Molecular Characterization and Geographic Distribution of a Mymonavirus in the Population of Botrytis cinerea
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Strains, Culture Conditions, and Biological Characterization
2.2. dsRNA Extraction and Purification
2.3. cDNA Cloning and Sequencing
2.4. Sequence Analysis
2.5. Detection of BcMyV1 in B. cinerea Population
3. Results
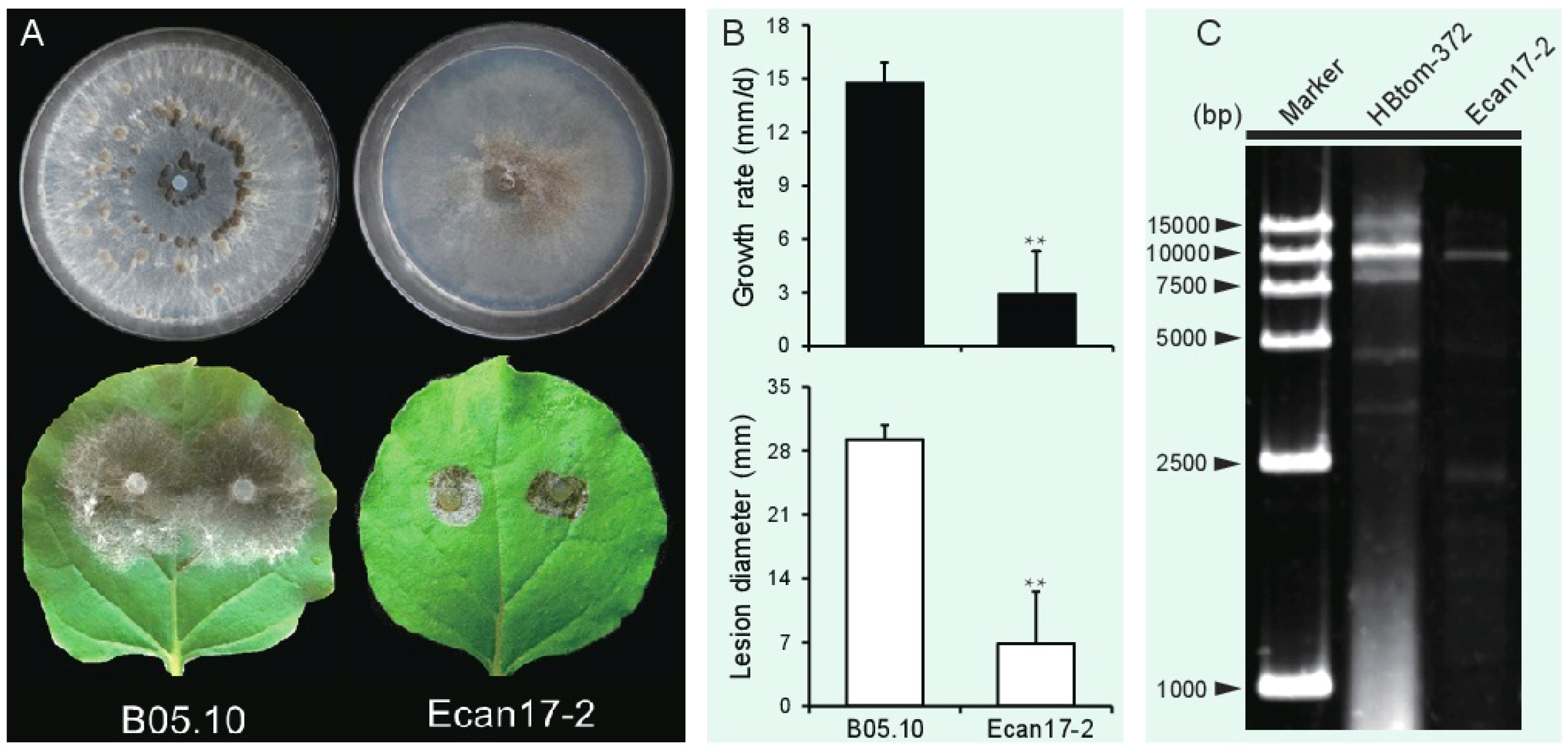
3.1. B. cinerea Strain Ecan17-2 Exhibits Hypovirulence Traits
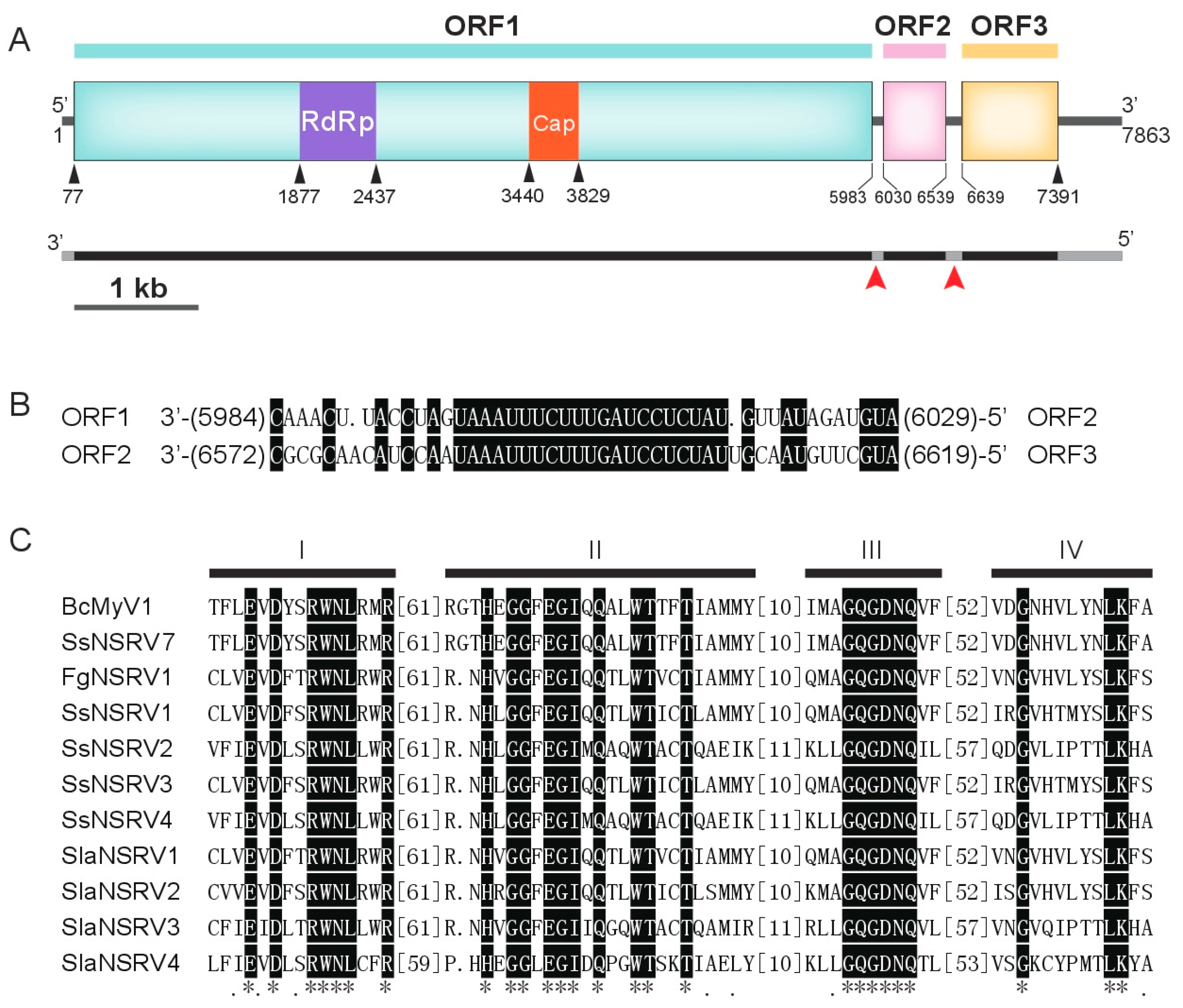
3.2. Genome Analysis of BcMyV1
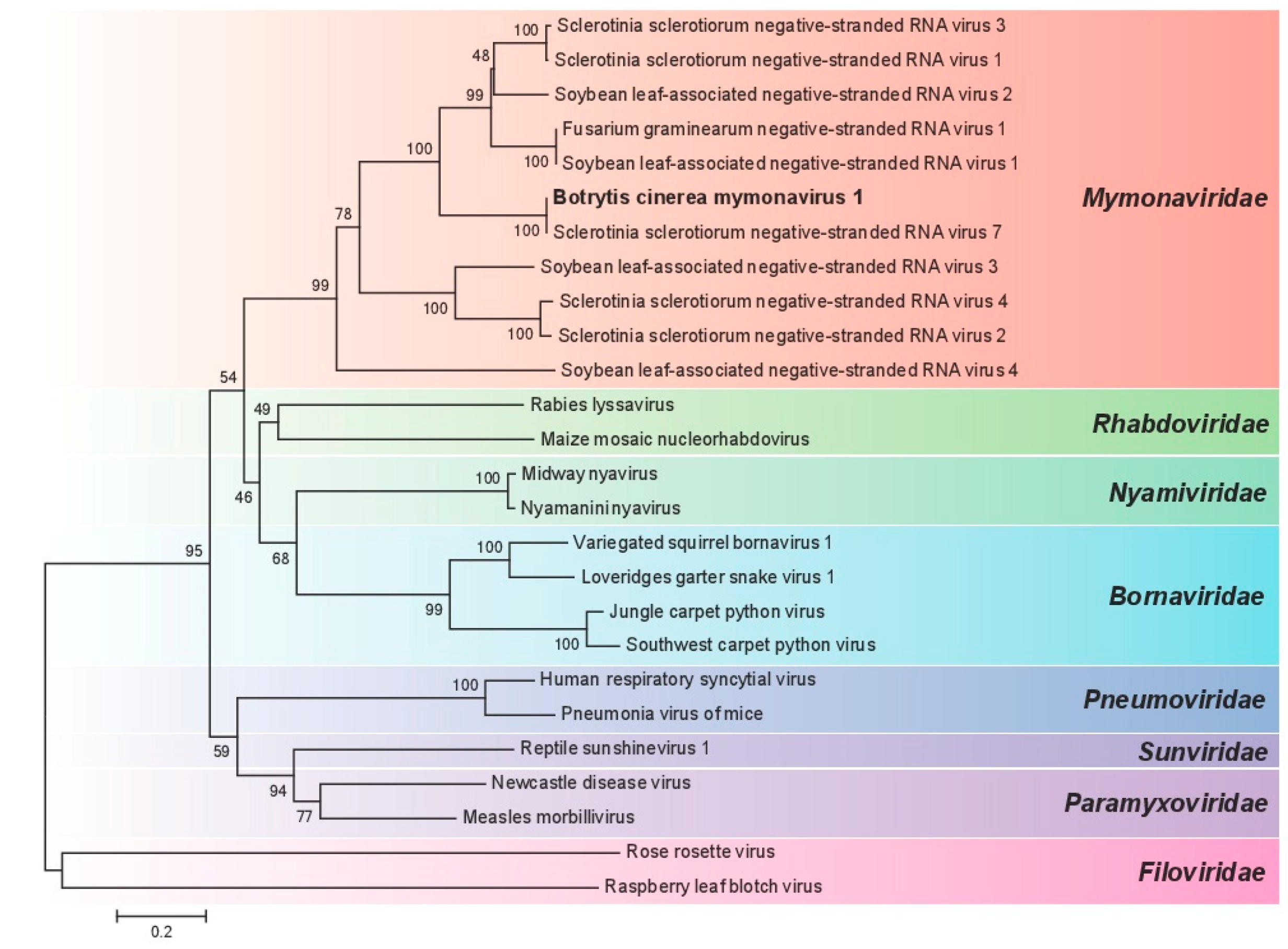
3.3. Phylogenetic Analysis of BcMyV1
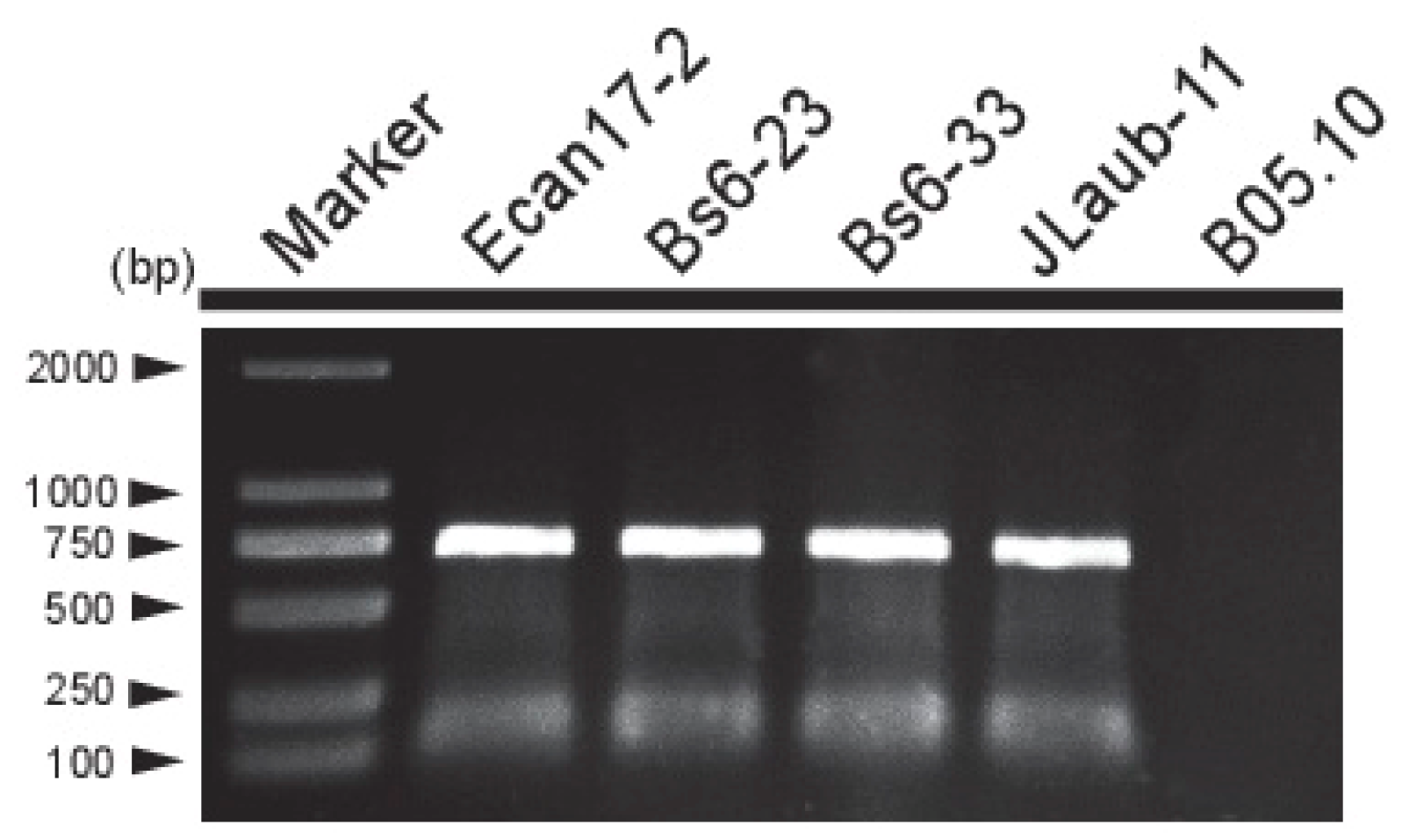
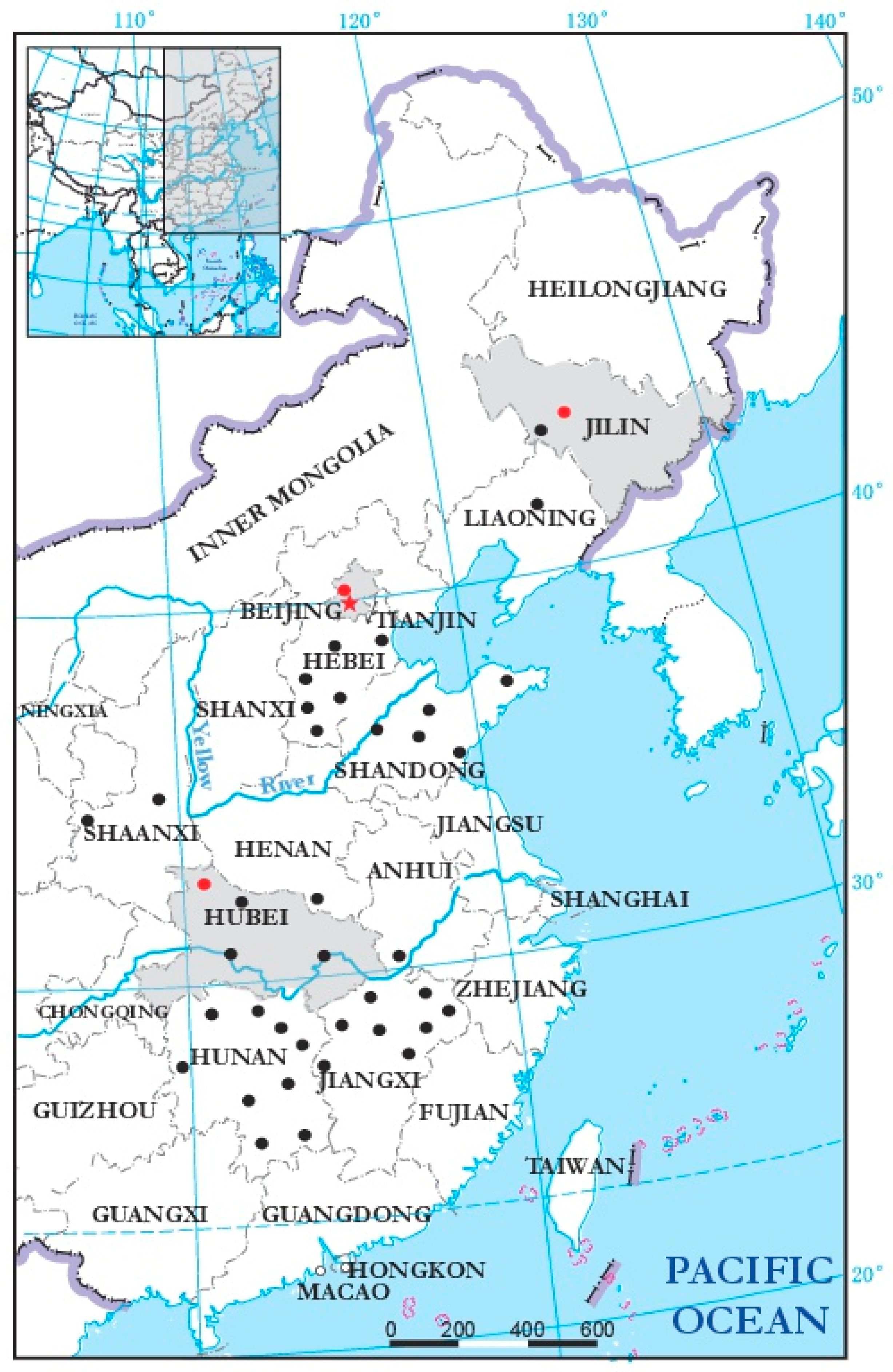
3.4. Incidence and Distribution of BcMyV1
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Elad, Y.; Pertot, I.; Marina, A.; Prado, A.M.; Stewart, A. Plant Hosts of Botrytis spp. In Botrytis—The Fungus, the Pathogen and Its Management in Agricultural Systems; Fillinger, S., Elad, Y., Eds.; Springer: Cham, Switzerland, 2016; pp. 413–486. [Google Scholar]
- Hao, F.M.; Zhou, Z.L.; Wu, M.D.; Li, G.Q. Molecular characterization of a novel endornavirus from the phytopathogenic fungus Botrytis cinerea. Arch. Virol. 2017, 162, 313–316. [Google Scholar] [CrossRef] [PubMed]
- Hao, F.M.; Ding, T.; Wu, M.D.; Zhang, J.; Yang, L.; Chen, W.D.; Li, G.Q. Two novel hypovirulence-associated mycoviruses in the phytopathogenic fungus Botrytis cinerea: Molecular characterization and suppression of infection cushion formation. Viruses 2018, 10, 254. