Viruses Infecting the Plant Pathogenic Fungus Rhizoctonia solani
Abstract
1. Introduction
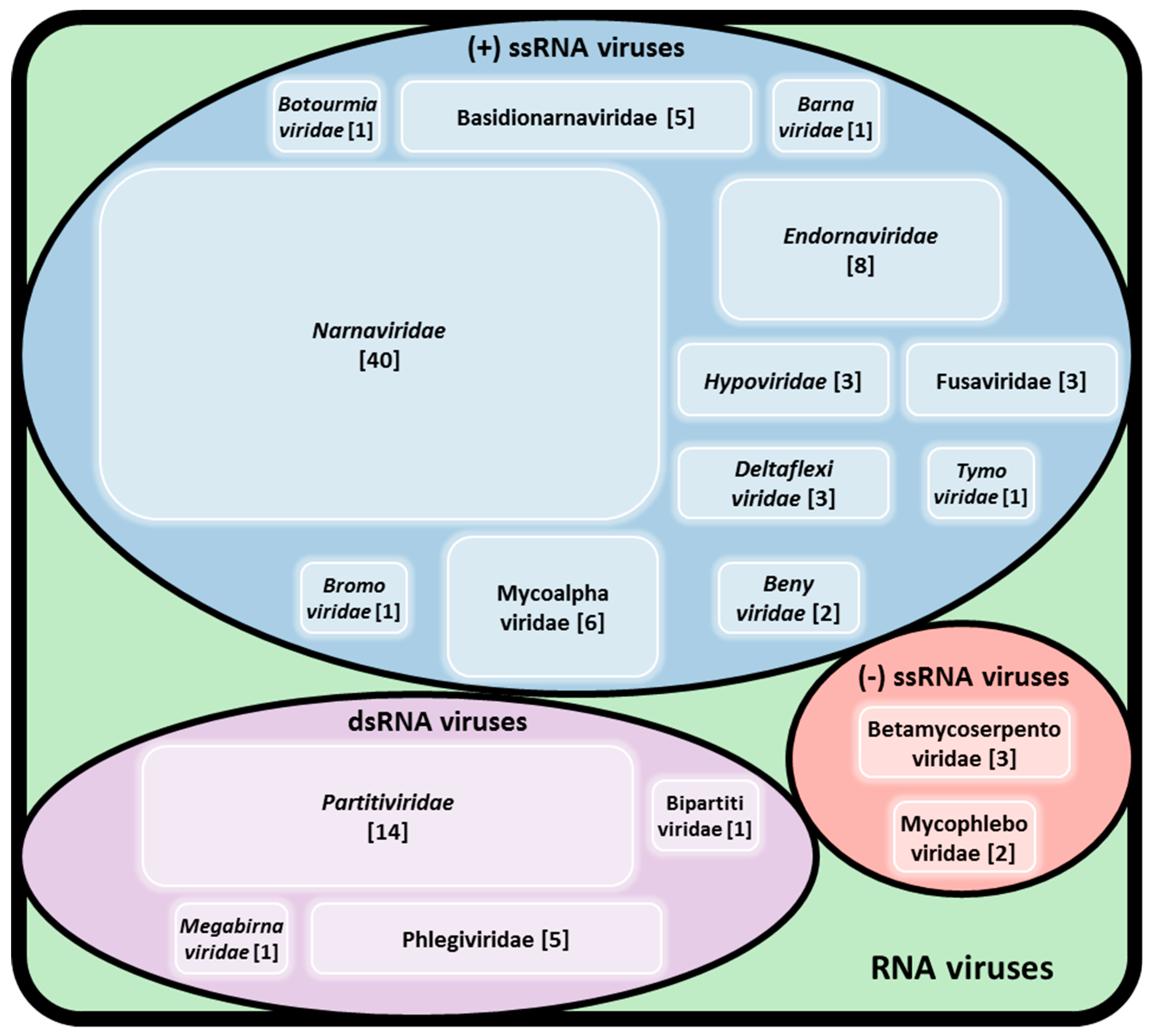
2. The Diversity of Viruses Infecting Rhizoctonia solani
2.1. Double-Stranded RNA Viruses
2.1.1. Megabirnaviridae and Phlegiviridae
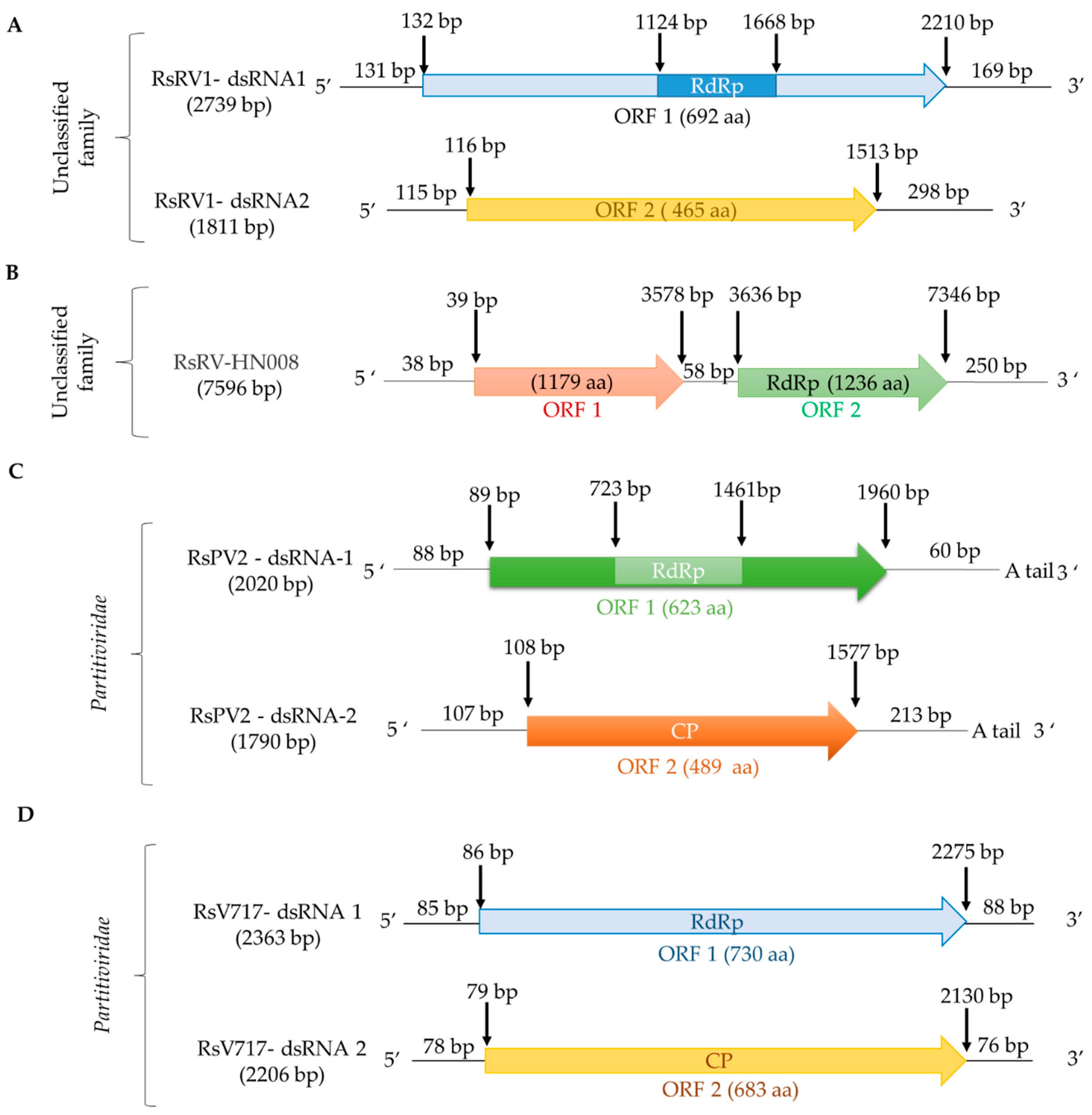
2.1.2. Partitiviridae and Bipartitiviridae
2.1.3. Unclassified dsRNA Viruses
2.2. Single-Stranded RNA Viruses
2.2.1. (+)ssRNA Viruses: Barnaviridae
2.2.2. (+)ssRNA Viruses: Benyviridae
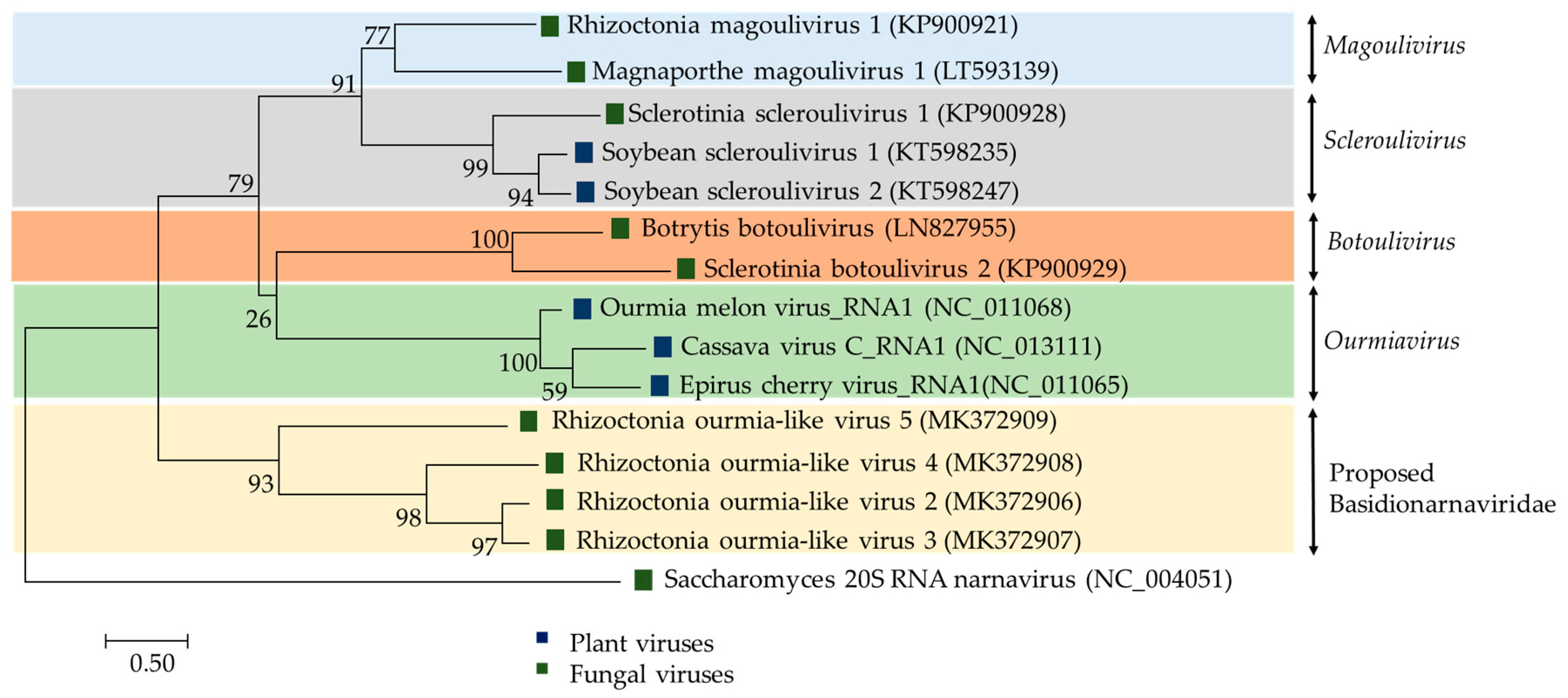
2.2.3. (+)ssRNA Viruses: Botourmiaviridae and Basidionarnaviridae
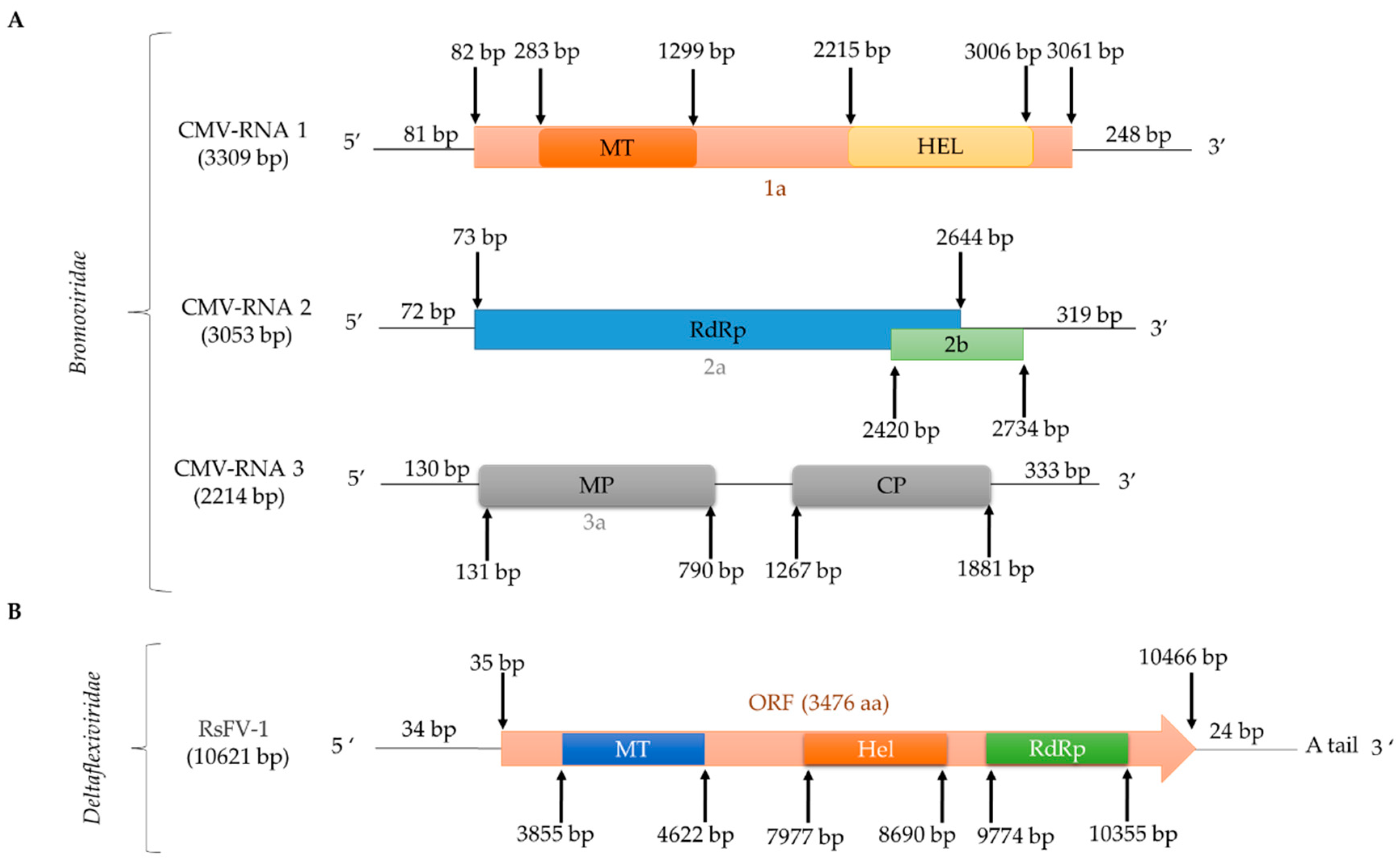
2.2.4. (+)ssRNA Viruses: Bromoviridae
2.2.5. (+)ssRNA Viruses: Deltaflexiviridae and Tymoviridae, Tymovirales
2.2.6. (+)ssRNA Viruses: Endornaviridae
2.2.7. (+)ssRNA Viruses: Hypoviridae and Fusariviridae
2.2.8. (+)ssRNA Viruses: Narnaviridae
2.2.9. (+)ssRNA Viruses: Mycoalphaviridae
2.2.10. (−)ssRNA Viruses: Betamycoserpentoviridae, Serpentovirales
2.2.11. (−)ssRNA Viruses: Mycophleboviridae, Bunyavirales
3. Transmission of Viruses Infecting Rhizoctonia solani
4. Effects of Virus Infection on Rhizoctonia solani
5. Conclusions and Future Prospects
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ram, R.M.; Singh, H. Rhizoctonia bataticola: A serious threat to chickpea production. Int. J. Chem. Stud. 2018, 6, 715–723. [Google Scholar]
- Hua, G.K.H.; Bertier, L.; Soltaninejad, S.; Höfte, M. Cropping systems and cultural practices determine the Rhizoctonia anastomosis groups associated with Brassica spp. in Vietnam. PLoS ONE 2014, 9, e111750. [Google Scholar] [CrossRef]
- Samsatly, J.; Copley, T.R.; Jabaji, S.H. Antioxidant genes of plants and fungal pathogens are distinctly regulated during disease development in different Rhizoctonia solani pathosystems. PLoS ONE 2018, 13, e0192682. [Google Scholar] [CrossRef]
- Xia, Y.; Fei, B.; He, J.; Zhou, M.; Zhang, D.; Pan, L.; Li, S.; Liang, Y.; Wang, L.; Zhu, J. Transcriptome analysis reveals the host selection fitness mechanisms of the Rhizoctonia solani AG1IA pathogen. Sci. Rep. 2017, 7, 10120. [Google Scholar] [CrossRef]
- Kouzai, Y.; Kimura, M.; Watanabe, M.; Kusunoki, K.; Osaka, D.; Suzuki, T.; Matsui, H.; Yamamoto, M.; Ichinose, Y.; Toyoda, K. Salicylic acid-dependent immunity contributes to resistance against Rhizoctonia solani, a necrotrophic fungal agent of sheath blight, in rice and Brachypodium distachyon. New Phytol. 2018, 217, 771–783. [Google Scholar] [CrossRef]
- Picarelli, M.A.S.; Forgia, M.; Rivas, E.B.; Nerva, L.; Chiapello, M.; Turina, M.; Colariccio, A. Extreme diversity of mycoviruses present in isolates of Rhizoctonia solani AG2-2 LP from Zoysia japonica from Brazil. Front. Cell. Infect. Microbiol. 2019, 9, 244. [Google Scholar] [CrossRef]
- Andika, I.B.; Wei, S.; Cao, C.; Salaipeth, L.; Kondo, H.; Sun, L. Phytopathogenic fungus hosts a plant virus: A naturally occurring cross-kingdom viral infection. Proc. Natl. Acad. Sci. USA 2017, 114, 12267–12272. [Google Scholar] [CrossRef]
- Agrios, G.N. Plant Pathology, 5th ed.; Department of Plant Pathology, University of Florida: Gainesville, FL, USA, 2005. [Google Scholar]
- Bartholomäus, A.; Wibberg, D.; Winkler, A.; Pühler, A.; Schlüter, A.; Varrelmann, M. Deep sequencing analysis reveals the mycoviral diversity of the virome of an avirulent isolate of Rhizoctonia solani AG-2-2 IV. PLoS ONE 2016, 11, e0165965. [Google Scholar] [CrossRef]
- Huang, X.; Zhang, N.; Yong, X.; Yang, X.; Shen, Q. Biocontrol of Rhizoctonia solani damping-off disease in cucumber with Bacillus pumilus SQR-N43. Microbiol. Res. 2012, 167, 135–143. [Google Scholar] [CrossRef]
- Hamid, M.; Xie, J.; Wu, S.; Maria, S.; Zheng, D.; Assane, H.A.; Wang, Q.; Cheng, J.; Fu, Y.; Jiang, D. A Novel Deltaflexivirus that Infects the Plant Fungal Pathogen, Sclerotinia sclerotiorum, Can Be Transmitted Among Host Vegetative Incompatible Strains. Viruses 2018, 10, 295. [Google Scholar] [CrossRef]
- Zhong, J.; Chen, C.-Y.; Gao, B.-D. Genome sequence of a novel mycovirus of Rhizoctonia solani, a plant pathogenic fungus. Virus Genes 2015, 51, 167–170. [Google Scholar] [CrossRef]
- Li, Z.; Chen, L.; Meiling, Z.; Mei, Y.; Erxun, Z. Diversity of dsRNA viruses infecting rice sheath blight fungus Rhizoctonia solani AG-1 IA. Rice Sci. 2018, 25, 57–60. [Google Scholar] [CrossRef]
- Das, S.; Falloon, R.E.; Stewart, A.; Pitman, A.R. Molecular characterisation of an endornavirus from Rhizoctonia solani AG-3PT infecting potato. Fungal Biol. 2014, 118, 924–934. [Google Scholar] [CrossRef]
