The Phylogeny and Pathogenesis of Sacbrood Virus (SBV) Infection in European Honey Bees, Apis mellifera
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Bee Sample Collection and SBV Inoculum Preparation
2.3. RNA Extraction
2.4. RT-PCR and RACE
2.5. Phylogenetic Analysis
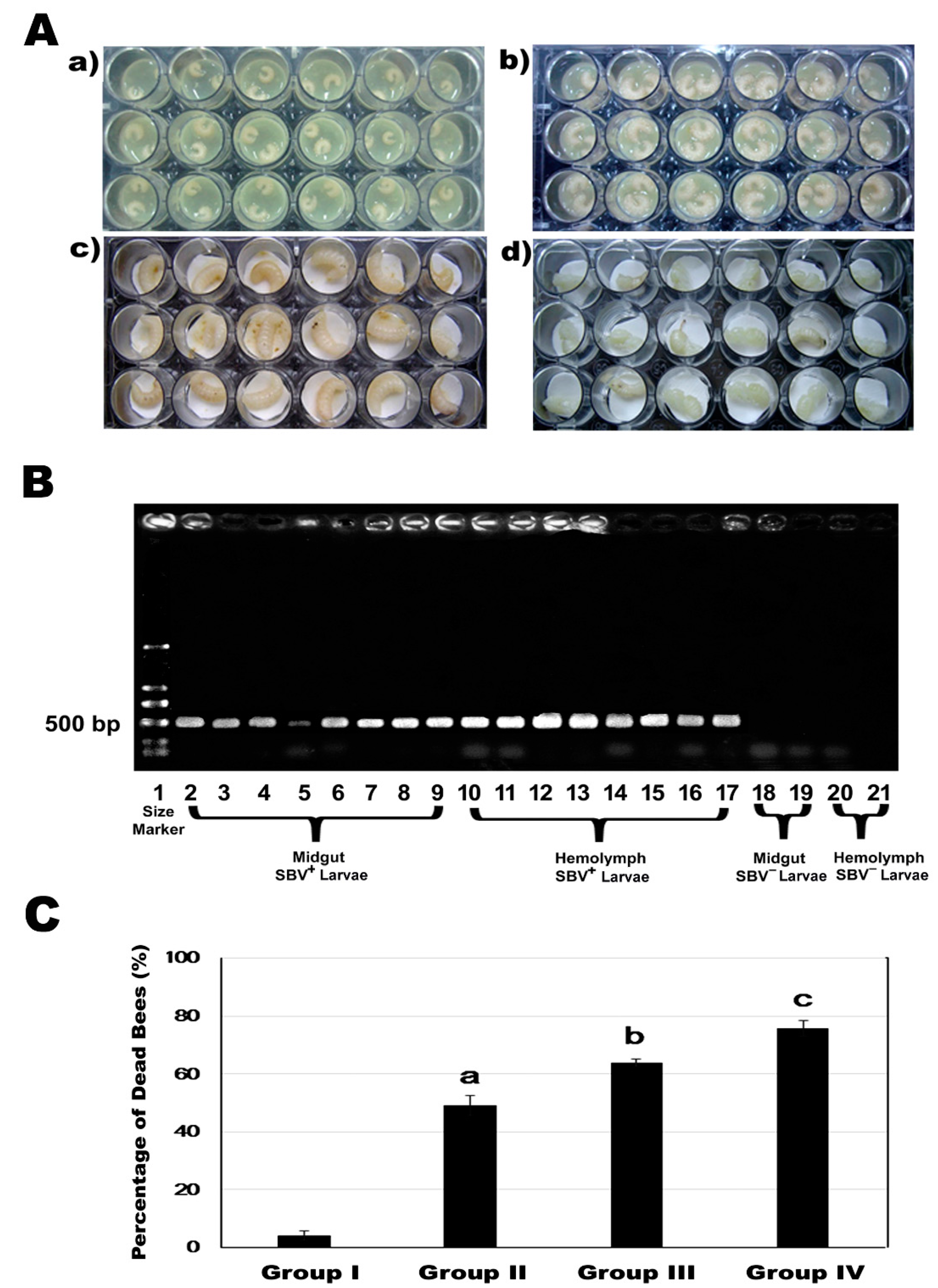
2.6. Laboratory Induced Sbv Pathogenic Infection
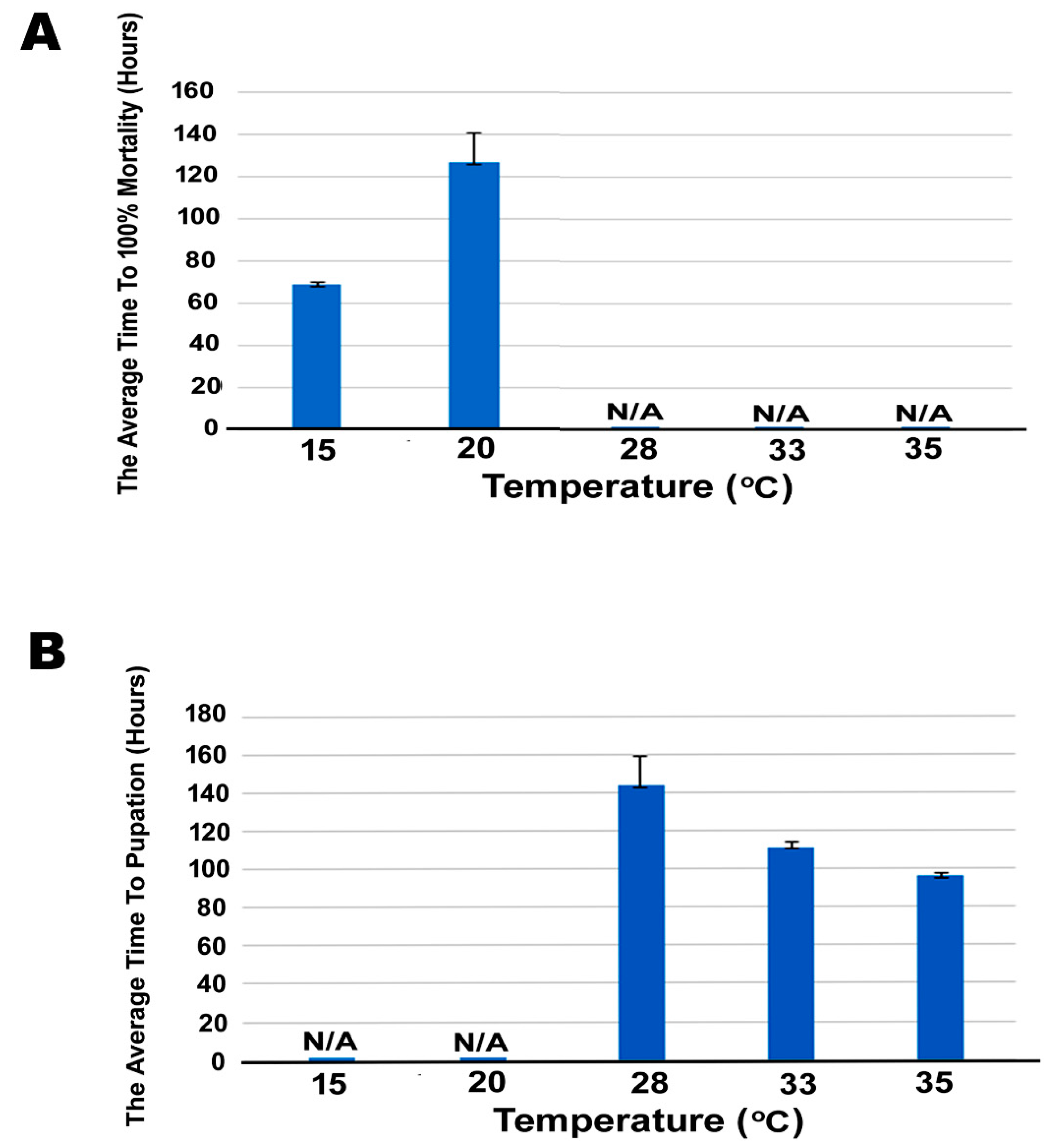
2.7. Cold Stress on Survivorship of SBV-Infected Larvae
3. Results
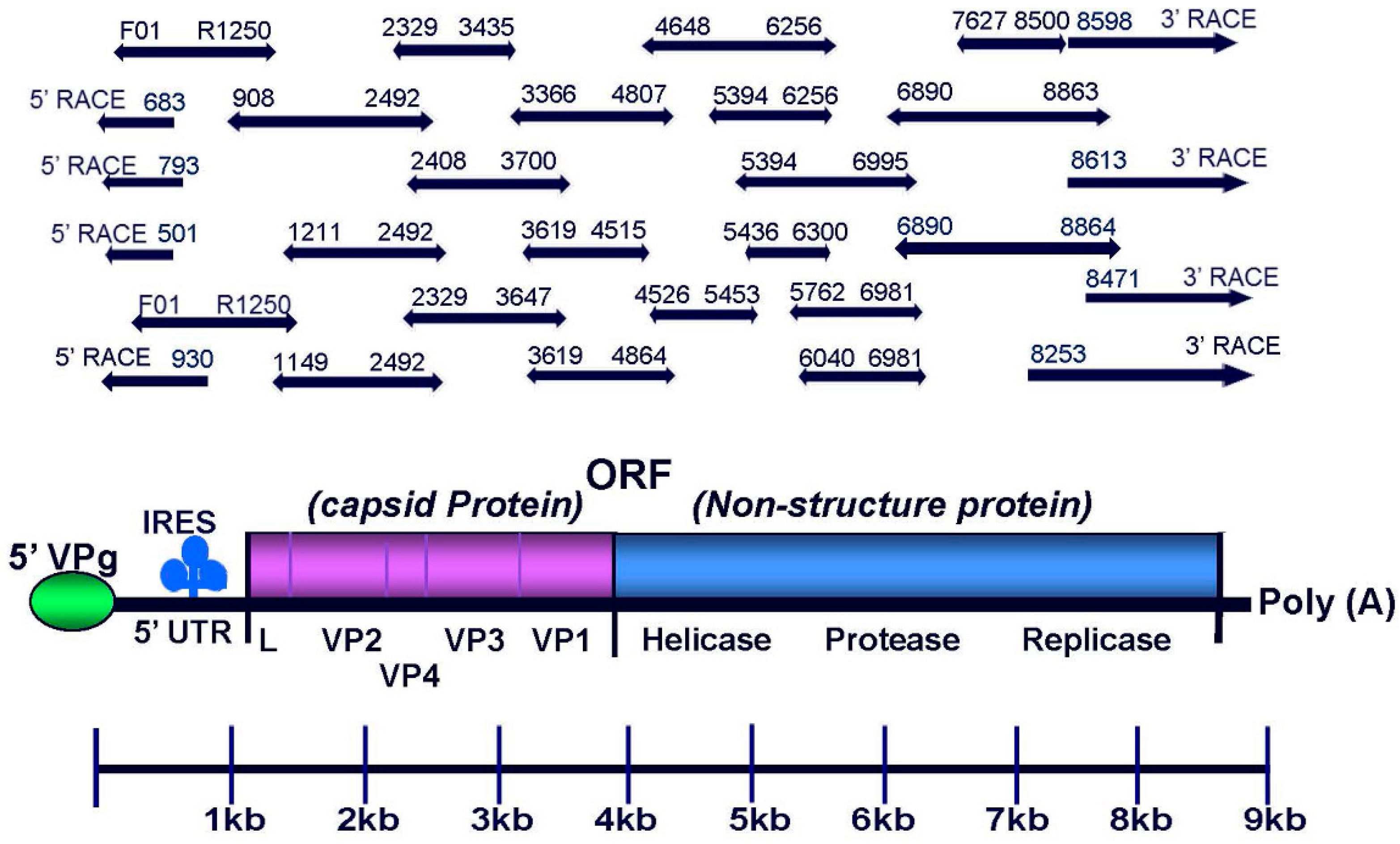
3.1. Complete Genome Sequences of U.S. Strains of SBV
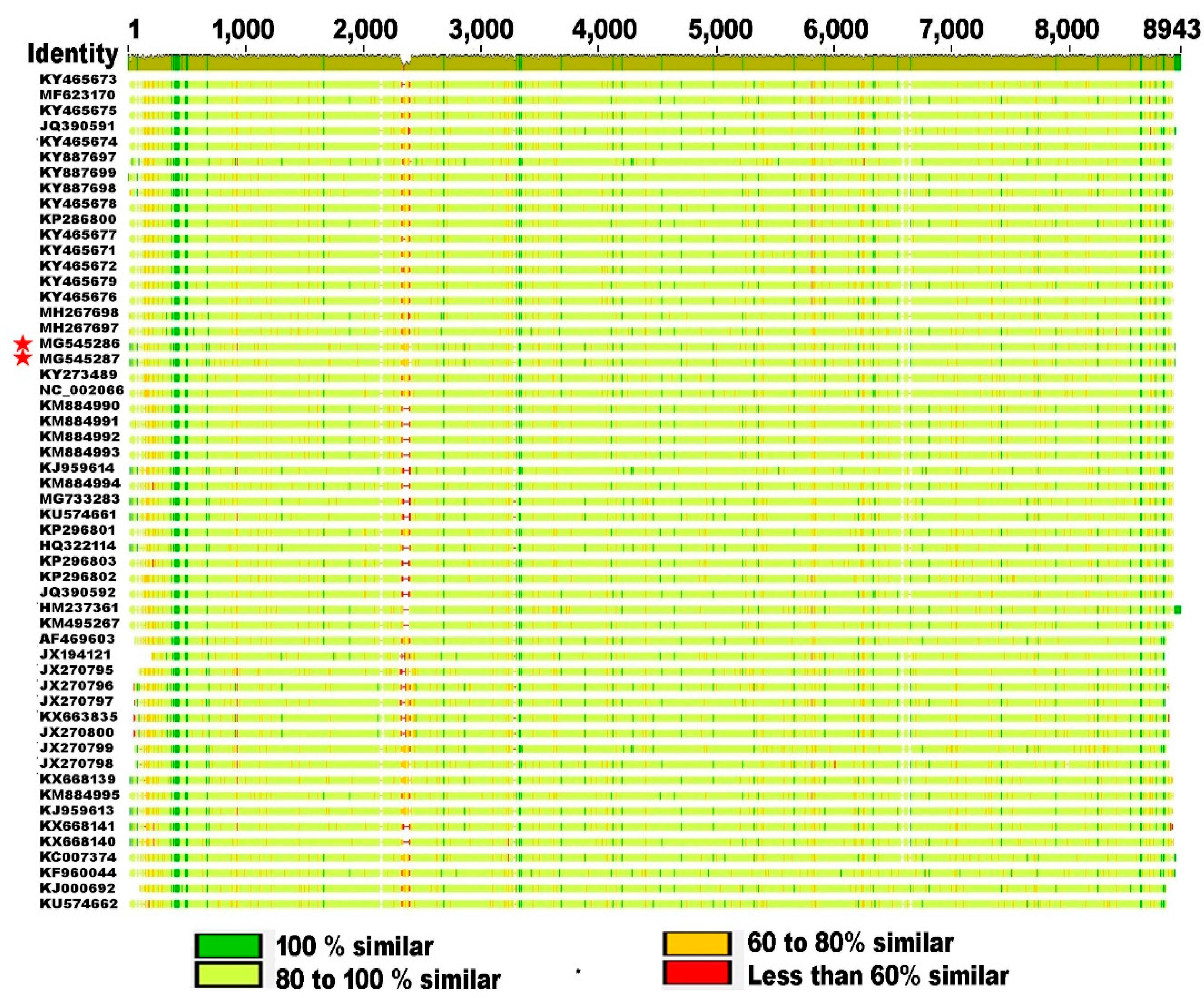
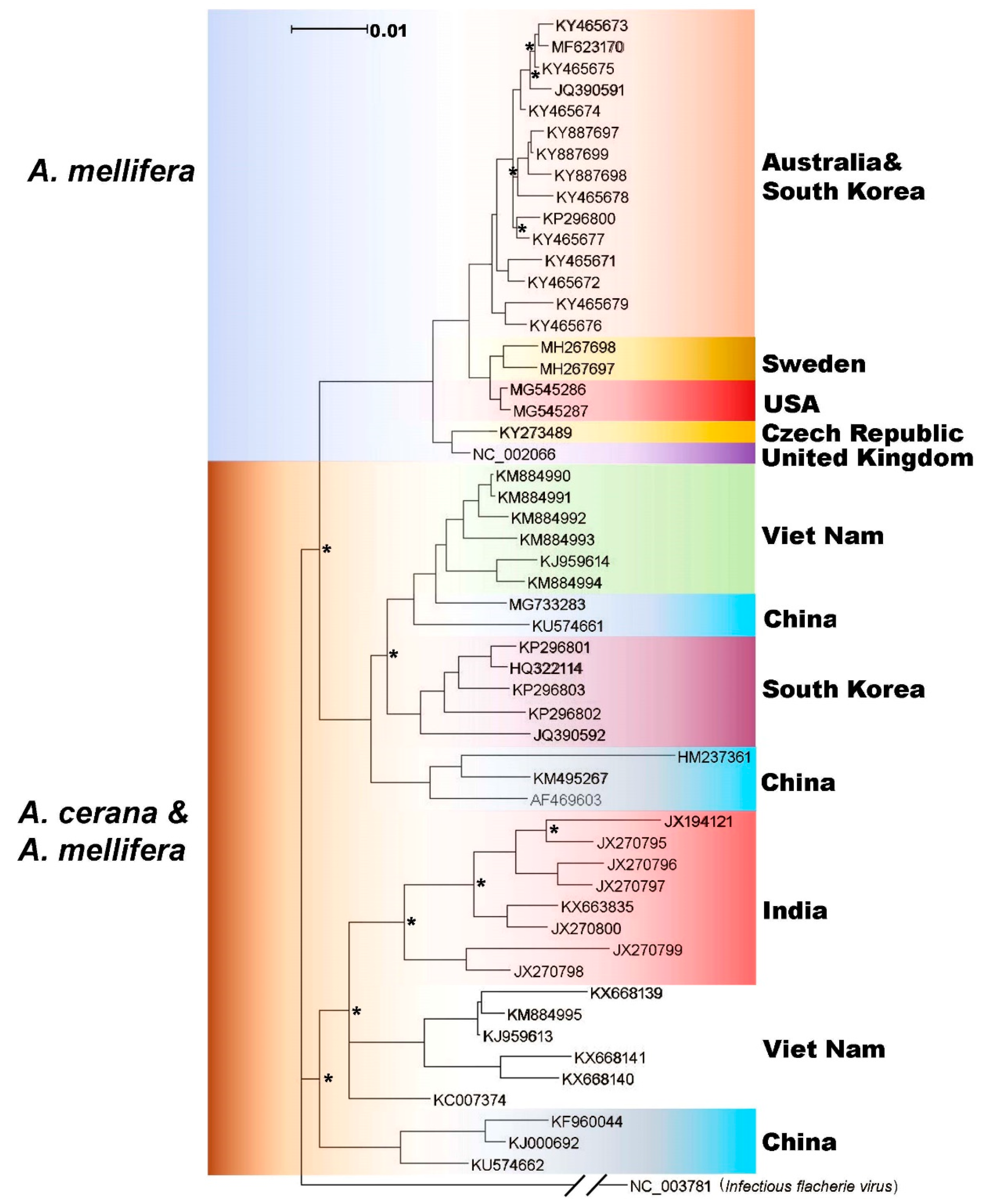
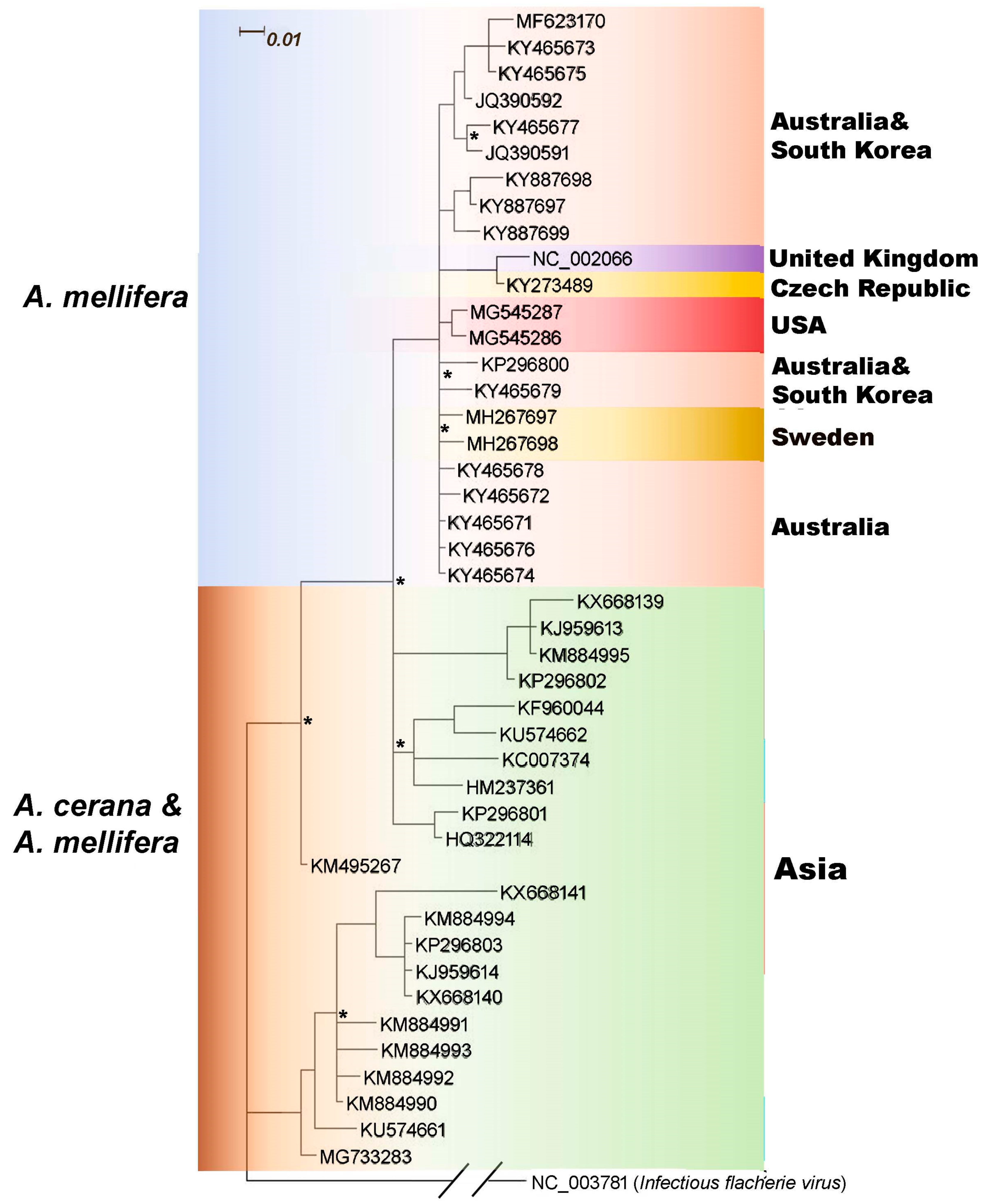
