1. Introduction
Flaviviruses are enveloped RNA viruses that are transmitted to vertebrate hosts by mosquito or tick bites. Several members of the flavivirus genus, such as dengue virus (DENV), West Nile virus (WNV), Japanese encephalitis virus (JEV), yellow fever virus (YFV) and Zika virus (ZIKV) are highly pathogenic to humans and constitute major global health problems. Despite differences in cell tropism and pathogenesis, flaviviruses share common genomic organization, replication strategies and virion structure. Their genome consists of a single-stranded, positive-sense RNA molecule of around 10.7 kb, encoding a polyprotein precursor that give rise to seven non-structural (NS) proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B and NS5) and three structural proteins (capsid C, membrane precursor prM and envelope E) upon processing by the viral protease NS3 and its cofactor NS2B, as well as by cellular proteases [
1]. The structural proteins constitute the viral particle, while NS proteins coordinate RNA replication, viral assembly and modulate innate immune responses. To promote their replication, flaviviruses rearrange membranes of the endoplasmic reticulum (ER) into novel organelle-like structures (also termed viral factories) [
2].
YFV, which is the prototypic flavivirus, is responsible for viral hemorrhagic fever resulting in up to 50% fatality [
3]. The virus circulates in tropical Africa and South America. The YFV vaccine 17 D is one of the most successful ever developed [
4]. However, due to poor vaccine coverage and vaccine shortage, YFV regularly resurges in the African and South American continents, as illustrated by recent outbreaks, and is now emerging in Asia [
5]. WNV is endemic throughout Africa, the Middle East, parts of Asia, and Europe. Since an outbreak in the USA in 1999, WNV has emerged as the most common cause of arboviral encephalitis in North and Middle America, causing millions of infections [
6]. ZIKV has recently emerged in the South Pacific, South and Central Americas. Although many cases of ZIKV are asymptomatic and the most common symptoms of infection are rash, fever, and joint pain, the recent outbreaks caused alarm because they were associated with severe fetal abnormalities, including stillbirth and microcephaly, or the Guillain-Barré syndrome in infected adults [
7]. ZIKV infection is now identified as a sexually-transmitted illness as well [
8,
9,
