Global Interactomics Connect Nuclear Mitotic Apparatus Protein NUMA1 to Influenza Virus Maturation
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Viruses
2.2. Virus Titration
2.3. Cytoplasmic and Nuclear Fractionation
2.4. Co-Immunoprecipitation (Co-IP)
2.5. Processing of IP Product for Western Blot Analysis and Mass Spectrometry
2.6. Transfection of Cells by siRNA
2.7. Western Blot Analysis
2.8. Immunofluorescent Microscopy
2.9. Structured Illumination Microscopy (SIM)
2.10. Electron Microscopy (EM)
2.11. Real-Time Polymerase Chain Reaction (PCR)
2.12. Bioinformatics and Statistical Analyses
3. Results
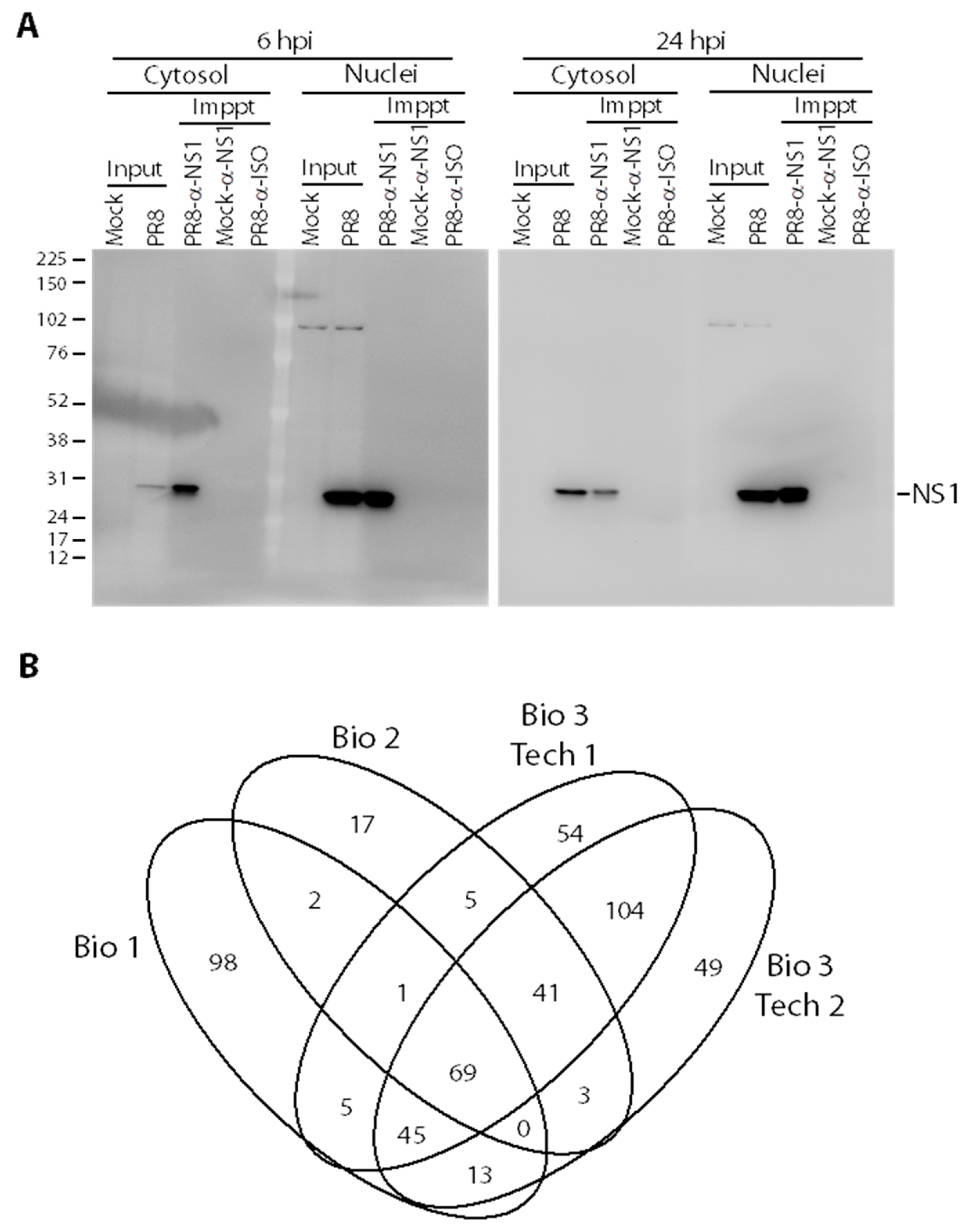
3.1. Co-Immunoprecipitation of Influenza A Virus (IAV) Non-Structural Protein 1 (NS1) from Infected Cells
3.2. Identifying NS1 Interacting Host Proteins by Mass Spectrometry (MS)
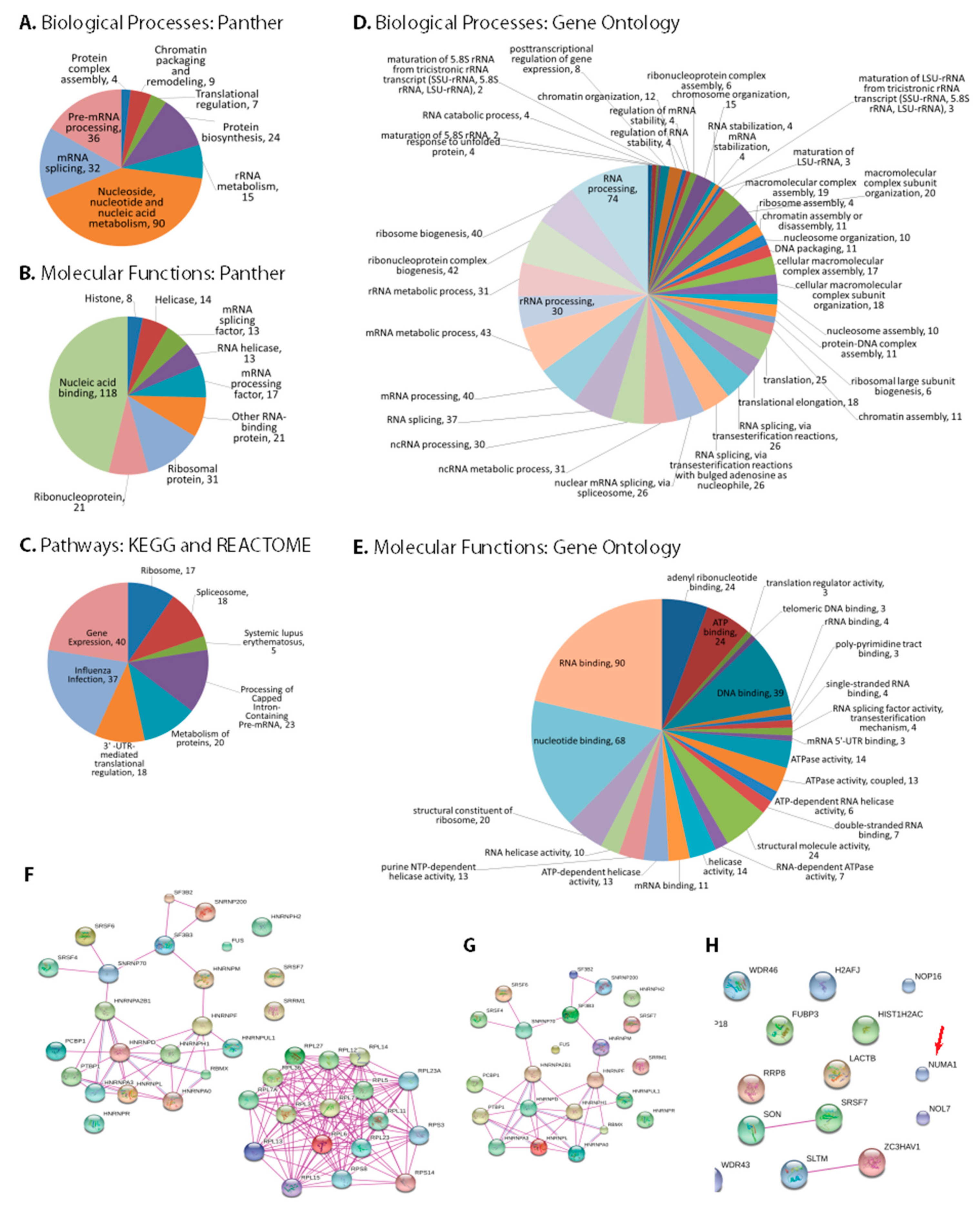
3.3. Bioinformatic Analysis of NS1 Interacting Proteins
3.4. Assessing the Necessity of NS1-Interacting Proteins for Viral Replication
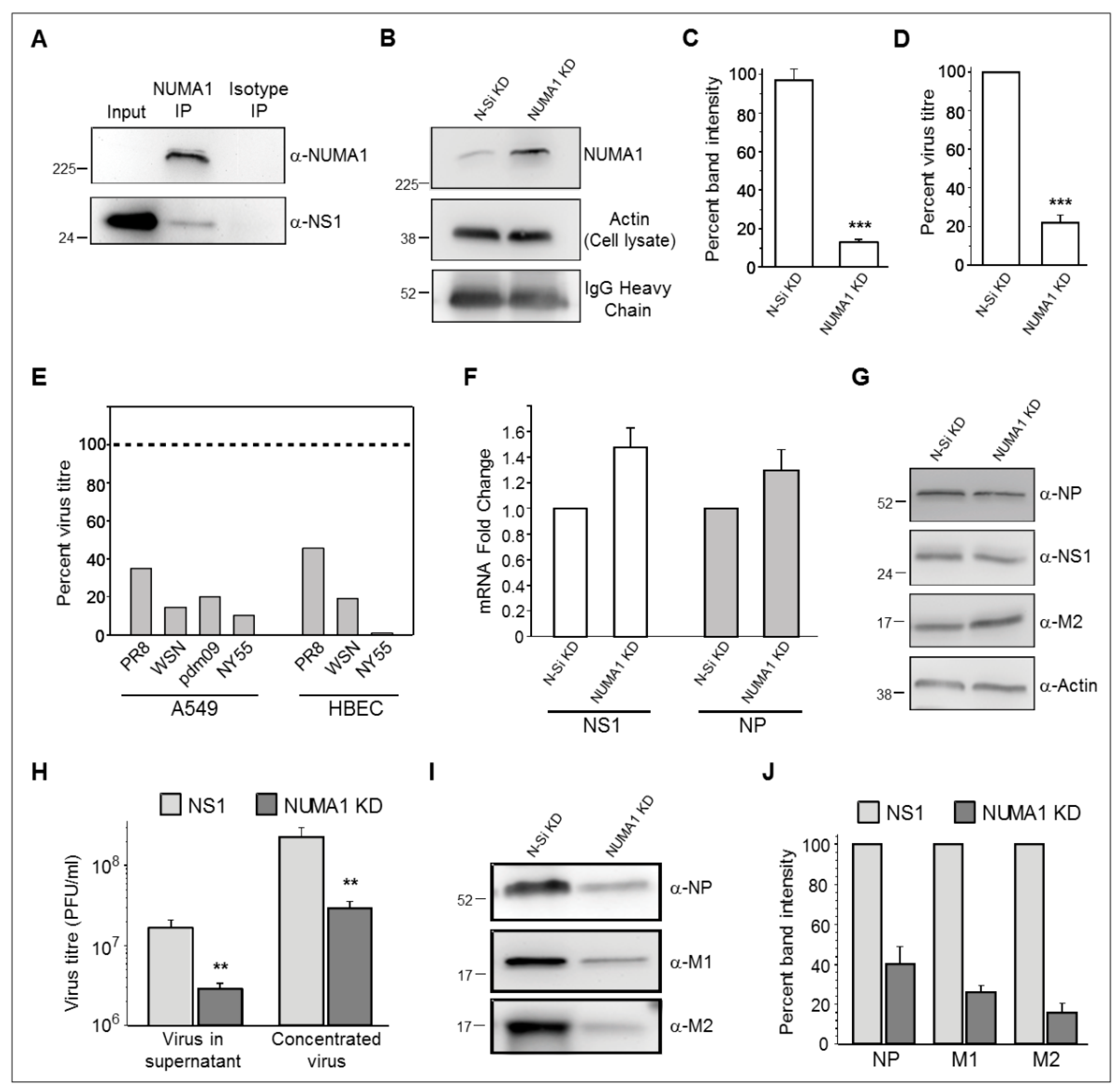
3.5. Validation of NS1-Host Factor Interaction by Reciprocal Immunoprecipitation (IP)
3.6. Role of Nuclear Mitotic Apparatus Protein 1 (NUMA1) in Influenza Virus Replication
3.7. NUMA1 Knockdown (KD) Does not Affect IAV Transcription and Translation