- Bartholomäus, A.; Wibberg, D.; Winkler, A.; Pühler, A.; Schlüter, A.; Varrelmann, M. Identification of a novel mycovirus isolated from Rhizoctonia solani (AG 2-2 IV) provides further information about genome plasticity within the order Tymovirales. Arch. Virol. 2017, 162, 555–559. [Google Scholar] [CrossRef]
- Zheng, L.; Liu, H.; Zhang, M.; Cao, X.; Zhou, E. The complete genomic sequence of a novel mycovirus from Rhizoctonia solani AG-1 IA strain B275. Arch. Virol. 2013, 158, 1609–1612. [Google Scholar] [CrossRef]
- Zheng, L.; Zhang, M.; Chen, Q.; Zhu, M.; Zhou, E. A novel mycovirus closely related to viruses in the genus Alphapartitivirus confers hypovirulence in the phytopathogenic fungus Rhizoctonia solani. Virology 2014, 456, 220–226. [Google Scholar] [CrossRef]
- Das, S.; Falloon, R.E.; Stewart, A.; Pitman, A.R. Novel mitoviruses in Rhizoctonia solani AG-3PT infecting potato. Fungal Biol. 2016, 120, 338–350. [Google Scholar] [CrossRef]
- Marzano, S.-Y.L.; Nelson, B.D.; Ajayi-Oyetunde, O.; Bradley, C.A.; Hughes, T.J.; Hartman, G.L.; Eastburn, D.M.; Domier, L.L. Identification of diverse mycoviruses through metatranscriptomics characterization of the viromes of five major fungal plant pathogens. J. Virol. 2016, 90, 6846–6863. [Google Scholar] [CrossRef]
- Strauss, E.E.; Lakshman, D.K.; Tavantzis, S.M. Molecular characterization of the genome of a partitivirus from the basidiomycete Rhizoctonia solani. J. Gen. Virol. 2000, 81, 549–555. [Google Scholar] [CrossRef]
- Das, S. Rhizoctonia Solani on Potato in New Zealand: Pathogen Characterisation and Identification of Double-Stranded RNA Viruses that may Affect their Virulence. Ph.D. Thesis, Lincoln University, Lincoln, New Zealand, 2013. [Google Scholar]
- Jian, J.; Lakshman, D.K.; Tavantzis, S.M. Association of distinct double-stranded RNAs with enhanced or diminished virulence in Rhizoctonia solani infecting potato. Mol. Plant-Microbe Interact 1997, 10, 1002–1009. [Google Scholar] [CrossRef]
- Son, M.; Yu, J.; Kim, K.-H. Five questions about mycoviruses. PLoS Pathog. 2015, 11, e1005172. [Google Scholar] [CrossRef] [PubMed]
- Abbas, A. A Review Paper on Mycoviruses. J. Plant Pathol. Microbiol. 2016, 7, 2. [Google Scholar] [CrossRef]
- Kotta-Loizou, I.; Coutts, R.H. Mycoviruses in Aspergilli: A comprehensive review. Front. Microbiol. 2017, 8, 1699. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Li, B.; Fu, Y.; Jiang, D.; Ghabrial, S.A.; Li, G.; Peng, Y.; Xie, J.; Cheng, J.; Huang, J. A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus. Proc. Natl. Acad. Sci. USA 2010, 107, 8387–8392. [Google Scholar] [CrossRef]
- Velasco, L.; Arjona-Girona, I.; Cretazzo, E.; López-Herrera, C. Viromes in Xylariaceae fungi infecting avocado in Spain. Virology 2019, 532, 11–21. [Google Scholar] [CrossRef]
- Lin, Y.-H.; Fujita, M.; Chiba, S.; Hyodo, K.; Andika, I.B.; Suzuki, N.; Kondo, H. Two novel fungal negative-strand RNA viruses related to mymonaviruses and phenuiviruses in the shiitake mushroom (Lentinula edodes). Virology 2019, 533, 125–136. [Google Scholar] [CrossRef]
- Nerva, L.; Forgia, M.; Ciuffo, M.; Chitarra, W.; Chiapello, M.; Vallino, M.; Varese, G.; Turina, M. The mycovirome of a fungal collection from the sea cucumber Holothuria polii. Virus Res. 2019, 273, 197737. [Google Scholar] [CrossRef]
- Hao, F.; Wu, M.; Li, G. Molecular characterization and geographic distribution of a mymonavirus in the population of Botrytis cinerea. Viruses 2018, 10, 432. [Google Scholar] [CrossRef]
- Mu, F.; Xie, J.; Cheng, S.; You, M.P.; Barbetti, M.J.; Jia, J.; Wang, Q.; Cheng, J.; Fu, Y.; Chen, T. Virome Characterization of a Collection of S. sclerotiorum from Australia. Front. Microbiol. 2018, 8, 2540. [Google Scholar] [CrossRef]
- Wang, L.; He, H.; Wang, S.; Chen, X.; Qiu, D.; Kondo, H.; Guo, L. Evidence for a novel negative-stranded RNA mycovirus isolated from the plant pathogenic fungus Fusarium graminearum. Virology 2018, 518, 232–240. [Google Scholar] [CrossRef]
- Donaire, L.; Pagán, I.; Ayllón, M.A. Characterization of Botrytis cinerea negative-stranded RNA virus 1, a new mycovirus related to plant viruses, and a reconstruction of host pattern evolution in negative-sense ssRNA viruses. Virology 2016, 499, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Osaki, H.; Sasaki, A.; Nomiyama, K.; Tomioka, K. Multiple virus infection in a single strain of Fusarium poae shown by deep sequencing. Virus Genes 2016, 52, 835–847. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zheng, L.; Liu, C.; Shu, C.; Zhou, E. Characterization of a novel dsRNA mycovirus isolated from strain A105 of Rhizoctonia solani AG-1 IA. Arch. Virol. 2018, 163, 427–430. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Zeng, M.; Zhang, M.; Shu, C.; Zhou, E. Complete nucleotide sequence of a partitivirus from Rhizoctonia solani AG-1 IA strain C24. Viruses 2018, 10, 703. [Google Scholar] [CrossRef]
