Novel T7 Phage Display Library Detects Classifiers for Active Mycobacterium Tuberculosis Infection
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Patient Selection
2.3. Serum Collection
2.4. Construction and Biopanning of T7 Phage Display cDNA Libraries
2.5. Microarray Construction and Immunoscreening
2.6. Sequencing of Phage cDNA Clones
2.7. Data Acquisition and Pre-Processing
2.8. Statistical Analyses
3. Results
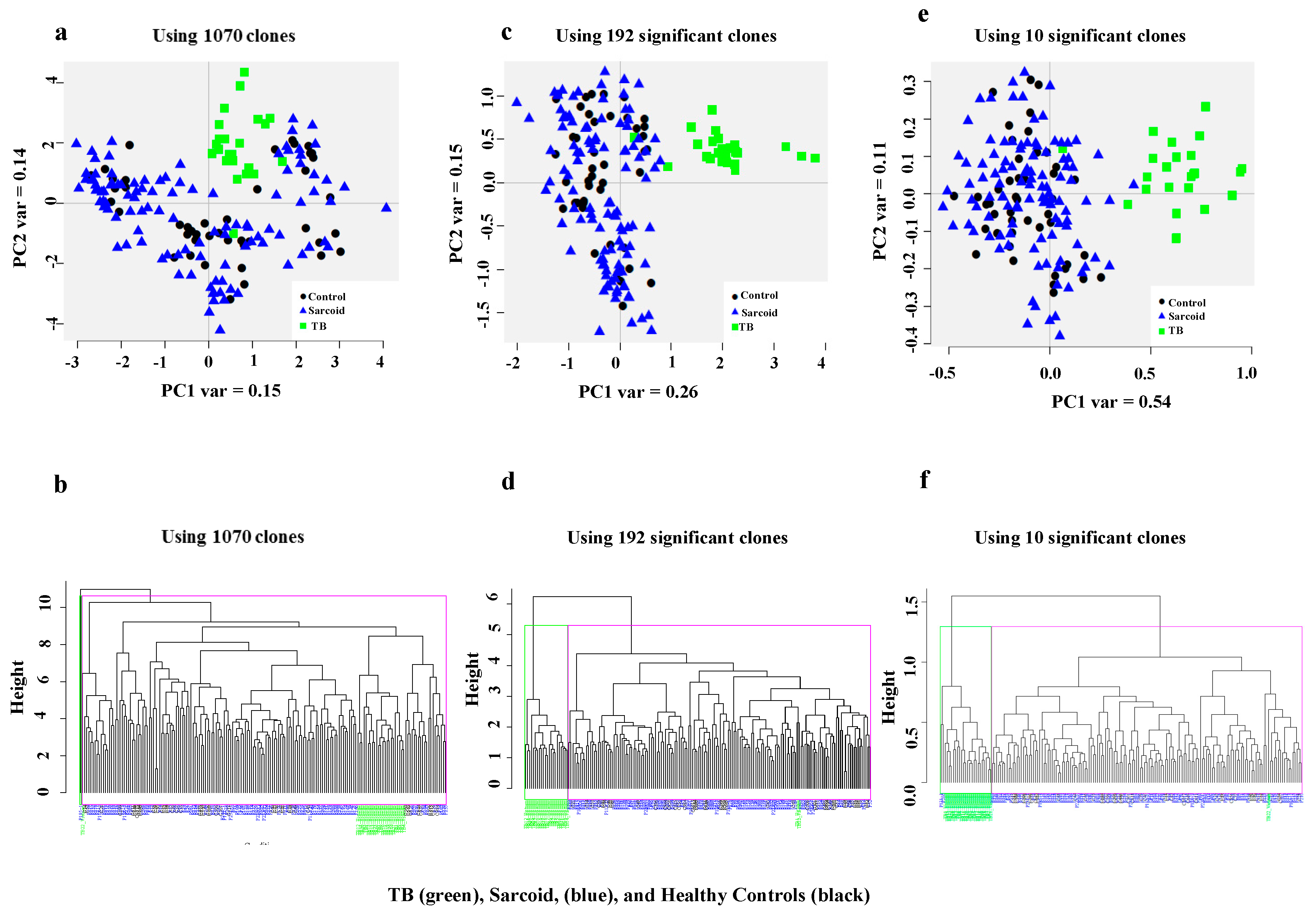
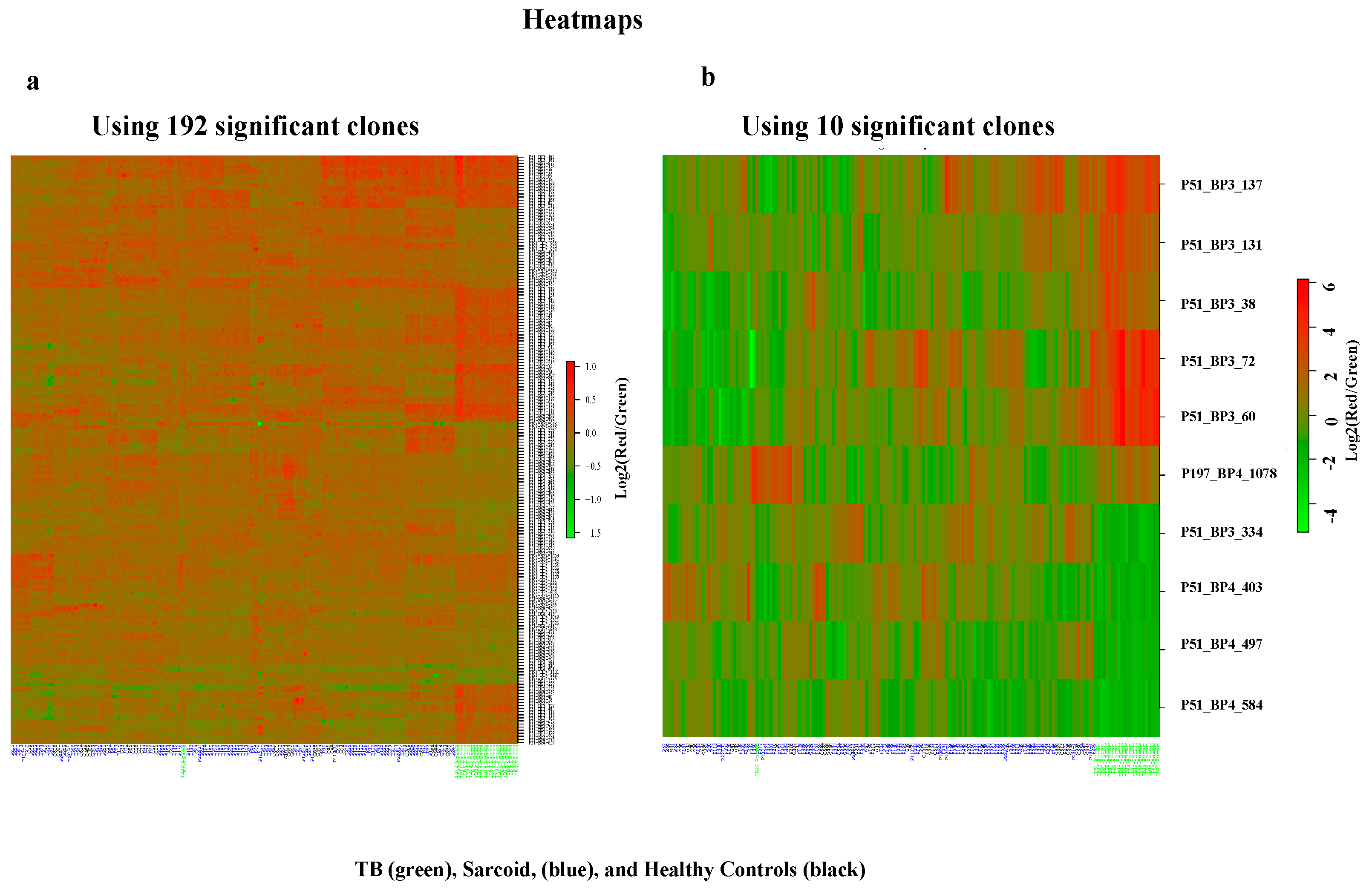
3.1. Complex Sarcoidosis (CSL) Library Detects Unique Antigens in the Sera of Active Tuberculosis Patients
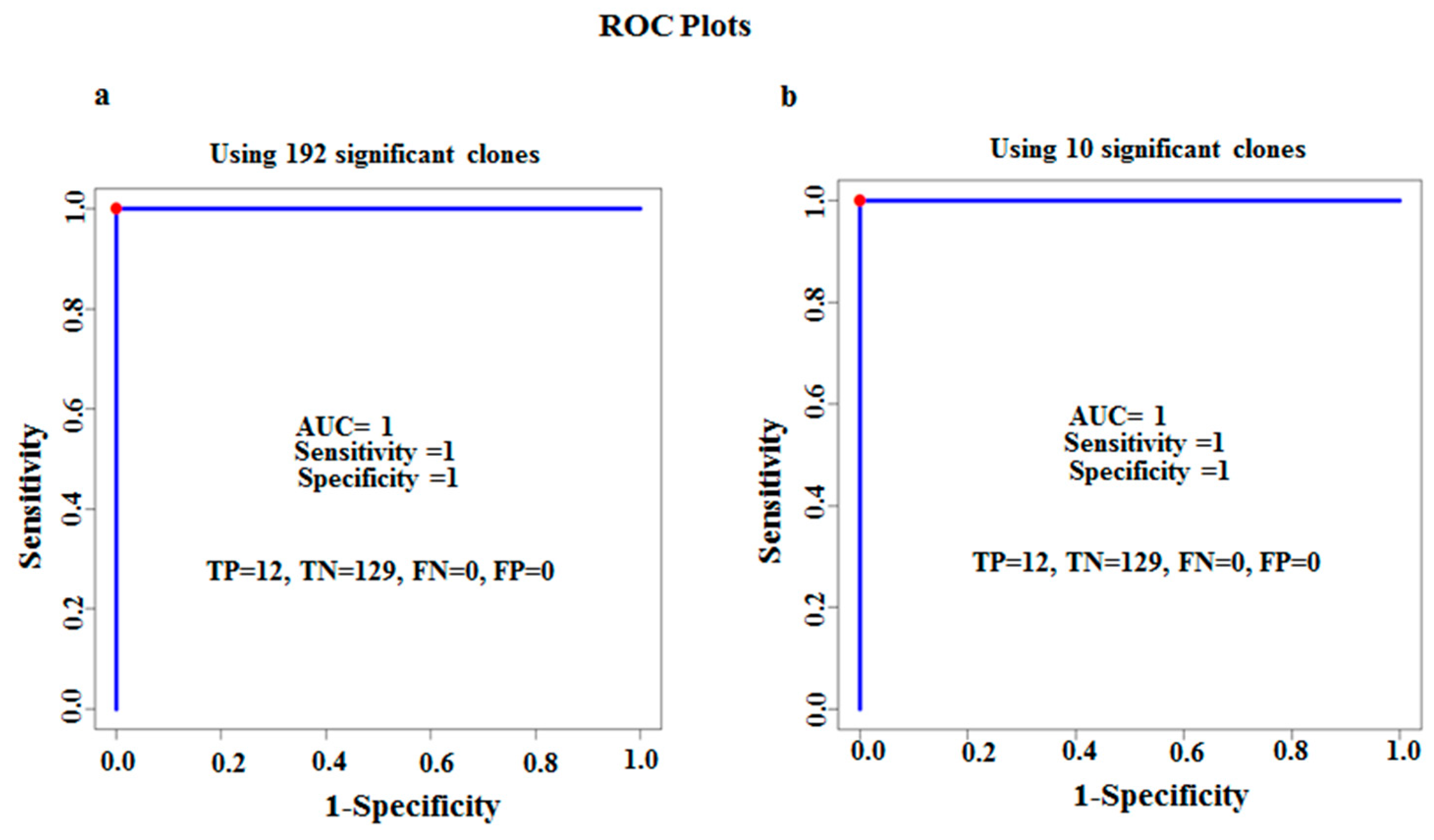
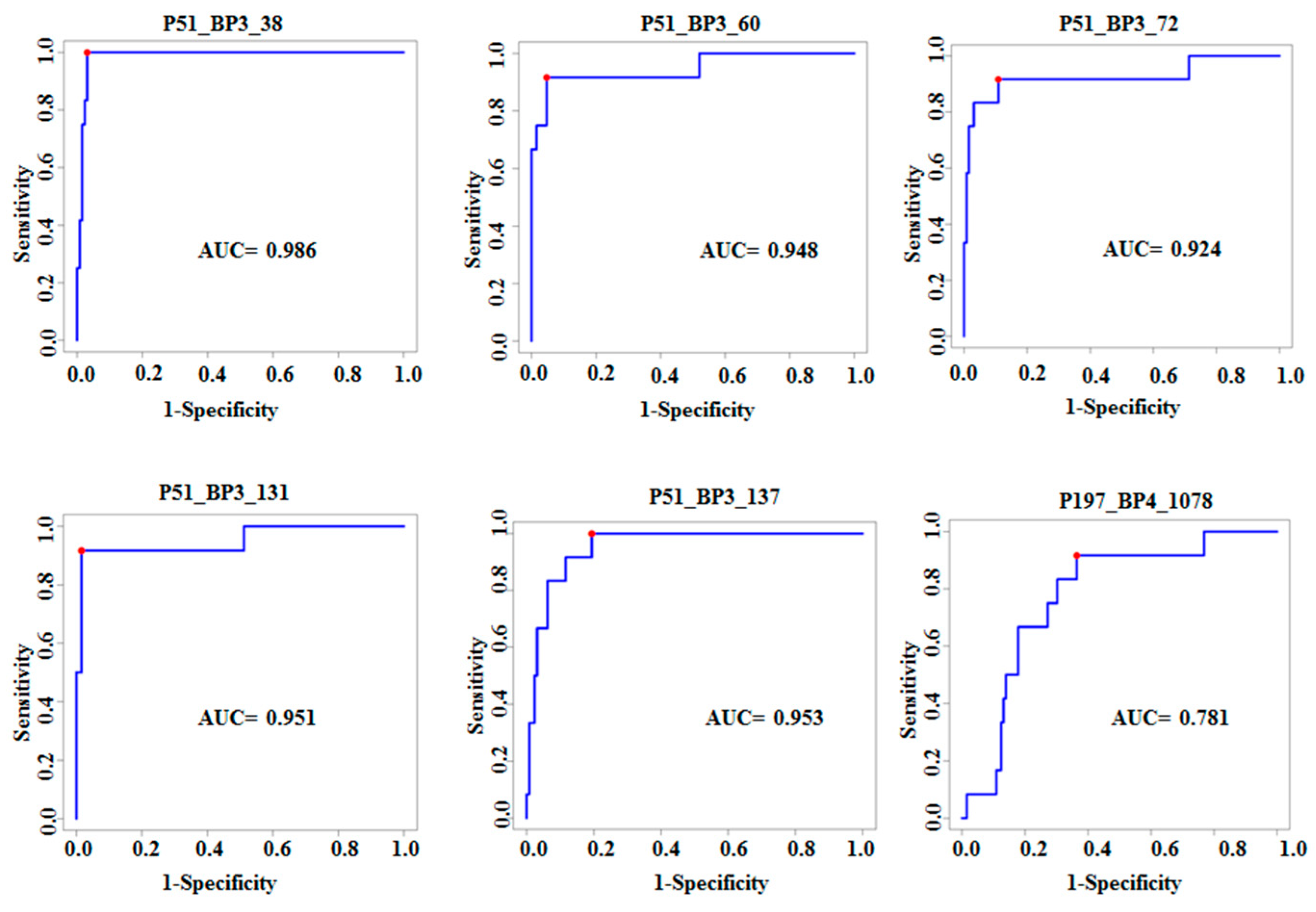
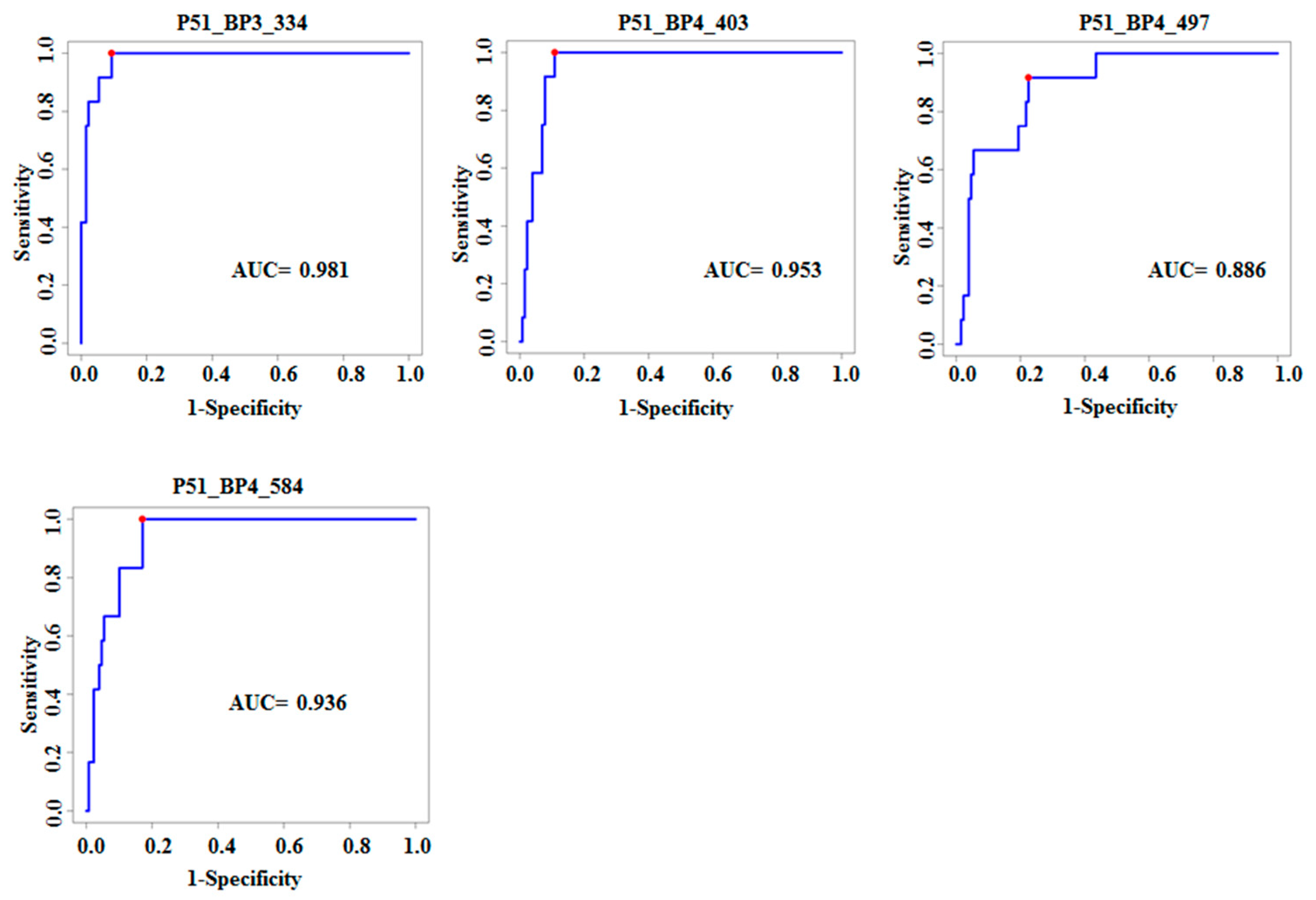
3.2. Characterization of Ten TB Classifiers
4. Discussion
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Lawn, S.D.; Zumla, A.I. Tuberculosis. Lancet 2011, 378, 57–72. [Google Scholar] [CrossRef]
- Nahid, P.; Saukkonen, J.; Mac Kenzie, W.R.; Johnson, J.L.; Phillips, P.P.; Andersen, J.; Bliven-Sizemore, E.; Belisle, J.T.; Boom, W.H.; Luetkemeyer, A.; et al. CDC/NIH Workshop. Tuberculosis biomarker and surrogate endpoint research roadmap. Am. J. Respir. Crit. Care Med. 2011, 184, 972–979. [Google Scholar] [CrossRef] [PubMed]
- Wallis, R.S.; Pai, M.; Menzies, D.; Doherty, T.M.; Walzl, G.; Perkins, M.D.; Zumla, A. Biomarkers and diagnostics for tuberculosis: Progress, needs, and translation into practice. Lancet 2010, 375, 1920–1937. [Google Scholar] [CrossRef]
- Dubaniewicz, A.; Dubaniewicz-Wybieralska, M.; Sternau, A.; Zwolska, Z.; Izycka-Swieszewska, E.; Augustynowicz-Kopec, E.; Skokowski, J.; Singh, M.; Zimnoch, L. Mycobacterium tuberculosis complex and mycobacterial heat shock proteins in lymph node tissue from patients with pulmonary sarcoidosis. J. Clin. Microbiol. 2006, 44, 3448–3451. [Google Scholar] [CrossRef] [PubMed]