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.D.; Zhang, J.; Yang, L.; Li, G.Q. RNA mycoviruses and their role in Botrytis Biology. In Botrytis—The Fungus, the Pathogen and Its Management in Agricultural Systems; Fillinger, S., Elad, Y., Eds.; Springer: Cham, Switzerland, 2016; pp. 71–90. [Google Scholar]
- Pearson, M.N.; Bailey, A.M. Viruses of botrytis. Arch. Virol. 2013, 86, 249–272. [Google Scholar]
- Yu, L.; Sang, W.; Wu, M.D.; Zhang, J.; Yang, L.; Zhou, Y.J.; Chen, W.D.; Li, G.Q. Novel hypovirulence-associated RNA mycovirus in the plant pathogenic fungus Botrytis cinerea. Appl. Environ. Microbiol. 2015, 81, 2299–2310. [Google Scholar] [CrossRef] [PubMed]
- Donaire, L.; Rozas, J.; María, A. Molecular characterization of Botrytis ourmia-like virus, a mycovirus close to the plant pathogenic genus Ourmiavirus. Virology 2016, 489, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Donaire, L.; Pagan, I.; Ayllon, M.A. Characterization of Botrytis cinerea negative-stranded RNA virus 1, a new mycovirus related to plant viruses, and a reconstruction of host pattern evolution in negative-sense ssRNA viruses. Virology 2016, 499, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Easton, A.J.; Pringle, C.R. Order Mononegavirales. In Virus Taxonomy: Classification and Nomenclature of Viruses, Ninth Report of the International Committee on Taxonomy of Viruses; King, M.Q., Adams, M.J., Carstens, E.B., Lefkowitz, E.J., Eds.; Elsevier, Academic Press: London, UK, 2011; pp. 653–657. [Google Scholar]
- Dietzgen, R.G.; Kondo, H.; Goodin, M.M.; Kurath, G.; Vasilakis, N. The family Rhabdoviridae: mono- and bipartite negative-sense RNA viruses with diverse genome organization and common evolutionary origins. Virus Res. 2017, 227, 158–170. [Google Scholar] [CrossRef] [PubMed]
