Phage Reduce Stability for Regaining Infectivity during Antagonistic Coevolution with Host Bacterium
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
2.2. Phage Propagation and Concentration Determination
2.3. Screening of Phage-Resistant Bacterial Mutants and Regained-Infectivity Phage Mutants
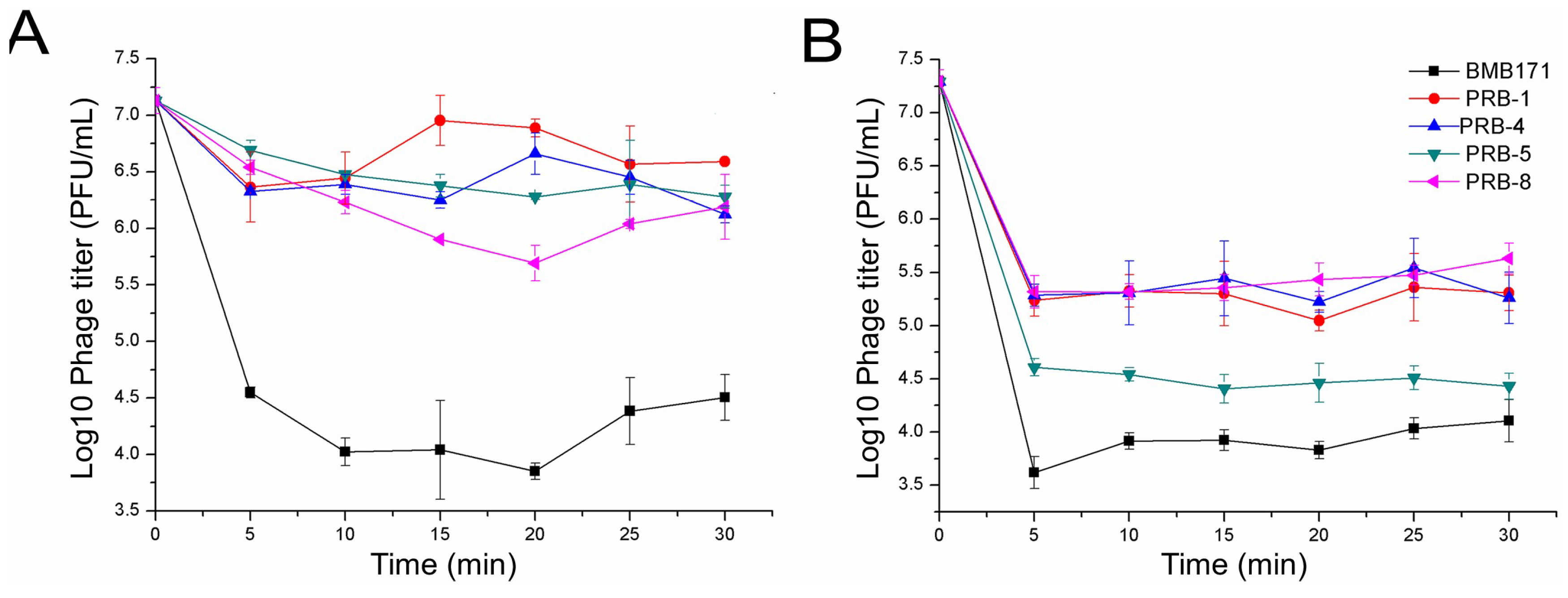
2.4. Binding Ability Assay of the Phage to Bacteria
2.5. Bacterial and Phage Genomic DNA Purification, Genome Sequencing, and Bioinformatic Analysis
3. Results
3.1. Phage-Resistant Bacterial Mutants and Regained-Infectivity Phage Mutants Appeared at High Frequency
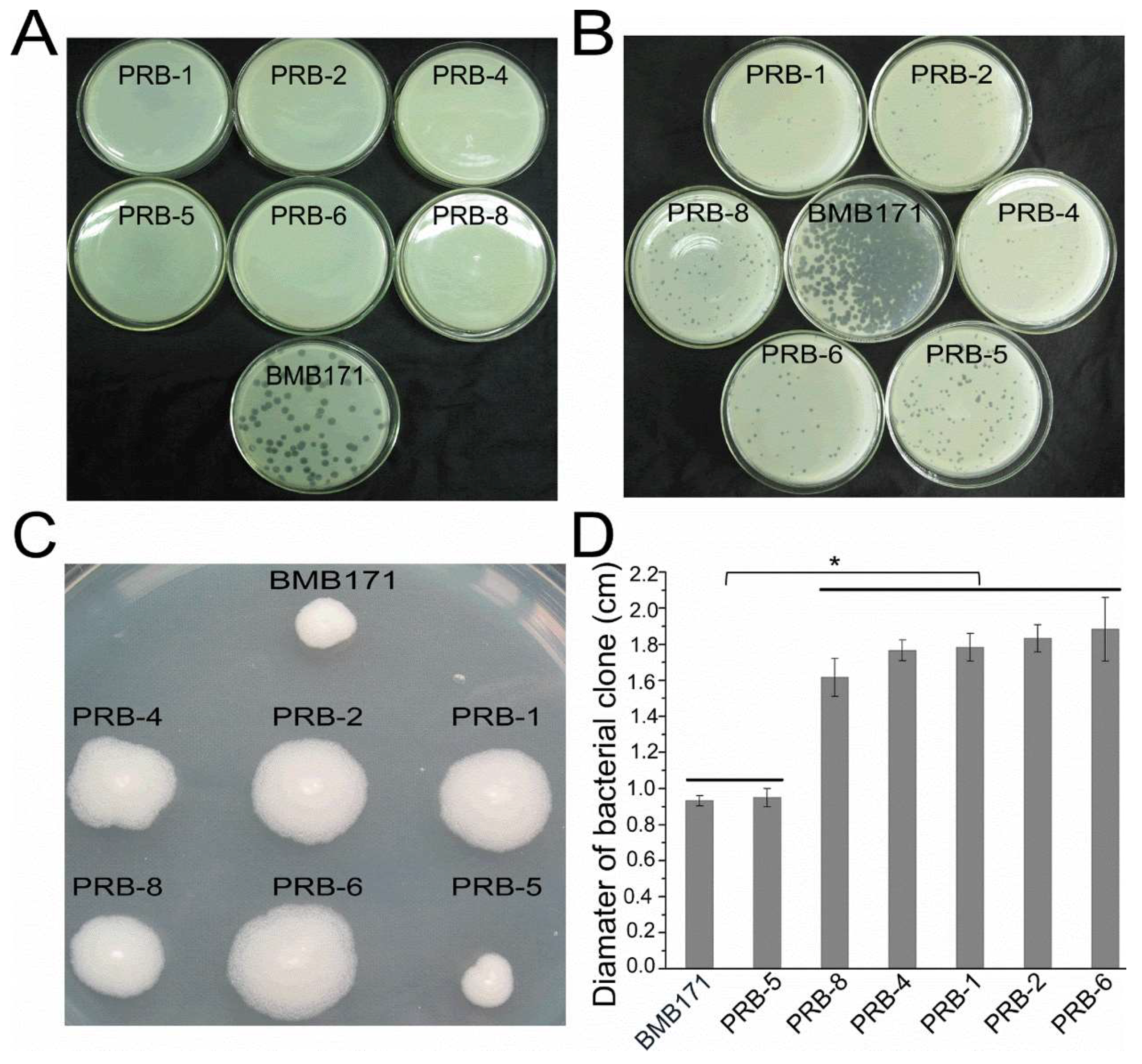
3.2. Phage-Resistant Bacterial Mutants Exhibiting Changed Features
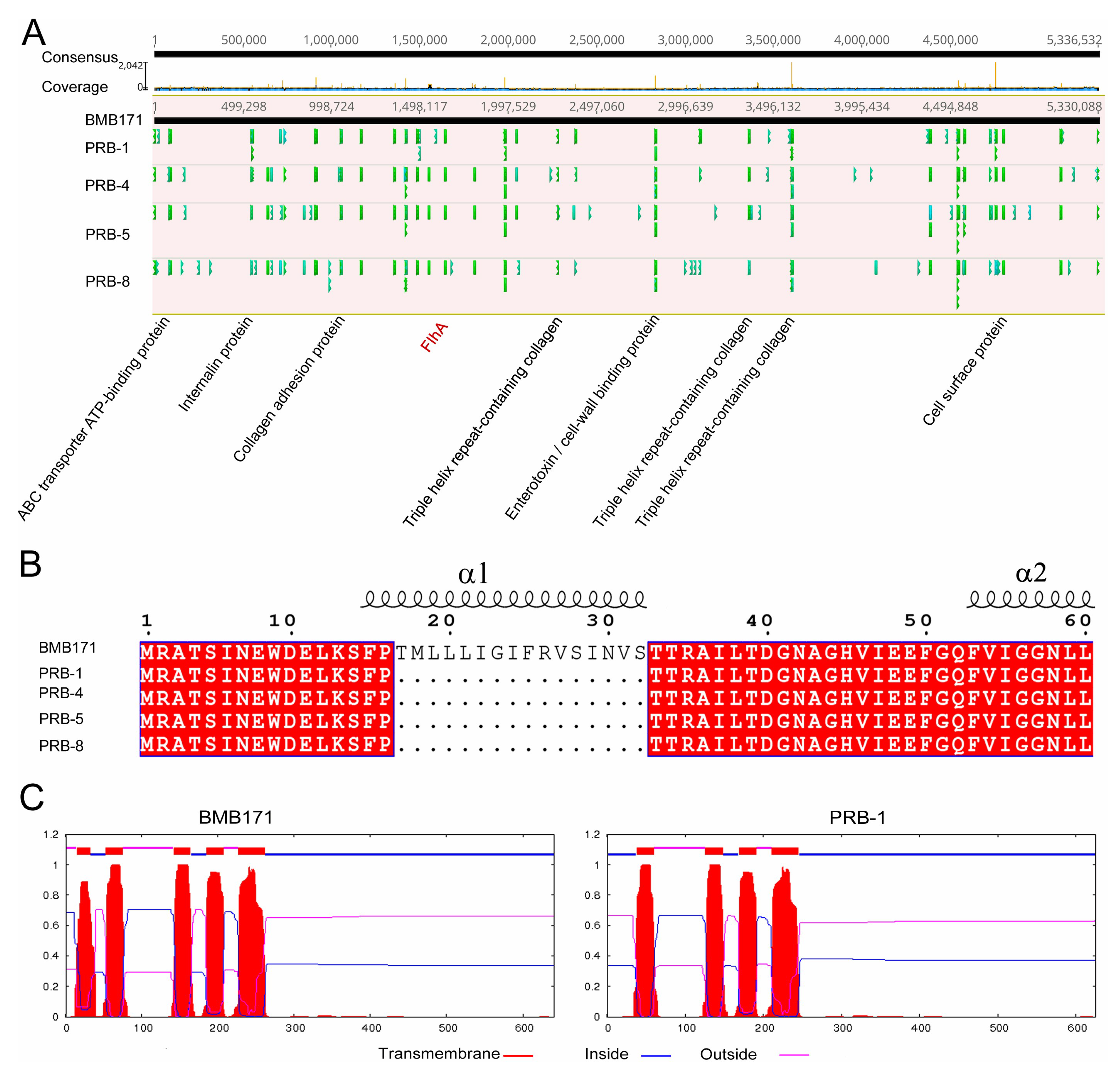
3.3. Phage-Resistant Bacterial Mutants Possessing Diverse Mutation Sites
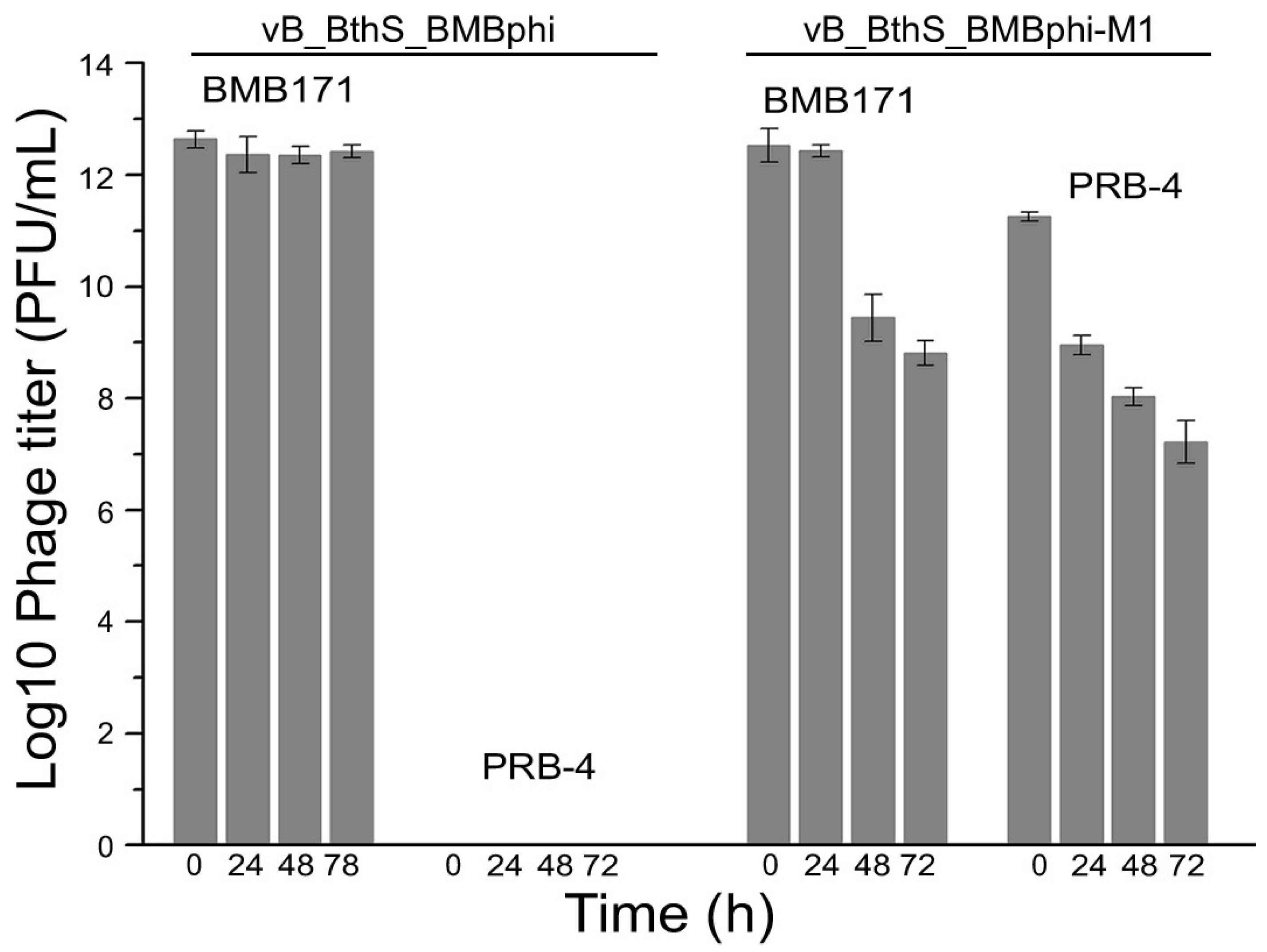
3.4. Regained-Infectivity Phage Mutants Showing Conserved Mutation Sites
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Rohwer, F. Global phage diversity. Cell 2003, 113, 141. [Google Scholar] [CrossRef]
- Koskella, B.; Brockhurst, M.A. Bacteria-phage coevolution as a driver of ecological and evolutionary processes in microbial communities. FEMS Microbiol. Rev. 2014, 38, 916–931. [Google Scholar] [CrossRef] [PubMed]
