Therapeutic Potential of an Endolysin Derived from Kayvirus S25-3 for Staphylococcal Impetigo
Abstract
1. Introduction
2. Materials and Methods
2.1. Production and Purification of Recombinant Protein
2.2. Bacterial Strains
2.3. Bacterial Isolation from Human Skin
2.4. Phylogenetic Classification of Bacteria
2.5. Measurement of the Bacteriolytic and Bactericidal Activities
2.6. Application of S25-3LYS in a Mouse Model of Impetigo
2.7. Enumeration of Bacterial Densities in the Mouse Pinnae
2.8. Histopathological and Immunofluorescence Analyses
2.9. Microbiota Analysis
2.10. Microbiome Analysis
2.11. Statistical Analysis
3. Results and Discussion
3.1. Characteristics of the Endolysin Derived from Kayvirus S25-3
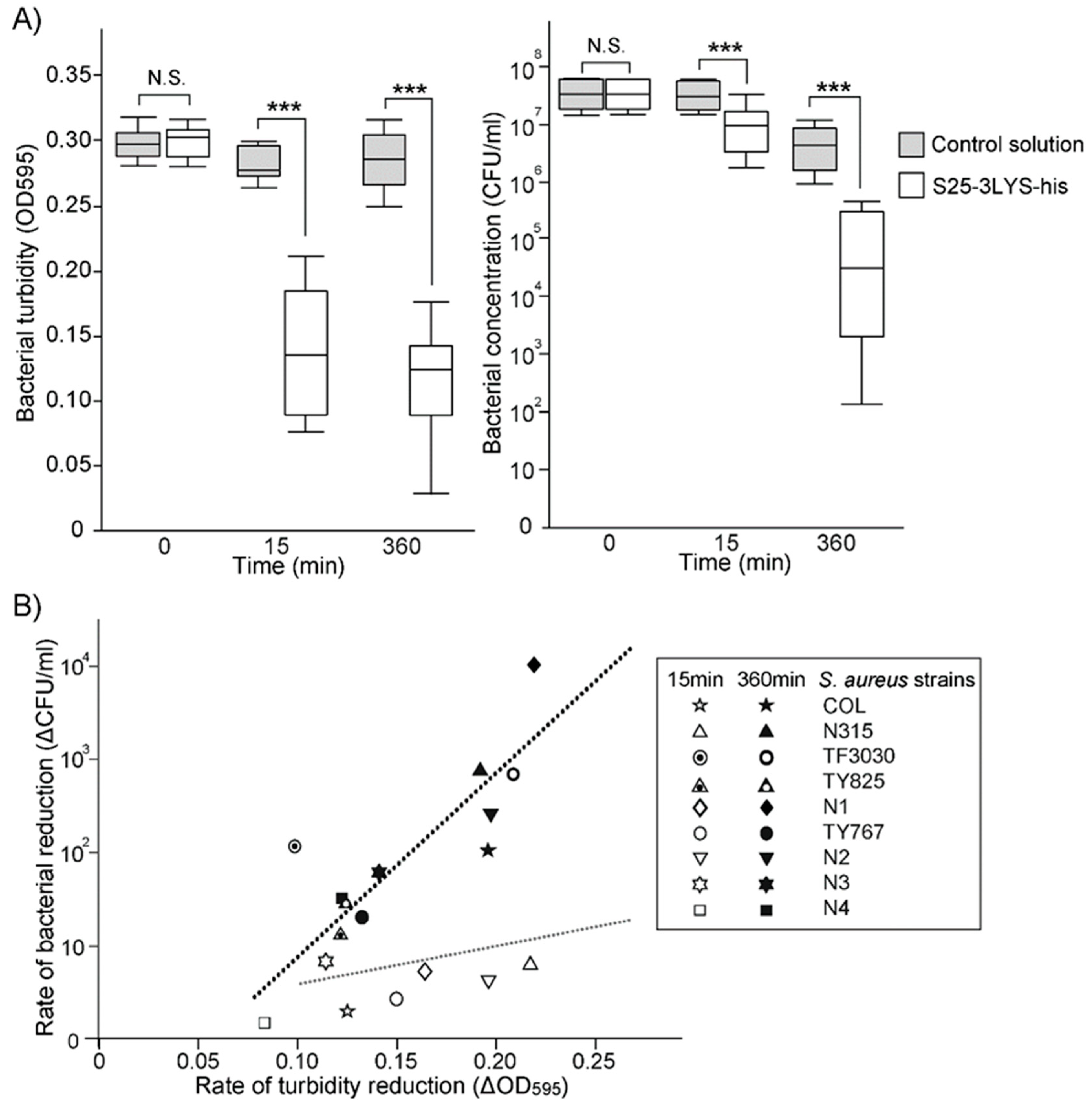
3.2. Prolonged Incubation of S25-3 Endolysin with S. aureus Increases the Bactericidal Effect
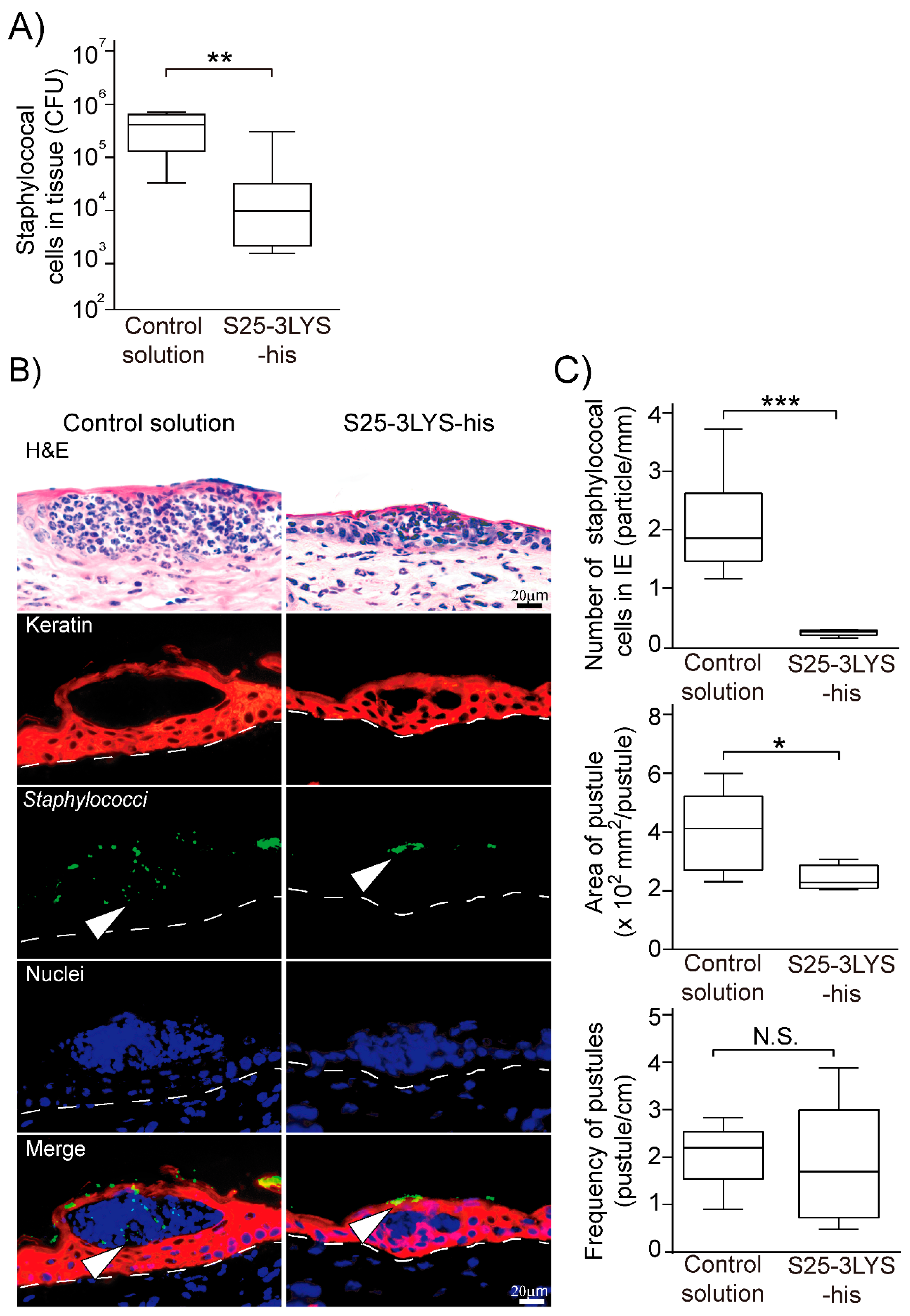
3.3. Topical Application of S25-3 Endolysin Inhibits Staphylococcal Epidermal Invasion in Experimental Staphylococcal Impetigo
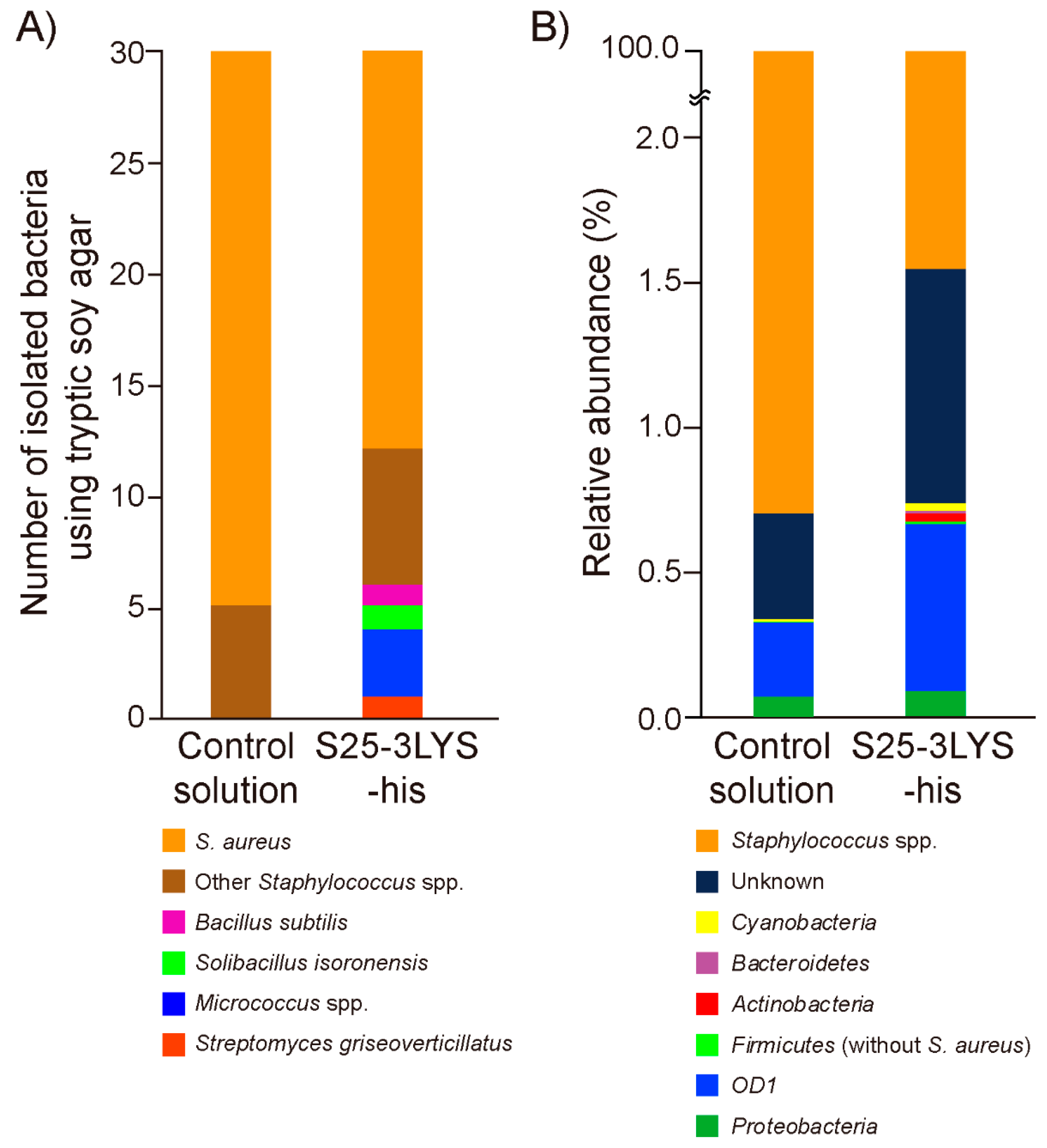
3.4. S25-3LYS-his Treatment Increases Bacterial Diversity in Mouse Skin
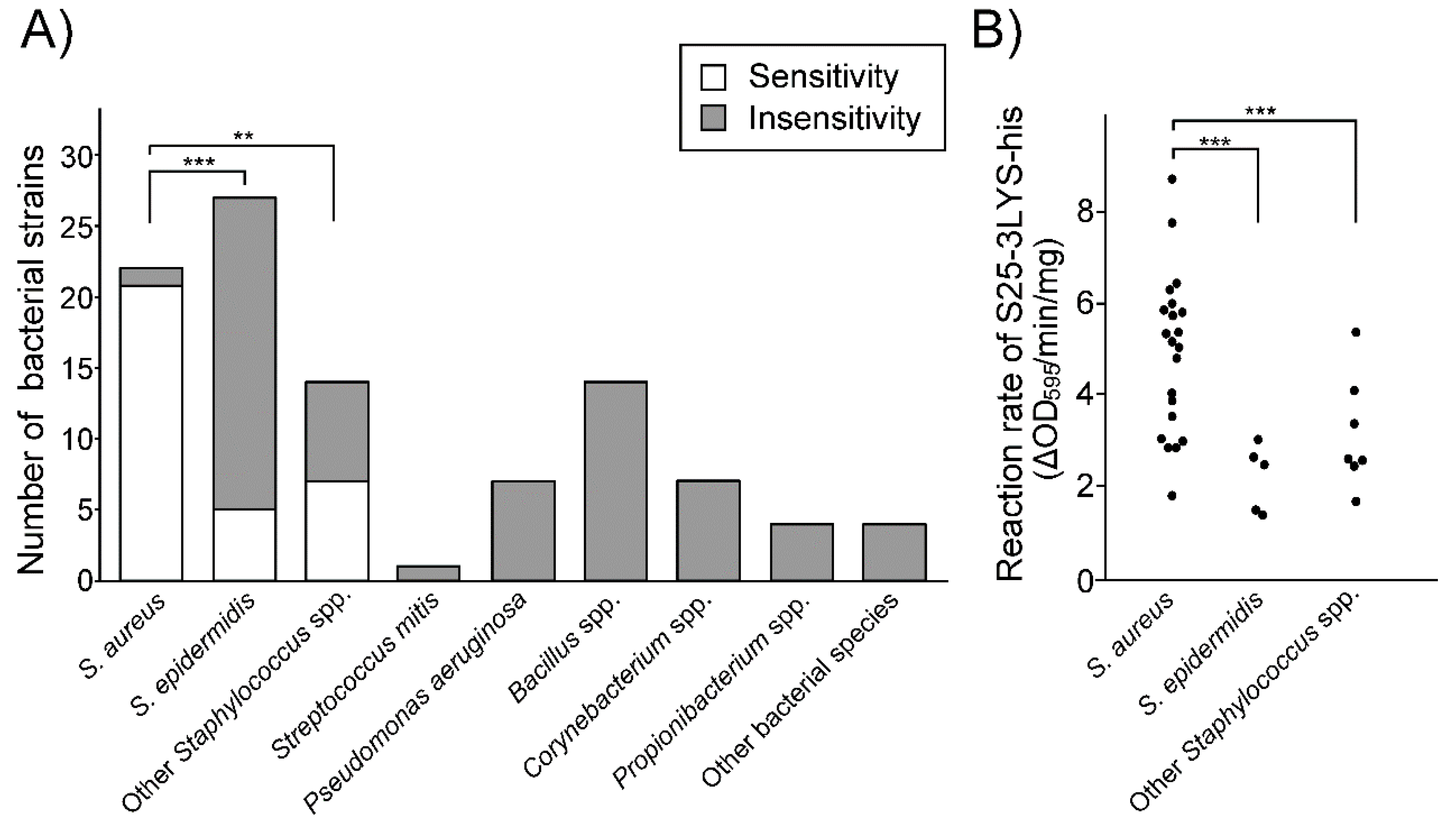
3.5. S25-3 Endolysin Preferentially Lyses S. aureus among Human Skin Bacteria
3.6. Future Prospects for Endolysin Application to Treat Staphylococcal Skin Infections