10]. DENV, by affecting an estimated 50–100 million people per year is the most prevalent and rapidly spreading arboviral disease in humans [
11]. Despite the high morbidity and mortality associated with flavivirus infections, antiviral therapies are missing. Thus, there is a pressing need to develop a deeper understanding of the interactions between flaviviruses and their cell host.
To carry out their replicative cycle, flaviviruses rely on hundreds of host gene products. Several genome-scale approaches, based on gene silencing techniques, such as siRNA-based screens [
12,
13,
14] and more recently, CRISPR/Cas9-based screens [
12,
15,
16,
17] have been used to identify flavivirus host factors. Here, we describe a lentiviral-based gain-of-function cDNA screening method to identify candidate host factors involved in flavivirus replication.
2. Materials and Methods
2.1. Cell Lines
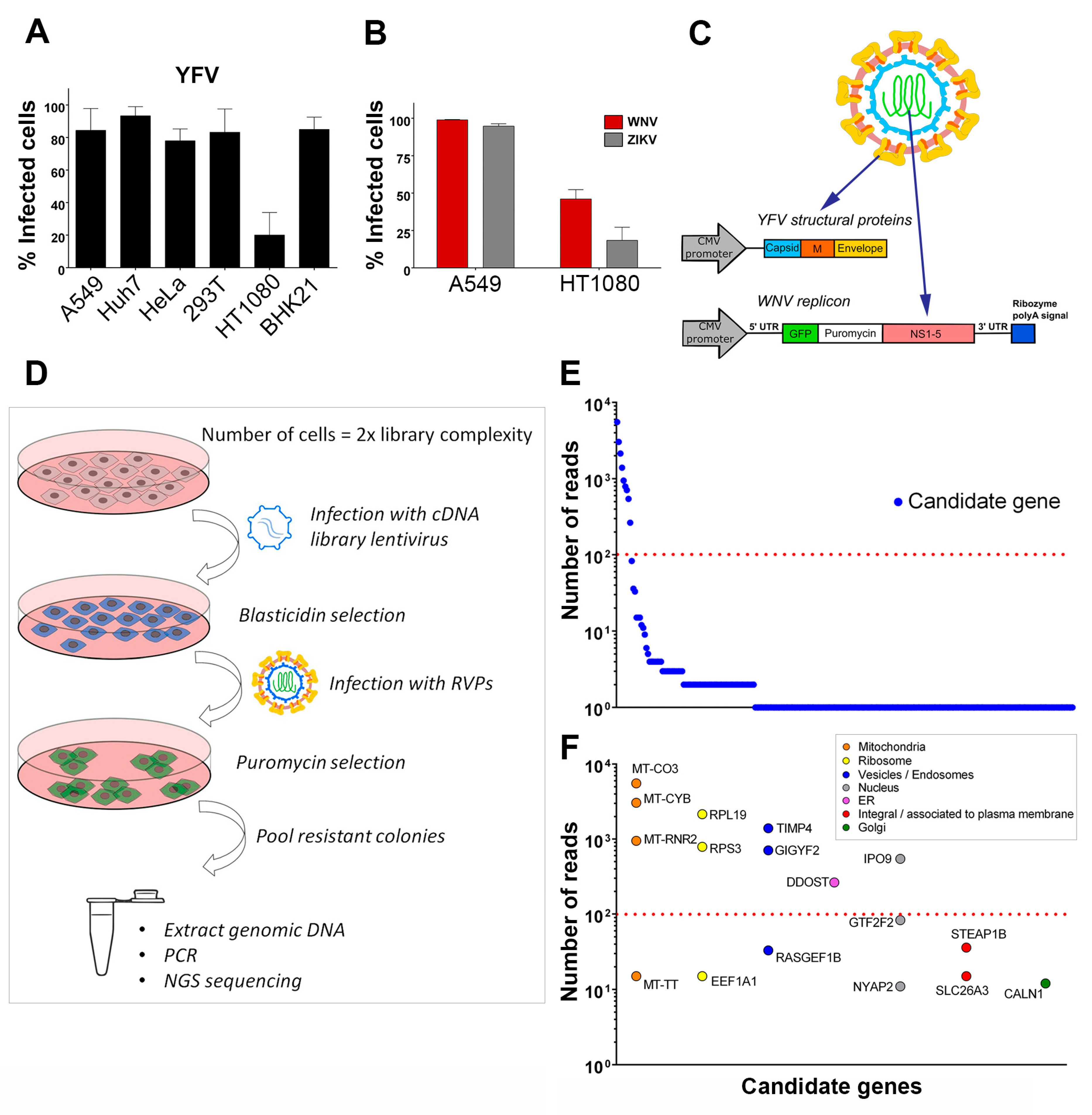
A549 human lung cancer cells, HeLa human cervical cancer cells, HEK293T human embryonic kidney cells, HT1080 human fibrosarcoma cells, BHK-21 baby hamster kidney cells and Vero African green monkey kidney cells were obtained from ATCC (Manassas, VT, USA). Huh7 human hepatocellular carcinoma cells were a gift from E. Meurs (Pasteur Institute, Paris, France). A549, HeLa, HEK293T and BHK-21 cell lines were cultured in Dulbecco’s Modified Eagle Medium (DMEM) (Thermo Fischer Scientific, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (P/S) (Thermo Fischer Scientific, Waltham, MA, USA). HT1080 cells were cultured in DMEM supplemented with 5% FBS and 1% P/S. Huh7 cells were cultured in DMEM supplemented with 10% FBS, 1% P/S and 1% non-essential amino acids (Sigma-Aldrich, St. Louis, MO, USA).
2.2. Virus Stocks and Infection
YFV strain 17D-204 (Stamaril vaccine; Sanofi Pasteur, Lyon, France) was provided by the Pasteur Institute Medical Center. WNV Israeli strain IS-98-STI was provided by the Biological Resource Center of Pasteur Institute. ZIKV strain MR766 was provided by ATCC (Manassas, VT, USA). YFV, WNV and ZIKV virus stocks were grown in Vero cells. Viruses were concentrated by polyethylene glycol 6000 precipitation and purified by centrifugation in a discontinued gradient of sucrose. Viruses were titrated on Vero cells by plaque assay as previously described [
18]. Cell infections were carried at a multiplicity of infection (MOI) of 1. The viral inoculum was replaced with fresh culture medium two hours post-infection.
