No Evidence for a Role for Antibodies during Vaccination-Induced Enhancement of Porcine Reproductive and Respiratory Syndrome
Abstract
1. Introduction
2. Materials and Methods
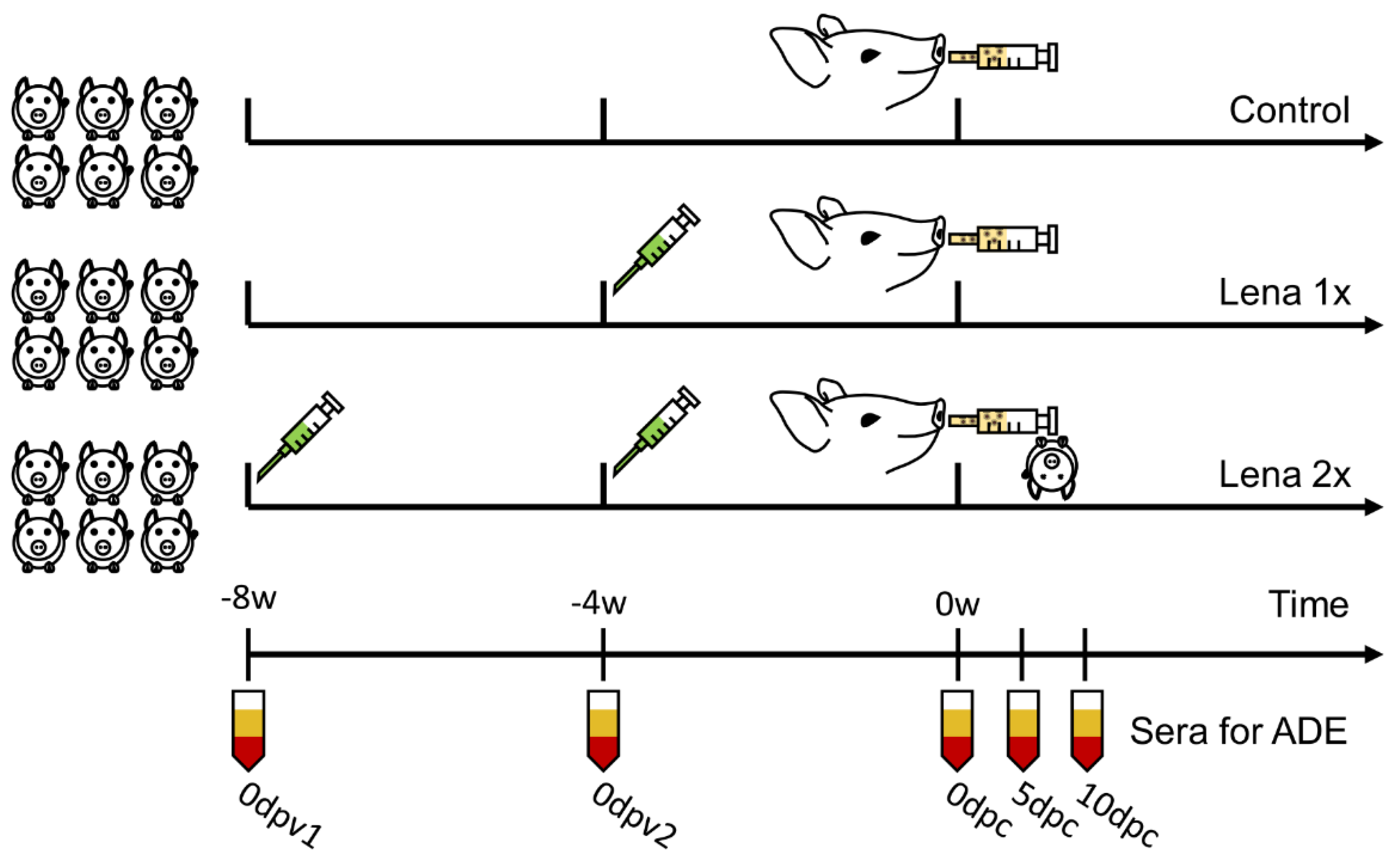
2.1. Animal Experiments
2.2. Monocyte-Derived Macrophages
2.3. Virus Preparation
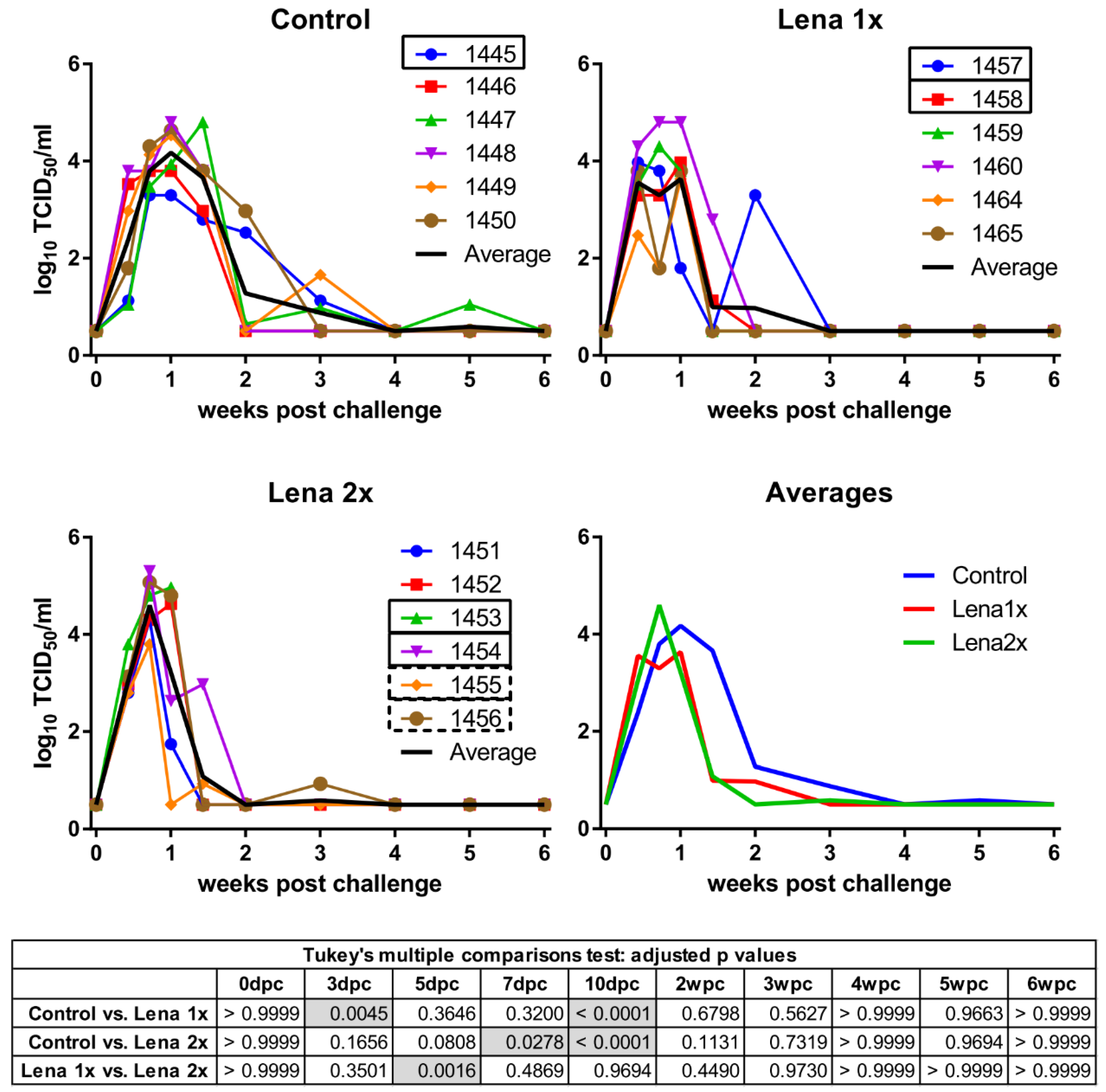
2.4. Quantification of Viremia and Viral Shedding In Vitro
2.5. Quantification of Neutralizing Antibodies in Positive Control Sera
2.6. Antibody Responses during Vaccination Experiment
2.7. ADE of Macrophage Infection
2.8. Flow Cytometry
2.9. ELISA
2.10. Statistical Analysis
3. Results
3.1. Viremia and PRRSV Neutralization Titers after Infection of Naïve Animals for Generation of the Reference Immune Serum
3.2. Vaccination with an Inactivated Virus and Homologous Challenge
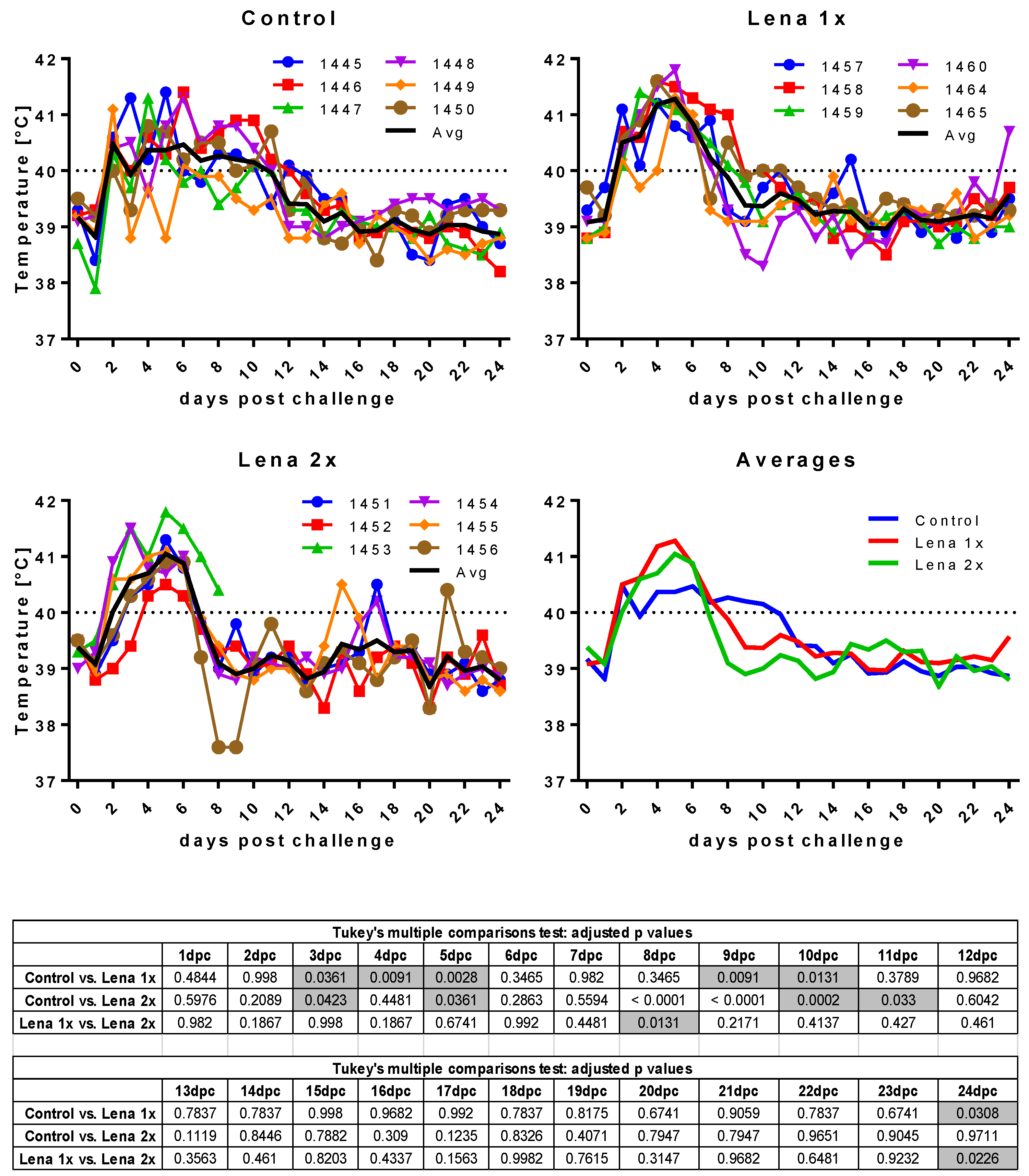
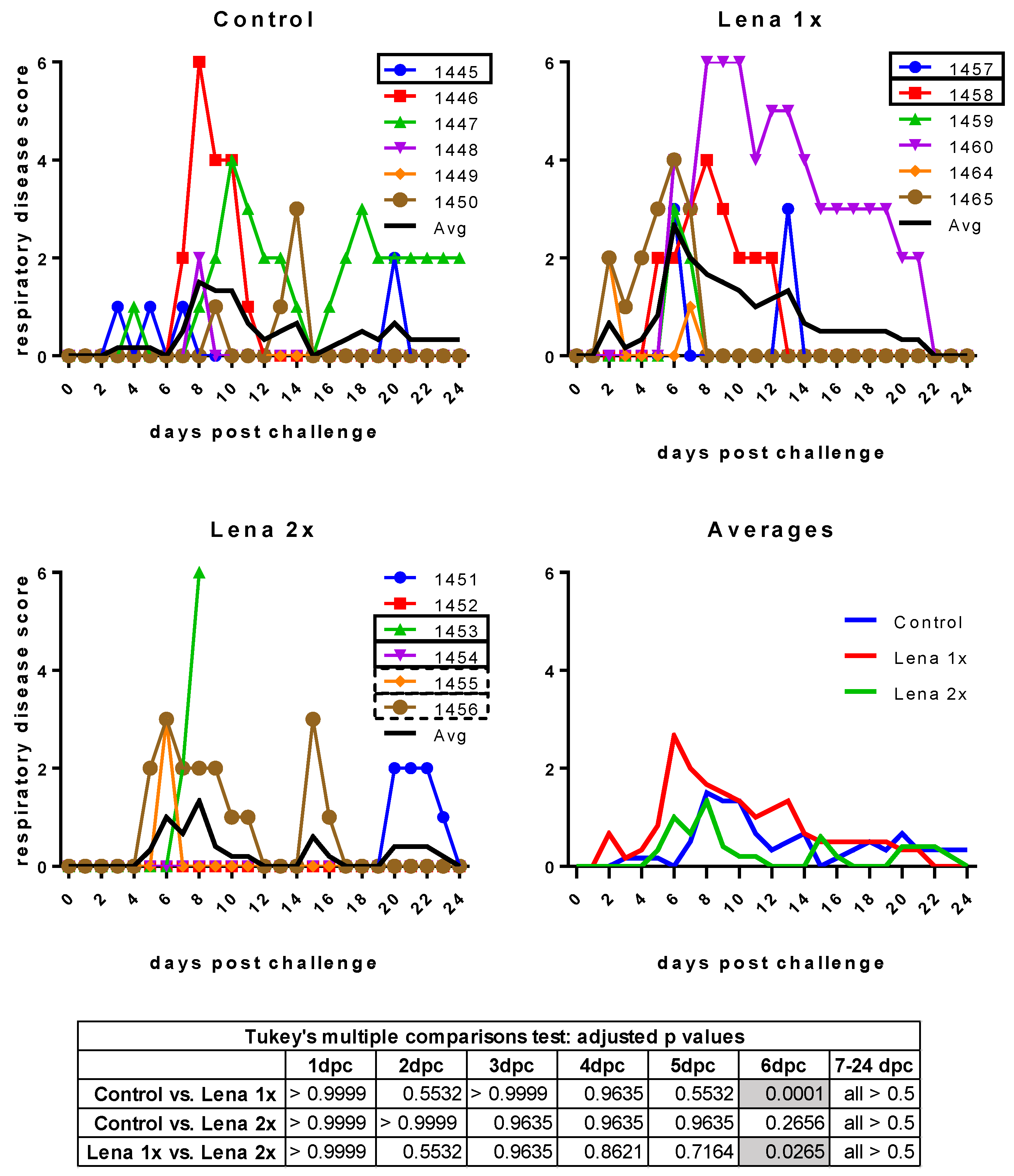
3.3. Body Temperature and Respiratory Disease Score
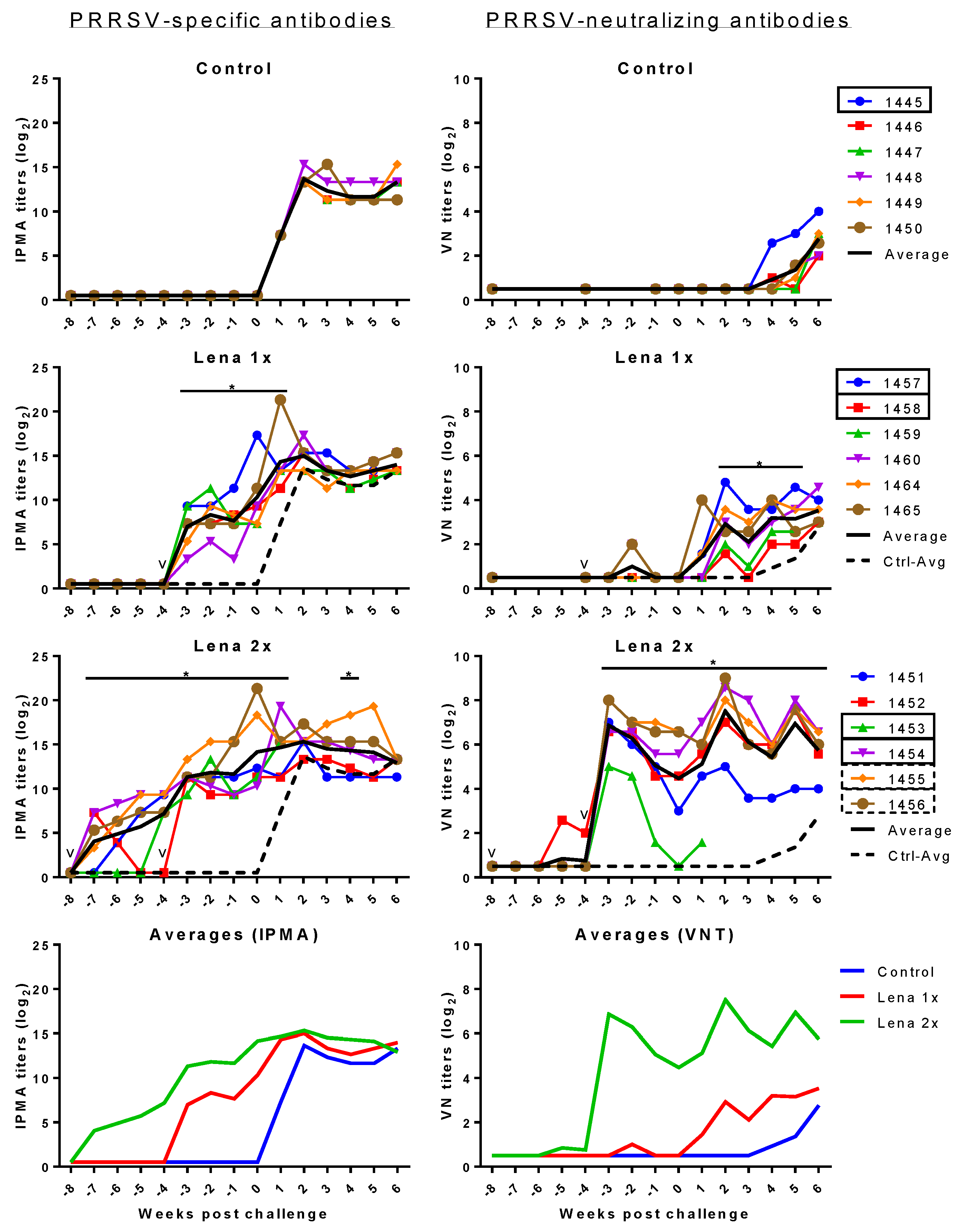
3.4. PRRSV-Specific and PRRSV-Neutralizing Antibody Titers
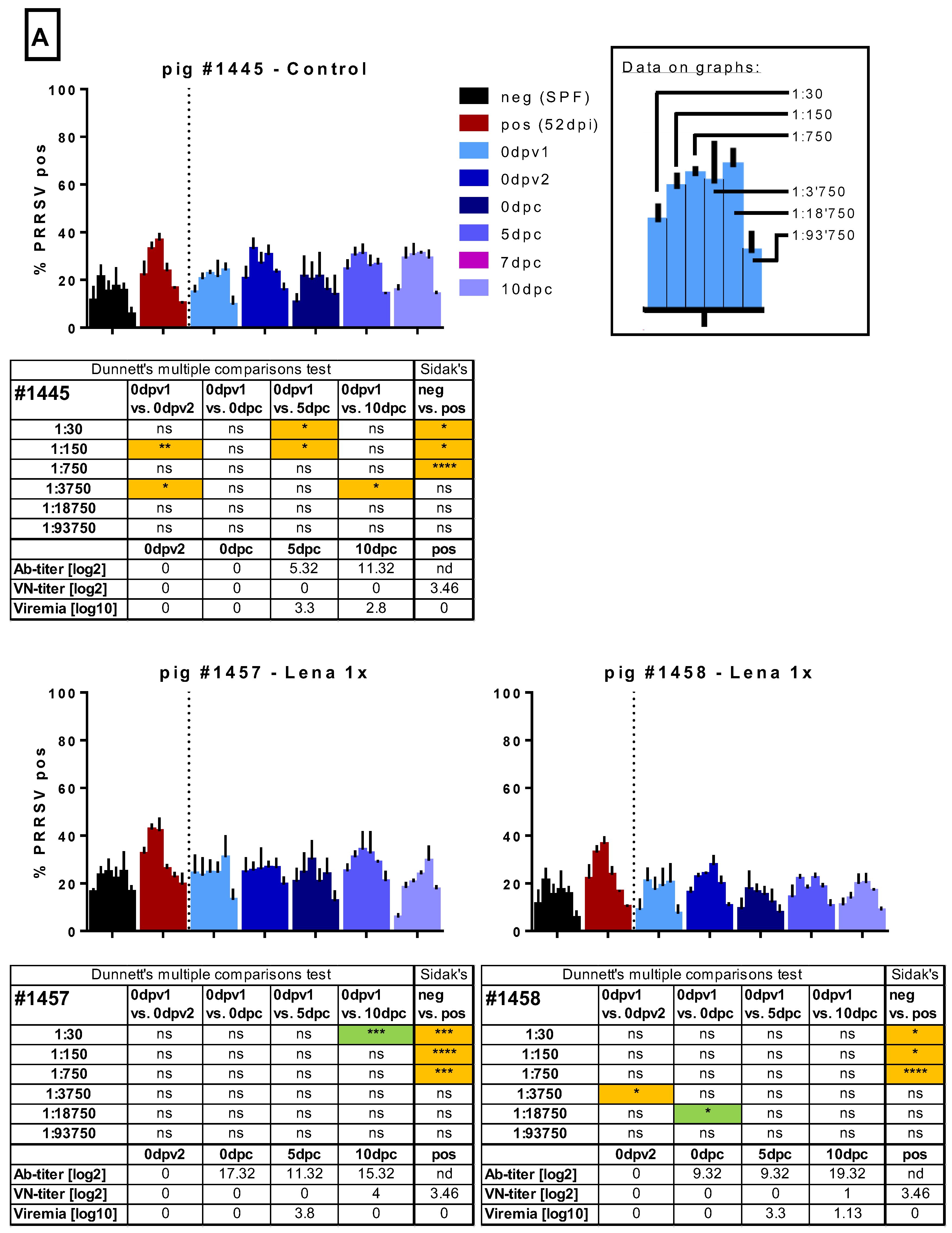
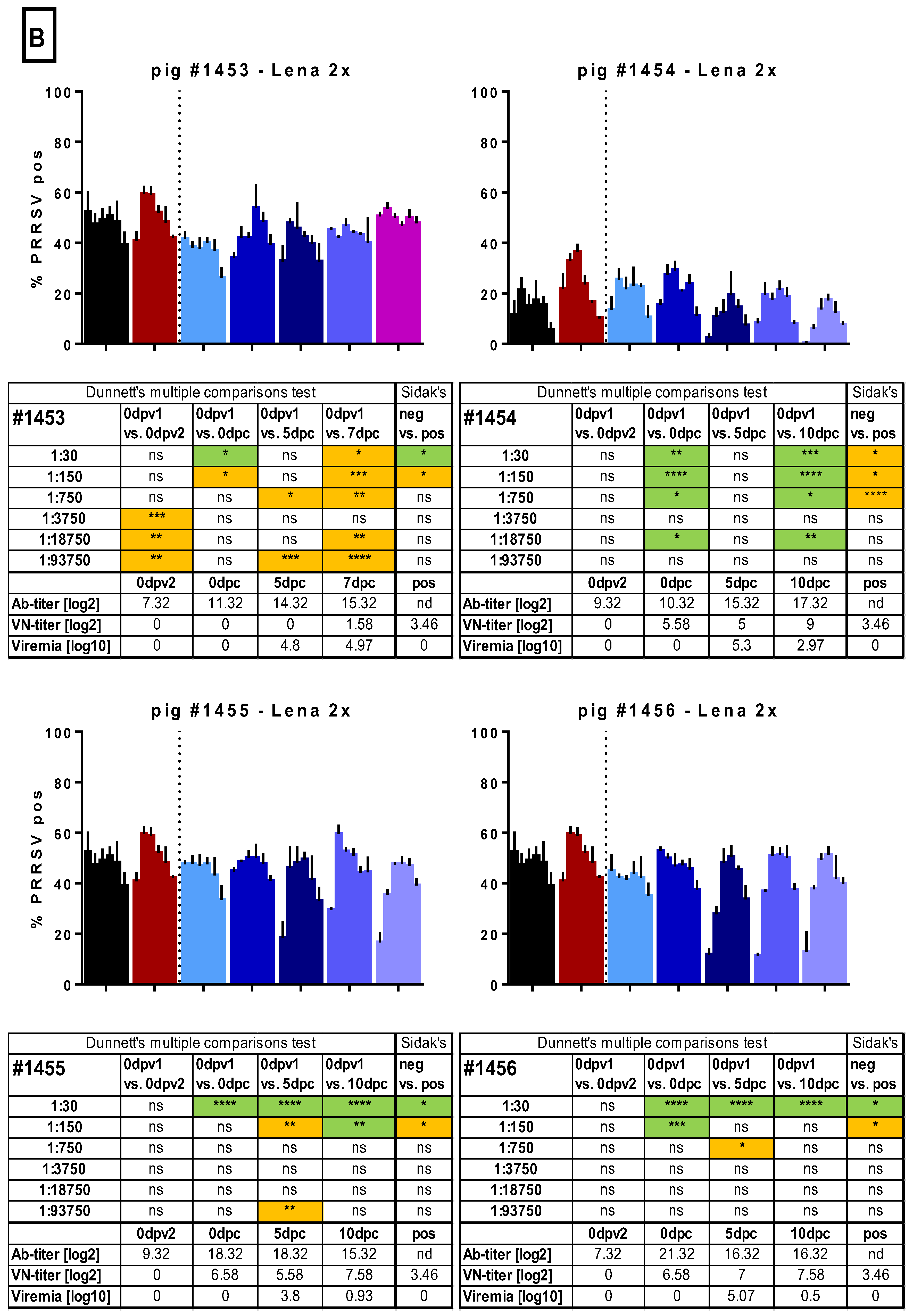
3.5. ADE of Macrophage Infection
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
References
- Cafruny, W.A.; Plagemann, P.G. Immune response to lactate dehydrogenase-elevating virus: Serologically specific rabbit neutralizing antibody to the virus. Infect. Immun. 1982, 37, 1007–1012. [Google Scholar]
- Takeda, A.; Tuazon, C.U.; Ennis, F.A. Antibody-enhanced infection by hiv-1 via fc receptor-mediated entry. Science 1988, 242, 580–583. [Google Scholar] [CrossRef]