- Chen, Y.; Gai, X.T.; Chen, R.X.; Li, C.X.; Zhao, G.K.; Xia, Z.Y.; Zou, C.M.; Zhong, J. Characterization of three novel betapartitiviruses co-infecting the phytopathogenic fungus Rhizoctonia solani. Virus Res. 2019, 270, 197649. [Google Scholar] [CrossRef]
- Lyu, R.; Zhang, Y.; Tang, Q.; Li, Y.; Cheng, J.; Fu, Y.; Chen, T.; Jiang, D.; Xie, J. Two alphapartitiviruses co-infecting a single isolate of the plant pathogenic fungus Rhizoctonia solani. Arch. Virol. 2018, 163, 515–520. [Google Scholar] [CrossRef]
- Zheng, L.; Shu, C.; Zhang, M.; Yang, M.; Zhou, E. Molecular Characterization of a Novel Endornavirus Conferring Hypovirulence in Rice Sheath Blight Fungus Rhizoctonia solani AG-1 IA Strain GD-2. Viruses 2019, 11, 178. [Google Scholar] [CrossRef]
- Lakshman, D.K.; Jian, J.; Tavantzis, S.M. A double-stranded RNA element from a hypovirulent strain of Rhizoctonia solani occurs in DNA form and is genetically related to the pentafunctional AROM protein of the shikimate pathway. Proc. Natl. Acad. Sci. USA 1998, 95, 6425–6429. [Google Scholar] [CrossRef]
- Lakshman, D.; Tavantzis, S. Spontaneous appearance of genetically distinct double-stranded RNA elements in Rhizoctonia solani. Phytopathol 1994, 84, 633–639. [Google Scholar] [CrossRef]
- Jia, H.; Dong, K.; Zhou, L.; Wang, G.; Hong, N.; Jiang, D.; Xu, W. A dsRNA virus with filamentous viral particles. Nat. Commun. 2017, 8, 168. [Google Scholar] [CrossRef]
- Ghabrial, S.A.; Castón, J.R.; Jiang, D.; Nibert, M.L.; Suzuki, N. 50-plus years of fungal viruses. Virology 2015, 479, 356–368. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.; Lo, G.C.; Chow, F.W.; Fan, R.Y.; Cai, J.J.; Yuen, K.-Y.; Woo, P.C. Novel Partitivirus Enhances Virulence of and Causes Aberrant Gene Expression in Talaromyces marneffei. mBio 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Depierreux, D.; Vong, M.; Nibert, M.L. Nucleotide sequence of Zygosaccharomyces bailii virus Z: Evidence for+ 1 programmed ribosomal frameshifting and for assignment to family Amalgaviridae. Virus Res. 2016, 217, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Gilmer, D.; Ratti, C.; Consortium, I.R. ICTV Virus taxonomy profile: Benyviridae. J. Gen. Virol. 2017, 98, 1571. [Google Scholar] [CrossRef]
- Wu, M.; Jin, F.; Zhang, J.; Yang, L.; Jiang, D.; Li, G. Characterization of a novel bipartite double-stranded RNA mycovirus conferring hypovirulence in the phytopathogenic fungus Botrytis porri. J. Virol. 2012, 86, 6605–6619. [Google Scholar] [CrossRef]
- Zhai, L.; Zhang, M.; Hong, N.; Xiao, F.; Fu, M.; Xiang, J.; Wang, G. Identification and characterization of a novel hepta-segmented dsRNA virus from the phytopathogenic fungus Colletotrichum fructicola. Front. Microbiol. 2018, 9, 754. [Google Scholar] [CrossRef]
- Kanhayuwa, L.; Kotta-Loizou, I.; Özkan, S.; Gunning, A.P.; Coutts, R.H. A novel mycovirus from Aspergillus fumigatus contains four unique dsRNAs as its genome and is infectious as dsRNA. Proc. Natl. Acad. Sci. USA 2015, 112, 9100–9105. [Google Scholar] [CrossRef]
- Kotta-Loizou, I.; Coutts, R.H. Studies on the virome of the entomopathogenic fungus Beauveria bassiana reveal novel dsRNA elements and mild hypervirulence. PLoS Pathog. 2017, 13, e1006183. [Google Scholar] [CrossRef]
- Xie, J.; Jiang, D. New insights into mycoviruses and exploration for the biological control of crop fungal diseases. Annu. Rev. Phytopathol. 2014, 52, 45–68. [Google Scholar] [CrossRef]
- Sato, Y.; Miyazaki, N.; Kanematsu, S.; Xie, J.; Ghabrial, S.A.; Hillman, B.I.; Suzuki, N.; Consortium, I.R. ICTV Virus Taxonomy Profile: Megabirnaviridae. J. Gen. Virol. 2019, 100, 1269–1270. [Google Scholar] [CrossRef]
- Chiba, S.; Salaipeth, L.; Lin, Y.-H.; Sasaki, A.; Kanematsu, S.; Suzuki, N. A novel bipartite double-stranded RNA mycovirus from the white root rot fungus Rosellinia necatrix: Molecular and biological characterization, taxonomic considerations, and potential for biological control. J. Virol. 2009, 83, 12801–12812. [Google Scholar] [CrossRef] [PubMed]
- Kanematsu, S.; Shimizu, T.; Salaipeth, L.; Yaegashi, H.; Sasaki, A.; Ito, T.; Suzuki, N. Genome rearrangement of a mycovirus Rosellinia necatrix megabirnavirus 1 affecting its ability to attenuate virulence of the host fungus. Virology 2014, 450, 308–315. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Wang, Y.; Sun, X.; Cheng, J.; Fu, Y.; Liu, H.; Jiang, D.; Ghabrial, S.A.; Xie, J. Characterization of a novel megabirnavirus from Sclerotinia sclerotiorum reveals horizontal gene transfer from single-stranded RNA virus to double-stranded RNA virus. J. Virol. 2015, 89, 8567–8579. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, A.; Nakamura, H.; Suzuki, N.; Kanematsu, S. Characterization of a new megabirnavirus that confers hypovirulence with the aid of a co-infecting partitivirus to the host fungus, Rosellinia necatrix. Virus Res. 2016, 219, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Nerva, L.; Ciuffo, M.; Vallino, M.; Margaria, P.; Varese, G.; Gnavi, G.; Turina, M. Multiple approaches for the detection and characterization of viral and plasmid symbionts from a collection of marine fungi. Virus Res. 2016, 219, 22–38. [Google Scholar] [CrossRef] [PubMed]
