessed on 18 May 2020).
- Ratnayake, J.T.B. The valuation of social and economic costs of mosquito-transmitted Ross River virus; Griffith University: Queensland, Australia, 2006. [Google Scholar]
- Tomerini, D.M. The impact of local government mosquito control programs on Ross River virus disease in Queensland, Australia; Griffith University: Queensland, Australia, 2008. [Google Scholar]
- Doherty, R.; Whitehead, R.; Gorman, B.; O’gower, A. The isolation of a third group A arbovirus in Australia, with preliminary observations on its relationship to epidemic polyarthritis. Aust. J. Sci. 1963, 26, 183–184. [Google Scholar]
- Russell, R. Ross River virus: Disease trends and vector ecology in Australia. Bull. Soc. Vector Ecol. 1994, 19, 73–81. [Google Scholar]
- Ritchie, S.A.; Phillips, D.A.; Standfast, H.A.; Fanning, I.D.; McGinn, D.; Kay, B.H. Ross River Virus in Mosquitoes (Diptera: Culicidae) During the 1994 Epidemic Around Brisbane, Australia. J. Med Entomol. 1997, 34, 156–159. [Google Scholar] [CrossRef]
- Kay, B.H.; Ryan, P.A.; Hall, R.A.; Boyd, A.M. Mosquito feeding patterns and natural infection of vertebrates with Ross River and Barmah Forest viruses in Brisbane, Australia. Am. J. Trop. Med. Hyg. 2007, 76, 417–423. [Google Scholar] [CrossRef]
- Boyd, A.M.; Hall, R.A.; Gemmell, R.T.; Kay, B.H. Experimental infection of Australian brushtail possums, Trichosurus vulpecula (Phalangeridae: Marsupialia), with Ross River and Barmah Forest viruses by use of a natural mosquito vector system. Am. J. Trop. Med. Hyg. 2001, 65, 777–782. [Google Scholar] [CrossRef]
- Holder, P.; Browne, G.; Bullians, M. The mosquitoes of New Zealand and their animal disease significance. Surveillance 1999, 26, 12–15. [Google Scholar]
- Derraik, J.G. A survey of the mosquito (Diptera: Culicidae) fauna of the Auckland Zoological Park. New Zealand Èntomol. 2004, 27, 51–55. [Google Scholar] [CrossRef]
- Kelly-Hope, L.A.; Kay, B.H.; Purdie, D.M.; Williams, G. Infectious Disease: The risk of Ross River and Barmah Forest virus disease in Queensland: Implications for New Zealand. Aust. New Zealand J. Public Heal. 2002, 26, 69–77. [Google Scholar] [CrossRef]
- Gubler, D.J. Transmission of Ross River virus by Aedes polynesiensis and Aedes aegypti. Am. J. Trop. Med. Hyg. 1981, 30, 1303–1306. [Google Scholar] [CrossRef] [PubMed]
- Nasci, R.S.; Mitchell, C.J. Larval Diet, Adult Size, and Susceptibility of Aedes aegypti (Diptera: Culicidae) to Infection with Ross River Virus. J. Med. Èntomol. 1994, 31, 123–126. [Google Scholar] [CrossRef] [PubMed]
- Whelan, P.I.; Merianos, A.; Patel, M.; Tai, K.; Currie, B. The epidemiology of arbovirus infection in the Northern Territory 1980–1992. Arbovirus Res. Aust. 1993, 6, 266–270. [Google Scholar]
- Taylor, W.P.; Marshall, I.D. Adaptation studies with Ross River virus: Retention of field level virulence. J. Gen. Virol. 1975, 28, 73–83. [Google Scholar] [CrossRef]
- Bhatt, S.; Gething, P.W.; Brady, O.J.; Messina, J.P.; Farlow, A.W.; Moyes, C.L.; Drake, J.M.; Brownstein, J.S.; Hoen, A.G.; Sankoh, O.; et al. The global distribution and burden of dengue. Nature 2013, 496, 504–507. [Google Scholar] [CrossRef] [PubMed]
- Nhan, D.M.T.-X.; Cao-Lormeau, V.-M.; Gubler, D.J. Zika virus: Following the path of dengue and chikungunya? Lancet 2015, 386, 243–244. [Google Scholar] [CrossRef]