- Homsy, J.; Meyer, M.; Tateno, M.; Clarkson, S.; Levy, J.A. The fc and not cd4 receptor mediates antibody enhancement of hiv infection in human cells. Science 1989, 244, 1357–1360. [Google Scholar] [CrossRef]
- Halstead, S.B.; Porterfield, J.S.; O’Rourke, E.J. Enhancement of dengue virus infection in monocytes by flavivirus antisera. Am. J. Trop. Med. Hyg. 1980, 29, 638–642. [Google Scholar] [CrossRef]
- Halstead, S.B. Dengue antibody-dependent enhancement: Knowns and unknowns. Microbiol. Spectr. 2014, 2. [Google Scholar] [CrossRef]
- Garcia-Nicolas, O.; Ricklin, M.E.; Liniger, M.; Vielle, N.J.; Python, S.; Souque, P.; Charneau, P.; Summerfield, A. A japanese encephalitis virus vaccine inducing antibodies strongly enhancing in vitro infection is protective in pigs. Viruses 2017, 9, 124. [Google Scholar] [CrossRef]
- Mason, P.W.; Baxt, B.; Brown, F.; Harber, J.; Murdin, A.; Wimmer, E. Antibody-complexed foot-and-mouth disease virus, but not poliovirus, can infect normally insusceptible cells via the fc receptor. Virology 1993, 192, 568–577. [Google Scholar] [CrossRef]
- Baxt, B.; Mason, P.W. Foot-and-mouth disease virus undergoes restricted replication in macrophage cell cultures following fc receptor-mediated adsorption. Virology 1995, 207, 503–509. [Google Scholar] [CrossRef]
- Lannes, N.; Python, S.; Summerfield, A. Interplay of foot-and-mouth disease virus, antibodies and plasmacytoid dendritic cells: Virus opsonization under non-neutralizing conditions results in enhanced interferon-alpha responses. Vet. Res. 2012, 43, 64. [Google Scholar] [CrossRef]