- Petrzik, K.; Sarkisova, T.; Starý, J.; Koloniuk, I.; Hrabáková, L.; Kubešová, O. Molecular characterization of a new monopartite dsRNA mycovirus from mycorrhizal Thelephora terrestris (Ehrh.) and its detection in soil oribatid mites (Acari: Oribatida). Virology 2016, 489, 12–19. [Google Scholar] [CrossRef]
- Vainio, E.J.; Chiba, S.; Ghabrial, S.A.; Maiss, E.; Roossinck, M.; Sabanadzovic, S.; Suzuki, N.; Xie, J.; Nibert, M. ICTV virus taxonomy profile: Partitiviridae. J. Gen. Virol. 2018, 99, 17. [Google Scholar] [CrossRef]
- Shiba, K.; Hatta, C.; Sasai, S.; Tojo, M.; Ohki, S.T.; Mochizuki, T. Genome sequence of a novel partitivirus identified from the oomycete Pythium nunn. Arch. Virol. 2018, 163, 2561–2563. [Google Scholar] [CrossRef]
- Xiao, X.; Cheng, J.; Tang, J.; Fu, Y.; Jiang, D.; Baker, T.S.; Ghabrial, S.A.; Xie, J. A novel partitivirus that confers hypovirulence on plant pathogenic fungi. J. Virol. 2014, 88, 10120–10133. [Google Scholar] [CrossRef]
- Jian, J.; Lakshman, D.K.; Tavantzis, S.M. A virulence-associated, 6.4-kb, double-stranded RNA from Rhizoctonia solani is phylogenetically related to plant bromoviruses and electron transport enzymes. Mol. Plant. Microbe Interact. 1998, 11, 601–609. [Google Scholar] [CrossRef]
- Zoll, J.; Verweij, P.E.; Melchers, W.J. Discovery and characterization of novel Aspergillus fumigatus mycoviruses. PLoS ONE 2018, 13, e0200511. [Google Scholar] [CrossRef] [PubMed]
- Usui, K.; Ichihashi, N.; Yomo, T. A design principle for a single-stranded RNA genome that replicates with less double-strand formation. Nucleic Acids Res. 2015, 43, 8033–8043. [Google Scholar] [CrossRef] [PubMed]
- Baltimore, D. Expression of animal virus genomes. Bacteriol. Rev. 1971, 35, 235. [Google Scholar] [PubMed]
- Koonin, E.V.; Dolja, V.V.; Morris, T.J. Evolution and taxonomy of positive-strand RNA viruses: Implications of comparative analysis of amino acid sequences. Crit. Rev. Biochem. Mol. Biol. 1993, 28, 375–430. [Google Scholar] [CrossRef] [PubMed]
- Reguera, J.; Gerlach, P.; Cusack, S. Towards a structural understanding of RNA synthesis by negative strand RNA viral polymerases. Curr. Opin. Struct. Biol. 2016, 36, 75–84. [Google Scholar] [CrossRef]
- Chen, X.; He, H.; Yang, X.; Zeng, H.; Qiu, D.; Guo, L. The complete genome sequence of a novel Fusarium graminearum RNA virus in a new proposed family within the order Tymovirales. Arch. Virol. 2016, 161, 2899–2903. [Google Scholar] [CrossRef]
- Davison, A.J. Journal of General Virology–Introduction to ‘ICTV Virus Taxonomy Profiles’. J. Gen. Virol. 2017, 98, 1. [Google Scholar] [CrossRef]
- Liu, L.; Xie, J.; Cheng, J.; Fu, Y.; Li, G.; Yi, X.; Jiang, D. Fungal negative-stranded RNA virus that is related to bornaviruses and nyaviruses. Proc. Natl. Acad. Sci. USA 2014, 111, 12205–12210. [Google Scholar] [CrossRef]
- Revill, P.A.; Davidson, A.D.; Wright, P.J. The nucleotide sequence and genome organization of mushroom bacilliform virus: A single-stranded RNA virus of Agaricus bisporus (Lange) Imbach. Virology 1994, 202, 904–911. [Google Scholar] [CrossRef]
- Saito, M.; Kiguchi, T.; Kusume, T.; Tamada, T. Complete nucleotide sequence of the Japanese isolate S of beet necrotic yellow vein virus RNA and comparison with European isolates. Arch. Virol. 1996, 141, 2163–2175. [Google Scholar] [CrossRef]
- Kondo, H.; Hirano, S.; Chiba, S.; Andika, I.B.; Hirai, M.; Maeda, T.; Tamada, T. Characterization of burdock mottle virus, a novel member of the genus Benyvirus, and the identification of benyvirus-related sequences in the plant and insect genomes. Virus Res. 2013, 177, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Turina, M.; Hillman, B.I.; Izadpanah, K.; Rastgou, M.; Rosa, C.; Consortium, I.R. ICTV virus taxonomy profile: Ourmiavirus. J. Gen. Virol. 2017, 98, 129. [Google Scholar] [CrossRef] [PubMed]
- Sztuba-Solińska, J.; Bujarski, J.J. Insights into the single-cell reproduction cycle of members of the family Bromoviridae: Lessons from the use of protoplast systems. J. Virol. 2008, 82, 10330–10340. [Google Scholar] [CrossRef] [PubMed]