- Powell, J.R. Mosquito-borne human viral diseases: Why Aedes aegypti? Am. J. Trop. Med. Hyg. 2018, 98, 1563–1565. [Google Scholar] [CrossRef]
- Tabachnick, W.J. Evolutionary genetics and arthropod-borne disease: The Yellow Fever mosquito. Am. Èntomol. 1991, 37, 14–26. [Google Scholar] [CrossRef]
- Sim, C.; Hong, Y.S.; van Landingham, D.L.; Harker, B.W.; Christophides, G.K.; Kafatos, F.C.; Higgs, S.; Collins, F.H. Modulation of Anopheles gambiae gene expression in response to o’nyong-nyong virus infection. Insect Mol. Biol. 2005, 14, 475–481. [Google Scholar] [CrossRef] [PubMed]
- Sim, C.; Hong, Y.S.; Ttsetsarkin, K.; van Landingham, D.L.; Higgs, S.; Collins, F.H. Anopheles gambiae heat shock protein cognate 70B impedes o’nyong-nyong virus replication. BMC Genom. 2007, 8, 231. [Google Scholar] [CrossRef] [PubMed]
- Xi, Z.; Ramirez, J.L.; Dimopoulos, G. The Aedes aegypti Toll pathway controls dengue virus infection. PLoS Pathog. 2008, 4, e1000098. [Google Scholar] [CrossRef] [PubMed]
- Sanders, H.R.; Foy, B.D.; Evans, A.M.; Ross, L.S.; Beaty, B.J.; Olson, K.E.; Gill, S.S. Sindbis virus induces transport processes and alters expression of innate immunity pathway genes in the midgut of the disease vector, Aedes aegypti. Insect Biochem. Mol. Biol. 2005, 35, 1293–1307. [Google Scholar] [CrossRef] [PubMed]
- Luplertlop, N.; Surasombatpattana, P.; Patramool, S.; Dumas, E.; Wasinpiyamongkol, L.; Sauné, L.; Hamel, R.; Bernard, E.; Sereno, D.; Thomas, F.; et al. Induction of a peptide with activity against a broad spectrum of pathogens in the Aedes aegypti salivary gland, following infection with dengue virus. PLoS Pathog. 2011, 7, e1001252. [Google Scholar] [CrossRef]
- McFarlane, M.; Arias-Goeta, C.; Martin, E.; O’Hara, Z.; Lulla, A.; Mousson, L.; Rainey, S.M.; Misbah, S.; Schnettler, E.; Donald, C.L.; et al. Characterization of Aedes aegypti Innate-Immune Pathways that Limit Chikungunya Virus Replication. PLoS Neglect. Trop. Dis. 2014, 8, e2994. [Google Scholar] [CrossRef]
- Jupatanakul, N.; Sim, S.; Angleró-Rodríguez, Y.I.; Souza-Neto, J.A.; Das, S.; Poti, K.E.; Rossi, S.L.; Bergren, N.; Vasilakis, N.; Dimopoulos, G. Engineered Aedes aegypti JAK/STAT pathway-mediated immunity to dengue virus. PLoS Neglect. Trop. Dis. 2017, 11, e0005187. [Google Scholar] [CrossRef]
- Angleró-Rodríguez, Y.I.; MacLeod, H.J.; Kang, S.; Carlson, J.S.; Jupatanakul, N.; Dimopoulos, G. Aedes aegypti molecular responses to Zika virus: Modulation of infection by the Toll and Jak/Stat immune pathways and virus host factors. Front. Microbiol. 2017, 8, 2050. [Google Scholar] [CrossRef]
- Hussain, M.; Etebari, K.; Asgari, S. Functions of small RNAs in mosquitoes. In Advances in Insect Physiology; Elsevier BV: Amsterdam, The Netherlands, 2016; Volume 51, pp. 189–222. [Google Scholar]
- Blair, C.D.; Olson, K.E. The Role of RNA interference (RNAi) in arbovirus-vector interactions. Viruses 2015, 7, 820–843. [Google Scholar] [CrossRef]
- Campbell, C.L.; Keene, K.M.; Brackney, D.E.; Olson, K.E.; Blair, C.D.; Wilusz, J.; Foy, B.D. Aedes aegypti uses RNA interference in defense against Sindbis virus infection. BMC Microbiol. 2008, 8, 47. [Google Scholar] [CrossRef]
- Cirimotich, C.; Scott, J.C.; Phillips, A.T.; Geiss, B.J.; Olson, K.E. Suppression of RNA interference increases alphavirus replication and virus-associated mortality in Aedes aegypti mosquitoes. BMC Microbiol. 2009, 9, 49. [Google Scholar] [CrossRef] [PubMed]