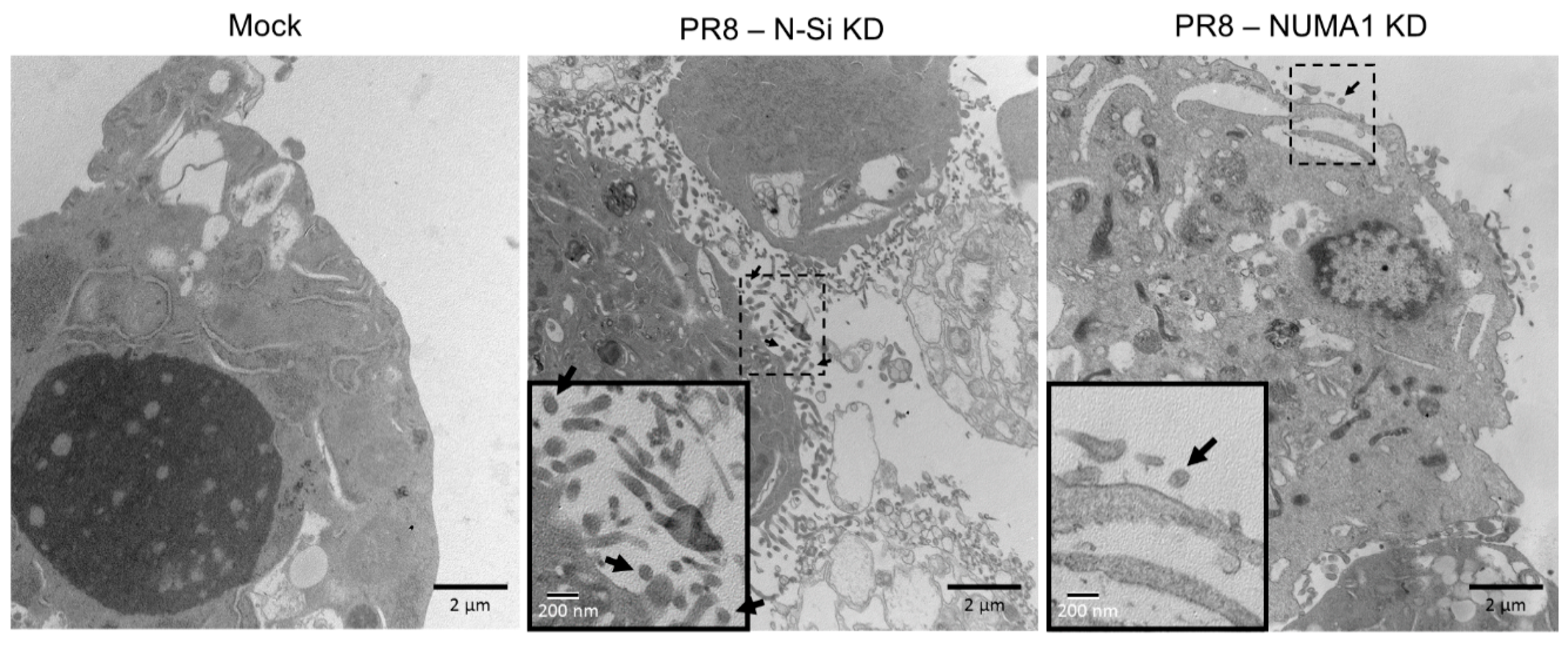
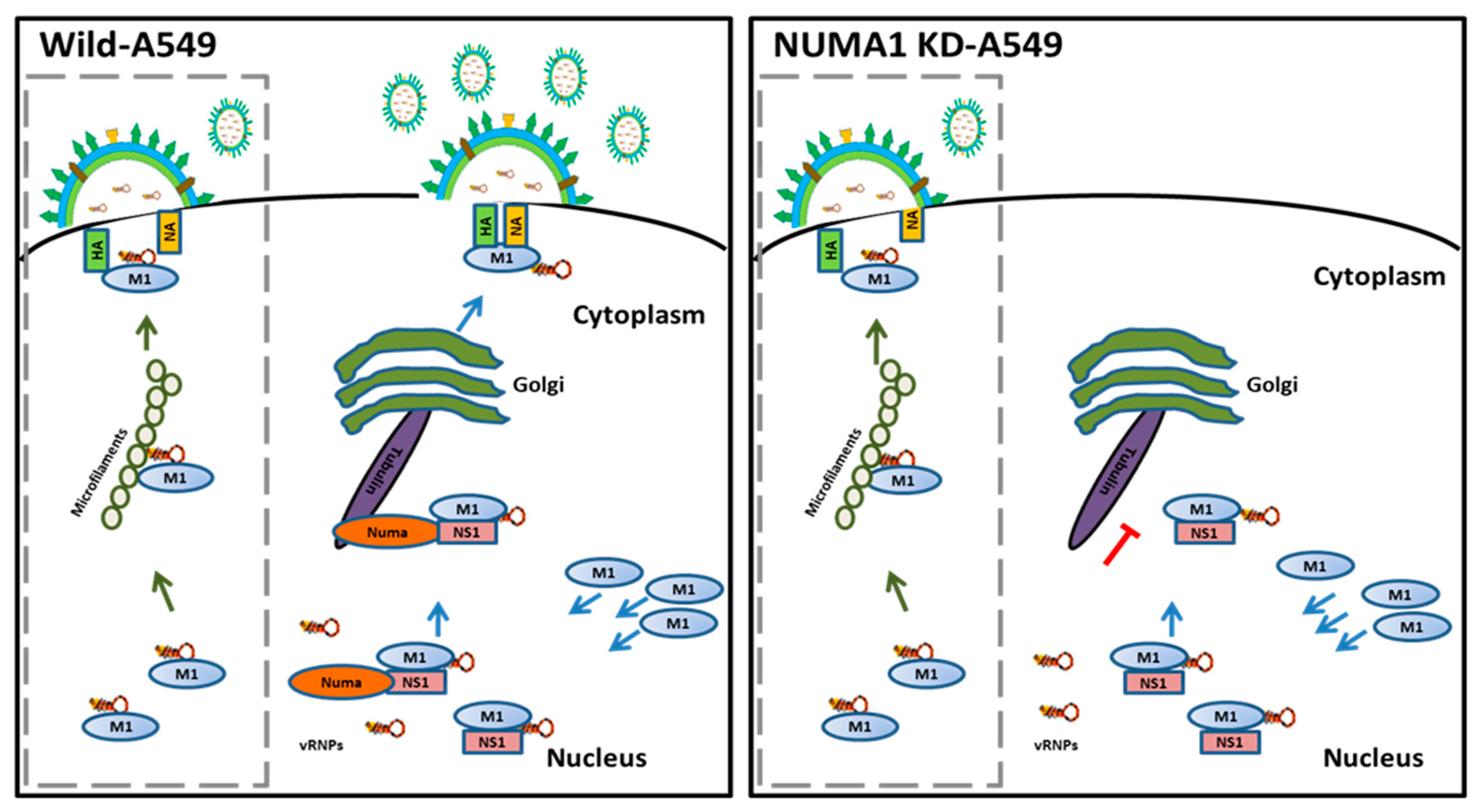
3.8. NUMA1 KD Affects IAV Maturation
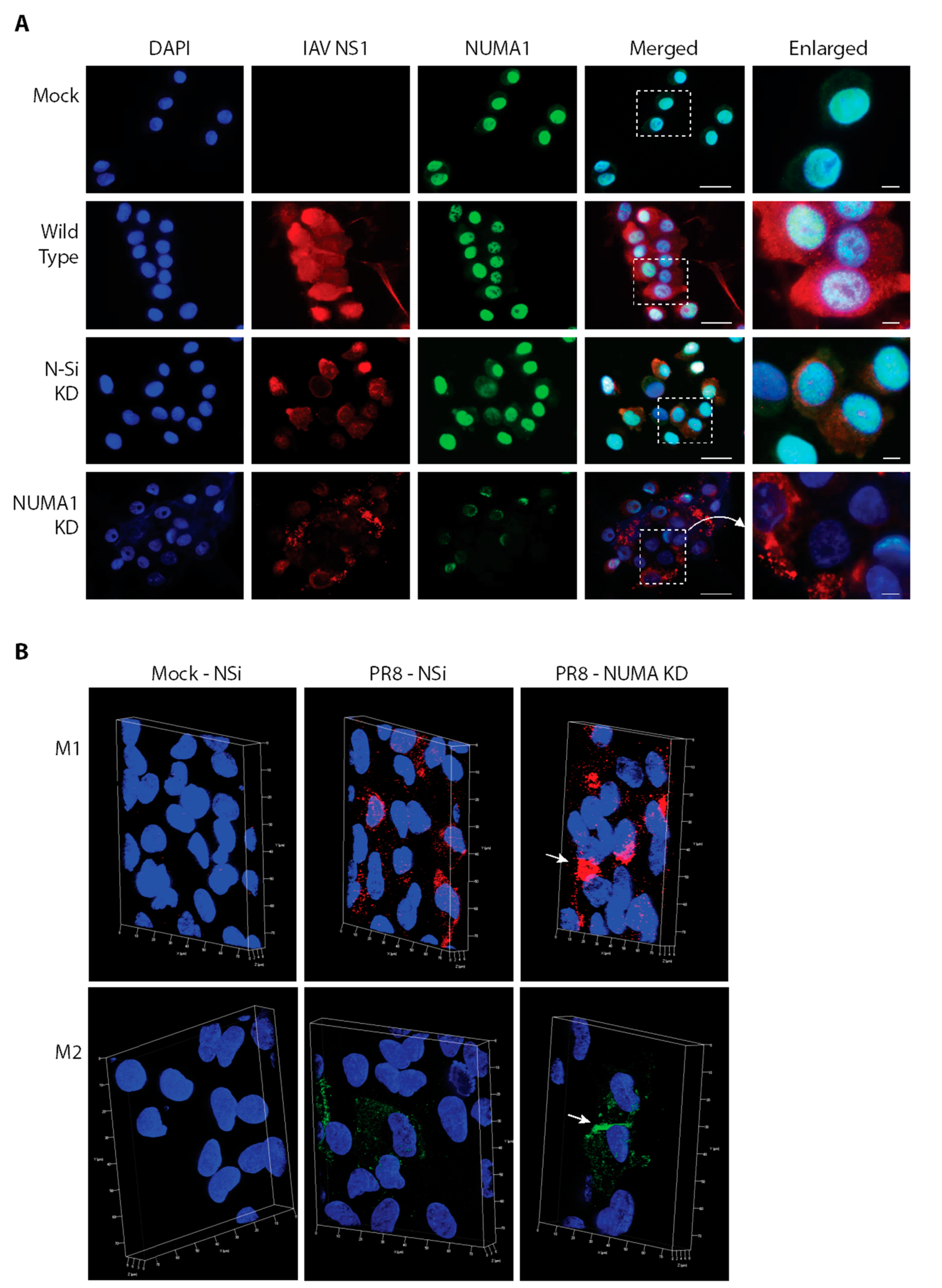
3.9. Localization of NUMA1 and Viral NS1, M1 and M2 Proteins in NUMA1-Deficient Cells
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pinto, L.H.; Lamb, R.A. Controlling influenza virus replication by inhibiting its proton channel. Mol. Biosyst. 2007, 3, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Hayden, F.G.; Hay, A.J. Emergence and transmission of influenza a viruses resistant to amantadine and rimantadine. Curr. Top. Microbiol. Immunol. 1992, 176, 119–130. [Google Scholar] [PubMed]
- Reece, P.A. Neuraminidase inhibitor resistance in influenza viruses. J. Med. Virol. 2007, 79, 1577–1586. [Google Scholar] [CrossRef]
- Lackenby, A.; Thompson, C.I.; Democratis, J. The potential impact of neuraminidase inhibitor resistant influenza. Curr. Opin. Inf. Dis. 2008, 21, 626–638. [Google Scholar]
- Coombs, K.M.; Berard, A.; Xu, W.; Krokhin, O.; Meng, X.; Cortens, J.P.; Kobasa, D.; Wilkins, J.; Brown, E.G. Quantitative proteomic analyses of influenza virus-infected cultured human lung cells. J. Virol. 2010, 84, 10888–10906. [Google Scholar] [CrossRef]