- King, A.M.; Lefkowitz, E.; Adams, M.J.; Carstens, E.B. Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses; Elsevier: Amsterdam, The Netherlands, 2011; Volume 9. [Google Scholar]
- Sabanadzovic, S.; Ghanem-Sabanadzovic, N.A.; Tzanetakis, I.E. Blackberry virus E: An unusual flexivirus. Arch. Virol. 2011, 156, 1665–1669. [Google Scholar] [CrossRef] [PubMed]
- Ghanem-Sabanadzovic, N.A.; Tzanetakis, I.E.; Sabanadzovic, S. Rubus canadensis virus 1, a novel betaflexivirus identified in blackberry. Arch. Virol. 2013, 158, 445–449. [Google Scholar] [CrossRef] [PubMed]
- Howitt, R.L.; Beever, R.E.; Pearson, M.N.; Forster, R.L. Genome characterization of Botrytis virus F, a flexuous rod-shaped mycovirus resembling plant ‘potex-like’viruses. J. Gen. Virol. 2001, 82, 67–78. [Google Scholar] [CrossRef]
- Li, K.; Zheng, D.; Cheng, J.; Chen, T.; Fu, Y.; Jiang, D.; Xie, J. Characterization of a novel Sclerotinia sclerotiorum RNA virus as the prototype of a new proposed family within the order Tymovirales. Virus Res. 2016, 219, 92–99. [Google Scholar] [CrossRef]
- Marzano, S.-Y.L.; Domier, L.L. Novel mycoviruses discovered from metatranscriptomics survey of soybean phyllosphere phytobiomes. Virus Res. 2016, 213, 332–342. [Google Scholar] [CrossRef]
- Charles, J.; Tangudu, C.S.; Hurt, S.L.; Tumescheit, C.; Firth, A.E.; Garcia-Rejon, J.E.; Machain-Williams, C.; Blitvich, B.J. Discovery of a novel Tymoviridae-like virus in mosquitoes from Mexico. Arch. Virol. 2019, 164, 649–652. [Google Scholar] [CrossRef]
- Valverde, R.A.; Khalifa, M.E.; Okada, R.; Fukuhara, T.; Sabanadzovic, S. ICTV Virus Taxonomy Profile: Endornaviridae. J. Gen. Virol. 2019, 100, 1204–1205. [Google Scholar] [CrossRef]
- Roossinck, M.J.; Sabanadzovic, S.; Okada, R.; Valverde, R.A. The remarkable evolutionary history of endornaviruses. J. Gen. Virol. 2011, 92, 2674–2678. [Google Scholar] [CrossRef] [PubMed]
- Okada, R.; Kiyota, E.; Moriyama, H.; Fukuhara, T.; Valverde, R.A. Molecular and biological properties of an endornavirus infecting winged bean (Psophocarpus tetragonolobus). Virus Genes 2017, 53, 141–145. [Google Scholar] [CrossRef] [PubMed]
- Du, Z.; Lin, W.; Qiu, P.; Liu, X.; Guo, L.; Wu, K.; Zhang, S.; Wu, Z. Complete sequence of a double-stranded RNA from the phytopathogenic fungus Erysiphe cichoracearum that might represent a novel endornavirus. Arch. Virol. 2016, 161, 2343–2346. [Google Scholar] [CrossRef] [PubMed]
- Okada, R.; Kiyota, E.; Moriyama, H.; Toshiyuki, F.; Valverde, R.A. A new endornavirus species infecting Malabar spinach, Basellaalba, L., Ed. Arch. Virol. 2014, 159, 807–809. [Google Scholar] [CrossRef] [PubMed]
- Okada, R.; Yong, C.K.; Valverde, R.A.; Sabanadzovic, S.; Aoki, N.; Hotate, S.; Kiyota, E.; Moriyama, H.; Fukuhara, T. Molecular characterization of two evolutionarily distinct endornaviruses co-infecting common bean (Phaseolus vulgaris). J. Gen. Virol. 2013, 94, 220–229. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, K.-i.; Nakamura, H.; Matsumoto, N. Hypovirulent strain of the violet root rot fungus Helicobasidium mompa. J. Gen. Plant Pathol. 2003, 69, 385–390. [Google Scholar] [CrossRef]
- Tuomivirta, T.T.; Kaitera, J.; Hantula, J. A novel putative virus of Gremmeniella abietina type B (Ascomycota: Helotiaceae) has a composite genome with endornavirus affinities. J. Gen. Virol. 2009, 90, 2299–2305. [Google Scholar] [CrossRef]
- Ong, J.W.; Li, H.; Sivasithamparam, K.; Dixon, K.W.; Jones, M.G.; Wylie, S.J. Novel Endorna-like viruses, including three with two open reading frames, challenge the membership criteria and taxonomy of the Endornaviridae. Virology 2016, 499, 203–211. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Lu, S.; Anderson, J.B.; Chitsaz, F.; Derbyshire, M.K.; DeWeese-Scott, C.; Fong, J.H.; Geer, L.Y.; Geer, R.C.; Gonzales, N.R. CDD: A Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res. 2010, 39, D225–D229. [Google Scholar] [CrossRef]
- Li, W.; Zhang, T.; Sun, H.; Deng, Y.; Zhang, A.; Chen, H.; Wang, K. Complete genome sequence of a novel endornavirus in the wheat sharp eyespot pathogen Rhizoctonia cerealis. Arch. Virol. 2014, 159, 1213–1216. [Google Scholar] [CrossRef]
- Suzuki, N.; Ghabrial, S.A.; Kim, K.-H.; Pearson, M.; Marzano, S.-Y.L.; Yaegashi, H.; Xie, J.; Guo, L.; Kondo, H.; Koloniuk, I. ICTV virus taxonomy profile: Hypoviridae. J. Gen. Virol. 2018, 99, 615. [Google Scholar] [CrossRef] [PubMed]
- Shapira, R.; Choi, G.H.; Nuss, D.L. Virus-like genetic organization and expression strategy for a double-stranded RNA genetic element associated with biological control of chestnut blight. EMBO J. 1991, 10, 731–739. [Google Scholar] [CrossRef] [PubMed]
