Comparative Pathogenesis of Asian and African-Lineage Zika Virus in Indian Rhesus Macaque’s and Development of a Non-Human Primate Model Suitable for the Evaluation of New Drugs and Vaccines
Abstract
:1. Introduction
2. Materials and Methods
2.1. Care and Use of Animals
2.2. Viruses, Cell Culture
2.3. Natural History Study
2.4. Primers and Probes
2.5. Biological Sample Collection
2.6. Viral Load by qRT-PCR
2.7. Viral Load by Plaque Assay
2.8. Indirect ELISA
2.9. Focus Reduction Neutralization Test (FRNT)
2.10. Efficacy and Immunogenicity Testing of Inactivated Vaccine
3. Results
3.1. Clinical Observations
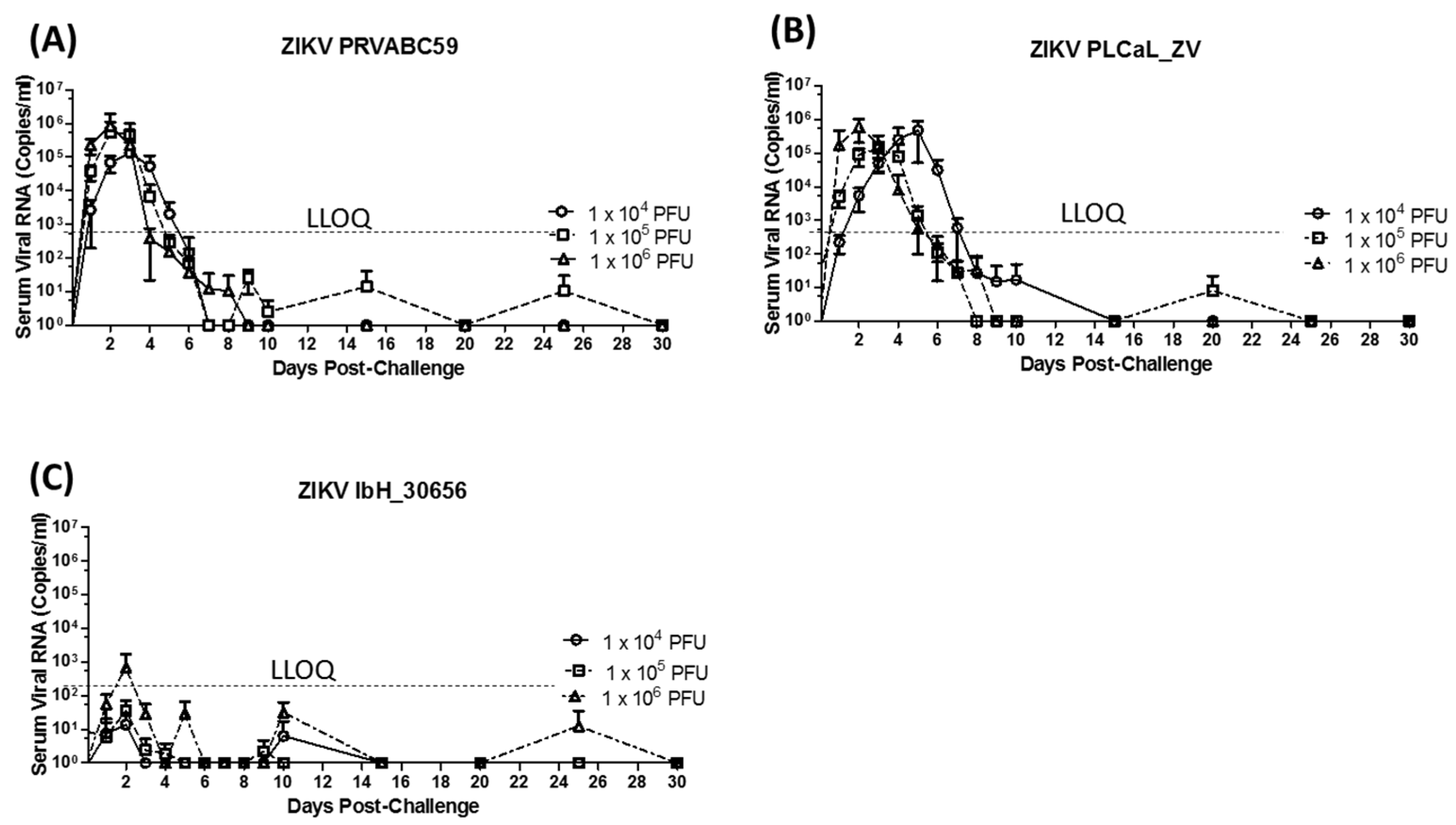
3.2. Virus Detection Following Primary Infection
3.3. Immune Response Following Primary Infection
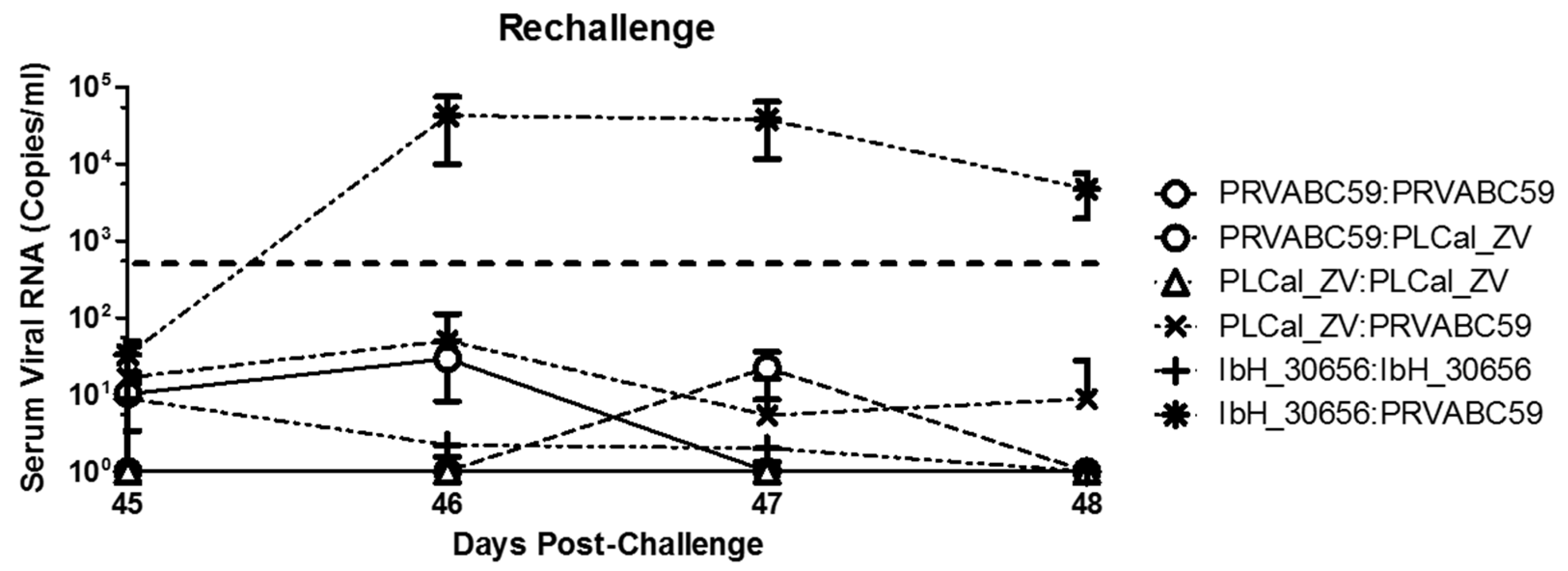
3.4. Virus Detection Following Secondary Challenge
3.5. Immune Response Following Secondary Challenge
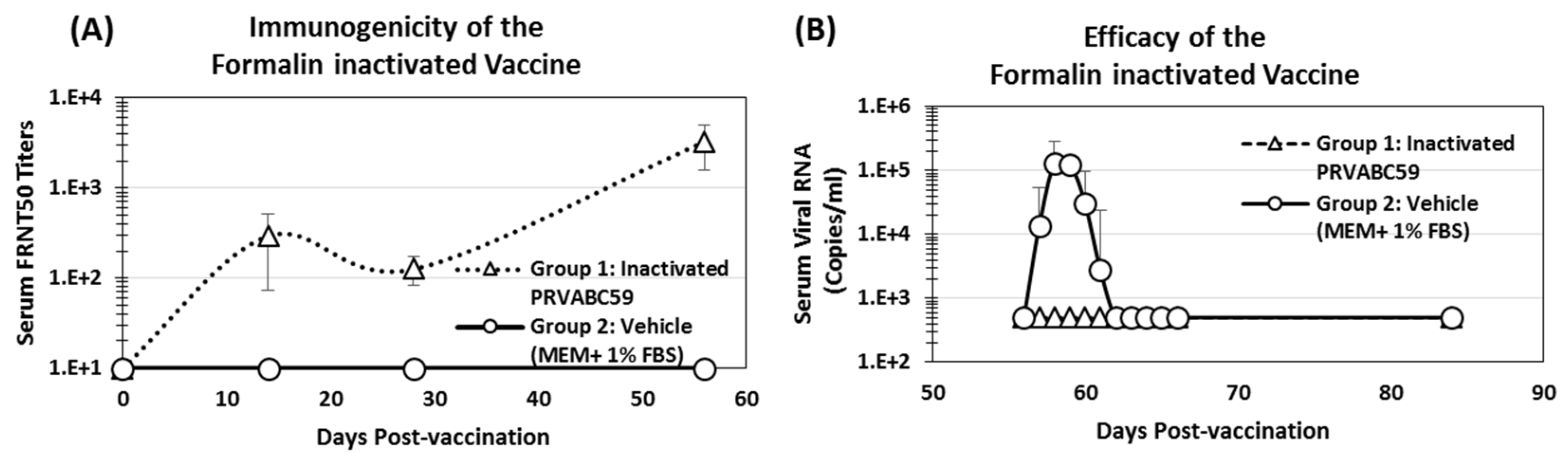
3.6. Immunogenicity and Efficacy of Inactivated ZIKV Vaccine
4. Discussion
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Schuler-Faccini, L.; Ribeiro, E.M.; Feitosa, I.M.; Horovitz, D.D.; Cavalcanti, D.P.; Pessoa, A.; Doriqui, M.J.; Neri, J.I.; Neto, J.M.; Wanderley, H.Y.; et al. Possible association between zika virus infection and microcephaly—Brazil, 2015. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 59–62. [Google Scholar] [CrossRef] [PubMed]
- Brasil, P.; Pereira, J.P., Jr.; Moreira, M.E.; Ribeiro Nogueira, R.M.; Damasceno, L.; Wakimoto, M.; Rabello, R.S.; Valderramos, S.G.; Halai, U.A.; Salles, T.S.; et al. Zika virus infection in pregnant women in Rio de Janeiro. N. Engl. J. Med. 2016, 375, 2321–2334. [Google Scholar] [CrossRef] [PubMed]
- Sarno, M.; Sacramento, G.A.; Khouri, R.; do Rosario, M.S.; Costa, F.; Archanjo, G.; Santos, L.A.; Nery, N., Jr.; Vasilakis, N.; Ko, A.I.; et al. Zika virus infection and stillbirths: A case of hydrops fetalis, hydranencephaly and fetal demise. PLoS Negl. Trop. Dis. 2016, 10, e0004517. [Google Scholar] [CrossRef] [PubMed]