- Betts, A.; Kaltz, O.; Hochberg, M.E. Contrasted coevolutionary dynamics between a bacterial pathogen and its bacteriophages. Proc. Natl. Acad. Sci. USA 2014, 111, 11109–11114. [Google Scholar] [CrossRef] [PubMed]
- Hall, A.R.; Scanlan, P.D.; Morgan, A.D.; Buckling, A. Host-parasite coevolutionary arms races give way to fluctuating selection. Ecol. Lett. 2011, 14, 635–642. [Google Scholar] [CrossRef] [PubMed]
- Laanto, E.; Hoikkala, V.; Ravantti, J.; Sundberg, L.R. Long-term genomic coevolution of host-parasite interaction in the natural environment. Nat. Commun. 2017, 8, 111. [Google Scholar] [CrossRef] [PubMed]
- Levin, B.R.; Bull, J.J. Population and evolutionary dynamics of phage therapy. Nat. Rev. Microbiol. 2004, 2, 166–173. [Google Scholar] [CrossRef] [PubMed]
- Scanlan, P.D.; Buckling, A. Co-evolution with lytic phage selects for the mucoid phenotype of Pseudomonas fluorescens SBW25. ISME J. 2012, 6, 1148–1158. [Google Scholar] [CrossRef]
- Lopez-Pascua, L.; Buckling, A. Increasing productivity accelerates host-parasite coevolution. J. Evol. Biol. 2008, 21, 853–860. [Google Scholar] [CrossRef] [PubMed]