- Franz, A.W.; Sanchez-Vargas, I.; Adelman, Z.N.; Blair, C.D.; Beaty, B.J.; James, A.A.; Olson, K.E. Engineering RNA interference-based resistance to dengue virus type 2 in genetically modified Aedes aegypti. Proc. Natl. Acad. Sci. USA 2006, 103, 4198–4203. [Google Scholar] [CrossRef] [PubMed]
- Siu, R.W.C.; Fragkoudis, R.; Simmonds, P.; Donald, C.L.; Chase-Topping, M.E.; Barry, G.; Attarzadeh-Yazdi, G.; Rodriguez-Andres, J.; Nash, A.A.; Merits, A.; et al. Antiviral RNA interference responses induced by Semliki Forest virus infection of mosquito cells: Characterization, origin, and frequency-dependent functions of virus-derived small interfering RNAs. J. Virol. 2010, 85, 2907–2917. [Google Scholar] [CrossRef] [PubMed]
- Bronkhorst, A.W.; van Rij, R.P. The long and short of antiviral defense: Small RNA-based immunity in insects. Curr. Opin. Virol. 2014, 7, 19–28. [Google Scholar] [CrossRef]
- Arensburger, P.; Hice, R.H.; Wright, J.A.; Craig, N.L.; Atkinson, P.W. The mosquito Aedes aegypti has a large genome size and high transposable element load but contains a low proportion of transposon-specific piRNAs. BMC Genom. 2011, 12, 606. [Google Scholar] [CrossRef]
- Hussain, M.; Walker, T.; O’Neill, S.L.; Asgari, S. Blood meal induced microRNA regulates development and immune associated genes in the Dengue mosquito vector, Aedes aegypti. Insect Biochem. Mol. Biol. 2013, 43, 146–152. [Google Scholar] [CrossRef]
- Liu, S.; Lucas, K.J.; Roy, S.; Ha, J.; Raikhel, A.S. Mosquito-specific microRNA-1174 targets serine hydroxymethyltransferase to control key functions in the gut. Proc. Natl. Acad. Sci. USA 2014, 111, 14460–14465. [Google Scholar] [CrossRef]
- Bryant, B.; Macdonald, W.; Raikhel, A.S. microRNA miR-275 is indispensable for blood digestion and egg development in the mosquito Aedes aegypti. Proc. Natl. Acad. Sci. USA 2010, 22391–22398. [Google Scholar] [CrossRef]
- Campbell, C.L.; Harrison, T.; Hess, A.M.; Ebel, G.D. MicroRNA levels are modulated in Aedes aegypti after exposure to D engue-2. Insect. Mol. Biol. 2014, 23, 132–139. [Google Scholar] [CrossRef]
- Maharaj, P.D.; Widen, S.G.; Huang, J.; Wood, T.G.; Thangamani, S. Discovery of mosquito saliva microRNAs during CHIKV infection. PLoS Neglect. Trop. Dis. 2015, 9, e0003386. [Google Scholar] [CrossRef] [PubMed]
- Arensburger, P.; Megy, K.; Waterhouse, R.M.; Abrudan, J.; Amedeo, P.; Antelo, B.; Bartholomay, L.; Bidwell, S.; Caler, E.; Camara, F.; et al. Sequencing of Culex quinquefasciatus establishes a platform for mosquito comparative genomics. Science 2010, 330, 86–88. [Google Scholar] [CrossRef] [PubMed]
- Kay, B.H.; Fanning, I.D.; Carley, J.G. Vector competence of Culex pipiens quinquefasciatus for Murray Valley Encephalitis, Kunjin, and Ross River viruses from Australia. Am. J. Trop. Med. Hyg. 1982, 31, 844–848. [Google Scholar] [CrossRef] [PubMed]
- Pearson, W.; Wood, T.; Zhang, Z.; Miller, W. Comparison of DNA sequences with protein sequences. Genomics 1997, 46, 24–36. [Google Scholar] [CrossRef]
- Dodt, M.; Roehr, J.T.; Ahmed, R.; Dieterich, C. FLEXBAR—Flexible barcode and adapter processing for next-generation sequencing platforms. Biology 2012, 1, 895–905. [Google Scholar] [CrossRef]
- Wickham, H.; Seidel, D. Scales: scale functions for visualization. Available online: https://CRAN.R-project.org/package=scales (accessed on 18 May 2020).