- Kondo, H.; Chiba, S.; Toyoda, K.; Suzuki, N. Evidence for negative-strand RNA virus infection in fungi. Virology 2013, 435, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Afonso, C.L.; Amarasinghe, G.K.; Bányai, K.; Bao, Y.; Basler, C.F.; Bavari, S.; Bejerman, N.; Blasdell, K.R.; Briand, F.; Briese, T.; et al. Taxonomy of the order Mononegavirales: Update. 2016. Arch. Virol. 2016, 161, 2351–2360. [Google Scholar] [CrossRef] [PubMed]
- Walker, P.J.; Firth, C.; Widen, S.G.; Blasdell, K.R.; Guzman, H.; Wood, T.G.; Paradkar, P.N.; Holmes, E.C.; Tesh, R.B.; Vasilakis, N. Evolution of genome size and complexity in the Rhabdoviridae. PLoS Pathog. 2015, 11, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Xie, J.; Cheng, J.; Fu, Y.; Li, G.; Yi, X.; Jiang, D. Fungal negative-stranded RNA virus that is related to bornaviruses and nyaviruses. Proc. Natl. Acad. Sci. USA 2014, 111, 12205–12210. [Google Scholar] [CrossRef] [PubMed]
- Marzano, S.Y.; Domier, L.L. Novel mycoviruses discovered from metatranscriptomics survey of soybean phyllosphere phytobiomes. Virus Res. 2016, 213, 332–342. [Google Scholar] [CrossRef] [PubMed]
- Marzano, L.S.Y.; Nelson, B.D.; Ajayi-Oyetunde, O.; Bradley, C.A.; Hughes, T.J.; Hartman, G.L.; Eastburn, D.M.; Domiera, L.L. Identification of diverse mycoviruses through metatranscriptomics characterization of the viromes of five major fungal plant pathogens. J. Virol. 2016, 90, 6846–6863. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; He, H.; Wang, S.; Chen, X.; Qiu, D.; Kondob, H.; Guo, L. Evidence for a novel negative-stranded RNA mycovirus isolated from the plant pathogenic fungus Fusarium graminearum. Virology 2018, 518, 232–240. [Google Scholar] [CrossRef] [PubMed]