2.3. Production and Titration of Reporter Virus Particles (RVPs)
The CMV promoter-driven WNV replicon constructs pWNVIIRep-G/Z [
19] was used as a template to generate pWNVIIRep-G/Puro. The GFP-puromycin fusion cassette was generated using the Puromycin coding sequence from pQCXIP vector (Clontech Laboratories, Mountain View, CA, USA) as a template, using forward 5′-GCCCTAGATCTATGACCGAGTACAAGCCCAC-3′ and reverse 5′-GCCCTGGATCCTCAGGCACCGGGCTTGCGGGTCATGCACCA-3′ primers. The PCR product was then ligated into the BglII/BamHI sites of pEGFP-C1 vector (Clontech Laboratories, Mountain View, CA, USA). The resulting GFP-puromycin fusion construct was amplified by PCR using forward 5′-GCCCTACGCGTATGGTGAGCAAGGGC-3′ and reverse 5′-GCCCTACGCGTGGCACCGGGCTTGC-3′ primers and ligated into the MluI site of pWNVIIRep. The codon-optimized sequence the YFV structural genes was synthesized and cloned in pcDNA3.1(+) vector (Life Technologies, Carlsbad, CA, USA). RVPs were produced by transfecting HEK293T cells with pWNVIIRep-G/Puro and pcDNA3.1(+)-YFV CprME vectors at a molar ratio of 1:1 by the calcium phosphate method. One to two hours prior transfection, the cells’ medium was changed with low-glucose DMEM medium (Life Technologies, Carlsbad, CA, USA) supplemented with 10% FBS and 1% P/S. Plasmid DNA was mixed with H
2O and 1 M CaCl
2 solution (Sigma-Aldrich, St. Louis, MO, USA) and then an equal volume of 2x HEPES-buffered saline (pH 7.1, 140 mM NaCl (Sigma-Aldrich, St. Louis, MO, USA), 1.5 mM Na
2HPO
4 (Sigma-Aldrich, St. Louis, MO, USA) and 50 mM HEPES (Sigma-Aldrich, St. Louis, MO, USA) was added to the mixture drop-wise with simultaneous vortexing. The final concentration of CaCl
2 in the transfection mix was 0.25 M. The resulting transfection mixture was incubated at room temperature for 20 min and then added to the cells. After overnight incubation, the medium of the transfected cells was replaced with fresh low-glucose medium. RVPs were harvested 48 and 72 h after transfection. The culture medium containing RVPs was centrifuged at 3000 rpm for 10 min at 4°C to remove cell debris, filtered through 0.45 μm sterile PVDF filter and stored at −80°C.
RVPs’ infectious titer was determined by infecting Huh7 cells for 48 h. The cells were fixed in 1% paraformaldehyde (PFA)-phosphate-buffered saline (PBS) solution for 10 min at room temperature, washed once with PBS and the percentage of GFP-positive cells was determined by flow cytometry. The RVPs titer (IU/mL, infectious units/mL) was calculated using the formula [(F × C)/V] × D, where F equals the frequency of GFP-positive cells (percentage obtained divided by 100), C equals the total number of cells in the well at the start of infection, V equals the volume of inoculum and D is the RVPs dilution [
20].
2.4. cDNA Library Screen
A pooled, uncut, three-reading frame cDNA library of human A549 cells was prepared and cloned into pLenti6/DEST vector (custom-made services, Life Technologies, Carlsbad, CA, USA). A total of 1.12 × 10
7 HT1080 cells were transduced with cDNA library lentiviruses at an MOI 0.2, resulting in two-fold representation of library complexity. Following selection with blasticidin (4 μg/mL) for 10 days, the cDNA library-transduced cells were harvested, 6.2 × 10
7 cells (five-fold representation of cDNA library complexity) were re-seeded and, 24 h later, infected with YFV/WNV chimeric RVPs. On the following day, the cells’ medium was replaced with fresh culture medium containing blasticidin. Selection of RVPs-infected cells with puromycin (2 μg/mL) was started 48 h after infection. After selection with puromycin for three weeks, the resistant colonies (77 in total) were pooled. An aliquot (~ 5 × 10
5 cells) was taken for genomic DNA extraction, which was done by lysing the cells in a buffer containing 100 mM NaCl, 10 mM TrisHCl, pH 8.0, 25 mM EDTA, pH 8.0, 0.5% SDS and 0.1 mg/mL proteinase K for 12 h at 50 °C, followed by phenol/chloroform extraction and ethanol precipitation. PCR for integrated cDNA clone isolation was done using forward 5′- CAGTACATCAATGGGCGTGG -3′ and reverse 5′-GGGGACTTTCCACACCCTAAC -3′ primers, 50–250 ng of genomic DNA, 1 U of Phusion DNA polymerase (Thermo Fischer Scientific, Waltham, MA, USA) and the following cycling parameters: 1× (98 °C, 30 s), 30× (98 °C, 10 s; 68°C, 20 s; 72 °C, 1 min), 1× (72 °C, 10 min; 4 °C, hold). Reactions with genomic DNA from HT1080 parental cells and HT1080 cells transduced with pLenti6-GFP were performed in parallel as negative and positive controls, respectively. NGS of PCR products from the pool of 77 clones was done at Institut Pasteur’s ‘Plateforme Transcriptome et Epigenome (PF2)’ using Illumina MiSEQ instrument. Amplification-free DNA library was prepared using NEXTflex® PCR-Free DNA Library Prep Kit (Bioo Scientific, Austin, TX, USA). The sequencing run was paired-end with a read length of 162 base pairs. Reads were mapped to the human reference genome USCS hg19 using STAR [
21] with parameters adjusted to detect the junction between the lentiviral vector and the cDNA clone, thus restricting the analysis to amplicons derived from the transduced vector and excluding hits derived from possible genomic DNA contaminants. Hits with more than 10 mapped reads were considered candidate genes. For gene classification according to sub-cellular localization, DAVID database, as well as manual content curation of literature for each gene was used.