- Wickham, H. The split-apply-combine strategy for data analysis. J. Stat. Softw. 2011, 40, 1–29. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: Berlin/Heidelberg, Germany, 2016. [Google Scholar]
- Shi, J.; Dong, M.; Li, L.; Luz-Madrigal, A.; Tsonis, P.A.; del Rio-Tsonis, K.; Liang, C. mirPRo—A novel standalone program for differential expression and variation analysis of miRNAs. Sci. Rep. 2015, 5, 14617. [Google Scholar] [CrossRef]
- Griffiths-Jones, S. miRBase: microRNA sequences, targets, and gene nomenclature. Nucleic Acids Res. 2006, 34, D140–D144. [Google Scholar] [CrossRef]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2014, 31, 166–169. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.R.; Durbin, R. The sequence alignment/map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Giraldo-Calderón, G.I.; Emrich, S.J.; Maccallum, R.M.; Maslen, G.; Dialynas, E.; Topalis, P.; Ho, N.; Gesing, S.; Madey, G.; Collins, F.H.; et al. VectorBase: An updated bioinformatics resource for invertebrate vectors and other organisms related with human diseases. Nucleic Acids Res. 2014, 43, D707–D713. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 002832. [Google Scholar] [CrossRef]
- Huber, W.; Carey, V.J.; Gentleman, R.; Anders, S.; Carlson, M.; Carvalho, B.S.; Bravo, H.C.; Davis, S.; Gatto, L.; Girke, T.; et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 2015, 12, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Anonymous. The R project for statistical computing. Available online: http://www.r-project.org/ (accessed on 13 February 2012).
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Zhu, A.; Ibrahim, J.G.; Love, M.I. Heavy-tailed prior distributions for sequence count data: Removing the noise and preserving large differences. Bioinformatics 2019, 35, 2084–2092. [Google Scholar] [CrossRef] [PubMed]
- Jones, A.; Lowry, K.; Aaskov, J.; Holmes, E.; Kitchen, A. Molecular evolutionary dynamics of Ross River virus and implications for vaccine efficacy. J. Gen. Virol. 2009, 91, 182–188. [Google Scholar] [CrossRef]
- Gel, B.; Serra, E. KaryoploteR: An R/Bioconductor package to plot customizable genomes displaying arbitrary data. Bioinformatics 2017, 33, 3088–3090. [Google Scholar] [CrossRef]
- Crooks, G.E.; Hon, G.; Chandonia, J.-M.; Brenner, S. WebLogo: A Sequence Logo Generator. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef]
- Afgan, E.; Baker, D.; Batut, B.; Beek, M.V.D.; Bouvier, D.; Čech, M.; Chilton, J.; Clements, D.; Coraor, N.; Grüning, B.A.; et al. The Galaxy platform for accessible, reproducible, and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 2018, 46, W537–W544. [Google Scholar] [CrossRef]
- Enright, A.J.; John, B.; Gaul, U.; Tuschl, T.; Sander, C.; Marks, D.S. MicroRNA targets in Drosophila. Genome Biol. 2003, 5, R1. [Google Scholar] [CrossRef]