- Boonnak, K.; Dambach, K.M.; Donofrio, G.C.; Tassaneetrithep, B.; Marovich, M.A. Cell type specificity and host genetic polymorphisms influence antibody-dependent enhancement of dengue virus infection. J. Virol. 2011, 85, 1671–1683. [Google Scholar] [CrossRef]
- Kou, Z.; Lim, J.Y.; Beltramello, M.; Quinn, M.; Chen, H.; Liu, S.; Martinez-Sobrido, L.; Diamond, M.S.; Schlesinger, J.J.; de Silva, A.; et al. Human antibodies against dengue enhance dengue viral infectivity without suppressing type i interferon secretion in primary human monocytes. Virology 2011, 410, 240–247. [Google Scholar] [CrossRef]
- Lidbury, B.A.; Mahalingam, S. Specific ablation of antiviral gene expression in macrophages by antibody-dependent enhancement of ross river virus infection. J. Virol. 2000, 74, 8376–8381. [Google Scholar] [CrossRef]
- Suhrbier, A.; La Linn, M. Suppression of antiviral responses by antibody-dependent enhancement of macrophage infection. Trends Immunol. 2003, 24, 165–168. [Google Scholar] [CrossRef]
- Yoon, K.J.; Wu, L.L.; Zimmerman, J.J.; Hill, H.T.; Platt, K.B. Antibody-dependent enhancement (ade) of porcine reproductive and respiratory syndrome virus (prrsv) infection in pigs. Viral Immunol. 1996, 9, 51–63. [Google Scholar] [CrossRef]
- Yoon, K.J.; Wu, L.L.; Zimmerman, J.J.; Platt, K.B. Field isolates of porcine reproductive and respiratory syndrome virus (prrsv) vary in their susceptibility to antibody dependent enhancement (ade) of infection. Vet. Microbiol. 1997, 55, 277–287. [Google Scholar] [CrossRef]