- Kroeker, A.L.; Ezzati, P.; Coombs, K.M.; Halayko, A.J. Influenza a infection of primary human airway epithelial cells up-regulates proteins related to purine metabolism and ubiquitin-related signaling. J. Proteome Res. 2013, 12, 3139–3151. [Google Scholar] [CrossRef]
- Kroeker, A.L.; Ezzati, P.; Halayko, A.J.; Coombs, K.M. Response of primary human airway epithelial cells to influenza infection—A quantitative proteomic study. J. Proteome Res. 2012, 11, 4132–4136. [Google Scholar] [CrossRef]
- Vester, D.; Rapp, E.; Gade, D.; Genzel, Y.; Reichl, U. Quantitative analysis of cellular proteome alterations in human influenza a virus-infected mammalian cell lines. Proteomics 2009, 9, 3316–3327. [Google Scholar] [CrossRef]
- Brass, A.L.; Huang, I.C.; Benita, Y.; John, S.P.; Krishnan, M.N.; Feeley, E.M.; Ryan, B.J.; Weyer, J.L.; van der Weyden, L.; Fikrig, E.; et al. The ifitm proteins mediate cellular resistance to influenza a h1n1 virus, west nile virus, and dengue virus. Cell 2009, 139, 1243–1254. [Google Scholar] [CrossRef]
- Karlas, A.; Machuy, N.; Shin, Y.; Pleissner, K.P.; Artarini, A.; Heuer, D.; Becker, D.; Khalil, H.; Ogilvie, L.A.; Hess, S.; et al. Genome-wide rnai screen identifies human host factors crucial for influenza virus replication. Nature 2010, 463, 818–822. [Google Scholar] [CrossRef]