- Krüger, J.; Rehmsmeier, M. RNAhybrid: microRNA target prediction easy, fast, and flexible. Nucleic Acids Res. 2006, 34, W451–W454. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, V.; Subtelny, A.O.; Thiru, P.; Ulitsky, I.; Bartel, B. Predicting microRNA targeting efficacy in Drosophila. Genome Biol. 2018, 19, 152. [Google Scholar] [CrossRef]
- Kertesz, M.; Iovino, N.; Unnerstall, U.; Gaul, U.; Segal, E. The role of site accessibility in microRNA target recognition. Nat. Genet. 2007, 39, 1278–1284. [Google Scholar] [CrossRef] [PubMed]
- Loher, P.; Rigoutsos, I. Interactive exploration of RNA22 microRNA target predictions. Bioinformatics 2012, 28, 3322–3323. [Google Scholar] [CrossRef] [PubMed]
- Quinlan, A.R.; Hall, I.M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, A.C.; Bovolenta, L.A.; Nachtigall, P.G.; Herkenhoff, M.; Lemke, N.; Pinhal, D. Combining results from distinct microrna target prediction tools enhances the performance of analyses. Front. Genet. 2017, 8, 59. [Google Scholar] [CrossRef] [PubMed]
- Götz, S.; García-Gómez, J.M.; Terol, J.; Williams, T.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talón, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, 45. [Google Scholar] [CrossRef]
- Trobaugh, D.W.; Klimstra, W.B. MicroRNA regulation of rna virus replication and pathogenesis. Trends Mol. Med. 2016, 23, 80–93. [Google Scholar] [CrossRef]
- Saldana, M.A.; Etebari, K.; Hart, C.E.; Widen, S.G.; Wood, T.G.; Thangamani, S.; Asgari, S.; Hughes, G.L. Zika virus alters the microRNA expression profile and elicits an RNAi response in Aedes aegypti mosquitoes. PLoS Neglect. Trop. Dis. 2017, 11, e0005760. [Google Scholar] [CrossRef]
- Shrinet, J.; Jain, S.; Jain, J.; Bhatnagar, R.K.; Sunil, S. Next generation sequencing reveals regulation of distinct Aedes microRNAs during chikungunya virus development. PLoS Neglect. Trop. Dis. 2014, 8, e2616. [Google Scholar] [CrossRef] [PubMed]
- Myles, K.M.; Wiley, M.R.; Morazzani, E.M.; Adelman, Z.N. Alphavirus-derived small RNAs modulate pathogenesis in disease vector mosquitoes. Proc. Natl. Acad. Sci. USA 2008, 105, 19938–19943. [Google Scholar] [CrossRef] [PubMed]
- Pakpour, N.; Riehle, M.A.; Luckhart, S. Effects of ingested vertebrate-derived factors on insect immune responses. Curr. Opin. Insect Sci. 2014, 3, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhou, Y.; Wu, J.; Zheng, P.; Li, Y.; Zheng, X.; Puthiyakunnon, S.; Tu, Z.; Chen, X.-G. The expression profile of Aedes albopictus miRNAs is altered by dengue virus serotype-2 infection. Cell Biosci. 2015, 5, 16. [Google Scholar] [CrossRef] [PubMed]
- Etebari, K.; Asgari, S. Accuracy of microrna discovery pipelines in non-model organisms using closely related species genomes. PLoS ONE 2014, 9, e84747. [Google Scholar] [CrossRef]
- Winter, F.; Edaye, S.; Hüttenhofer, A.; Brunel, C. Anopheles gambiae miRNAs as actors of defence reaction against Plasmodium invasion. Nucleic Acids Res. 2007, 35, 6953–6962. [Google Scholar] [CrossRef]