- Qiao, S.; Jiang, Z.; Tian, X.; Wang, R.; Xing, G.; Wan, B.; Bao, D.; Liu, Y.; Hao, H.; Guo, J.; et al. Porcine fcgammariib mediates enhancement of porcine reproductive and respiratory syndrome virus (PRRSV) infection. PLoS ONE 2011, 6, e28721. [Google Scholar] [CrossRef]
- Gu, W.; Guo, L.; Yu, H.; Niu, J.; Huang, M.; Luo, X.; Li, R.; Tian, Z.; Feng, L.; Wang, Y. Involvement of cd16 in antibody-dependent enhancement of porcine reproductive and respiratory syndrome virus infection. J. Gen. Virol. 2015, 96, 1712–1722. [Google Scholar] [CrossRef]
- Shi, P.; Su, Y.; Li, Y.; Zhang, L.; Lu, D.; Li, R.; Zhang, L.; Huang, J. The alternatively spliced porcine fcgammari regulated prrsv-ade infection and proinflammatory cytokine production. Dev. Comp. Immunol. 2019, 90, 186–198. [Google Scholar] [CrossRef]
- Cancel-Tirado, S.M.; Evans, R.B.; Yoon, K.J. Monoclonal antibody analysis of porcine reproductive and respiratory syndrome virus epitopes associated with antibody-dependent enhancement and neutralization of virus infection. Vet. Immunol. Immunopathol. 2004, 102, 249–262. [Google Scholar] [CrossRef]
- Loving, C.L.; Osorio, F.A.; Murtaugh, M.P.; Zuckermann, F.A. Innate and adaptive immunity against porcine reproductive and respiratory syndrome virus. Vet. Immunol. Immunopathol. 2015, 167, 1–14. [Google Scholar] [CrossRef]
- Rahe, M.C.; Murtaugh, M.P. Mechanisms of adaptive immunity to porcine reproductive and respiratory syndrome virus. Viruses 2017, 9, 148. [Google Scholar] [CrossRef]