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- D’Cunha, N.M.; Peterson, G.M.; Baby, K.E.; Thomas, J. Impetigo: A need for new therapies in a world of increasing antimicrobial resistance. J. Clin. Pharm. Ther. 2018, 43, 150–153. [Google Scholar] [CrossRef] [PubMed]
- Bowen, A.C.; Mahe, A.; Hay, R.J.; Andrews, R.M.; Steer, A.C.; Tong, S.Y.; Carapetis, J.R. The global epidemiology of impetigo: A systematic review of the population prevalence of impetigo and pyoderma. PLoS ONE 2015, 10, e0136789. [Google Scholar] [CrossRef] [PubMed]
- Hartman-Adams, H.; Banvard, C.; Juckett, G. Impetigo: Diagnosis and treatment. Am. Fam. Physician 2014, 90, 229–235. [Google Scholar] [PubMed]
- Pereira, L.B. Impetigo—Review. Anais Brasileiros de Dermatologia 2014, 89, 293–299. [Google Scholar] [CrossRef] [PubMed]
- Occelli, P.; Blanie, M.; Sanchez, R.; Vigier, D.; Dauwalder, O.; Darwiche, A.; Provenzano, B.; Dumartin, C.; Parneix, P.; Venier, A.G. Outbreak of Staphylococcal bullous impetigo in a maternity ward linked to an asymptomatic healthcare worker. J. Hosp. Infect. 2007, 67, 264–270. [Google Scholar] [CrossRef] [PubMed]
- Williamson, D.A.; Carter, G.P.; Howden, B.P. Current and emerging topical antibacterials and antiseptics: Agents, action, and resistance patterns. Clin. Microbiol. Rev. 2017, 30, 827–860. [Google Scholar] [CrossRef]