- Moller, D.R. Potential etiologic agents in sarcoidosis. Proc. Am. Thorac. Soc. 2007, 4, 465–468. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Marzilli, L.; Greenlee, B.M.; Chen, E.S.; Silver, R.F.; Askin, F.B.; Teirstein, A.S.; Zhang, Y.; Cotter, R.J.; Moller, D.R. Mycobacterial catalase-peroxidase is a tissue antigen and target of the adaptive immune response in systemic sarcoidosis. J. Exp. Med. 2005, 201, 755–767. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, R.; Du, W.; Ju, D.; Pirockinaite, G.; Liu, Y.; Nunez, G.; Samavati, L. Dysregulation of p38 and MKP-1 in response to NOD1/TLR4 stimulation in sarcoid bronchoalveolar cells. Am. J. Respir. Crit. Care Med. 2011, 183, 500–510. [Google Scholar] [CrossRef] [PubMed]
- Carlisle, J.; Evans, W.; Hajizadeh, R.; Nadaf, M.; Shepherd, B.; Ott, R.D.; Richter, K.; Drake, W. Multiple Mycobacterium antigens induce interferon-gamma production from sarcoidosis peripheral blood mononuclear cells. Clin. Exp. Immunol. 2007, 150, 460–468. [Google Scholar] [CrossRef] [PubMed]
- Drake, W.P.; Dhason, M.S.; Nadaf, M.; Shepherd, B.E.; Vadivelu, S.; Hajizadeh, R.; Newman, L.S.; Kalams, S.A. Cellular recognition of Mycobacterium tuberculosis ESAT-6 and KatG peptides in systemic sarcoidosis. Infect. Immun. 2007, 75, 527–530. [Google Scholar] [CrossRef] [PubMed]
- Hajizadeh, R.; Sato, H.; Carlisle, J.; Nadaf, M.T.; Evans, W.; Shepherd, B.E.; Miller, R.F.; Kalams, S.A.; Drake, W.P. Mycobacterium tuberculosis Antigen 85A induces Th-1 immune responses in systemic sarcoidosis. J. Clin. Immunol. 2007, 27, 445–454. [Google Scholar] [CrossRef] [PubMed]