2.5. siRNA-Mediated Gene Silencing
siRNA oligos (
Table S1) were transfected in HeLa or A549 cells (40–50% confluent) at 30 nM final concentration using Lipofectamine RNAiMax reagent (Thermo Fischer Scientific, Waltham, MA, USA) and following the manufacturer’s protocol. Transfected cells were harvested or used for infection assays 48 h after transfection.
2.6. Flow Cytometry Analyses
Infected cells were fixed with cytofix/cytoperm kit (BD Biosciences, Franklin Lakes, NJ, USA) and stained using the pan-flavivirus anti-Env 4G2 antibody and secondary antibodies. Non-infected, antibody-stained samples served as controls for the signal background. Data were acquired using Attune NxT Acoustic Focusing Cytometer (Life Technologies, Carlsbad, CA, USA) and analyzed using FlowJo software (version 10.1).
2.7. Lentiviral Production and Generation of HT1080 Stable Cell Lines
Lentiviral constructs expressing RPL19, RPS3, DDOST or TIM-1 were generated using the Gateway cloning technology (Life Technologies, Carlsbad, CA, USA). Entry clones for DDOST were obtained from Yves Jacob (Pasteur Institute, Paris, France). RPL19 and RPS3 were cloned into pDONR221 vector (Life Technologies, Carlsbad, CA, USA) using as template HEK293T or HeLa cells cDNA, respectively. TIM-1 was cloned into pDONR221 vector using pTRIP-TIM-1 vector (Ali Amara, IUH, Paris, France) as a template. The primers used for gene amplification by PCR are listed in
Table S2. Stable H1080 cell lines over-expressing our selected genes were generated by lentivirus transduction. Gateway LR recombination reaction was performed to insert the cloned genes into pSCRPSY-TagRFP-DEST (Paul Bieniasz, Rockefeller University, New York, US) or pLenti-CMV-Puro-DEST (# 17452, Addgene, Cambridge, MA, USA) lentiviral vectors. pLenti6-GFP (# 35637, Addgene Cambridge, MA, USA) was used as a positive control for cDNA library lentivirus infection efficiency. Lentiviruses were produced by calcium phosphate-mediated transfection of HEK293T cells with pCMV-VSV-G codon-optimized, pCMVR8.74 (both plasmids were given by Pierre Charneau Pasteur Institute, Paris, France) and lentiviral vector at a mass ratio of 1:4:4. Lentivirus titration was done on HT1080 cells by flow cytometry (pSCRPSY-TagRFP vector) or colony formation (pLenti6 and Lenti-CMV-Puro-DEST vectors) assays. Cells stably expressing the gene of interest were selected and maintained with culture medium containing puromycin at 1 μg/mL final concentration. Cells transduced with empty vector lentivirus were used as controls.
2.8. RNA Purification, Reverse Transcription, and Quantitative Real-Time PCR
Total RNA was extracted from cells using NucleoSpin RNA kit (Macherey-Nagel, Düren, Germany), following the manufacturer’s protocol. cDNA was synthetized by reverse transcription of equal amounts of purified total RNA using random hexamers (Thermo Fischer Scientific, Waltham, MA, USA) and RevertAid H Minus Reverse Transcriptase (EP0451, Thermo Fischer Scientific, Waltham, MA, USA), according to the manufacturers’ instructions. Gene expression was determined by real-time qPCR using primers described in
Table S3, FastStart Universal SYBR Green Master Mix (Roche, Basel, Switzerland), and a Quant Studio 6 Flex PCR instrument (Life Technologies, Carlsbad, CA, USA). Gene expression was quantified according to the ΔΔ
CT method [
22] or the ΔΔ
CT method with correction for efficiency [
23], using GAPDH as endogenous reference control. Standard curves for cellular genes were established using 10-fold serial dilutions of plasmids containing the entire cDNA of the gene of interest. The measured amounts of each mRNA were normalized to the amounts of GAPDH mRNA. The amounts of RNA were expressed as number of copies/μg total RNA. Viral genome copy number per μg RNA was determined by absolute quantification using a standard curve obtained with serial dilutions of the pYFVRneo [
24], pIRES-Hyg2-WNVCprME [
19] or pcDNA6.2 ZIKV [
25] plasmids.