- Rijnders, M.I.; Wolffs, P.F.; Hopstaken, R.M.; den Heyer, M.; Bruggeman, C.A.; Stobberingh, E.E. Spread of the epidemic European fusidic acid-resistant impetigo clone (eefic) in general practice patients in the south of the Netherlands. J. Antimicrob. Chemother. 2012, 67, 1176–1180. [Google Scholar] [CrossRef]
- SanMiguel, A.J.; Meisel, J.S.; Horwinski, J.; Zheng, Q.; Grice, E.A. Topical antimicrobial treatments can elicit shifts to resident skin bacterial communities and reduce colonization by Staphylococcus aureus competitors. Antimicrob. Agents Chemother. 2017, 61, e00774–e00817. [Google Scholar] [CrossRef]
- Gutiérrez, D.F.L.; Rodríguez, A.; García, P. Are phage lytic proteins the secret weapon to kill Staphylococcus aureus? mBio 2018, 9, e01923–e02017. [Google Scholar] [CrossRef]
- Haddad Kashani, H.; Schmelcher, M.; Sabzalipoor, H.; Seyed Hosseini, E.; Moniri, R. Recombinant endolysins as potential therapeutics against antibiotic-resistant Staphylococcus aureus: Current status of research and novel delivery strategies. Clin. Microbiol. Rev. 2018, 31, e00071–e00117. [Google Scholar] [CrossRef]
- Szweda, P.; Schielmann, M.; Kotlowski, R.; Gorczyca, G.; Zalewska, M.; Milewski, S. Peptidoglycan hydrolases-potential weapons against Staphylococcus aureus. Appl. Microbiol. Biotechnol. 2012, 96, 1157–1174. [Google Scholar] [CrossRef] [PubMed]