3.2. Genetic Relationship of SBV Strains Worldwide
3.3. Effect of Cold Stress on Mortality Of SBV-Infected Larvae
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Potts, S.G.; Petanidou, T.; Roberts, S.; O’Toole, C.; Hulbert, A.; Willmer, P. Plant-pollinator biodiversity and pollination services in a complex Mediterranean landscape. Biol. Conserv. 2006, 129, 519–529. [Google Scholar] [CrossRef]
- Potts, S.G.; Imperatriz-Fonseca, V.; Ngo, H.T.; Aizen, M.A.; Biesmeijer, J.C.; Breeze, T.D.; Dicks, L.V.; Garibaldi, L.A.; Hill, R.; Settele, J.; et al. Safeguarding pollinators and their values to human well-being. Nature 2016, 540, 220–229. [Google Scholar] [CrossRef] [PubMed]
- Gallai, N.; Salles, J.M.; Settele, J.; Vaissière, B.E. Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol. Econ. 2009, 68, 810–821. [Google Scholar] [CrossRef]
- Aizen, M.A.; Harder, L.D. The global stock of domesticated honey bees is growing slower than agricultural demand for pollination. Curr. Biol. 2009, 19, 915–918. [Google Scholar] [CrossRef] [PubMed]
- Levy, S. The pollinator crisis: What’s best for bees. Nature 2011, 479, 164–165. [Google Scholar] [CrossRef] [PubMed]
- Cox-Foster, D.L.; Conlan, S.; Holmes, E.C.; Palacios, G.; Evans, J.D.; Moran, N.A.; Quan, P.L.; Briese, T.; Hornig, M.; Geiser, D.M.; et al. A metagenomic survey of microbes in honey bee colony collapse disorder. Science 2007, 318, 283–287. [Google Scholar] [CrossRef]