- Oswald-Richter, K.; Sato, H.; Hajizadeh, R.; Shepherd, B.E.; Sidney, J.; Sette, A.; Newman, L.S.; Drake, W.P. Mycobacterial ESAT-6 and katG are recognized by sarcoidosis CD4+ T cells when presented by the American sarcoidosis susceptibility allele, DRB1*. J. Clin. Immunol. 2010, 30, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Richmond, B.W.; Ploetze, K.; Isom, J.; Chambers-Harris, I.; Braun, N.A.; Taylor, T.; Abraham, S.; Mageto, Y.; Culver, D.A.; Oswald-Richter, K.A.; et al. Sarcoidosis Th17 cells are ESAT-6 antigen specific but demonstrate reduced IFN-gamma expression. J. Clin. Immunol. 2013, 33, 446–455. [Google Scholar] [CrossRef] [PubMed]
- Smith-Rohrberg, D.; Sharma, S. Tuberculin skin test among pulmonary sarcoidosis patients with and without tuberculosis: Its utility for the screening of the two conditions in tuberculosis-endemic regions. Sarcoidosis Vasc. Diffuse Lung Dis. 2006, 23, 130–134. [Google Scholar] [PubMed]
- Murray, P.J.; Wynn, T.A. Protective and pathogenic functions of macrophage subsets. Nat. Rev. Immunol. 2011, 11, 723–737. [Google Scholar] [CrossRef] [PubMed]
- Peyron, P.; Vaubourgeix, J.; Poquet, Y.; Levillain, F.; Botanch, C.; Bardou, F.; Daffé, M.; Emile, J.-F.; Marchou, B.; Cardona, P.-J.; et al. Foamy macrophages from tuberculous patients’ granulomas constitute a nutrient-rich reservoir for M. tuberculosis persistence. PLoS Pathog. 2008, 4, e1000204. [Google Scholar] [CrossRef] [PubMed]
- Talreja, J.; Talwar, H.; Ahmad, N.; Rastogi, R.; Samavati, L. Dual Inhibition of Rip2 and IRAK1/4 Regulates IL-1beta and IL-6 in Sarcoidosis Alveolar Macrophages and Peripheral Blood Mononuclear Cells. J. Immunol. 2016, 197, 1368–1378. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.; Summer, R.; Wilson, A.A.; Kotton, D.N.; Fine, A. The prolonged life-span of alveolar macrophages. Am. J. Respir. Cell Mol. Biol. 2008, 38, 380–385. [Google Scholar] [CrossRef] [PubMed]