- Gomez, P.; Ashby, B.; Buckling, A. Population mixing promotes arms race host-parasite coevolution. Proc. R. Soc. B Biol. Sci. 2015, 282, 20142297. [Google Scholar] [CrossRef] [PubMed]
- Labrie, S.J.; Samson, J.E.; Moineau, S. Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 2010, 8, 317–327. [Google Scholar] [CrossRef]
- Vale, P.F.; Little, T.J. CRISPR-mediated phage resistance and the ghost of coevolution past. Proc. R. Soc. B Biol. Sci. 2010, 277, 2097–2103. [Google Scholar] [CrossRef]
- Koskella, B.; Taylor, T.B.; Bates, J.; Buckling, A. Using experimental evolution to explore natural patterns between bacterial motility and resistance to bacteriophages. ISME J. 2011, 5, 1809–1817. [Google Scholar] [CrossRef] [PubMed]
- Yen, M.; Cairns, L.S.; Camilli, A. A cocktail of three virulent bacteriophages prevents Vibrio cholerae infection in animal models. Nat. Commun. 2017, 8, 14187. [Google Scholar] [CrossRef] [PubMed]
- Nobrega, F.L.; Costa, A.R.; Kluskens, L.D.; Azeredo, J. Revisiting phage therapy: New applications for old resources. Trends Microbiol. 2015, 23, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, C.; Venail, P.; Holguin, A.V.; Vives, M.J. Co-Evolutionary Dynamics of the Bacteria Vibrio sp. CV1 and Phages V1G, V1P1, and V1P2: Implications for Phage Therapy. Microb. Ecol. 2013, 66, 897–905. [Google Scholar] [CrossRef]