- Mu, F.; Xie, J.T.; Cheng, S.F.; You, M.P.; Barbetti, M.J.; Jia, J.C.; Wang, Q.Q.; Cheng, J.S.; Fu, Y.P.; Chen, T.; et al. Virome characterization of a collection of Sclerotinia sclerotiorum from Australia. Front. Microbial. 2018, 8, 2540. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.D.; Zhang, L.; Li, G.Q. Genome characterization of a debilitation-associated mitovirus infecting the phytopathogenic fungus Botrytis cinerea. Virology 2010, 406, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.D.; Zhang, L.; Li, G.Q.; Jiang, D.H.; Hou, M.S.; Huang, H.C. Hypovirulence and double-stranded RNA in Botrytis cinerea. Phytopathology 2007, 97, 1590–1599. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.D.; Jin, F.Y.; Zhang, J.; Yang, L.; Jiang, D.H.; Li, G.Q. Characterization of a novel bipartite double-stranded RNA mycovirus conferring hypovirulence in the phytopathogenic fungus Botrytis porri. J. Virol. 2012, 86, 6605–6619. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Blol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef] [PubMed]
- Choi, G.H.; Nuss, D.L. Hypovirulence of chestnut blight fungus conferred by an infectious viral cDNA. Science 1992, 257, 800–803. [Google Scholar] [CrossRef] [PubMed]