- Talwar, H.; Rosati, R.; Li, J.; Kissner, D.; Ghosh, S.; Madrid, F.F.; Samavati, L. Development of a T7 Phage Display Library to Detect Sarcoidosis and Tuberculosis by a Panel of Novel Antigens. EBioMedicine 2015, 2, 341–350. [Google Scholar] [CrossRef] [PubMed]
- Talwar, H.; Hanoudi, S.N.; Geamanu, A.; Kissner, D.; Draghici, S.; Samavati, L. Detection of Cystic Fibrosis Serological Biomarkers Using a T7 Phage Display Library. Sci. Rep. 2017, 7, 17745. [Google Scholar] [CrossRef] [PubMed]
- Talwar, H.; Talreja, J.; Samavati, L. T7 Phage Display Library a Promising Strategy to Detect Tuberculosis Specific Biomarkers. Mycobact. Dis. 2016, 6, 214. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2015.
- Ritchie, M.E.; Silver, J.; Oshlack, A.; Holmes, M.; Diyagama, D.; Holloway, A.; Smyth, G.K. A comparison of background correction methods for two-colour microarrays. Bioinformatics 2007, 23, 2700–2707. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.H.; Dudoit, S.; Luu, P.; Lin, D.M.; Peng, V.; Ngai, J.; Speed, T.P. Normalization for cDNA microarray data: A robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 2002, 30, e15. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 289–300. [Google Scholar]
- Wang, X.; Yu, J.; Sreekumar, A.; Varambally, S.; Shen, R.; Giacherio, D.; Mehra, R.; Montie, J.E.; Pienta, K.J.; Sanda, M.G.; et al. Autoantibody signatures in prostate cancer. N. Engl. J. Med. 2005, 353, 1224–1235. [Google Scholar] [CrossRef] [PubMed]
- Kunnath-Velayudhan, S.; Salamon, H.; Wang, H.Y.; Davidow, A.L.; Molina, D.M.; Huynh, V.T.; Cirillo, D.M.; Michel, G.; Talbot, E.A.; Perkins, M.D.; et al. Dynamic antibody responses to the Mycobacterium tuberculosis proteome. Proc. Natl. Acad. Sci. USA 2010, 107, 14703–14708. [Google Scholar] [CrossRef] [PubMed]
- De Groote, M.A.; Nahid, P.; Jarlsberg, L.; Johnson, J.L.; Weiner, M.; Muzanyi, G.; Janjic, N.; Sterling, D.G.; Ochsner, U.A. Elucidating novel serum biomarkers associated with pulmonary tuberculosis treatment. PLoS ONE 2013, 8, e61002. [Google Scholar] [CrossRef] [PubMed]