- Love, M.J.; Bhandari, D.; Dobson, R.C.J.; Billington, C. Potential for bacteriophage endolysins to supplement or replace antibiotics in food production and clinical care. Antibiotics 2018, 7, 17. [Google Scholar] [CrossRef] [PubMed]
- Fischetti, V.A. Bacteriophage endolysins: A novel anti-infective to control gram-positive pathogens. Int. J. Med. Microbiol. 2010, 300, 357–362. [Google Scholar] [CrossRef] [PubMed]
- Cisek, A.A.; Dabrowska, I.; Gregorczyk, K.P.; Wyzewski, Z. Phage therapy in bacterial infections treatment: One hundred years after the discovery of bacteriophages. Curr. Microbiol. 2017, 74, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Li, D.; Li, X.; Hu, L.; Cheng, M.; Xia, F.; Gong, P.; Wang, B.; Ge, J.; Zhang, H.; et al. Lysgh15 kills Staphylococcus aureus without being affected by the humoral immune response or inducing inflammation. Sci. Rep. 2016, 6, 29344. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Zhang, H.; Wang, J.; Yu, J.; Wei, H. A novel chimeric lysin with robust antibacterial activity against planktonic and biofilm methicillin-resistant Staphylococcus aureus. Sci. Rep. 2017, 7, 40182. [Google Scholar] [CrossRef] [PubMed]