- Akbari, O.S.; Antoshechkin, I.; Amrhein, H.; Williams, B.; Diloreto, R.; Sandler, J.; Hay, B.A. The developmental transcriptome of the mosquito Aedes aegypti, an invasive species and major arbovirus vector. G3 Genes/Genomes/Genetics 2013, 3, 1493–1509. [Google Scholar] [CrossRef]
- Mead, E.A.; Tu, Z. Cloning, characterization, and expression of microRNAs from the Asian malaria mosquito, Anopheles stephensi. BMC Genom. 2008, 9, 244. [Google Scholar] [CrossRef]
- Skalsky, R.L.; van Landingham, D.L.; Scholle, F.; Higgs, S.; Cullen, B.R. Identification of microRNAs expressed in two mosquito vectors, Aedes albopictus and Culex quinquefasciatus. BMC Genom. 2010, 11, 119. [Google Scholar] [CrossRef]
- Etebari, K.; Osei-Amo, S.; Blomberg, S.P.; Asgari, S. Dengue virus infection alters post-transcriptional modification of microRNAs in the mosquito vector Aedes aegypti. Sci. Rep. 2015, 5, 15968. [Google Scholar] [CrossRef]
- Mayoral, J.G.; Etebari, K.; Hussain, M.; Khromykh, A.A.; Asgari, S. Wolbachia infection modifies the profile, shuttling and structure of microRNAs in a mosquito cell line. PLoS ONE 2014, 9, e96107. [Google Scholar] [CrossRef] [PubMed]
- Hedges, L.M.; Brownlie, J.C.; O’Neill, S.L.; Johnson, K.; Pörtner, H.-O.; Farrell, A.P. Wolbachia and virus protection in insects. Science 2008, 322, 702. [Google Scholar] [CrossRef] [PubMed]
- Asad, S.; Hussain, M.; Hugo, L.E.; Osei-Amo, S.; Zhang, G.; Watterson, D.; Asgari, S. Suppression of the pelo protein by Wolbachia and its effect on dengue virus in Aedes aegypti. PLoS Neglect. Trop. Dis. 2018, 12, e0006405. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Hussain, M.; O’Neill, S.L.; Asgari, S. Wolbachia uses a host microRNA to regulate transcripts of a methyltransferase, contributing to dengue virus inhibition in Aedes aegypti. Proc. Natl. Acad. Sci. USA 2013, 110, 10276–10281. [Google Scholar] [CrossRef]
- Bick, M.J.; Carroll, J.-W.N.; Gao, G.; Goff, S.P.; Rice, C.M.; Macdonald, M.R. expression of the zinc-finger antiviral protein inhibits alphavirus replication. J. Virol. 2003, 77, 11555–11562. [Google Scholar] [CrossRef]
- Nakamoto, M.; Moy, R.; Xu, J.; Bambina, S.; Yasunaga, A.; Shelly, S.S.; Gold, B.; Cherry, S. Virus recognition by Toll-7 activates antiviral autophagy in Drosophila. Immunity 2012, 36, 658–667. [Google Scholar] [CrossRef]
- Patel, R.K.; Hardy, R.W. Role for the phosphatidylinositol 3-Kinase-Akt-TOR pathway during Sindbis virus Replication in arthropods. J. Virol. 2012, 86, 3595–3604. [Google Scholar] [CrossRef]
- Mazzon, M.; Castro, C.; Thaa, B.; Liu, L.; Mutso, M.; Liu, X.; Mahalingam, S.; Griffin, J.L.; Marsh, M.; McInerney, G. Alphavirus-induced hyperactivation of PI3K/AKT directs pro-viral metabolic changes. PLoS Pathog. 2018, 14, e1006835. [Google Scholar] [CrossRef]
- Gulia-Nuss, M.; Robertson, A.E.; Brown, M.R.; Strand, M.R. insulin-like peptides and the target of rapamycin pathway coordinately regulate blood digestion and egg maturation in the mosquito Aedes aegypti. PLoS ONE 2011, 6, e20401. [Google Scholar] [CrossRef]