- Rahe, M.C.; Murtaugh, M.P. Effector mechanisms of humoral immunity to porcine reproductive and respiratory syndrome virus. Vet. Immunol. Immunopathol. 2017, 186, 15–18. [Google Scholar] [CrossRef]
- Karniychuk, U.U.; Geldhof, M.; Vanhee, M.; Van Doorsselaere, J.; Saveleva, T.A.; Nauwynck, H.J. Pathogenesis and antigenic characterization of a new east european subtype 3 porcine reproductive and respiratory syndrome virus isolate. BMC Vet. Res. 2010, 6, 30. [Google Scholar] [CrossRef]
- Vanhee, M.; Delputte, P.L.; Delrue, I.; Geldhof, M.F.; Nauwynck, H.J. Development of an experimental inactivated prrsv vaccine that induces virus-neutralizing antibodies. Vet. Res. 2009, 40, 63. [Google Scholar] [CrossRef]
- Garcia-Nicolas, O.; Auray, G.; Sautter, C.A.; Rappe, J.C.; McCullough, K.C.; Ruggli, N.; Summerfield, A. Sensing of porcine reproductive and respiratory syndrome virus-infected macrophages by plasmacytoid dendritic cells. Front. Microbiol. 2016, 7, 771. [Google Scholar] [CrossRef]
- Sautter, C.A.; Auray, G.; Python, S.; Liniger, M.; Summerfield, A. Phenotypic and functional modulations of porcine macrophages by interferons and interleukin-4. Dev. Comp. Immunol. 2018, 84, 181–192. [Google Scholar] [CrossRef]
- Van Breedam, W.; Costers, S.; Vanhee, M.; Gagnon, C.A.; Rodriguez-Gomez, I.M.; Geldhof, M.; Verbeeck, M.; Van Doorsselaere, J.; Karniychuk, U.; Nauwynck, H.J. Porcine reproductive and respiratory syndrome virus (prrsv)-specific mabs: Supporting diagnostics and providing new insights into the antigenic properties of the virus. Vet. Immunol. Immunopathol. 2011, 141, 246–257. [Google Scholar] [CrossRef]