- Hillman, B.I.; Halpern, B.T.; Brown, M.P. A viral dsRNA element of the chestnut blight fungus with a distinct genetic organization. Virology 1994, 201, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Smart, C.; Yuan, W.; Foglia, R.; Nuss, D.; Fulbright, D.; Hillman, B. Cryphonectria hypovirus 3, a virus species in the family Hypoviridae with a single open reading frame. Virology 1999, 265, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Linder-Basso, D.; Dynek, J.N.; Hillman, B.I. Genome analysis of Cryphonectria hypovirus 4, the most common hypovirus species in North America. Virology 2005, 337, 192–203. [Google Scholar] [CrossRef]
- Xie, J.; Xiao, X.; Fu, Y.; Liu, H.; Cheng, J.; Ghabrial, S.A.; Li, G.; Jiang, D. A novel mycovirus closely related to hypoviruses that infects the plant pathogenic fungus Sclerotinia sclerotiorum. Virology 2011, 418, 49–56. [Google Scholar] [CrossRef]
- Aulia, A.; Andika, I.B.; Kondo, H.; Hillman, B.I.; Suzuki, N. A symptomless hypovirus, CHV4, facilitates stable infection of the chestnut blight fungus by a coinfecting reovirus likely through suppression of antiviral RNA silencing. Virology 2019, 533, 99–107. [Google Scholar] [CrossRef]
- Khalifa, M.E.; Pearson, M.N. Characterisation of a novel hypovirus from Sclerotinia sclerotiorum potentially representing a new genus within the Hypoviridae. Virology 2014, 464, 441–449. [Google Scholar] [CrossRef]
- Zhang, R.; Liu, S.; Chiba, S.; Kondo, H.; Kanematsu, S.; Suzuki, N. A novel single-stranded RNA virus isolated from a phytopathogenic filamentous fungus, Rosellinia necatrix, with similarity to hypo-like viruses. Front. Microbiol. 2014, 5, 360. [Google Scholar] [CrossRef]
- Niu, Y.; Yuan, Y.; Mao, J.; Yang, Z.; Cao, Q.; Zhang, T.; Wang, S.; Liu, D. Characterization of two novel mycoviruses from Penicillium digitatum and the related fungicide resistance analysis. Sci. Rep. 2018, 8, 5513. [Google Scholar] [CrossRef]
- Bruenn, J.A.; Warner, B.E.; Yerramsetty, P. Widespread mitovirus sequences in plant genomes. PeerJ 2015, 3, e876. [Google Scholar] [CrossRef] [PubMed]
- Nerva, L.; Vigani, G.; Di Silvestre, D.; Ciuffo, M.; Forgia, M.; Chitarra, W.; Turina, M. Biological and Molecular Characterization of Chenopodium quinoa Mitovirus 1 Reveals a Distinct Small RNA Response Compared to Those of Cytoplasmic RNA Viruses. J. Virol. 2019, 93, e01998-18. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.-J.; Chan, D.; Xiang, Y.; Williams, H.; Li, X.-R.; Sniezko, R.A.; Sturrock, R.N. Characterization of Five Novel Mitoviruses in the White Pine Blister Rust Fungus Cronartium ribicola. PLoS ONE 2016, 11, e0154267. [Google Scholar] [CrossRef] [PubMed]
- Marais, A.; Nivault, A.; Faure, C.; Theil, S.; Comont, G.; Candresse, T.; Corio-Costet, M.-F. Determination of the complete genomic sequence of Neofusicoccum luteum mitovirus 1 (NLMV1), a novel mitovirus associated with a phytopathogenic Botryosphaeriaceae. Arch. Virol. 2017, 162, 2477–2480. [Google Scholar] [CrossRef] [PubMed]
- Turina, M.; Ghignone, S.; Astolfi, N.; Silvestri, A.; Bonfante, P.; Lanfranco, L. The virome of the arbuscular mycorrhizal fungus Gigaspora margarita reveals the first report of DNA fragments corresponding to replicating non-retroviral RNA viruses in fungi. Environ. Microbiol. 2018, 20, 2012–2025. [Google Scholar] [CrossRef] [PubMed]
- Ran, H.; Liu, L.; Li, B.; Cheng, J.; Fu, Y.; Jiang, D.; Xie, J. Co-infection of a hypovirulent isolate of Sclerotinia sclerotiorum with a new botybirnavirus and a strain of a mitovirus. Virol. J. 2016, 13, 92. [Google Scholar] [CrossRef] [PubMed]
- Nibert, M.L.; Vong, M.; Fugate, K.K.; Debat, H.J. Evidence for contemporary plant mitoviruses. Virology 2018, 518, 14–24. [Google Scholar] [CrossRef]
- Volchkov, V.; Volchkova, V.; Netesov, S. Complete nucleotide sequence of the Eastern equine encephalomyelitis virus genome. Mol. Genet. Mikrobiol. Virusol. 1991, 5, 8–15. [Google Scholar]
- Netolitzky, D.J.; Schmaltz, F.L.; Parker, M.D.; Rayner, G.A.; Fisher, G.R.; Trent, D.W.; Bader, D.E.; Nagata, L.P. Complete genomic RNA sequence of western equine encephalitis virus and expression of the structural genes. J. Gen. Virol. 2000, 81, 151–159. [Google Scholar] [CrossRef]
- Kinney, R.M.; Tsuchiya, K.R.; Sneider, J.M.; Trent, D.W. Genetic evidence that epizootic Venezuelan equine encephalitis (VEE) viruses may have evolved from enzootic VEE subtype ID virus. Virology 1992, 191, 569–580. [Google Scholar] [CrossRef]
- Strauss, E.G.; Rice, C.M.; Strauss, J.H. Complete nucleotide sequence of the genomic RNA of Sindbis virus. Virology 1984, 133, 92–110. [Google Scholar] [CrossRef]
- Faragher, S.; Meek, A.; Rice, C.; Dalgarno, L. Genome sequences of a mouse-avirulent and a mouse-virulent strain of Ross River virus. Virology 1988, 163, 509–526. [Google Scholar] [CrossRef]