- Lucas, K.J.; Roy, S.; Ha, J.; Gervaise, A.L.; Kokoza, V.A.; Raikhel, A.S. MicroRNA-8 targets the Wingless signaling pathway in the female mosquito fat body to regulate reproductive processes. Proc. Natl. Acad. Sci. USA 2015, 112, 1440–1445. [Google Scholar] [CrossRef]
- Schwenke, R.A.; Lazzaro, B.P.; Wolfner, M.F. Reproduction-Immunity Trade-Offs in Insects. Annu. Rev. Entomol. 2015, 61, 239–256. [Google Scholar] [CrossRef]
- Kuss-Duerkop, S.K.; Wang, J.; Mena, I.; White, K.M.; Metreveli, G.; Sakthivel, R.; Mata, M.A.; Muñoz-Moreno, R.; Chen, X.; Krammer, F.; et al. Influenza virus differentially activates mTORC1 and mTORC2 signaling to maximize late stage replication. PLoS Pathog. 2017, 13, e1006635. [Google Scholar] [CrossRef] [PubMed]
- Liang, Q.; Luo, Z.; Zeng, J.; Chen, W.; Foo, S.-S.; Lee, S.-A.; Ge, J.; Wang, S.; Goldman, S.A.; Zlokovic, B.V.; et al. Zika virus NS4A and NS4B proteins deregulate Akt-mTOR signaling in human fetal neural stem cells to inhibit neurogenesis and induce autophagy. Cell Stem Cell 2016, 19, 663–671. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Behura, S.K.; Franz, A.W.E. The midgut transcriptome of Aedes aegypti fed with saline or protein meals containing chikungunya virus reveals genes potentially involved in viral midgut escape. BMC Genom. 2017, 18, 382. [Google Scholar] [CrossRef]
- Etebari, K.; Hegde, S.; Saldaña, M.A.; Widen, S.G.; Wood, T.G.; Asgari, S.; Hughes, G.L. global transcriptome analysis of Aedes aegypti mosquitoes in response to zika virus infection. mSphere 2017, 2, e00456-17. [Google Scholar] [CrossRef] [PubMed]
- Raquin, V.; Merkling, S.H.; Gausson, V.; Moltini-Conclois, I.; Frangeul, L.; Varet, H.; Dillies, M.-A.; Saleh, M.C.; Lambrechts, L. Individual co-variation between viral RNA load and gene expression reveals novel host factors during early dengue virus infection of the Aedes aegypti midgut. PLoS Neglect. Trop. Dis. 2017, 11, e0006152. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Alto, B.W.; Jiang, Y.; Yu, F.; Zhang, Y. Transcriptomic analysis of Aedes aegypti innate immune system in response to ingestion of Chikungunya virus. Int. J. Mol. Sci. 2019, 20, 3133. [Google Scholar] [CrossRef] [PubMed]
- Khoo, C.C.H.; Piper, J.; Sanchez-Vargas, I.; Olson, K.E.; Franz, A.W. The RNA interference pathway affects midgut infection- and escape barriers for Sindbis virus in Aedes aegypti. BMC Microbiol. 2010, 10, 130. [Google Scholar] [CrossRef]
- Joosten, J.; Miesen, P.; Taşköprü, E.; Pennings, B.; Jansen, P.W.T.C.; Huynen, M.A.; Vermeulen, M.; van Rij, R.P. The Tudor protein Veneno assembles the ping-pong amplification complex that produces viral piRNAs in Aedes mosquitoes. Nucleic Acids Res. 2019, 47, 2546–2559. [Google Scholar] [CrossRef]
- Miesen, P.; Girardi, E.; van Rij, R.P. Distinct sets of PIWI proteins produce arbovirus and transposon-derived piRNAs in Aedes aegypti mosquito cells. Nucleic Acids Res. 2015, 43, 6545–6556. [Google Scholar] [CrossRef]