- Schnepf, E.; Crickmore, N.; Van Rie, J.; Lereclus, D.; Baum, J.; Feitelson, J.; Zeigler, D.R.; Dean, D.H. Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol. Mol. Biol. Rev. 1998, 62, 775–806. [Google Scholar] [PubMed]
- Vilas-Boas, G.T.; Peruca, A.P.; Arantes, O.M. Biology and taxonomy of Bacillus cereus, Bacillus anthracis, and Bacillus thuringiensis. Can. J. Microbiol. 2007, 53, 673–687. [Google Scholar] [CrossRef]
- Bottone, E.J. Bacillus cereus, a volatile human pathogen. Clin. Microbiol. Rev. 2010, 23, 382–398. [Google Scholar] [CrossRef] [PubMed]
- Koehler, T.M. Bacillus anthracis genetics and virulence gene regulation. Curr. Top. Microbiol. Immunol. 2002, 271, 143–164. [Google Scholar]
- Schuch, R.; Fischetti, V.A. Detailed genomic analysis of the Wbeta and gamma phages infecting Bacillus anthracis: Implications for evolution of environmental fitness and antibiotic resistance. J. Bacteriol. 2006, 188, 3037–3051. [Google Scholar] [CrossRef]
- He, J.; Shao, X.; Zheng, H.; Li, M.; Wang, J.; Zhang, Q.; Li, L.; Liu, Z.; Sun, M.; Wang, S.; et al. Complete genome sequence of Bacillus thuringiensis mutant strain BMB171. J. Bacteriol. 2010, 192, 4074–4075. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Peng, Q.; Yang, S.; Zhang, S.; Fu, Y.; Wu, Y.; Gao, M. Isolation of A Novel Bacillus thuringiensis Phage Representing A New Phage Lineage and Characterization of Its Endolysin. Viruses 2018, 10, 611. [Google Scholar] [CrossRef]
- Yuan, Y.H.; Peng, Q.; Wu, D.D.; Kou, Z.; Wu, Y.; Liu, P.M.; Gao, M.Y. Effects of Actin-Like Proteins Encoded by Two Bacillus pumilus Phages on Unstable Lysogeny, Revealed by Genomic Analysis. Appl. Environ. Microb. 2015, 81, 339–350. [Google Scholar] [CrossRef]
- Yuan, Y.H.; Gao, M.Y. Genomic analysis of a ginger pathogen Bacillus pumilus providing the understanding to the pathogenesis and the novel control strategy. Sci. Rep. 2015, 5, 10259. [Google Scholar] [CrossRef] [PubMed]
- Zerbino, D.R.; Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008, 18, 821–829. [Google Scholar] [CrossRef]
- Stothard, P.; Wishart, D.S. Circular genome visualization and exploration using CGView. Bioinformatics 2005, 21, 537–539. [Google Scholar] [CrossRef]
- Geneious. Available online: https://www.geneious.com/ (accessed on 1 October 2018).