- Delrue, I.; Van Gorp, H.; Van Doorsselaere, J.; Delputte, P.L.; Nauwynck, H.J. Susceptible cell lines for the production of porcine reproductive and respiratory syndrome virus by stable transfection of sialoadhesin and cd163. BMC Biotechnol. 2010, 10, 48. [Google Scholar] [CrossRef]
- Decorte, I.; Van Breedam, W.; Van der Stede, Y.; Nauwynck, H.J.; De Regge, N.; Cay, A.B. Detection of total and prrsv-specific antibodies in oral fluids collected with different rope types from prrsv-vaccinated and experimentally infected pigs. BMC Vet. Res. 2014, 10, 134. [Google Scholar] [CrossRef]
- Labarque, G.G.; Nauwynck, H.J.; Van Reeth, K.; Pensaert, M.B. Effect of cellular changes and onset of humoral immunity on the replication of porcine reproductive and respiratory syndrome virus in the lungs of pigs. J. Gen. Virol. 2000, 81, 1327–1334. [Google Scholar] [CrossRef]
- Geldhof, M.F.; Vanhee, M.; Van Breedam, W.; Van Doorsselaere, J.; Karniychuk, U.U.; Nauwynck, H.J. Comparison of the efficacy of autogenous inactivated porcine reproductive and respiratory syndrome virus (prrsv) vaccines with that of commercial vaccines against homologous and heterologous challenges. BMC Vet. Res. 2012, 8, 182. [Google Scholar] [CrossRef]
- Osorio, F.A.; Galeota, J.A.; Nelson, E.; Brodersen, B.; Doster, A.; Wills, R.; Zuckermann, F.; Laegreid, W.W. Passive transfer of virus-specific antibodies confers protection against reproductive failure induced by a virulent strain of porcine reproductive and respiratory syndrome virus and establishes sterilizing immunity. Virology 2002, 302, 9–20. [Google Scholar] [CrossRef]