2.9. Immunoblot
Cells were lysed in RIPA buffer (Sigma-Aldrich, St. Louis, MO, USA) supplemented with protease and phosphatase inhibitors cocktail for 30 min at 4 °C with rotation. Lysates were cleared by centrifugation at 20,000× g and 4°C for 15 minutes. Protein extracts were mixed with LDS sample buffer (Thermo Fischer Scientific, Waltham, MA, USA) and NuPAGE reducing agent (Thermo Fischer Scientific, Waltham, MA, USA), boiled for 5 min at 95 °C and separated by SDS-PAGE on 4–12% NuPAGE bis-tris pre-cast polyacrylamide gels (Thermo Fischer Scientific, Waltham, MA, USA). Proteins were transferred onto nitrocellulose membrane (BioRad, Hercules, CA, USA) using Bio-Rad trans-blot protein transfer system. Membranes were blocked in PBS containing 0.1% Tween-20 and 5% non-fat milk for 1 h at room temperature. Membranes were subsequently incubated overnight at 4°C with primary antibodies diluted in blocking buffer. Membranes were washed with PBS containing 0.1% Tween-20 and further incubated with secondary antibodies. Membranes were scanned using Odyssey CLx imaging system (Li-COR Biosciences, Lincoln, NE, USA). The following primary antibodies were used in the study: mouse anti-RPL19 (clone K-12, sc-100830, Santa Cruz Biotechnology, Dallas, TX, USA) diluted 1:500, mouse anti-DENV NS1 17A12 [
26] diluted 1:1000, anti-Env 4G2 antibody diluted 1:2000, rabbit YFV-NS4B [
27] diluted 1:1000, and mouse anti-βActin (reference A5316, Sigma-Aldrich, St. Louis, MO, USA) diluted 1:5000.
2.10. Immunofluorescence Analysis
pDONR-223-DDOST was used as a template to clone DDOST by in vitro recombination into a Gateway compatible peGFP-N1 vector (both plasmids were provided by Yves Jacob, Institut Pasteur Paris) to generate DDOST-GFP. HeLa cells were transfected with Lipofectamine LTX (Thermo Fischer Scientific, Waltham, MA, USA) using four times less DNA and transfection reagents than recommended by the manufacturer. Following two days of transfection, cells were infected at an MOI of 1 with YFV, WNV or ZIKV. Twenty-four hours post-infection, cells were fixed with PFA 4% for 30 min at room temperature (RT). Cells were then permeabilized with PBS, 0,5% Triton X-100 for 10 min at RT, blocked with PBS containing 0,05% Tween-20, 5% BSA (PBSTA) for 30 min and stained with anti-NS1 MAb 6B8 (provided by Marie Flamand, Pasteur Institute, Paris, France) in PBSTA for one hour. Cells were then stained with secondary antibodies Alexa-Fluor 488 in PBSTA for 30 min. Nuclei were stained with NucBlue (Thermo Fischer Scientific, Waltham, MA, USA) for 15 min at RT. Images were acquired with a LSM 720 laser scanning confocal microscope equipped with an ×63 objective (Zeiss, Oberkochen, Germany).