- Vodovar, N.; Bronkhorst, A.W.; van Cleef, K.W.R.; Miesen, P.; Blanc, H.; van Rij, R.P.; Saleh, M.C. Arbovirus-derived piRNAs exhibit a ping-pong signature in mosquito cells. PLoS ONE 2012, 7, e30861. [Google Scholar] [CrossRef] [PubMed]
- Brennecke, J.; Aravin, A.A.; Stark, A.; Dus, M.; Kellis, M.; Sachidanandam, R.; Hannon, G.J. Discrete Small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 2007, 128, 1089–1103. [Google Scholar] [CrossRef] [PubMed]
- Gunawardane, L.S.; Saito, K.; Nishida, K.M.; Miyoshi, K.; Kawamura, Y.; Nagami, T.; Siomi, H.; Siomi, M.C. A slicer-mediated mechanism for repeat-associated siRNA 5’ end formation in Drosophila. Science 2007, 315, 1587–1590. [Google Scholar] [CrossRef] [PubMed]
- Miesen, P.; Joosten, J.; van Rij, R.P. PIWIs Go Viral: Arbovirus-derived piRNAs in vector mosquitoes. PLoS Pathog. 2016, 12, e1006017. [Google Scholar] [CrossRef] [PubMed]

| Tissue 1 | dpi | miRNA | Control 2 | RRV 2 | FC 3 | p-Value | FDR |
|---|---|---|---|---|---|---|---|
| FB | 2 | miR-9b-5p | 4752 | 10,939 | 2.30 | 9.10 10−10 | 9.55 10−8 |
| FB | 2 | miR-317-3p | 22,084 | 13,362 | −1.65 | 4.81 10−6 | 2.52 10−4 |
| FB | 2 | miR-275-3p | 12,465 | 6290 | −1.98 | 3.74 10−5 | 1.31 10−3 |
| FB | 2 | miR-275-5p | 2208 | 752 | −2.94 | 4.39 10−4 | 9.22 10−3 |
| FB | 2 | miR-8-5p | 9926 | 5071 | −1.96 | 2.41 10−4 | 6.32 10−3 |
| FB | 2 | miR-2945-3p | 9070 | 14,292 | 1.58 | 1.40 10−3 | 0.02 |
| FB | 2 | miR-9a-5p | 4779 | 8276 | 1.73 | 1.56 10−3 | 0.02 |
| FB | 2 | miR-988-3p | 1340 | 2062 | −1.53 | 1.68 10−3 | 0.02 |
| FB | 2 | miR-34-3p | 852 | 523 | −1.63 | 1.69 10−3 | 0.02 |
| FB | 2 | miR-184-3p | 149,602 | 93,549 | −1.60 | 1.83 10−3 | 0.02 |
| FB | 2 | miR-2940-5p | 29,034 | 16,911 | −1.72 | 1.89 10−3 | 0.02 |
| FB | 2 | miR-989-3p | 11,468 | 67,138 | 5.85 | 2.18 10−3 | 0.02 |
| FB | 12 | miR-71-3p | 498 | 366 | −2.01 | 3.85 10−5 | 9.58 10−3 |
| FB | 12 | miR-2940-3p | 11,495 | 22,203 | 1.93 | 4.19 10−4 | 0.05 |
| MG | 2 | miR-9b-5p | 9130 | 15,236 | 1.66 | 1.67 10−4 | 0.02 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sinclair, J.B.; Asgari, S. Ross River Virus Provokes Differentially Expressed MicroRNA and RNA Interference Responses in Aedes aegypti Mosquitoes. Viruses 2020, 12, 695. https://doi.org/10.3390/v12070695
Sinclair JB, Asgari S. Ross River Virus Provokes Differentially Expressed MicroRNA and RNA Interference Responses in Aedes aegypti Mosquitoes. Viruses. 2020; 12(7):695. https://doi.org/10.3390/v12070695
Chicago/Turabian StyleSinclair, James B., and Sassan Asgari. 2020. "Ross River Virus Provokes Differentially Expressed MicroRNA and RNA Interference Responses in Aedes aegypti Mosquitoes" Viruses 12, no. 7: 695. https://doi.org/10.3390/v12070695
APA StyleSinclair, J. B., & Asgari, S. (2020). Ross River Virus Provokes Differentially Expressed MicroRNA and RNA Interference Responses in Aedes aegypti Mosquitoes. Viruses, 12(7), 695. https://doi.org/10.3390/v12070695