- Kulhanek, K.; Steinhauer, N.; Rennich, N.K.; Caron, D.M.; Sagili, R.R.; Pettis, J.P.; Ellis, J.D.; Wilson, M.J.; Wilkes, J.T.; Tarpy, D.R.; et al. A national survey of managed honey bee 2015–2016 annual colony losses in the USA. J. Apic. Res. 2017, 56, 328–340. [Google Scholar] [CrossRef]
- Potts, S.G.; Biesmeijer, J.C.; Kremen, C.; Neumann, P.; Schweiger, O.; Kunin, W.E. Global pollinator declines: Trends, impacts and drivers. Trends Ecol. Evol. 2010, 25, 345–353. [Google Scholar] [CrossRef]
- Goulson, D.; Nicholls, E.; Botias, C.; Rotheray, E.L. Bee declines driven by combined stress from parasites, pesticides, and lack of flowers. Science 2015, 347, 1255957. [Google Scholar] [CrossRef]
- Vanbergen, A.J.; Insect Pollinators Initiative. Threats to an ecosystem service: Pressures on pollinators. Front. Ecol. Environ. 2013, 11, 251–259. [Google Scholar] [CrossRef]
- Cornman, R.S.; Tarpy, D.R.; Chen, Y.; Jeffreys, L.; Lopez, D.; Pettis, J.S.; vanEngelsdorp, D.; Evans, J.D. Pathogen webs in collapsing honey bee colonies. PLoS ONE 2012, 7, e43562. [Google Scholar] [CrossRef] [PubMed]
- Spivak, M.; Mader, E.; Vaughan, M.; Euliss, N.H., Jr. The plight of the bees. Environ. Sci. Technol. 2011, 45, 34–38. [Google Scholar] [CrossRef] [PubMed]
- White, G.F. Sacbrood, a Disease of Bees; US Department of Agriculture, Bureau of Entomology: Washington, DC, USA, 1913.