2.11. Cell Viability Assays
Cell viability was determined using a trypan blue exclusion assay.
2.12. Statistical Analysis
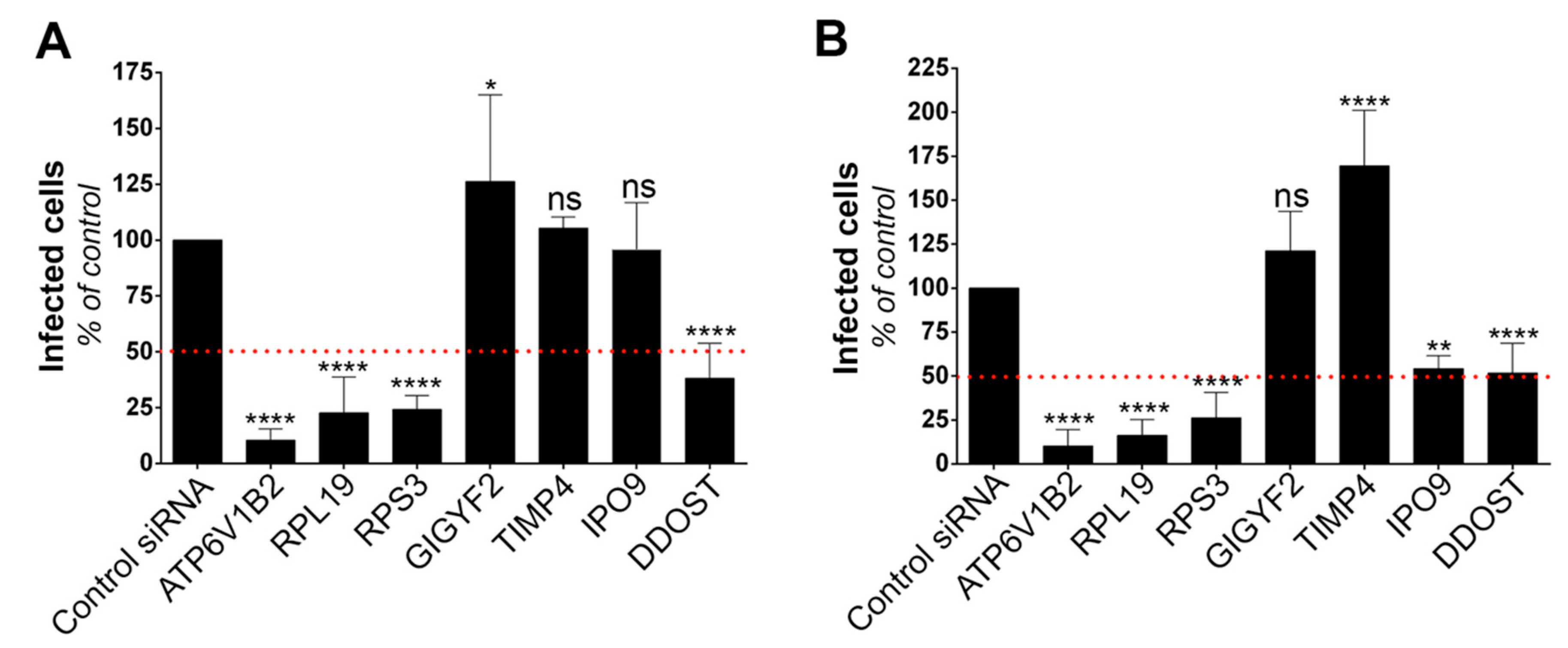
Data were analyzed using GraphPad Prism 7. Statistical analysis was performed with One-way ANOVA or unpaired t tests. Data are presented as means ± SD of at least two independent experiments. Statistically significant differences are indicated as follows: * p < 0.05, ** p < 0.01, *** p < 0.001, and **** p < 0.0001; ns, not significant.
4. Discussion
All the large-scale genetic perturbation screens that have been performed so far in the context of flaviviral infection consisted of loss-of-function screens based on either siRNA or CRISPR approaches [
12,
13,
14,
15,
16,
17]. Limitations of these loss-of-function screens include lack of sensitivity due to incomplete knockdown efficiency, false-negative results because of certain siRNA or gRNAs lacking activity and false-positive candidate genes due to potential off-target effects. We performed a genome-wide gain-of-function screen by over-expressing into HT1080 cells a library of cDNA generated from A459 cells. This approach, based on the generation of RVPs expressing antibiotic selection cassettes, can be adapted to any flavivirus. Our screen allowed the recovery of 17 potential flavivirus host dependency factors, including the previously known factor DDOST. The role of four candidates in YFV and WNV replication (RPL19, RPS3, DDOST and IPO9) was validated using siRNA-silencing assays, demonstrating the utility and relevance of our lentiviral overexpression screening strategy. However, as with any other genetic screens, our approach has its limitations, including the identification of false positive genes. GIGYF2 may be one of them since its reduced expression did not perturb WNV replication. GIGYF2 could have been selected in the screen because its integration in the genome of HT1080 cells disrupted the expression of a gene with antiviral properties. False positive genes may also arise during amplification by PCR of genomic DNA that was not the result of lentiviral integrative events. To limit this, we discarded non-integrated genes from the analysis. Another caveat of genome-wide gain-of-function screens is the potential loss of ORFs during the generation of the cDNA library, especially long ones. In sum, exploiting both gain- and loss-of-function screening strategies in the context of flavivirus infection should provide complementary information.