- Van Diepeningen, A.D.; Debets, A.J.M.; Hoekstra, R.F. Dynamics of dsRNA mycoviruses in black Aspergillus populations. Fungal. Genet. Biol. 2006, 43, 446–452. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.M.; Jung, J.E.; Park, J.A.; Park, S.M.; Cha, B.J.; Kim, D.H. Biological function of a novel chrysovirus, CnV1-BS122, in the Korean Cryphonectria nitschkei BS122 strain. J. Biosci. Bioeng. 2013, 115, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Fulbright, D.W. Effect of eliminating dsRNA in hypovirulent Endothia parasitica. Phytopathology 1984, 74, 722–724. [Google Scholar] [CrossRef]
- Elias, K.S.; Cotty, P.J. Incidence and stability of infection by double-stranded RNA genetic elements in Aspergillus section Flavi and effects on aflatoxigenicity. Can. J. Bot. 1996, 74, 716–725. [Google Scholar] [CrossRef]
- Kwon, Y.C.; Jeong, D.W.; Gim, S.I.; Ro, H.S.; Lee, H.S. Curing viruses in Pleurotus ostreatus by growth on a limited nutrient medium containing cAMP and rifamycin. J. Virol. Methods 2012, 185, 156–159. [Google Scholar] [CrossRef] [PubMed]
- Amselem, J.; Cuomo, C.A.; van Kan, J.A.L.; Viaud, M.; Benito, E.P.; Couloux, A.; Coutinho, P.M.; de Vries, R.P.; Dyer, P.S.; Fillinger, S.; et al. Genomic analysis of the necrotrophic fungal pathogens Sclerotinia sclerotiorum and Botrytis cinerea. PLoS Genet. 2011, 7, e1002230. [Google Scholar] [CrossRef] [PubMed]
- Bolton, M.D.; Thomma, B.P.; Nelson, B.D. Sclerotinia sclerotiorum (Lib.) de Bary: Biology and molecular traits of a cosmopolitan pathogen. Mol. Plant. Pathol. 2006, 7, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Coenen, A.; Kevei, F.; Hoekstra, R.F. Factors affecting the spread of double-stranded RNA viruses in Aspergillus nidulans. Genet. Res. 1997, 69, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Melzer, M.S.; Ikeda, S.S.; Boland, G.J. Interspecific transmission of double-stranded RNA and hypovirulence from Sclerotinia sclerotiorum to S. minor. Phytopathology 2002, 92, 780–784. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.C.; Hillman, B.I.; Linder-Basso, D.; Kaneko, S.; Milgroom, M.G. Evidence for interspecies transmission of viruses in natural populations of filamentous fungi in the genus Cryphonectria. Mol. Ecol. 2003, 12, 1619–1628. [Google Scholar] [CrossRef] [PubMed]
- Bouneb, M.; Turchetti, T.; Nannelli, R.; Roversi, P.F.; Paoli, F.; Danti, R.; Simoni, S. Occurrence and transmission of mycovirus Cryphonectria hypovirus 1 from dejecta of Thyreophagus corticalis (Acari, Acaridae). Fungal Biol. 2016, 120, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Xie, J.; Cheng, J.; Li, B.; Chen, T.; Fu, Y. Fungal DNA virus infects a mycophagous insect and utilizes it as a transmission vector. Proc. Natl. Acad. Sci. USA 2016, 113, 12803–12808. [Google Scholar] [CrossRef] [PubMed]
- Korinsak, S.; Tangphatsornruang, S.; Pootakham, W.; Wanchana, S.; Plabpla, A.; Jantasuriyarat, C.; Patarapuwadol, S.; Vanavichit, A.; Toojinda, T. Genome-wide association mapping of virulence gene in rice blast fungus Magnaporthe oryzae using a genotyping by sequencing approach. Genomics 2018. [Google Scholar] [CrossRef] [PubMed]
- Castiblanco, V.; Marulanda, J.J.; Würschum, T.; Miedaner, T. Candidate gene based association mapping in Fusarium culmorum for field quantitative pathogenicity and mycotoxin production in wheat. BMC Genet. 2017, 18, 49. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-García, C.; Medina, V.; Alonso, A.; Ayllón, M.A. Mycoviruses of Botrytis cinerea isolates from different hosts. Ann. Appl. Biol. 2013, 164, 46–61. [Google Scholar] [CrossRef]