- Tzipilevich, E.; Habusha, M.; Ben-Yehuda, S. Acquisition of Phage Sensitivity by Bacteria through Exchange of Phage Receptors. Cell 2017, 168, 186.e12–199.e12. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Li, L.; Zhao, Z.; Peng, D.; Zhou, X. Polar flagella rotation in Vibrio parahaemolyticus confers resistance to bacteriophage infection. Sci. Rep. 2016, 6, 26147. [Google Scholar] [CrossRef] [PubMed]
- Lavelle, K.; Murphy, J.; Fitzgerald, B.; Lugli, G.A.; Zomer, A.; Neve, H.; Ventura, M.; Franz, C.M.; Cambillau, C.; van Sinderen, D.; et al. A decade of Streptococcus thermophilus phage evolution in an Irish dairy plant. Appl. Environ. Microbiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Beres, S.B.; Richter, E.W.; Nagiec, M.J.; Sumby, P.; Porcella, S.F.; DeLeo, F.R.; Musser, J.M. Molecular genetic anatomy of inter- and intraserotype variation in the human bacterial pathogen group A Streptococcus. Proc. Natl. Acad. Sci. USA 2006, 103, 7059–7064. [Google Scholar] [CrossRef]
- Terahara, N.; Inoue, Y.; Kodera, N.; Morimoto, Y.V.; Uchihashi, T.; Imada, K.; Ando, T.; Namba, K.; Minamino, T. Insight into structural remodeling of the FlhA ring responsible for bacterial flagellar type III protein export. Sci. Adv. 2018, 4, eaao7054. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.S.; Choi, S.; Shin, H.; Lee, J.H.; Choi, S.H. Vibrio vulnificus Bacteriophage SSP002 as a Possible Biocontrol Agent. Appl. Environ. Microb. 2014, 80, 515–524. [Google Scholar] [CrossRef]
- Legrand, P.; Collins, B.; Blangy, S.; Murphy, J.; Spinelli, S.; Gutierrez, C.; Richet, N.; Kellenberger, C.; Desmyter, A.; Mahony, J.; et al. The Atomic Structure of the Phage Tuc2009 Baseplate Tripod Suggests that Host Recognition Involves Two Different Carbohydrate Binding Modules. mBio 2016, 7, e01781-15. [Google Scholar] [CrossRef] [PubMed]
- Stockdale, S.R.; Mahony, J.; Courtin, P.; Chapot-Chartier, M.P.; van Pijkeren, J.P.; Britton, R.A.; Neve, H.; Heller, K.J.; Aideh, B.; Vogensen, F.K.; et al. The Lactococcal Phages Tuc2009 and TP901-1 Incorporate Two Alternate Forms of Their Tail Fiber into Their Virions for Infection Specialization. J. Biol. Chem. 2013, 288, 5581–5590. [Google Scholar] [CrossRef] [PubMed]
- Spinelli, S.; Campanacci, V.; Blangy, S.; Moineau, S.; Tegoni, M.; Cambillau, C. Modular structure of the receptor binding proteins of Lactococcus lactis phages. The RBP structure of the temperate phage TP901-1. J. Biol. Chem. 2006, 281, 14256–14262. [Google Scholar] [CrossRef]
- Veesler, D.; Robin, G.; Lichiere, J.; Auzat, I.; Tavares, P.; Bron, P.; Campanacci, V.; Cambillau, C. Crystal structure of bacteriophage SPP1 distal tail protein (gp19.1): A baseplate hub paradigm in gram-positive infecting phages. J. Biol. Chem. 2010, 285, 36666–36673. [Google Scholar] [CrossRef]
- Dieterle, M.E.; Bowman, C.; Batthyany, C.; Lanzarotti, E.; Turjanski, A.; Hatfull, G.; Piuri, M. Exposing the secrets of two well-known Lactobacillus casei phages, J-1 and PL-1, by genomic and structural analysis. Appl. Environ. Microbiol. 2014, 80, 7107–7121. [Google Scholar] [CrossRef]
- Doron, S.; Melamed, S.; Ofir, G.; Leavitt, A.; Lopatina, A.; Keren, M.; Amitai, G.; Sorek, R. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 2018, 359, eaar4120. [Google Scholar] [CrossRef]