- Totte, J.E.E.; van Doorn, M.B.; Pasmans, S. Successful treatment of chronic Staphylococcus aureus-related dermatoses with the topical endolysin staphefekt sa.100: A report of 3 cases. Case Rep. Dermatol. 2017, 9, 19–25. [Google Scholar] [CrossRef]
- De Wit, J.; Totte, J.E.E.; van Mierlo, M.M.F.; van Veldhuizen, J.; van Doorn, M.B.A.; Schuren, F.H.J.; Willemsen, S.P.; Pardo, L.M.; Pasmans, S. Endolysin treatment against Staphylococcus aureus in adults with atopic dermatitis: A randomized controlled trial. J. Allergy Clin. Immunol. 2019, 27, 30691–30698. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, H.; Melo, L.; Santos, S.; Nóbrega, F.L.; Ferreira, E.; Cerca, N.; Azeredo, J.; Kluskens, L.D. Molecular aspects and comparative genomics of bacteriophage endolysins. J. Virol. 2013, 87, 4558–4570. [Google Scholar] [CrossRef]
- Totté, J.; de Wit, J.; Pardo, L.; Schuren, F.; van Doorn, M.; Pasmans, S. Targeted anti-staphylococcal therapy with endolysins in atopic dermatitis and the effect on steroid use, disease severity and the microbiome: Study protocol for a randomized controlled trial (maas trial). Trials 2017, 18, 404. [Google Scholar] [CrossRef]
- Kim, H.B.; Park, B.W. Phase iia Clinical Study of n-rephasin® sal200; ClinicalTrials.Gov Identifier: NCT03089697. Available online: https://clinicaltrials.gov/ct2/show/NCT03089697 (accessed on 10 July 2018).