- Konig, R.; Stertz, S.; Zhou, Y.; Inoue, A.; Hoffmann, H.H.; Bhattacharyya, S.; Alamares, J.G.; Tscherne, D.M.; Ortigoza, M.B.; Liang, Y.; et al. Human host factors required for influenza virus replication. Nature 2010, 463, 813–817. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Kawakami, E.; Shoemaker, J.E.; Lopes, T.J.; Matsuoka, Y.; Tomita, Y.; Kozuka-Hata, H.; Gorai, T.; Kuwahara, T.; Takeda, E.; et al. Influenza virus-host interactome screen as a platform for antiviral drug development. Cell Host Microbe 2014, 16, 795–805. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.C.; Kuo, R.L.; Lin, J.Y.; Huang, P.N.; Huang, Y.; Liu, H.; Arnold, J.J.; Chen, S.J.; Wang, R.Y.; Cameron, C.E.; et al. Cytoplasmic viral rna-dependent rna polymerase disrupts the intracellular splicing machinery by entering the nucleus and interfering with prp8. PLoS Pathog. 2014, 10, e1004199. [Google Scholar] [CrossRef] [PubMed]
- Jorba, N.; Juarez, S.; Torreira, E.; Gastaminza, P.; Zamarreno, N.; Albar, J.P.; Ortin, J. Analysis of the interaction of influenza virus polymerase complex with human cell factors. Proteomics 2008, 8, 2077–2088. [Google Scholar] [CrossRef] [PubMed]