| Family | Virus | Acronym | aa Identity (%) | Accession no. | |
|---|---|---|---|---|---|
| Full Sequence | RdRp | ||||
| Mymonaviridae | Sclerotinia sclerotiorum negative-stranded RNA virus 7 | SsNSRV7 | 99.85 | 100 | MF444285 |
| Sclerotinia sclerotiorum negative-stranded RNA virus 1 | SsNSRV1 | 33.12 | 56.68 | NC_025383.1 | |
| Sclerotinia sclerotiorum negative-stranded RNA virus 2 | SsNSRV2 | 22.28 | 41.45 | KP900931.1 | |
| Sclerotinia sclerotiorum negative-stranded RNA virus 3 | SsNSRV3 | 33.57 | 56.15 | NC_026732.1 | |
| Sclerotinia sclerotiorum negative-stranded RNA virus 4 | SsNSRV4 | 21.67 | 39.38 | KP900930.1 | |
| Soybean leaf-associated negative-stranded RNA virus 1 | SlaNSRV1 | 32.75 | 59.36 | KT598225.1 | |
| Soybean leaf-associated negative-stranded RNA virus 2 | SlaNSRV2 | 33.12 | 61.5 | KT598227.1 | |
| Soybean leaf-associated negative-stranded RNA virus 3 | SlaNSRV3 | 21.43 | 37.31 | KT598228.1 | |
| Soybean leaf-associated negative-stranded RNA virus 4 | SlaNSRV4 | 17.82 | 34.57 | KT598229.1 | |
| Fusarium graminearum negative-stranded RNA virus 1 | FgNSRV1 | 32.75 | 59.36 | MF276904.1 | |
| Bornaviridae | Jungle carpet python virus | JCPV | 14.84 | 24.6 | MF135780 |
| Southwest carpet python virus | SWCPV | 14.33 | 21.93 | MF135781 | |
| Loveridge’s garter snake virus 1 | LGSV1 | 14.28 | 23.53 | KM114265 | |
| Variegated squirrel bornavirus 1 | VSBV1 | 14.16 | 25.13 | LN713681 | |