- Lopez, O.J.; Oliveira, M.F.; Garcia, E.A.; Kwon, B.J.; Doster, A.; Osorio, F.A. Protection against porcine reproductive and respiratory syndrome virus (prrsv) infection through passive transfer of prrsv-neutralizing antibodies is dose dependent. Clin. Vaccine Immunol. 2007, 14, 269–275. [Google Scholar] [CrossRef]
- Robinson, S.R.; Rahe, M.C.; Gray, D.K.; Martins, K.V.; Murtaugh, M.P. Porcine reproductive and respiratory syndrome virus neutralizing antibodies provide in vivo cross-protection to prrsv1 and prrsv2 viral challenge. Virus Res. 2018, 248, 13–23. [Google Scholar] [CrossRef]
- Geldhof, M.F.; Van Breedam, W.; De Jong, E.; Lopez Rodriguez, A.; Karniychuk, U.U.; Vanhee, M.; Van Doorsselaere, J.; Maes, D.; Nauwynck, H.J. Antibody response and maternal immunity upon boosting prrsv-immune sows with experimental farm-specific and commercial prrsv vaccines. Vet. Microbiol. 2013, 167, 260–271. [Google Scholar] [CrossRef]
- Cardosa, M.J.; Porterfield, J.S.; Gordon, S. Complement receptor mediates enhanced flavivirus replication in macrophages. J. Exp. Med. 1983, 158, 258–263. [Google Scholar] [CrossRef]
- Prohaszka, Z.; Nemes, J.; Hidvegi, T.; Toth, F.D.; Kerekes, K.; Erdei, A.; Szabo, J.; Ujhelyi, E.; Thielens, N.; Dierich, M.P.; et al. Two parallel routes of the complement-mediated antibody-dependent enhancement of hiv-1 infection. AIDS 1997, 11, 949–958. [Google Scholar] [CrossRef]