- Allen, M.; Ball, B. The incidence and world distribution of honey bee viruses. Bee World 1996, 77, 141–162. [Google Scholar] [CrossRef]
- Chen, Y.P.; Siede, R. Honey bee viruses. Adv. Virus Res. 2007, 70, 33–80. [Google Scholar] [PubMed]
- Bailey, L.; Gibbs, A.J.; Woods, R.D. Sacbrood virus of the larval honeybee (Apis mellifera Linnaeus). Virology 1964, 23, 425–429. [Google Scholar] [CrossRef]
- Shen, M.; Cui, L.; Ostiguy, N.; Cox-Foster, D. Intricate transmission routes and interactions between picorna-like viruses (Kashmir bee virus and sacbrood virus) with the honeybee host and the parasitic varroa mite. J. Gen. Virol. 2005, 86, 2281–2289. [Google Scholar] [CrossRef]
- Grabensteiner, E.; Ritter, W.; Carter, M.J.; Davison, S.; Pechhacker, H.; Kolodziejek, J.; Boecking, O.; Derakhshifar, I.; Moosbeckhofer, R.; Licek, E.; et al. Sacbrood virus of the honeybee (Apis mellifera): Rapid identification and phylogenetic analysis using reverse transcription-PCR. Clin. Diagn. Lab. Immunol. 2001, 8, 93–104. [Google Scholar] [CrossRef]
- Anderson, D.L.; Gibbs, A.J. Inapparent virus infections and their interactions in pupae of the honeybee (Apis mellifera L.) in Australia. J. Gen. Virol. 1988, 69, 1617–1625. [Google Scholar] [CrossRef]
- Berenyi, O.; Bakonyi, T.; Derakhshifar, I.; Koglberger, H.; Nowotny, N. Occurrence of six honeybee viruses in diseased Austrian apiaries. Appl. Environ. Microbiol. 2006, 72, 2414–2420. [Google Scholar] [CrossRef]
- Tentcheva, D.; Gauthier, L.; Zappulla, N.; Dainat, B.; Cousserans, F.; Colin, M.E.; Bergoin, M. Prevalence and seasonal variations of six bee viruses in Apis mellifera L. and Varroa destructor mite populations in France. Appl. Environ. Microbiol. 2004, 70, 7185–7191. [Google Scholar] [CrossRef]
- De Miranda, J.R.; Gauthier, L.; Ribière, M.; Chen, Y.P. Honey bee viruses and their effects on bee and colony health. In Honey Bee Colony Health: Challenges and Sustainable Solution; Sammataro, D., Yoder, J.A., Eds.; CRC Press Taylor&Francis Group: Boca Raton, FL, USA, 2011; pp. 71–102. [Google Scholar]
- Oldroyd, B.P. What’s killing American honey bees? PLoS Biol. 2007, 5, e168. [Google Scholar] [CrossRef] [PubMed]
- Martin, S.J.; Highfield, A.C.; Brettell, L.; Villalobos, E.M.; Budge, G.E.; Powell, M.; Nikaido, S.; Schroeder, D.C. Global honey bee viral landscape altered by a parasitic mite. Science 2012, 336, 1304–1306. [Google Scholar] [CrossRef] [PubMed]
- Rosenkranz, P.; Aumeier, P.; Ziegelmann, B. Biology and control of Varroa destructor. J. Invertebr. Pathol. 2010, 103 (Suppl. 1), S96–S119. [Google Scholar] [CrossRef] [PubMed]
- De Miranda, J.R.; Genersch, E. Deformed wing virus. J. Invertebr. Pathol. 2010, 103 (Suppl. 1), S48–S61. [Google Scholar] [CrossRef] [PubMed]
- Woolhouse, M.E.; Gowtage-Sequeria, S. Host range and emerging and reemerging pathogens. Emerg. Infect. Dis. 2005, 11, 1842–1847. [Google Scholar] [CrossRef]
- Xia, X.; Zhou, B.; Wei, T. Complete genome of Chinese sacbrood virus from Apis cerana and analysis of the 3C-like cysteine protease. Virus Genes 2015, 50, 277–285. [Google Scholar] [CrossRef]
- Ghosh, R.C.; Ball, B.V.; Willcocks, M.M.; Carter, M.J. The nucleotide sequence of sacbrood virus of the honey bee: An insect picorna-like virus. J. Gen. Virol. 1999, 80 Pt 6, 1541–1549. [Google Scholar] [CrossRef]