- Takemura-Uchiyama, I.; Uchiyama, J.; Kato, S.-I.; Ujihara, T.; Daibata, M.; Matsuzaki, S. Genomic and phylogenetic traits of Staphylococcus phages s25-3 and s25-4 (family myoviridae, genus twort-like viruses). Ann. Microbiol. 2013, 64, 1453–1456. [Google Scholar] [CrossRef]
- Takemura-Uchiyama, I.; Uchiyama, J.; Kato, S.; Inoue, T.; Ujihara, T.; Ohara, N.; Daibata, M.; Matsuzaki, S. Evaluating efficacy of bacteriophage therapy against Staphylococcus aureus infections using a silkworm larval infection model. FEMS Microbiol. Lett. 2013, 347, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.R.; Fouts, D.E.; Archer, G.L.; Mongodin, E.F.; Deboy, R.T.; Ravel, J.; Paulsen, I.T.; Kolonay, J.F.; Brinkac, L.; Beanan, M.; et al. Insights on evolution of virulence and resistance from the complete genome analysis of an early methicillin-resistant Staphylococcus aureus strain and a biofilm-producing methicillin-resistant Staphylococcus epidermidis strain. J. Bacteriol. 2005, 187, 2426–2438. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, M.; Ohta, T.; Uchiyama, I.; Baba, T.; Yuzawa, H.; Kobayashi, I.; Cui, L.; Oguchi, A.; Aoki, K.; Nagai, Y.; et al. Whole genome sequencing of meticillin-resistant Staphylococcus aureus. Lancet 2001, 357, 1225–1240. [Google Scholar] [CrossRef]
- Oyama, K.; Kawada-Matsuo, M.; Oogai, Y.; Hayashi, T.; Nakamura, N.; Komatsuzawa, H. Antibacterial effects of glycyrrhetinic acid and its derivatives on Staphylococcus aureus. PloS ONE 2016, 11, e0165831. [Google Scholar] [CrossRef] [PubMed]
- Hisatsune, J.; Hirakawa, H.; Yamaguchi, T.; Fudaba, Y.; Oshima, K.; Hattori, M.; Kato, F.; Kayama, S.; Sugai, M. Emergence of Staphylococcus aureus carrying multiple drug resistance genes on a plasmid encoding exfoliative toxin b. Antimicrob Agents Chemother 2013, 57, 6131–6140. [Google Scholar] [CrossRef] [PubMed]
- Imanishi, I.; Hattori, S.; Hisatsune, J.; Ide, K.; Sugai, M.; Nishifuji, K. Staphylococcus aureus penetrate the interkeratinocyte spaces created by skin-infiltrating neutrophils in a mouse model of impetigo. Vet. Dermatol. 2017, 28, e126–e127. [Google Scholar] [CrossRef]
- Kawamura, Y.; Whiley, R.A.; Shu, S.E.; Ezaki, T.; Hardie, J.M. Genetic approaches to the identification of the mitis group within the genus Streptococcus. Microbiology 1999, 145, 2605–2713. [Google Scholar] [CrossRef][Green Version]
- Uchiyama, J.; Suzuki, M.; Nishifuji, K.; Kato, S.; Miyata, R.; Nasukawa, T.; Yamaguchi, K.; Takemura-Uchiyama, I.; Ujihara, T.; Shimakura, H.; et al. Analyses of short-term antagonistic evolution of Pseudomonas aeruginosa strain pao1 and phage kpp22 (myoviridae family, pb1-like virus genus). Appl. Environ. Microbiol. 2016, 82, 4482–4491. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Haritani, M.; Nakazawa, M.; Hashimoto, K.; Narita, M.; Tagawa, Y.; Nakagawa, M. Immunoperoxidase evaluation of the relationship between necrotic lesions and causative bacteria in lungs of calves with naturally acquired pneumonia. Am J Vet Res. 1990, 51, 1975–1981. [Google Scholar] [PubMed]