- Agranoff, D.; Fernandez-Reyes, D.; Papadopoulos, M.C.; Rojas, S.A.; Herbster, M.; Loosemore, A.; Tarelli, E.; Sheldon, J.; Schwenk, A.; Pollok, R. Identification of diagnostic markers for tuberculosis by proteomic fingerprinting of serum. Lancet 2006, 368, 1012–1021. [Google Scholar] [CrossRef]
- Gavalda, S.; Bardou, F.; Laval, F.; Bon, C.; Malaga, W.; Chalut, C.; Guilhot, C.; Mourey, L.; Daffé, M.; Quémard, A. The polyketide synthase Pks13 catalyzes a novel mechanism of lipid transfer in mycobacteria. Chem. Biol. 2014, 21, 1660–1669. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, A.; Parai, M.K.; Shetty, N.; Wallis, D.; Woolhiser, L.; Hastings, C.; Dutta, N.K.; Galaviz, S.; Dhakal, R.C.; Shrestha, R.; et al. Development of a novel lead that targets M. tuberculosis polyketide synthase. Cell 2017, 170, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Ouellet, H.; Johnston, J.B.; de Montellano, P.R.O. The Mycobacterium tuberculosis cytochrome P450 system. Arch. Biochem. Biophys. 2010, 493, 82–95. [Google Scholar] [CrossRef] [PubMed]
- Ollinger, J.; O’Malley, T.; Ahn, J.; Odingo, J.; Parish, T. Inhibition of the sole type I signal peptidase of Mycobacterium tuberculosis is bactericidal under replicating and nonreplicating conditions. J. Bacteriol. 2012, 194, 2614–2619. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.; Chandra, D.; Srivastava, B.S.; Srivastava, R. Downregulation of Rv0189c, encoding a dihydroxyacid dehydratase, affects growth of Mycobacterium tuberculosis in vitro and in mice. Microbiology 2011, 157, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Kolly, G.S.; Sala, C.; Vocat, A.; Cole, S.T. Assessing essentiality of transketolase in Mycobacterium tuberculosis using an inducible protein degradation system. FEMS Microbiol. Lett. 2014, 358, 30–35. [Google Scholar] [CrossRef] [PubMed]