- Engelhardt, O.G.; Smith, M.; Fodor, E. Association of the influenza a virus rna-dependent rna polymerase with cellular rna polymerase ii. J. Virol. 2005, 79, 5812–5818. [Google Scholar] [CrossRef] [PubMed]
- Deng, T.; Engelhardt, O.G.; Thomas, B.; Akoulitchev, A.V.; Brownlee, G.G.; Fodor, E. Role of ran binding protein 5 in nuclear import and assembly of the influenza virus rna polymerase complex. J. Virol. 2006, 80, 11911–11919. [Google Scholar] [CrossRef] [PubMed]
- Tafforeau, L.; Chantier, T.; Pradezynski, F.; Pellet, J.; Mangeot, P.E.; Vidalain, P.O.; Andre, P.; Rabourdin-Combe, C.; Lotteau, V. Generation and comprehensive analysis of an influenza virus polymerase cellular interaction network. J. Virol. 2011, 85, 13010–13018. [Google Scholar] [CrossRef]
- Huarte, M.; Sanz-Ezquerro, J.J.; Roncal, F.; Ortin, J.; Nieto, A. Pa subunit from influenza virus polymerase complex interacts with a cellular protein with homology to a family of transcriptional activators. J. Virol. 2001, 75, 8597–8604. [Google Scholar] [CrossRef]
- Salvatore, M.; Basler, C.F.; Parisien, J.P.; Horvath, C.M.; Bourmakina, S.; Zheng, H.; Muster, T.; Palese, P.; Garcia-Sastre, A. Effects of influenza a virus ns1 protein on protein expression: The ns1 protein enhances translation and is not required for shutoff of host protein synthesis. J. Virol. 2002, 76, 1206–1212. [Google Scholar] [CrossRef]