- Tenaillon, O.; Rodriguez-Verdugo, A.; Gaut, R.L.; McDonald, P.; Bennett, A.F.; Long, A.D.; Gaut, B.S. The Molecular Diversity of Adaptive Convergence. Science 2012, 335, 457–461. [Google Scholar] [CrossRef]
- Horvath, P.; Barrangou, R. CRISPR/Cas, the immune system of bacteria and archaea. Science 2010, 327, 167–170. [Google Scholar] [CrossRef]
- Ghosh, S.; Marintcheva, B.; Takahashi, M.; Richardson, C.C. C-terminal Phenylalanine of Bacteriophage T7 Single-stranded DNA-binding Protein Is Essential for Strand Displacement Synthesis by T7 DNA Polymerase at a Nick in DNA. J. Biol. Chem. 2009, 284, 30339–30349. [Google Scholar] [CrossRef] [PubMed]
- Paterson, S.; Vogwill, T.; Buckling, A.; Benmayor, R.; Spiers, A.J.; Thomson, N.R.; Quail, M.; Smith, F.; Walker, D.; Libberton, B.; et al. Antagonistic coevolution accelerates molecular evolution. Nature 2010, 464, 275–278. [Google Scholar] [CrossRef] [PubMed]
- Scanlan, P.D.; Hall, A.R.; Lopez-Pascua, L.D.; Buckling, A. Genetic basis of infectivity evolution in a bacteriophage. Mol. Ecol. 2011, 20, 981–989. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Shin, H.; Lee, J.H.; Ryu, S. Identification and characterization of a novel flagellum-dependent Salmonella-infecting bacteriophage, iEPS5. Appl. Environ. Microbiol. 2013, 79, 4829–4837. [Google Scholar] [CrossRef]
- Fukumura, T.; Makino, F.; Dietsche, T.; Kinoshita, M.; Kato, T.; Wagner, S.; Namba, K.; Imada, K.; Minamino, T. Assembly and stoichiometry of the core structure of the bacterial flagellar type III export gate complex. PLoS Biol. 2017, 15, e2002281. [Google Scholar] [CrossRef] [PubMed]
- Erhardt, M.; Wheatley, P.; Kim, E.A.; Hirano, T.; Zhang, Y.; Sarkar, M.K.; Hughes, K.T.; Blair, D.F. Mechanism of type-III protein secretion: Regulation of FlhA conformation by a functionally critical charged-residue cluster. Mol. Microbiol. 2017, 104, 234–249. [Google Scholar] [CrossRef] [PubMed]
- Dieterle, M.E.; Spinelli, S.; Sadovskaya, I.; Piuri, M.; Cambillau, C. Evolved distal tail carbohydrate binding modules of Lactobacillus phage J-1: A novel type of anti-receptor widespread among lactic acid bacteria phages. Mol. Microbiol. 2017, 104, 608–620. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yuan, Y.; Peng, Q.; Zhang, S.; Liu, T.; Yang, S.; Yu, Q.; Wu, Y.; Gao, M. Phage Reduce Stability for Regaining Infectivity during Antagonistic Coevolution with Host Bacterium. Viruses 2019, 11, 118. https://doi.org/10.3390/v11020118
Yuan Y, Peng Q, Zhang S, Liu T, Yang S, Yu Q, Wu Y, Gao M. Phage Reduce Stability for Regaining Infectivity during Antagonistic Coevolution with Host Bacterium. Viruses. 2019; 11(2):118. https://doi.org/10.3390/v11020118
Chicago/Turabian StyleYuan, Yihui, Qin Peng, Shaowen Zhang, Tingting Liu, Shuo Yang, Qiuhan Yu, Yan Wu, and Meiying Gao. 2019. "Phage Reduce Stability for Regaining Infectivity during Antagonistic Coevolution with Host Bacterium" Viruses 11, no. 2: 118. https://doi.org/10.3390/v11020118
APA StyleYuan, Y., Peng, Q., Zhang, S., Liu, T., Yang, S., Yu, Q., Wu, Y., & Gao, M. (2019). Phage Reduce Stability for Regaining Infectivity during Antagonistic Coevolution with Host Bacterium. Viruses, 11(2), 118. https://doi.org/10.3390/v11020118