- Franca, G.V.; Schuler-Faccini, L.; Oliveira, W.K.; Henriques, C.M.; Carmo, E.H.; Pedi, V.D.; Nunes, M.L.; Castro, M.C.; Serruya, S.; Silveira, M.F.; et al. Congenital zika virus syndrome in brazil: A case series of the first 1501 livebirths with complete investigation. Lancet 2016, 388, 891–897. [Google Scholar] [CrossRef]
- Melo, A.S.; Aguiar, R.S.; Amorim, M.M.; Arruda, M.B.; Melo, F.O.; Ribeiro, S.T.; Batista, A.G.; Ferreira, T.; Dos Santos, M.P.; Sampaio, V.V.; et al. Congenital zika virus infection: Beyond neonatal microcephaly. JAMA Neurol. 2016, 73, 1407–1416. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, W.K.; Carmo, E.H.; Henriques, C.M.; Coelho, G.; Vazquez, E.; Cortez-Escalante, J.; Molina, J.; Aldighieri, S.; Espinal, M.A.; Dye, C. Zika virus infection and associated neurologic disorders in Brazil. N. Engl. J. Med. 2017, 376, 1591–1593. [Google Scholar] [CrossRef] [PubMed]
- Brasil, P.; Sequeira, P.C.; Freitas, A.D.; Zogbi, H.E.; Calvet, G.A.; de Souza, R.V.; Siqueira, A.M.; de Mendonca, M.C.; Nogueira, R.M.; de Filippis, A.M.; et al. Guillain-barre syndrome associated with zika virus infection. Lancet 2016, 387, 1482. [Google Scholar] [CrossRef]
- Musso, D.; Gubler, D.J. Zika virus. Clin. Microbiol. Rev. 2016, 29, 487–524. [Google Scholar] [CrossRef] [PubMed]
- Dick, G.W.; Kitchen, S.F.; Haddow, A.J. Zika virus. I. Isolations and serological specificity. Trans. R. Soc. Trop. Med. Hyg. 1952, 46, 509–520. [Google Scholar] [CrossRef]
- Duffy, M.R.; Chen, T.-H.; Hancock, W.T.; Powers, A.M.; Kool, J.L.; Lanciotti, R.S.; Pretrick, M.; Marfel, M.; Holzbauer, S.; Dubray, C.; et al. Zika virus outbreak on yap island, federated states of micronesia. N. Engl. J. Med. 2009, 360, 2536–2543. [Google Scholar] [CrossRef] [PubMed]
- Musso, D.; Nilles, E.J.; Cao-Lormeau, V.M. Rapid spread of emerging zika virus in the Pacific area. Clin. Microbiol. Infect. 2014, 20, O595–O596. [Google Scholar] [CrossRef] [PubMed]