- Balhana, R.J.; Singla, A.; Sikder, M.H.; Withers, M.; Kendall, S.L. Global analyses of TetR family transcriptional regulators in mycobacteria indicates conservation across species and diversity in regulated functions. BMC Genom. 2015, 16, 479. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Yang, M.; He, Z.-G. Novel TetR family transcriptional factor regulates expression of multiple transport-related genes and affects rifampicin resistance in Mycobacterium smegmatis. Sci. Rep. 2016, 6, 27489. [Google Scholar] [CrossRef] [PubMed]
- Cuthbertson, L.; Nodwell, J.R. The TetR family of regulators. Microbiol. Mol. Biol. Rev. 2013, 77, 440–475. [Google Scholar] [CrossRef] [PubMed]
- Van Pinxteren, L.A.; Ravn, P.; Agger, E.M.; Pollock, J.; Andersen, P. Diagnosis of tuberculosis based on the two specific antigens ESAT-6 and CFP. Clin. Diagn. Lab. Immunol. 2000, 7, 155–160. [Google Scholar] [PubMed]
- Xu, J.-N.; Chen, J.-P.; Chen, D.-L. Serodiagnosis efficacy and immunogenicity of the fusion protein of Mycobacterium tuberculosis composed of the 10-kilodalton culture filtrate protein, ESAT-6, and the extracellular domain fragment of PPE. Clin. Vaccine Immunol. 2012, 19, 536–544. [Google Scholar] [CrossRef] [PubMed]
- Kumar, G.; Dagur, P.K.; Singh, P.K.; Shankar, H.; Yadav, V.S.; Katoch, V.M.; Bajaj, B.; Gupta, R.; Sengupta, U.; Joshi, B. Serodiagnostic efficacy of Mycobacterium tuberculosis 30/32-kDa mycolyl transferase complex, ESAT-6, and CFP-10 in patients with active tuberculosis. Arch. Immunol. Ther. Exp. 2010, 58, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.-H.; Liu, I.-J.; Lu, R.-M.; Wu, H.-C. Advancement and applications of peptide phage display technology in biomedical science. J. Biomed. Sci. 2016, 23, 8. [Google Scholar] [CrossRef] [PubMed]