- Castelino, M.; Eyre, S.; Moat, J.; Fox, G.; Martin, P.; Ho, P.; Upton, M.; Barton, A. Optimisation of methods for bacterial skin microbiome investigation: Primer selection and comparison of the 454 versus miseq platform. BMC Microbiology 2017, 17, 23. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Qiime 2: Reproducible, interactive, scalable, and extensible microbiome data science. PeerJ Preprints 2018, 6, e27295. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. Dada2: High-resolution sample inference from illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Flaherty, S.; Coffey, A.; Edwards, R.; Meaney, W.; Fitzgerald, G.F.; Ross, R.P. Genome of staphylococcal phage k: A new lineage of myoviridae infecting gram-positive bacteria with a low g + c content. J. Bacteriol. 2004, 186, 2862–2871. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Xu, W.; Lei, L.; Huang, J.; Feng, X.; Sun, C.; Du, C.; Zuo, J.; Li, Y.; Du, T.; et al. Lysgh15, a novel bacteriophage lysin, protects a murine bacteremia model efficiently against lethal methicillin-resistant Staphylococcus aureus infection. J. Clin. Microbiol. 2011, 49, 111–117. [Google Scholar] [CrossRef] [PubMed]
- Jun, S.Y.; Jung, G.M.; Yoon, S.J.; Oh, M.D.; Choi, Y.J.; Lee, W.J.; Kong, J.C.; Seol, J.G.; Kang, S.H. Antibacterial properties of a pre-formulated recombinant phage endolysin, sal-1. Int. J. Antimicrob. Agents 2013, 41, 156–161. [Google Scholar] [CrossRef] [PubMed]
- Kawai, Y.; Mickiewicz, K.; Errington, J. Lysozyme counteracts β-lactam antibiotics by promoting the emergence of l-form bacteria. Cell 2018, 172, 1038–1049. [Google Scholar] [CrossRef]
- Monteiro, R.; Pires, D.P.; Costa, A.R.; Azeredo, J. Phage therapy: Going temperate? Trends Microbiol. 2019, 27, 368–378. [Google Scholar] [CrossRef]
- Cogen, A.L.; Nizet, V.; Gallo, R.L. Skin microbiota: A source of disease or defence? Br. J. Dermatol. 2008, 158, 442–455. [Google Scholar] [CrossRef]
- Scharschmidt, T.C.; Fischbach, M.A. What lives on our skin: Ecology, genomics and therapeutic opportunities of the skin microbiome. Drug Discov. Today Dis. Mech. 2013, 10, e83–e89. [Google Scholar] [CrossRef] [PubMed]
- Elias, P.M. The skin barrier as an innate immune element. Semin. Immunopathol. 2007, 29, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Alkilani, A.Z.; McCrudden, M.T.C.; Donnelly, R.F. Transdermal drug delivery: Innovative pharmaceutical developments based on disruption of the barrier properties of the stratum corneum. Pharmaceutics 2015, 7, 438–470. [Google Scholar] [CrossRef] [PubMed]
- Clokie, M.R.; Millard, A.D.; Letarov, A.V.; Heaphy, S. Phages in nature. Bacteriophage 2011, 1, 31–45. [Google Scholar] [CrossRef] [PubMed]
- Filatova, L.Y.; Becker, S.C.; Donovan, D.M.; Gladilin, A.K.; Klyachko, N.L. Lysk, the enzyme lysing Staphylococcus aureus cells: Specific kinetic features and approaches towards stabilization. Biochimie 2010, 92, 507–513. [Google Scholar] [CrossRef] [PubMed]
- Garg, T.; Rath, G.; Goyal, A.K. Comprehensive review on additives of topical dosage forms for drug delivery. Drug Deliv. 2015, 22, 969–987. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Barros, M.; Vennemann, T.; Gallagher, D.T.; Yin, Y.; Linden, S.B.; Heselpoth, R.D.; Spencer, D.J.; Donovan, D.M.; Moult, J.; et al. A bacteriophage endolysin that eliminates intracellular streptococci. eLife 2016, 5, e13152. [Google Scholar] [CrossRef] [PubMed]
- Fischetti, V.A. Lysin Therapy Offers New Hope For Fighting Drug-Resistant Bacteria. Available online: https://www.rockefeller.edu/news/24920-lysin-therapy-offers-new-hope-fighting-drug-resistant-bacteria/ (accessed on 4 August 2019).