- Hale, B.G.; Randall, R.E.; Ortin, J.; Jackson, D. The multifunctional ns1 protein of influenza a viruses. J. Gen. Virol. 2008, 89, 2359–2376. [Google Scholar] [CrossRef]
- Kreijtz, J.H.; Fouchier, R.A.; Rimmelzwaan, G.F. Immune responses to influenza virus infection. Virus Res. 2011, 162, 19–30. [Google Scholar] [CrossRef] [PubMed]
- Hale, B.G.; Albrecht, R.A.; Garcia-Sastre, A. Innate immune evasion strategies of influenza viruses. Future Microbiol. 2010, 5, 23–41. [Google Scholar] [CrossRef] [PubMed]
- Solorzano, A.; Webby, R.J.; Lager, K.M.; Janke, B.H.; Garcia-Sastre, A.; Richt, J.A. Mutations in the ns1 protein of swine influenza virus impair anti-interferon activity and confer attenuation in pigs. J. Virol. 2005, 79, 7535–7543. [Google Scholar] [CrossRef] [PubMed]
- Quinlivan, M.; Zamarin, D.; Garcia-Sastre, A.; Cullinane, A.; Chambers, T.; Palese, P. Attenuation of equine influenza viruses through truncations of the ns1 protein. J. Virol. 2005, 79, 8431–8439. [Google Scholar] [CrossRef]
- Donelan, N.R.; Basler, C.F.; Garcia-Sastre, A. A recombinant influenza a virus expressing an rna-binding-defective ns1 protein induces high levels of beta interferon and is attenuated in mice. J. Virol. 2003, 77, 13257–13266. [Google Scholar] [CrossRef] [PubMed]