- Kääriäinen, L.; Takkinen, K.; Keränen, S.; Söderlund, H. Replication of the genome of alphaviruses. J. Cell Sci. 1987, 1987, 231–250. [Google Scholar] [CrossRef]
- Khan, A.H.; Morita, K.; del Carmen Parquet, M.; Hasebe, F.; Mathenge, E.G.; Igarashi, A. Complete nucleotide sequence of chikungunya virus and evidence for an internal polyadenylation site. J. Gen. Virol. 2002, 83, 3075–3084. [Google Scholar] [CrossRef]
- Contigiani, M.S.; Diaz, L.A. Togaviridae. In Arthropod Borne Diseases; Springer: Berlin/Heidelberg, Germany, 2017; pp. 115–135. [Google Scholar]
- Chen, R.; Mukhopadhyay, S.; Merits, A.; Bolling, B.; Nasar, F.; Coffey, L.L.; Powers, A.; Weaver, S.C. ICTV virus taxonomy profile: Togaviridae. J. Gen. Virol. 2018, 99, 761–762. [Google Scholar] [CrossRef]
- Li, L.; Jose, J.; Xiang, Y.; Kuhn, R.J.; Rossmann, M.G. Structural changes of envelope proteins during alphavirus fusion. Nature 2010, 468, 705. [Google Scholar] [CrossRef]
- García, M.L.; Dal Bó, E.; da Graça, J.V.; Gago-Zachert, S.; Hammond, J.; Moreno, P.; Natsuaki, T.; Pallás, V.; Navarro, J.A.; Reyes, C.A. ICTV Virus Taxonomy Profile: Ophioviridae. J. Gen. Virol. 2017, 98, 1161. [Google Scholar] [CrossRef]
- Feau, N.; Dutech, C.; Brusini, J.; Rigling, D.; Robin, C. Multiple introductions and recombination in C ryphonectria hypovirus 1: Perspective for a sustainable biological control of chestnut blight. Evol. Appl. 2014, 7, 580–596. [Google Scholar] [CrossRef]
- Brusini, J.; Robin, C. Mycovirus transmission revisited by in situ pairings of vegetatively incompatible isolates of Cryphonectria parasitica. J. Virol. Methods 2013, 187, 435–442. [Google Scholar] [CrossRef]
- Zilio, G.; Thiévent, K.; Koella, J.C. Host genotype and environment affect the trade-off between horizontal and vertical transmission of the parasite Edhazardia aedis. BMC Evol. Biol. 2018, 18, 59. [Google Scholar] [CrossRef]
- Frank, A.; Saldierna Guzmán, J.; Shay, J. Transmission of bacterial endophytes. Microorganisms 2017, 5, 70. [Google Scholar] [CrossRef] [PubMed]
- Dawe, A.L.; Nuss, D.L. Hypovirus molecular biology: From Koch’s postulates to host self-recognition genes that restrict virus transmission. In Advances in Virus Research; Elsevier: Amsterdam, The Netherlands, 2013; Volume 86, pp. 109–147. [Google Scholar]
- Moleleki, N.; van Heerden, S.W.; Wingfield, M.J.; Wingfield, B.D.; Preisig, O. Transfection of Diaporthe perjuncta with Diaporthe RNA virus. Appl. Environ. Microbiol. 2003, 69, 3952–3956. [Google Scholar] [CrossRef] [PubMed]
- Ihrmark, K.; Stenström, E.; Stenlid, J. Double-stranded RNA transmission through basidiospores of Heterobasidion annosum. Mycol. Res. 2004, 108, 149–153. [Google Scholar] [CrossRef] [PubMed]
- Pearson, M.N.; Beever, R.E.; Boine, B.; Arthur, K. Mycoviruses of filamentous fungi and their relevance to plant pathology. Mol. Plant Pathol. 2009, 10, 115–128. [Google Scholar] [CrossRef] [PubMed]
| Anastomosis Groups (AGs) | Anastomosis Subgroups | Host or Habitat |
|---|---|---|
| AG-1 | 1-IA, 1-IB, 1-IC, 1-ID, 1-IE, 1-IF | Rice, maize, soybean Brassica crops, Sudan grass |
| AG-2 | 2-1, 2-1 II, 2-2 IIIB, 2-2 LP, 2-3 | Tobacco, Brassica crops, six-rowed barley, wheat, rice, grass |
| AG-3 | 3 IV | Tobacco, potato, Brassica crops |
| AG-4 | 4-HGI, 4-HGII, 4-HGIII, AGIIIA | Potato, Brassica crops, cauliflower |
| AG-5 | Brassica crops, soil | |
| AG-6 | Brassica crops, soil | |
| AG-7 | Brassica crops, radish field soil | |
| AG-8 | Brassica crops | |
| AG-9 | Brassica crops | |
| AG-10 | Brassica crops | |
| AG-11 | Brassica crops | |
| AG-12 | Brassica crops | |
| AG-13 | Brassica crops | |
| AGBI | Soil |
| Name (abbr.) Rhizoctonia solani | Classification (Family) | Host Strain | Segment | Genome Size (bp/nt) | 5′ UTR (bp/nt) | 3′ UTR (bp/nt) | ORF Length (bp/nt) | Protein Length (aa) | Molecular Mass (kDa) | Accession Number |
|---|---|---|---|---|---|---|---|---|---|---|
| barnavirus 1 (RsBarV1) | Barnaviridae | DV-8 | (+)ssRNA | 3914 partial | ≥69 | ≥176 | 2033 | 677 | 75.9 | KP900904 |
| 1424 | 474 | 53.1 | ||||||||
| 557 | 185 | 20.7 | ||||||||
| beny-like virus 1 42304-9a (RsBenV1/42304-9a) | Benyviridae | 42304-9a | (+)ssRNA | 1306 partial | ≥1 | ≥1 | ≥1306 | ≥435 | ≥48.6 | KP900902 |
| beny-like virus 1 BR2 (RsBenV1/BR2) | Benyviridae | AG-2.2 LP BR2 | (+)ssRNA | 11666 partial | ≥622 | ≥289 | 10755 | 3584 | 403.9 | MK507778 |
| ourmia-like virus 1 (RsOLV1) | Botourmiaviridae | RsAG2 | (+)ssRNA | 2792 partial | ≥1 | ≥689 | ≥2103 | ≥700 | ≥79.0 | KP900921 |
| Cucumber mosaic virus (CMV) | Bromoviridae | AG-3 | (+)ssRNA | 3309 | 81 | 248 | 2959 | 992 | 111.3 | MG025947 |
| (+)ssRNA | 3053 | 72 | 316 | 2573 | 856 | 96.5 | MG025948 | |||