| Rhabdoviridae | Rabies virus | RabV | 14.97 | 23.62 | AB517659 |
| Maize mosaic virus | MMV | 15.19 | 23.12 | NC_005975.1 | |
| Paramyxoviridae | Newcastle disease virus | NDV | 14.94 | 27.69 | JF827026.1 |
| Measles virus | MV | 15.44 | 30.77 | NC_001498.1 | |
| Nyamiviridae | Midway nyavirus | MIDMV | 15.81 | 26.42 | NC_012702.1 |
| Nyamanini nyavirus | NYMV | 16.01 | 26.42 | NC_012703.1 | |
| Filoviridae | Rose rosette virus | RRV | 10.57 | 11.4 | HQ871942 |
| Raspberry leaf blotch virus | RLBV | 10.71 | 10.27 | FR823299 | |
| Sunviridae | Reptile sunshinevirus 1 | RSV-1 | 14.26 | 24.26 | NC_025345 |
| Pneumoviridae | Human respiratory syncytial virus | HRSV | 13.43 | 22.68 | NC_001781 |
| Pneumonia virus of mice | PVM | 14.22 | 22.8 | NC_006579 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hao, F.; Wu, M.; Li, G. Molecular Characterization and Geographic Distribution of a Mymonavirus in the Population of Botrytis cinerea. Viruses 2018, 10, 432. https://doi.org/10.3390/v10080432
Hao F, Wu M, Li G. Molecular Characterization and Geographic Distribution of a Mymonavirus in the Population of Botrytis cinerea. Viruses. 2018; 10(8):432. https://doi.org/10.3390/v10080432
Chicago/Turabian StyleHao, Fangmin, Mingde Wu, and Guoqing Li. 2018. "Molecular Characterization and Geographic Distribution of a Mymonavirus in the Population of Botrytis cinerea" Viruses 10, no. 8: 432. https://doi.org/10.3390/v10080432
APA StyleHao, F., Wu, M., & Li, G. (2018). Molecular Characterization and Geographic Distribution of a Mymonavirus in the Population of Botrytis cinerea. Viruses, 10(8), 432. https://doi.org/10.3390/v10080432