- Cao-Lormeau, V.M.; Roche, C.; Teissier, A.; Robin, E.; Berry, A.L.; Mallet, H.P.; Sall, A.A.; Musso, D. Zika virus, French Polynesia, South Pacific, 2013. Emerg. Infect. Dis. 2014, 20, 1085–1086. [Google Scholar] [CrossRef] [PubMed]
- Cao-Lormeau, V.M.; Blake, A.; Mons, S.; Lastere, S.; Roche, C.; Vanhomwegen, J.; Dub, T.; Baudouin, L.; Teissier, A.; Larre, P.; et al. Guillain-barre syndrome outbreak associated with zika virus infection in French Polynesia: A case-control study. Lancet 2016, 387, 1531–1539. [Google Scholar] [CrossRef]
- Besnard, M.; Eyrolle-Guignot, D.; Guillemette-Artur, P.; Lastere, S.; Bost-Bezeaud, F.; Marcelis, L.; Abadie, V.; Garel, C.; Moutard, M.L.; Jouannic, J.M.; et al. Congenital cerebral malformations and dysfunction in fetuses and newborns following the 2013 to 2014 zika virus epidemic in French Polynesia. Eur. Surveill. 2016, 21. [Google Scholar] [CrossRef] [PubMed]
- Campos, G.S.; Bandeira, A.C.; Sardi, S.I. Zika virus outbreak, bahia, brazil. Emerg. Infect. Dis. 2015, 21, 1885–1886. [Google Scholar] [CrossRef] [PubMed]
- Zanluca, C.; Melo, V.C.; Mosimann, A.L.; Santos, G.I.; Santos, C.N.; Luz, K. First report of autochthonous transmission of zika virus in Brazil. Mem. Inst. Oswaldo. Cruz. 2015, 110, 569–572. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Zika Virus, Microcephaly and Guillain-Barre Syndrome; WHO: Geneva, Switzerland, 2017. [Google Scholar]
- Morrison, T.E.; Diamond, M.S. Animal models of zika virus infection, pathogenesis, and immunity. J. Virol. 2017, 91, e00009-17. [Google Scholar] [CrossRef] [PubMed]
- Lazear, H.M.; Govero, J.; Smith, A.M.; Platt, D.J.; Fernandez, E.; Miner, J.J.; Diamond, M.S. A mouse model of zika virus pathogenesis. Cell Host Microbe. 2016, 19, 720–730. [Google Scholar] [CrossRef] [PubMed]