| 332 | 62 | 7.27 | ||||||||
| (+)ssRNA | 2214 | 97 | 321 | 839 | 278 | 30.3 | MG025949 | |||
| 656 | 218 | 23.8 | ||||||||
| flexivirus 1 (RsFV1) | Deltaflexiviridae | AG2-2 IV DC17 | (+)ssRNA | 10644 | 34 | 176 | 10,433 | 3476 | 381.0 | KX349055 |
| flexi-like virus 1 (RsFLV1) | Deltaflexiviridae | AG-2.2 LP BR9 | (+)ssRNA | 2982 partial | ≥15 | ≥178 | 2888 | 962 | 110.8 | MK507787 |
| endornavirus RS002 (RsEV-RS002) | Endornaviridae | AG-3PT RS002 | (+)ssRNA | 14694 partial | ≥13 | ≥1 | ≥14,680 | ≥4893 | ≥555.6 | KC792590 |
| endornavirus 2 Illinois1 (RsEV2/Illinois1) | Endornaviridae | Illinois1 | (+)ssRNA | 15850 partial | ≥35 | ≥26 | 15,783 | 5262 | 597.0 | KT823701 |
| hypovirus 1 (RsHV1) | Hypoviridae | AG-2.2 LP BR20 | (+)ssRNA | 18371 partial | ≥752 | ≥1584 | 16,033 | 5344 | 363.0 | MK558259 |
| megabirnavirus 1 (RsMBV1) | Megabirnaviridae | AG2-2 IV DC17 | dsRNA | 975 partial | ≥1 | ≥1 | ≥975 | ≥325 | ≥36.2 | KX349071 |
| mitovirus 1 RS002 (RMV1-RS002) | Narnaviridae | AG-3PT RS002 | (+)ssRNA | 2797 partial | ≥192 | ≥126 | 2475 | 825 | 92.7 | KC792591 |
| virus 717 (RsV-717) | Partitiviridae | AG-2 Rhs 717 | dsRNA1 | 2363 | 85 | 88 | 2189 | 730 | 86.0 | AF133290 |
| dsRNA2 | 2206 | 78 | 76 | 2051 | 683 | 76.0 | AF133291 | |||
| partitivirus 1 OA-1 (RsPV1/OA-1) | Partitiviridae | OA-1 | dsRNA1 | 1810 partial | ≥1 | ≥1 | ≥1810 | ≥603 | ≥67.3 | KU299048 |
| partitivirus 2 GD-11 (RsPV2/GD-11) | Partitiviridae | AG-1 IA GD-11 | dsRNA1 | 2020 | 88 | 60 | 1871 | 623 | 72.6 | KF372436 |
| dsRNA2 | 1790 | 107 | 213 | 1469 | 489 | 53.3 | KF372437 | |||
| dsRNA virus 2 A (RsDSRV2/A) | Partitiviridae | AG-2.2 LP A | dsRNA1 | 1942 partial | ≥58 | ≥11 | 1869 | 622 | 76.6 | MK400668 |
| dsRNA2 | 1727 partial | ≥79 | ≥181 | 1467 | 488 | 53.3 | MK400669 | |||
| positive-stranded RNA virus 1 (RsPSV1) | Tymoviridae | Illinois1 | (+)ssRNA | 3492 partial | ≥1 | ≥248 | ≥2265 | ≥754 | ≥85.0 | KT823702 |
| 542 | 180 | 20.2 | ||||||||
| 596 | 198 | 22.2 | ||||||||
| bipartite-like virus 1 (RsBLV1) | Bipartitiviridae | AG-2.2 LP BR1 | dsRNA | 1827 partial | ≥39 | ≥1 | 1787 | 595 | 68.5 | MK492913 |
| 1888 partial | ≥126 | ≥151 | 972 | 323 | 37.0 | MK492914 | ||||
| 552 | 183 | 21.1 | ||||||||
| negative-stranded RNA virus 1 (RsNSRV1) | Betamycoserpentoviridae | DK13-1 | (−)ssRNA | 5593 partial | ≥148 | ≥1 | ≥7237 | ≥2411 | ≥271.0 | KP900919 |
| negative-stranded RNA virus 2 (RsNSRV2) | Betamycoserpentoviridae | 248-36 | (−)ssRNA | 7335 partial | ≥136 | ≥192 | 7145 | 2381 | 267.6 | KP900920 |
| negative-stranded RNA virus 3 (RsNSRV2) | Betamycoserpentoviridae | DK13-3 | (−)ssRNA | 7335 partial | ≥127 | ≥65 | 7142 | 2380 | 267.5 | KP900903 |
| fusarivirus 1 BR18 (RsFV1/BR18) | Fusariviridae | AG-2.2 LP BR18 | (+)ssRNA | 10776 partial | ≥161 | ≥235 | 2194 | 731 | 49.7 | MK558257 |
| 1577 | 525 | 35.7 | ||||||||
| 4682 | 1560 | 106.0 | ||||||||
| alphavirus-like 1 BR15 (RsALV1/BR15) | Mycoalphaviridae | AG-2.2 LP BR15 | (+)ssRNA | 2414 partial | ≥61 | ≥1 | ≥2352 | ≥784 | ≥90.3 | MK507793 |
| bunya/phlebo-like virus 1 (RsBPLV1) | Mycophleboviridae | AG-2.2 LP BR3 | (−)ssRNA | 7804 partial | ≥150 | ≥112 | 7542 | 2513 | 295.6 | MK507779 |
| RNA virus HN008 (RsRV-HN008) | Unclassified | HN008 | dsRNA | 7596 | 38 | 250 | 3539 | 1179 | 128.0 | KP861921 |
| 3710 | 1236 | 140.0 | ||||||||
| dsRNA virus 1 B275 (RsDSRV1/B275) | Unclassified | AG-1 IA B275 | dsRNA1 | 2379 | 131 | 169 | 2078 | 692 | 78.7 | JX976612 |
| dsRNA2 | 1811 | 115 | 298 | 1397 | 465 | 51.8 | JX976613 | |||
| putative virus 1 BR4 (RsV1/BR4) | Unclassified | AG-2.2 LP BR4 | RNA | 6311 partial | ≥48 | ≥374 | 5887 | 1962 | 133.3 | MK507780 |
| M1 dsRNA | Unclassified | AG-3 Rhs 1A | dsRNA | 6398 partial | ≥9 | 419 | 639 | 212 | 24.0 | AF020042 |
| 5172 | 1723 | 196.5 | ||||||||
| M2 dsRNA | Unclassified | AG-3 Rhs 1A | dsRNA | 3570 | 421 | 884 | 2265 | 754 | 84.4 | U51331 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdoulaye, A.H.; Foda, M.F.; Kotta-Loizou, I. Viruses Infecting the Plant Pathogenic Fungus Rhizoctonia solani. Viruses 2019, 11, 1113. https://doi.org/10.3390/v11121113
Abdoulaye AH, Foda MF, Kotta-Loizou I. Viruses Infecting the Plant Pathogenic Fungus Rhizoctonia solani. Viruses. 2019; 11(12):1113. https://doi.org/10.3390/v11121113
Chicago/Turabian StyleAbdoulaye, Assane Hamidou, Mohamed Frahat Foda, and Ioly Kotta-Loizou. 2019. "Viruses Infecting the Plant Pathogenic Fungus Rhizoctonia solani" Viruses 11, no. 12: 1113. https://doi.org/10.3390/v11121113
APA StyleAbdoulaye, A. H., Foda, M. F., & Kotta-Loizou, I. (2019). Viruses Infecting the Plant Pathogenic Fungus Rhizoctonia solani. Viruses, 11(12), 1113. https://doi.org/10.3390/v11121113