- Robinson, W.E., Jr.; Montefiori, D.C.; Mitchell, W.M. Complement-mediated antibody-dependent enhancement of hiv-1 infection requires cd4 and complement receptors. Virology 1990, 175, 600–604. [Google Scholar] [CrossRef]
- Bordet, E.; Blanc, F.; Tiret, M.; Crisci, E.; Bouguyon, E.; Renson, P.; Maisonnasse, P.; Bourge, M.; Leplat, J.J.; Giuffra, E.; et al. Porcine reproductive and respiratory syndrome virus type 1.3 lena triggers conventional dendritic cells 1 activation and t helper 1 immune response without infecting dendritic cells. Front. Immunol. 2018, 9, 2299. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sautter, C.A.; Trus, I.; Nauwynck, H.; Summerfield, A. No Evidence for a Role for Antibodies during Vaccination-Induced Enhancement of Porcine Reproductive and Respiratory Syndrome. Viruses 2019, 11, 829. https://doi.org/10.3390/v11090829
Sautter CA, Trus I, Nauwynck H, Summerfield A. No Evidence for a Role for Antibodies during Vaccination-Induced Enhancement of Porcine Reproductive and Respiratory Syndrome. Viruses. 2019; 11(9):829. https://doi.org/10.3390/v11090829
Chicago/Turabian StyleSautter, Carmen A., Ivan Trus, Hans Nauwynck, and Artur Summerfield. 2019. "No Evidence for a Role for Antibodies during Vaccination-Induced Enhancement of Porcine Reproductive and Respiratory Syndrome" Viruses 11, no. 9: 829. https://doi.org/10.3390/v11090829
APA StyleSautter, C. A., Trus, I., Nauwynck, H., & Summerfield, A. (2019). No Evidence for a Role for Antibodies during Vaccination-Induced Enhancement of Porcine Reproductive and Respiratory Syndrome. Viruses, 11(9), 829. https://doi.org/10.3390/v11090829