- Dowall, S.D.; Graham, V.A.; Rayner, E.; Atkinson, B.; Hall, G.; Watson, R.J.; Bosworth, A.; Bonney, L.C.; Kitchen, S.; Hewson, R. A susceptible mouse model for zika virus infection. PLoS Negl. Trop. Dis. 2016, 10, e0004658. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, S.; Balasubramaniam, V.R.M.T.; Brown, J.A.; Mena, I.; Grant, A.; Bardina, S.V.; Maringer, K.; Schwarz, M.C.; Maestre, A.M.; Sourisseau, M.; et al. A novel zika virus mouse model reveals strain specific differences in virus pathogenesis and host inflammatory immune responses. PLOS Pathog. 2017, 13, e1006258. [Google Scholar] [CrossRef] [PubMed]
- Aliota, M.T.; Caine, E.A.; Walker, E.C.; Larkin, K.E.; Camacho, E.; Osorio, J.E. Characterization of lethal zika virus infection in ag129 mice. PLoS Negl. Trop. Dis. 2016, 10, e0004682. [Google Scholar] [CrossRef] [PubMed]
- Rossi, S.L.; Tesh, R.B.; Azar, S.R.; Muruato, A.E.; Hanley, K.A.; Auguste, A.J.; Langsjoen, R.M.; Paessler, S.; Vasilakis, N.; Weaver, S.C. Characterization of a novel murine model to study zika virus. Am. J. Trop. Med. Hyg. 2016, 94, 1362–1369. [Google Scholar] [CrossRef] [PubMed]
- Larocca, R.A.; Abbink, P.; Peron, J.P.; Zanotto, P.M.; Iampietro, M.J.; Badamchi-Zadeh, A.; Boyd, M.; Nganga, D.; Kirilova, M.; Nityanandam, R.; et al. Vaccine protection against zika virus from brazil. Nature 2016, 536, 474–478. [Google Scholar] [CrossRef] [PubMed]
- Duggal, N.K.; Ritter, J.M.; McDonald, E.M.; Romo, H.; Guirakhoo, F.; Davis, B.S.; Chang, G.-J.J.; Brault, A.C. Differential neurovirulence of african and asian genotype zika virus isolates in outbred immunocompetent mice. Am. J. Trop. Med. Hyg. 2017, 97, 1410–1417. [Google Scholar] [CrossRef] [PubMed]
- Li, X.F.; Dong, H.L.; Huang, X.Y.; Qiu, Y.F.; Wang, H.J.; Deng, Y.Q.; Zhang, N.N.; Ye, Q.; Zhao, H.; Liu, Z.Y.; et al. Characterization of a 2016 clinical isolate of zika virus in non-human primates. EBioMedicine 2016, 12, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Dudley, D.M.; Aliota, M.T.; Mohr, E.L.; Weiler, A.M.; Lehrer-Brey, G.; Weisgrau, K.L.; Mohns, M.S.; Breitbach, M.E.; Rasheed, M.N.; Newman, C.M.; et al. A rhesus macaque model of asian-lineage zika virus infection. Nat. Commun. 2016, 7, 12204. [Google Scholar] [CrossRef] [PubMed]