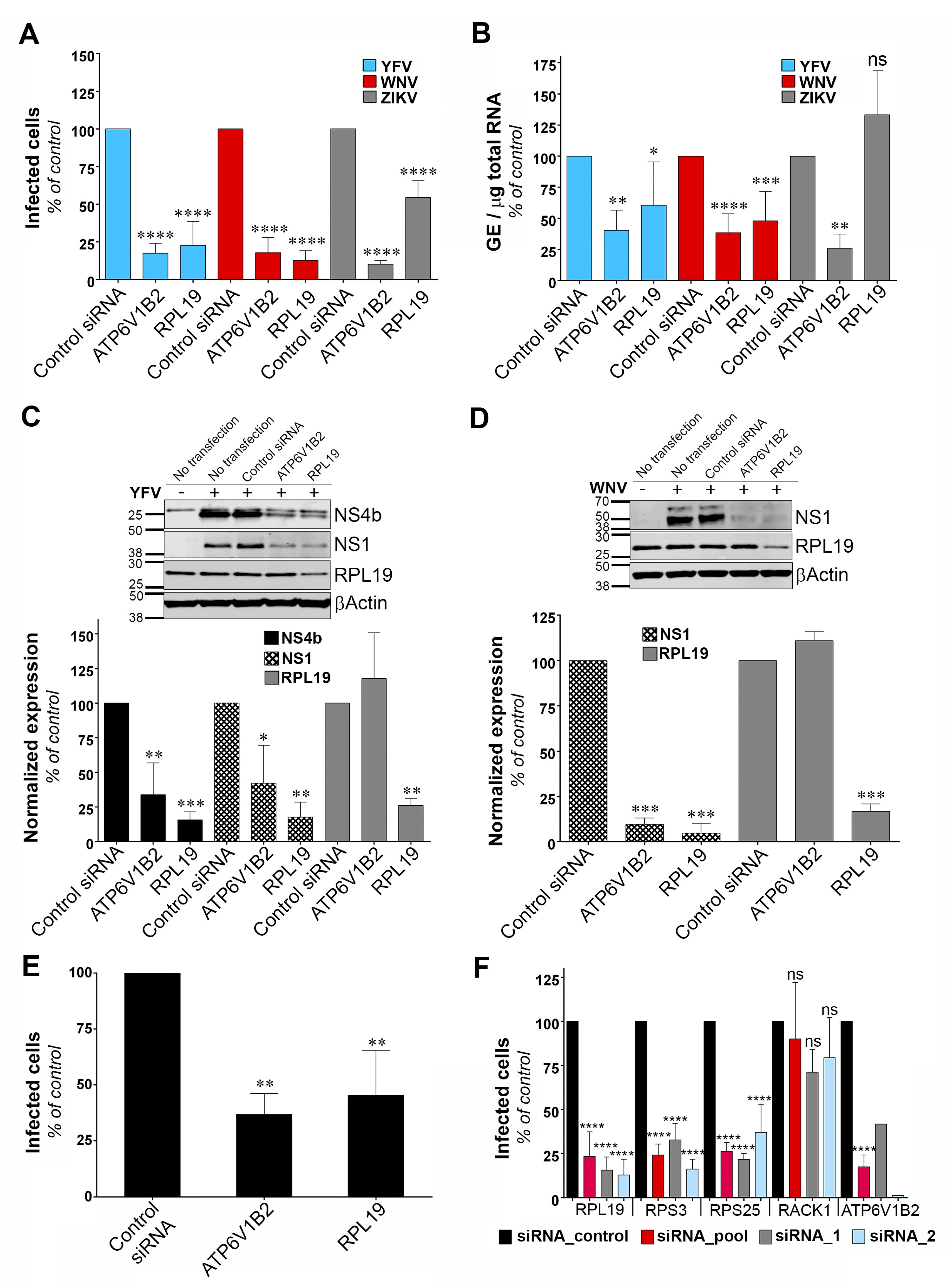
Two of our main hits were the ribosomal proteins RPL19 and RPS3. The screen also recovered EEF1A1, a ribosome associated translation elongation factor. We showed that RPL19 is required for efficient YFV and WNV protein production. ZIKV seemed less dependent on RPL19 than YFV and WNV for efficient replication. Other ribosomal proteins, such as RPS25, RPL18, RPL7, RPL1 and RPLP2 were previously recovered in genome-wide loss-of-function screens performed in the context of DENV, YFV and WNV infections [
13,
15,
36]. Moreover, several ribosomal proteins, including RPS3 and RPL19, were identified as DENV NS1 binding partners using affinity chromatography and immunoprecipitation assays, both in cells over-expressing NS1 [
37] or replicating a DENV replicon [
30]. Among the dozen ribosomal proteins identified as DENV NS1 partners, RPL18 and RPS25 were chosen for further investigations and shown to be required for DENV protein translation [
30,
37]. Similarly, knockdown of the ribosomal proteins RPLP1/2 strongly reduced DENV protein accumulation, suggesting a requirement for RPLP1/2 in viral protein translation [
36]. These data, together with ours, suggest that flaviviruses depend on a subset of ribosomal proteins for efficient replication. One can wonder why, among the 80 ribosomal proteins that exist, only a small subset was recovered in our screen and others [
13,
15,
36]. RPL19 and RPS3 may have been the most abundant ribosomal proteins in the cDNA library that we generated. Alternatively, as suggested [
30,
37], viral NS proteins may recruit and/or anchor ribosomes to the ER via direct (or indirect) interaction with a subset of ribosomal proteins, including RPL19 and RPS3. Finally, one could also envisage that RPL19 and RPS3 have pro-viral functions that are not linked to their ribosomal activities [
38]. Indeed, RPL19 has been proposed to inhibit IRF3 activation in HEK293 cells over-expressing TLR3 [
39]. However, despite our efforts, we were unable to establish a link between RPL19 expression and antiviral IRF3-mediated interferon signaling.