- Choe, S.E.; Nguyen, L.T.K.; Noh, J.H.; Kweon, C.H.; Reddy, K.E.; Koh, H.B.; Chang, K.Y.; Kang, S.W. Analysis of the complete genome sequence of two Korean sacbrood viruses in the Honey bee, Apis mellifera. Virology 2012, 432, 155–161. [Google Scholar] [CrossRef]
- Roberts, J.M.; Anderson, D.L. A novel strain of sacbrood virus of interest to world apiculture. J. Invertebr. Pathol. 2014, 118, 71–74. [Google Scholar] [CrossRef]
- Roy, C.; Vidal-Naquet, N.; Provost, B. A severe sacbrood virus outbreak in a honeybee (Apis mellifera L.) colony: A case report. Vet. Med. 2015, 60, 330–335. [Google Scholar] [CrossRef]
- Bailey, L.; Carpenter, J.M.; Woods, R.D. A strain of sacbrood virus from Apis cerana. J. Invertebr. Pathol. 1982, 39, 264–265. [Google Scholar] [CrossRef]
- Rana, B.S.; Garg, I.D.; Khurana, S.P.; Verma, L.R.; Agrawal, H.O. Thai sacbrood virus of honeybees (Apis cerana indica F) in North-west Himalayas. Indian J. Virol. 1986, 2, 127–131. [Google Scholar]
- Verma, L.R.; Rana, B.S.; Verma, S. Observations on Apis cerana colonies surviving from Thai sacbrood virus infestation. Apidologie 1990, 21, 169–174. [Google Scholar] [CrossRef]
- Zhang, J.; Feng, J.; Liang, Y.; Cheng, D.; Zhou, Z.H.; Zhang, Q.; Lu, X. Three-dimensional structure of the Chinese sacbrood bee virus. Sci. China 2001, 44, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Mingxiao, M.; Ming, L.; Jian, C.; Song, Y.; Shude, W.; Pengfei, L. Molecular and Biological Characterization of Chinese Sacbrood Virus LN Isolate. Comp. Funct. Genom. 2011, 2011, 409386. [Google Scholar] [CrossRef]
- Choe, S.E.; Nguyen, L.T.K.; Noh, J.H.; Koh, H.B.; Jean, Y.H.; Kweon, C.H.; Kang, S.W. Prevalence and distribution of six bee viruses in Korean Apis cerana populations. J. Invertebr. Pathol. 2012, 109, 330–333. [Google Scholar] [CrossRef]
- Nguyen, N.T.; Le, T.H. Complete Genome Sequence of Sacbrood Virus Strain SBM2, Isolated from the Honeybee Apis cerana in Vietnam. Genome Announc. 2013, 1, e00076-12. [Google Scholar] [CrossRef]
- Rao, K.M.; Katna, S.; Rana, B.S.; Rana, R. Thai sacbrood and sacbrood viruses versus European foulbrood of hive bees in India—A review. J. Apic. Res. 2015, 54, 192–199. [Google Scholar]
- Li, Y.; Zeng, Z.J.; Wang, Z.L. Phylogenetic analysis of the honeybee Sacbrood virus. J. Apic. Sci. 2016, 60, 31–38. [Google Scholar] [CrossRef]
- Gong, H.R.; Chen, X.X.; Chen, Y.P.; Hu, F.L.; Zhang, J.L.; Lin, Z.G.; Yu, J.W.; Zheng, H.Q. Evidence of Apis cerana Sacbrood virus Infection in Apis mellifera. Appl. Environ. Microbiol. 2016, 82, 2256–2262. [Google Scholar] [CrossRef]
- Li, J.L.; Cornman, R.S.; Evans, J.D.; Pettis, J.S.; Zhao, Y.; Murphy, C.; Peng, W.J.; Wu, J.; Hamilton, M.; Boncristiani, H.F., Jr.; et al. Systemic spread and propagation of a plant-pathogenic virus in European honeybees, Apis mellifera. mBio 2014, 5, e00898-13. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 2004, 5, 113. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Vandenberg, J.D.; Shimanuki, H. Technique for Rearing Worker Honeybees in the Laboratory. J. Apic. Res. 1987, 26, 90–97. [Google Scholar] [CrossRef]
- LME4: Linear Mixed-Effects Models Using S4 Classes. Available online: http://cran.R-project.org/package=lme4 (accessed on 5 January 2019).