- Krug, R.M. Functions of the influenza a virus ns1 protein in antiviral defense. Curr. Opin. Virol. 2015, 12, 1–6. [Google Scholar] [CrossRef]
- Talon, J.; Horvath, C.M.; Polley, R.; Basler, C.F.; Muster, T.; Palese, P.; Garcia-Sastre, A. Activation of interferon regulatory factor 3 is inhibited by the influenza a virus ns1 protein. J. Virol. 2000, 74, 7989–7996. [Google Scholar] [CrossRef]
- Wang, X.Y.; Li, M.; Zheng, H.Y.; Muster, T.; Palese, P.; Beg, A.A.; Garcia-Sastre, A. Influenza a virus ns1 protein prevents activation of nf-kappa b and induction of alpha/beta interferon. J. Virol. 2000, 74, 11566–11573. [Google Scholar] [CrossRef] [PubMed]
- Silverman, R.H. Viral encounters with 2’,5’-oligoadenylate synthetase and rnase l during the interferon antiviral response. J. Virol. 2007, 81, 12720–12729. [Google Scholar] [CrossRef]
- Iwai, A.; Shiozaki, T.; Kawai, T.; Akira, S.; Kawaoka, Y.; Takada, A.; Kida, H.; Miyazaki, T. Influenza a virus polymerase inhibits type i interferon induction by binding to interferon beta promoter stimulator 1. J. Biol. Chem. 2010, 285, 32064–32074. [Google Scholar] [CrossRef]
- VirHostNet 2.0. Available online: http://virhostnet.prabi.fr/ (accessed on 17 May 2016).