DDOST was among the top-ranked hits of our screen. The human OST complex, which catalyzes the transfer of a preassembled oligosaccharide to selected asparagine residues of polypeptide chains, consists of seven subunits: Ribophorin I (RPN1), ribophorin II (RPN2), DDOST/OST48, DAD1, STT3 (isoform A or B), OST4 and N33/Tusc3 or IAP/MAGT1 [
40]. Our results showed that DDOST is required for efficient replication of YFV, WNV and ZIKV in HeLa cells. This is in line with data showing that STT3A and OSTC expression is required for ZIKV, JEV, DENV and YFV replication in HEK293T cells [
17], as well as with data showing the dependency of DENV, YFV, WNV and ZIKV replication on STT3B in HAP1 cells [
15]. However, surprisingly, replication of YFV and WNV was not dependent on STT3A or STT3B in Huh7.5.1 cells [
16]. Together, these results suggest that different flaviviruses rely on different OST subunits for replication. The use of different cell types may also account for some of the discrepancies described for flavivirus OST-dependency. Immunofluorescence and electron microscopy analysis have revealed co-localization of HA-tagged MAGT1 and STT3B with DENV replication compartments [
15,
16], which is in line with our experiments showing recruitment of DDOST-GFP at viral replication sites during YFV, WNV and ZIKV infection. Direct interactions between STT3A- and DENV-NS2B and -NS3 proteins have also been documented in the context of viral infection [
15] and between DDOST and ectopically expressed DENV-NS1, -NS3 and -NS4B [
30]. Glycosylation of the flaviviral proteins M, E, NS1 and NS4B is key for viral replication and pathogenesis, as well as for viral dissemination in mammalian hosts and mosquito vectors [
34,
41,
42,
43]. Whether the OST complex favors flavivirus replication in a glycosylation-dependent or -independent manner is still a matter of debate. Since DENV replication is drastically inhibited in OST-silenced cells [
15,
16], the assessment of viral protein expression and glycosylation were performed in cells exogenously expressing DENV NS1 and NS4B [
16,
30]. These experiments concluded that the OST complex participates in DENV NS1 and NS4B glycosylation. Our Western blot analysis, which were performed in cells infected with YFV, WNV and ZIKV and silenced for DDOST expression, suggest that DDOST could be involved, at least in part, in YFV-NS1 glycosylation, but not in YFV-NS4B, WNV-NS1 or ZIKV-NS1 glycosylation. However, reduced expression of YFV-NS4B, WNV-NS1, and ZIKV-NS1 in cells silenced for DDOST was observed, suggesting an effect of the OST machinery on the stability of these NS proteins. We propose that the OST complex acts on flavivirus replication by glycosylating a specific subset of viral proteins, such as YFV-NS1, but also by non-canonical functions that stabilize NS proteins, such as YFV-NS4B, WNV-NS1, and ZIKV-NS1. Glycosylation-independent activities of the OST complex may include structural scaffolding function [
15], the catalytic oxidoreductase activity of MAGT1 [
16] and/or promotion of viral protein stability. Alternatively, the OST complex could also favor flavivirus replication via the glycosylation of a proviral cellular factor.
Four mitochondrial genes were identified during the screen (MT-CO3, MT-CYB, MT-RNR2 and MT-TT). It will be of interest to investigate further their potential role in flavivirus replication. A recent proteomic analysis identified a dozen mitochondrial proteins as partners of DENV replication complexes [
30]. Moreover, DENV infection disrupts mitochondria-associated membranes (MAMs), which are ER-connected interfaces critical for innate immune signaling [
2,
44]. Manipulation of specific mitochondrial proteins or MAMs by flaviviruses may thus create a favorable replicative environment by protecting flaviviruses from innate immunity and/or by promoting cell survival.
Another main hit from the screen is TIMP4. However, silencing its expression had no effect on YFV and WNV replication in Hela cells, suggesting that it is dispensable for efficient replication of these two flaviviruses. TIMP4-mediated enhancement of viral replication maybe cell-type specific. Of note, TIMP4 was identified in a yeast-hybrid screen as a partner of the protein 2A of the enterovirus coxsackievirus B3, a positive-strand RNA virus unrelated to flaviviruses [
45] and may be involved in coxsackievirus pathogenesis [
46].
Finally, our data showed that IPO9, which functions mainly in nuclear protein import, was necessary for optimal WNV replication but not required for YFV replication. Its precise role in viral replication, as well as its specificity towards WNV remains to be explored.