- Rana, S.; Bajaj, A.; Mout, R.; Rotello, V.M. Monolayer coated gold nanoparticles for delivery applications. Adv. Drug Deliv. Rev. 2012, 64, 200–216. [Google Scholar] [CrossRef] [PubMed]
- Koth, L.L.; Solberg, O.D.; Peng, J.C.; Bhakta, N.R.; Nguyen, C.P.; Woodruff, P.G. Sarcoidosis blood transcriptome reflects lung inflammation and overlaps with tuberculosis. Am. J. Respir. Crit. Care Med. 2011, 184, 1153–1163. [Google Scholar] [CrossRef] [PubMed]
- Maertzdorf, J.; Weiner, J.; Mollenkopf, H.J.; Network, T.B.; Bauer, T.; Prasse, A.; Muller-Quernheim, J.; Kaufmann, S.H. Common patterns and disease-related signatures in tuberculosis and sarcoidosis. Proc. Natl. Acad. Sci. USA 2012, 109, 7853–7858. [Google Scholar] [CrossRef] [PubMed]
| Characteristic | Control Subjects | Sarcoidosis Subjects | TB Subjects |
|---|---|---|---|
| Age (Mean ± SEM) | 40.3 ± 7.5 | 30.6 ± 11.8 | 40.5 ± 8.5 |
| Gender, N (%) | |||
| Male | 11 (25) | 27 (25) | 18 (64) |
| Female | 34 (75) | 80 (75) | 10 (36) |
| Race, N (%) | |||
| African American | 31 (69) | 95 (89) | |
| African | - | 4 (25) | |
| Caucasian | - | 12 (11) | |
| Asians | 14 (31) | 20 (75) | |
| BMI (Mean ± SEM) | 27 ± 3.8 | 28 ± 10.5 | 28 ± 6.9 |
| Organ involvement | |||
| Neuro-ophthalmologic | NA | 31 (29) | - |
| Lung | NA | 101 (94) | 24 (100) |
| Skin | NA | 46 (43) | - |
| Multiorgan | NA | 65 (61) | - |
| PPD a | NA | Negative | - |
| TB smear b | NA | Negative | Positive |
| Clone | Increased in Tuberculosis (TB) | Gene Name | p Value | FDR Corrected p Value | AUC 95% CI | Sensitivity, 95% CI | Specificity, 95% CI |
|---|---|---|---|---|---|---|---|
| P51_BP3_38 | Polyketide synthase | Pks13 Rv3800c | 4.7 × 10−7 | 2.79 × 10−5 | 0.98 | 1 | 0.97 |
| P51_BP3_60 | Hydrolase | Rv1723 | 1.62 × 10−8 | 3.46 × 10−6 | 0.95 | 0.92 | 0.95 |