- Osuna, C.E.; Lim, S.Y.; Deleage, C.; Griffin, B.D.; Stein, D.; Schroeder, L.T.; Omange, R.; Best, K.; Luo, M.; Hraber, P.T.; et al. Zika viral dynamics and shedding in rhesus and cynomolgus macaques. Nat. Med. 2016, 22, 1448–1455. [Google Scholar] [CrossRef] [PubMed]
- Haddow, A.D.; Nalca, A.; Rossi, F.D.; Miller, L.J.; Wiley, M.R.; Perez-Sautu, U.; Washington, S.C.; Norris, S.L.; Wollen-Roberts, S.E.; Shamblin, J.D.; et al. High infection rates for adult macaques after intravaginal or intrarectal inoculation with zika virus. Emerg. Infect. Dis. 2017, 23, 1274–1281. [Google Scholar] [CrossRef] [PubMed]
- Dowd, K.A.; Ko, S.-Y.; Morabito, K.M.; Yang, E.S.; Pelc, R.S.; DeMaso, C.R.; Castilho, L.R.; Abbink, P.; Boyd, M.; Nityanandam, R.; et al. Rapid development of a DNA vaccine for zika virus. Science 2016, 354, 237–240. [Google Scholar] [CrossRef] [PubMed]
- Abbink, P.; Larocca, R.A.; De La Barrera, R.A.; Bricault, C.A.; Moseley, E.T.; Boyd, M.; Kirilova, M.; Li, Z.; Ng’ang’a, D.; Nanayakkara, O.; et al. Protective efficacy of multiple vaccine platforms against zika virus challenge in rhesus monkeys. Science 2016, 353, 1129–1132. [Google Scholar] [CrossRef] [PubMed]
- Koide, F.; Goebel, S.; Snyder, B.; Walters, K.B.; Gast, A.; Hagelin, K.; Kalkeri, R.; Rayner, J. Development of a zika virus infection model in cynomolgus macaques. Front. Microbiol. 2016, 7, 2028. [Google Scholar] [CrossRef] [PubMed]
- Adams Waldorf, K.M.; Stencel-Baerenwald, J.E.; Kapur, R.P.; Studholme, C.; Boldenow, E.; Vornhagen, J.; Baldessari, A.; Dighe, M.K.; Thiel, J.; Merillat, S.; et al. Fetal brain lesions after subcutaneous inoculation of zika virus in a pregnant nonhuman primate. Nat. Med. 2016, 22, 1256. [Google Scholar] [CrossRef] [PubMed]
- Aliota, M.T.; Dudley, D.M.; Newman, C.M.; Mohr, E.L.; Gellerup, D.D.; Breitbach, M.E.; Buechler, C.R.; Rasheed, M.N.; Mohns, M.S.; Weiler, A.M.; et al. Heterologous protection against asian zika virus challenge in Rhesus macaques. PLoS Negl. Trop. Dis. 2016, 10, e0005168. [Google Scholar] [CrossRef] [PubMed]