- Jun, S.Y.; Jung, G.M.; Yoon, S.J.; Choi, Y.-J.; Koh, W.S.; Moon, K.S.; Kang, S.H. Preclinical safety evaluation of intravenously administered sal200 containing the recombinant phage endolysin sal-1 as a pharmaceutical ingredient. Antimicrob. Agents Chemother. 2014, 58, 2084–2088. [Google Scholar] [CrossRef] [PubMed]
- Naik, S.; Larsen, S.B.; Gomez, N.C.; Alaverdyan, K.; Sendoel, A.; Yuan, S.; Polak, L.; Kulukian, A.; Chai, S.; Fuchs, E. Inflammatory memory sensitizes skin epithelial stem cells to tissue damage. Nature 2017, 550, 475–480. [Google Scholar] [CrossRef]
- Pastagia, M.; Euler, C.; Chahales, P.; Fuentes-Duculan, J.; Krueger, J.G.; Fischetti, V.A. A novel chimeric lysin shows superiority to mupirocin for skin decolonization of methicillin-resistant and -sensitive Staphylococcus aureus strains. Antimicrob. Agents Chemother. 2011, 55, 738–744. [Google Scholar] [CrossRef]
- Shimakura, H.; Uchiyama, J.; Saito, T.; Miyaji, K.; Fujimura, M.; Masuda, K.; Okamoto, N.; DeBoer, D.J.; Sakaguchi, M. Ige reactivity to hen egg white allergens in dogs with cutaneous adverse food reactions. Vet. Immunol. Immunopathol. 2016, 177, 52–57. [Google Scholar] [CrossRef] [PubMed]
- Caubet, J.-C.; Wang, J. Current understanding of egg allergy. Pediatr. Clin. North Am. 2011, 58, 427–443. [Google Scholar] [CrossRef] [PubMed]
- Piechowicz, L.; Garbacz, K.; Budzynska, A.; Dabrowska-Szponar, M. Outbreak of bullous impetigo caused by Staphylococcus aureus strains of phage type 3c/71 in a maternity ward linked to nasal carriage of a healthcare worker. Eur. J. Dermatol. 2012, 22, 252–255. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.A.; Wang, A.S.; Eichenfield, L.F.; Barrio, V. “Eb, or not eb?” Neonatal desquamative impetigo in a degloving pattern. Pediatr. Dermatol. 2016, 33, e147–e148. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Imanishi, I.; Uchiyama, J.; Tsukui, T.; Hisatsune, J.; Ide, K.; Matsuzaki, S.; Sugai, M.; Nishifuji, K. Therapeutic Potential of an Endolysin Derived from Kayvirus S25-3 for Staphylococcal Impetigo. Viruses 2019, 11, 769. https://doi.org/10.3390/v11090769
Imanishi I, Uchiyama J, Tsukui T, Hisatsune J, Ide K, Matsuzaki S, Sugai M, Nishifuji K. Therapeutic Potential of an Endolysin Derived from Kayvirus S25-3 for Staphylococcal Impetigo. Viruses. 2019; 11(9):769. https://doi.org/10.3390/v11090769
Chicago/Turabian StyleImanishi, Ichiro, Jumpei Uchiyama, Toshihiro Tsukui, Junzo Hisatsune, Kaori Ide, Shigenobu Matsuzaki, Motoyuki Sugai, and Koji Nishifuji. 2019. "Therapeutic Potential of an Endolysin Derived from Kayvirus S25-3 for Staphylococcal Impetigo" Viruses 11, no. 9: 769. https://doi.org/10.3390/v11090769
APA StyleImanishi, I., Uchiyama, J., Tsukui, T., Hisatsune, J., Ide, K., Matsuzaki, S., Sugai, M., & Nishifuji, K. (2019). Therapeutic Potential of an Endolysin Derived from Kayvirus S25-3 for Staphylococcal Impetigo. Viruses, 11(9), 769. https://doi.org/10.3390/v11090769