| P51_BP3_72 | Ferredoxin | fdxA Rv2007c | 1.36 × 10−9 | 7.27 × 10−7 | 0.92 | 0.91 | 0.89 |
| P51_BP3_131 | Dihydroxy acid dehydratase | ilvD Rv0189c | 2.15 × 10−8 | 3.84 × 10−6 | 0.95 | 0.92 | 0.98 |
| P51_BP3_137 | Transketolase | TKT Rv1449c | 7.14 × 10−8 | 9.72 × 10−6 | 0.95 | 1 | 0.81 |
| P197_BP4_1078 | Signal peptidase | lepB Rv2903 | 1.44 × 10−7 | 1.36 × 10−5 | 0.78 | 0.92 | 0.64 |
| Decreased in Tuberculosis (TB) | |||||||
| P51_BP3_334 | TetR family transcriptional regulator | MRA_2532 | 4.02 × 10−10 | 4.3 × 10−7 | 0.98 | 1 | 0.91 |
| P51_BP4_403 | Menaquinone biosynthesis protein | menD Rv0555 | 7.27 × 10−8 | 9.71 × 10−6 | 0.95 | 1 | 0.87 |
| P51_BP4_497 | Cobalamin biosynthesis protein | CobN Rv2062c | 1.11 × 10−8 | 2.96 × 10−6 | 0.88 | 0.92 | 0.78 |
| P51_BP4_584 | 5-oxoprolinase | OplA Rv0266c | 5.82 × 10−9 | 2.10 × 10−6 | 0.94 | 1 | 0.83 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Talwar, H.; Hanoudi, S.N.; Draghici, S.; Samavati, L. Novel T7 Phage Display Library Detects Classifiers for Active Mycobacterium Tuberculosis Infection. Viruses 2018, 10, 375. https://doi.org/10.3390/v10070375
Talwar H, Hanoudi SN, Draghici S, Samavati L. Novel T7 Phage Display Library Detects Classifiers for Active Mycobacterium Tuberculosis Infection. Viruses. 2018; 10(7):375. https://doi.org/10.3390/v10070375
Chicago/Turabian StyleTalwar, Harvinder, Samer Najeeb Hanoudi, Sorin Draghici, and Lobelia Samavati. 2018. "Novel T7 Phage Display Library Detects Classifiers for Active Mycobacterium Tuberculosis Infection" Viruses 10, no. 7: 375. https://doi.org/10.3390/v10070375
APA StyleTalwar, H., Hanoudi, S. N., Draghici, S., & Samavati, L. (2018). Novel T7 Phage Display Library Detects Classifiers for Active Mycobacterium Tuberculosis Infection. Viruses, 10(7), 375. https://doi.org/10.3390/v10070375