- Lanciotti, R.S.; Kosoy, O.L.; Laven, J.J.; Velez, J.O.; Lambert, A.J.; Johnson, A.J.; Stanfield, S.M.; Duffy, M.R. Genetic and serologic properties of zika virus associated with an epidemic, yap state, Micronesia, 2007. Emerg. Infect. Dis. 2008, 14, 1232–1239. [Google Scholar] [CrossRef] [PubMed]
- Goebel, S.; Snyder, B.; Sellati, T.; Saeed, M.; Ptak, R.; Murray, M.; Bostwick, R.; Rayner, J.; Koide, F.; Kalkeri, R. A sensitive virus yield assay for evaluation of antivirals against zika virus. J. Virol. Methods 2016, 238, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Modjarrad, K.; Lin, L.; George, S.L.; Stephenson, K.E.; Eckels, K.H.; de la Barrera, R.A.; Jarman, R.G.; Sondergaard, E.; Tennant, J.; Ansel, J.L.; et al. Preliminary aggregate safety and immunogenicity results from three trials of a purified inactivated zika virus vaccine candidate: Phase 1, randomised, double-blind, placebo-controlled clinical trials. Lancet 2017, 391, 563–571. [Google Scholar] [CrossRef]
- Rayner, J.O.; Kalkeri, R.; Goebel, S.; Cai, Z.; Green, B.; Lin, S.; Snyder, B.; Hagelin, K.; Walters, K.B.; Koide, F. Viral RNA Was Only Detected in 4 Urine Samples from Animals Challenged with IbH_30656 While None of the Saliva Samples Had Detectable RNA Levels; Southern Research: Frederick, MD, USA, 2016. [Google Scholar]
- Gong, Z.; Xu, X.; Han, G.Z. The diversification of zika virus: Are there two distinct lineages? Genome Biol. Evol. 2017, 9, 2940–2945. [Google Scholar] [CrossRef] [PubMed]
- Haddow, A.D.; Schuh, A.J.; Yasuda, C.Y.; Kasper, M.R.; Heang, V.; Huy, R.; Guzman, H.; Tesh, R.B.; Weaver, S.C. Genetic characterization of zika virus strains: Geographic expansion of the asian lineage. PLoS Negl. Trop. Dis. 2012, 6, e1477. [Google Scholar] [CrossRef] [PubMed]
- Styer, L.M.; Kent, K.A.; Albright, R.G.; Bennett, C.J.; Kramer, L.D.; Bernard, K.A. Mosquitoes inoculate high doses of west nile virus as they probe and feed on live hosts. PLoS Pathog. 2007, 3, 1262–1270. [Google Scholar] [CrossRef] [PubMed]