- Fares, A. Factors influencing the seasonal patterns of infectious diseases. Int. J. Prev. Med. 2013, 4, 128–132. [Google Scholar] [PubMed]
- Gammon, D.B.; Mello, C.C. RNA interference-mediated antiviral defense in insects. Curr. Opin. Insect Sci. 2015, 8, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Brutscher, L.M.; Flenniken, M.L. RNAi and Antiviral Defense in the Honey Bee. J. Immunol. Res. 2015, 2015, 941897. [Google Scholar] [CrossRef]
- Nylin, S.; Gotthard, K. Plasticity in Life-History Traits. Annu. Rev. Entomol. 1998, 43, 63–83. [Google Scholar] [CrossRef]
- Dodson, B.L.; Kramer, L.D.; Rasgon, J.L. Effects of larval rearing temperature on immature development and West Nile virus vector competence of Culex tarsalis. Parasites Vectors 2012, 5, 199. [Google Scholar] [CrossRef]
- Kronenberg, F.; Heller, H.C. Colonial thermoregulation in honey bees (Apis mellifera). J. Comp. Physiol. B 1982, 148, 65–76. [Google Scholar] [CrossRef]
- Wang, Q.; Xu, X.; Zhu, X.; Chen, L.; Zhou, S.; Huang, Z.Y.; Zhou, B. Low-Temperature Stress during Capped Brood Stage Increases Pupal Mortality, Misorientation and Adult Mortality in Honey Bees. PLoS ONE 2016, 11, e0154547. [Google Scholar] [CrossRef]
| Primer Name | Primer Sequences (5′ to 3′) |
|---|---|
| F01 | TACGAATCGTGATTCGATTCATT |
| F908 | ACGCTAAGTGTGCGCCTAAT |
| F1149 | GATATCATCGCGCCTTTGTT |
| F1211 | GGAAGTTTGCTAGTATTTACGTG |
| F2329 | TGGAGGTAAGGGACAACCTG |
| F2408 | TGGACACTGGTGCTAAAGAAGATG |
| F3083 | GAAGCTGGGGATGATTTTGA |
| F3366 | CGCTATAGGTGGCATGCAGA |
| F3619 | GGCATGGATTGATCGAAGTT |
| F4526 | CCGGAAATGGCTCATTTAGA |
| F4648 | TACCGTAGGAGAGATGTGTTATTGT |
| F5394 | TCGAGAAGTGGTTCAGTGCC |
| F5436 | AAGTAGTCCAGTGCCCGATG |
| F5762 | TCGGATGGTGAAGATGATGA |
| F6040 | CCAGCGGTACAAAGAGGAAA |
| F6155 | CACTGGATGAGAGCGAATGA |
| F6890 | CGGTATTTTATGCGAATGATGT |
| F7627 | CCCAGCGTTCTGGAGGAAAT |
| F8256 | AACGAGGGCAAACTTGGGAA |
| F8471 | CATGGGTTTCATCCCCACGA |
| F8598 | GTCGAGCCGCTCTGTATCAA |
| F8613 | ATCAAGCGCATGGTCATGGA |
| R501 | ACTGCGCGTCTAACATTCCA |
| R683 | AACTCTGCTGTGTAGCGTCC |
| R793 | TTGTTGCGTTGGTTCGGAAG |
| R930 | GTGTTGGGTGCACACTTAGC |
| R1250 | AACGGTGACAGCATTTGCAC |
| R2450 | CCACAAGCTCCTTGTTGTGA |
| R2492 | GAATCCAAAGACTGAAAACCC |
| R3435 | CCCGTAACGGTATTCTGCAT |
| R3647 | CTGCAGTCCAACTTCGATCA |
| R3700 | GCTGCCCAAAAGTTGTAGCC |
| R3936 | ACCCCCACCAAATAAGAAGG |
| R4515 | TGCATTAAAGCTTGGGGTTC |
| R4807 | CAGGTTTAGATGTACACGAGGATG |
| R4884 | AACTTCGCAACCAACCAATC |
| R5453 | TCGGGCACTGGACTACTTCT |
| R5871 | ATTTCCCTCTCTCGCATCAA |
| R6256 | CCACTGCCGTCACTAAACCT |
| R6300 | TCATTCGCTCTCATCCAGTG |
| R6609 | ATACTCCCCACTGCCATCAC |
| R6981 | TCCTTAATGGCACGCACATA |
| R6995 | TGCATTCCCACAATAGGTCTTT |
| R8500 | TAGAATTCCAAGCCAGCGCA |
| R8863 | TAAAATGCCATATATTGATATTAATCCA |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, J.; Wang, T.; Evans, J.D.; Rose, R.; Zhao, Y.; Li, Z.; Li, J.; Huang, S.; Heerman, M.; Rodríguez-García, C.; et al. The Phylogeny and Pathogenesis of Sacbrood Virus (SBV) Infection in European Honey Bees, Apis mellifera. Viruses 2019, 11, 61. https://doi.org/10.3390/v11010061
Li J, Wang T, Evans JD, Rose R, Zhao Y, Li Z, Li J, Huang S, Heerman M, Rodríguez-García C, et al. The Phylogeny and Pathogenesis of Sacbrood Virus (SBV) Infection in European Honey Bees, Apis mellifera. Viruses. 2019; 11(1):61. https://doi.org/10.3390/v11010061
Chicago/Turabian StyleLi, Jianghong, Tingyun Wang, Jay D. Evans, Robyn Rose, Yazhou Zhao, Zhiguo Li, Jilian Li, Shaokang Huang, Matthew Heerman, Cristina Rodríguez-García, and et al. 2019. "The Phylogeny and Pathogenesis of Sacbrood Virus (SBV) Infection in European Honey Bees, Apis mellifera" Viruses 11, no. 1: 61. https://doi.org/10.3390/v11010061
APA StyleLi, J., Wang, T., Evans, J. D., Rose, R., Zhao, Y., Li, Z., Li, J., Huang, S., Heerman, M., Rodríguez-García, C., Banmeke, O., Brister, J. R., Hatcher, E. L., Cao, L., Hamilton, M., & Chen, Y. (2019). The Phylogeny and Pathogenesis of Sacbrood Virus (SBV) Infection in European Honey Bees, Apis mellifera. Viruses, 11(1), 61. https://doi.org/10.3390/v11010061