- Dudley, D.M.; Newman, C.M.; Lalli, J.; Stewart, L.M.; Koenig, M.R.; Weiler, A.M.; Semler, M.R.; Barry, G.L.; Zarbock, K.R.; Mohns, M.S.; et al. Infection via mosquito bite alters zika virus tissue tropism and replication kinetics in Rhesus macaques. Nat. Commun. 2017, 8, 2096. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, A.J.; Smith, J.L.; Haese, N.N.; Broeckel, R.M.; Parkins, C.J.; Kreklywich, C.; DeFilippis, V.R.; Denton, M.; Smith, P.P.; Messer, W.B.; et al. Zika virus infection of rhesus macaques leads to viral persistence in multiple tissues. PLoS Pathog. 2017, 13, e1006219. [Google Scholar] [CrossRef] [PubMed]
- Aid, M.; Abbink, P.; Larocca, R.A.; Boyd, M.; Nityanandam, R.; Nanayakkara, O.; Martinot, A.J.; Moseley, E.T.; Blass, E.; Borducchi, E.N.; et al. Zika virus persistence in the central nervous system and lymph nodes of rhesus monkeys. Cell 2017, 169, 610–620. [Google Scholar] [CrossRef] [PubMed]
- Magnani, D.M.; Rogers, T.F.; Beutler, N.; Ricciardi, M.J.; Bailey, V.K.; Gonzalez-Nieto, L.; Briney, B.; Sok, D.; Le, K.; Strubel, A.; et al. Neutralizing human monoclonal antibodies prevent zika virus infection in macaques. Sci. Transl. Med. 2017, 9, eaan8184. [Google Scholar] [CrossRef] [PubMed]
| Group Number | Animal Number | Isolate | Target Dose (PFU/Animal) | Delivered Dose (PFU/Animal) |
|---|---|---|---|---|
| 1 | 4 (2M/2F) | PRVABC59 | 1 × 104 | 5.5 × 103 |
| 2 | 4 (2M/2F) | PRVABC59 | 1 × 105 | 9.1 × 104 |
| 3 | 4 (2M/2F) | PRVABC59 | 1 × 106 | 7.6 × 105 |
| 4 | 4 (2M/2F) | PLCal_ZV | 1 × 104 | 3.0 × 103 |
| 5 | 4 (2M/2F) | PLCal_ZV | 1 × 105 | 9.0 × 104 |
| 6 | 4 (2M/2F) | PLCal_ZV | 1 × 106 | 6.5 × 105 |
| 7 | 4 (2M/2F) | IbH_30656 | 1 × 104 | 2.8 × 103 |
| 8 | 4 (2M/2F) | IbH_30656 | 1 × 105 | 3.4 × 104 |
| 9 | 4 (2M/2F) | IbH_30656 | 1 × 106 | 3.9 × 105 |
| Group Number | Animal Number | Initial Challenge | Secondary Challenge | Target Dose (PFU/Animal) | Delivered Dose (PFU/Animal) |
|---|---|---|---|---|---|
| 1 | 6 (3M/3F) | PRVABC59 | PRVABC59 | 1 × 106 | 5.9 × 105 |
| 2 | 6 (3M/3F) | PRVABC59 | PLCal_ZV | 1 × 106 | 1.4 × 106 |
| 3 | 6 (3M/3F) | PLCal_ZV | PLCal_ZV | 1 × 106 | 1.4 × 106 |
| 4 | 6 (3M/3F) | PLCal_ZV | PRVABC59 | 1 × 106 | 5.9 × 105 |
| 5 | 6 (3M/3F) | IbH_30656 | IbH_30656 | 1 × 106 | 1.0 × 106 |
| 6 | 6 (3M/3F) | IbH_30656 | PRVABC59 | 1 × 106 | 5.9 × 105 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rayner, J.O.; Kalkeri, R.; Goebel, S.; Cai, Z.; Green, B.; Lin, S.; Snyder, B.; Hagelin, K.; Walters, K.B.; Koide, F. Comparative Pathogenesis of Asian and African-Lineage Zika Virus in Indian Rhesus Macaque’s and Development of a Non-Human Primate Model Suitable for the Evaluation of New Drugs and Vaccines. Viruses 2018, 10, 229. https://doi.org/10.3390/v10050229
Rayner JO, Kalkeri R, Goebel S, Cai Z, Green B, Lin S, Snyder B, Hagelin K, Walters KB, Koide F. Comparative Pathogenesis of Asian and African-Lineage Zika Virus in Indian Rhesus Macaque’s and Development of a Non-Human Primate Model Suitable for the Evaluation of New Drugs and Vaccines. Viruses. 2018; 10(5):229. https://doi.org/10.3390/v10050229
Chicago/Turabian StyleRayner, Jonathan O., Raj Kalkeri, Scott Goebel, Zhaohui Cai, Brian Green, Shuling Lin, Beth Snyder, Kimberly Hagelin, Kevin B. Walters, and Fusataka Koide. 2018. "Comparative Pathogenesis of Asian and African-Lineage Zika Virus in Indian Rhesus Macaque’s and Development of a Non-Human Primate Model Suitable for the Evaluation of New Drugs and Vaccines" Viruses 10, no. 5: 229. https://doi.org/10.3390/v10050229
APA StyleRayner, J. O., Kalkeri, R., Goebel, S., Cai, Z., Green, B., Lin, S., Snyder, B., Hagelin, K., Walters, K. B., & Koide, F. (2018). Comparative Pathogenesis of Asian and African-Lineage Zika Virus in Indian Rhesus Macaque’s and Development of a Non-Human Primate Model Suitable for the Evaluation of New Drugs and Vaccines. Viruses, 10(5), 229. https://doi.org/10.3390/v10050229