- Guirimand, T.; Delmotte, S.; Navratil, V. Virhostnet 2.0: Surfing on the web of virus/host molecular interactions data. Nucleic Acids Res. 2015, 43, D583–D587. [Google Scholar] [CrossRef] [PubMed]
- de Chassey, B.; Aublin-Gex, A.; Ruggieri, A.; Meyniel-Schicklin, L.; Pradezynski, F.; Davoust, N.; Chantier, T.; Tafforeau, L.; Mangeot, P.E.; Ciancia, C.; et al. The interactomes of influenza virus ns1 and ns2 proteins identify new host factors and provide insights for adar1 playing a supportive role in virus replication. PLoS Pathog. 2013, 9, e1003440. [Google Scholar] [CrossRef] [PubMed]
- Falcon, A.M.; Fortes, P.; Marion, R.M.; Beloso, A.; Ortin, J. Interaction of influenza virus ns1 protein and the human homologue of staufen in vivo and in vitro. Nucleic Acids Res. 1999, 27, 2241–2247. [Google Scholar] [CrossRef] [PubMed]
- Mok, B.W.; Song, W.; Wang, P.; Tai, H.; Chen, Y.; Zheng, M.; Wen, X.; Lau, S.Y.; Wu, W.L.; Matsumoto, K.; et al. The ns1 protein of influenza a virus interacts with cellular processing bodies and stress granules through rna-associated protein 55 (rap55) during virus infection. J. Virol. 2012, 86, 12695–12707. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhou, J.; Du, Y. Hnrnp a2/b1 interacts with influenza a viral protein ns1 and inhibits virus replication potentially through suppressing ns1 rna/protein levels and ns1 mrna nuclear export. Virology 2014, 449, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Li, Y.; Pyo, H.M.; Lu, X.; Raman, S.N.; Liu, Q.; Brown, E.G.; Zhou, Y. Identification of rna helicase a as a cellular factor that interacts with influenza a virus ns1 protein and its role in the virus life cycle. J. Virol. 2012, 86, 1942–1954. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Liu, C.H.; Zhou, L.; Krug, R.M. Cellular ddx21 rna helicase inhibits influenza a virus replication but is counteracted by the viral ns1 protein. Cell Host Microbe 2014, 15, 484–493. [Google Scholar] [CrossRef] [PubMed]
- Tawaratsumida, K.; Phan, V.; Hrincius, E.R.; High, A.A.; Webby, R.; Redecke, V.; Hacker, H. Quantitative proteomic analysis of the influenza a virus nonstructural proteins ns1 and ns2 during natural cell infection identifies pact as an ns1 target protein and antiviral host factor. J. Virol. 2014, 88, 9038–9048. [Google Scholar] [CrossRef]
- Kuo, R.L.; Li, Z.H.; Li, L.H.; Lee, K.M.; Tam, E.H.; Liu, H.M.; Liu, H.P.; Shih, S.R.; Wu, C.C. Interactome analysis of the ns1 protein encoded by influenza a h1n1 virus reveals a positive regulatory role of host protein prp19 in viral replication. J. Proteome Res. 2016, 15, 1639–1648. [Google Scholar] [CrossRef]
- Thulasi Raman, S.N.; Zhou, Y. Networks of host factors that interact with ns1 protein of influenza a virus. Front. Microbiol. 2016, 7, 654. [Google Scholar] [CrossRef]
- Rahim, M.N.; Selman, M.; Sauder, P.J.; Forbes, N.E.; Stecho, W.; Xu, W.; Lebar, M.; Brown, E.G.; Coombs, K.M. Generation and characterization of a new panel of broadly-reactive monoclonal anti-ns1 antibodies for detection of influenza a virus. J. Gen. Virol. 2013, 94, 592–604. [Google Scholar] [CrossRef]
- Brown, E.G. Increased virulence of a mouse-adapted variant of influenza a/fm/1/47 virus is controlled by mutations in genome segments 4, 5, 7, and 8. J. Virol. 1990, 64, 4523–4533. [Google Scholar]
- Yang, M.; Berhane, Y.; Salo, T.; Li, M.; Hole, K.; Clavijo, A. Development and application of monoclonal antibodies against avian influenza virus nucleoprotein. J. Virol. Meth. 2008, 147, 265–274. [Google Scholar] [CrossRef]
- Righolt, C.H.; Guffei, A.; Knecht, H.; Young, I.T.; Stallinga, S.; van Vliet, L.J.; Mai, S. Differences in nuclear DNA organization between lymphocytes, hodgkin and reed-sternberg cells revealed by structured illumination microscopy. J. Cell. Biochem. 2014, 115, 1441–1448. [Google Scholar] [CrossRef]
- Database for Annotation, Visualization and Integrated Discovery (DAVID). Available online: https://david.ncifcrf.gov/ (accessed on 30 March 2015).
- Consensus Path Database (CPDB). Available online: http://consensuspathdb.org/ (accessed on 3 September 2016).
- Search Tool for the Retrieval of Interacting Genes/Proteins (STRING). Available online: http://string-db.org/ (accessed on 22 October 2015).
- Fant, X.; Merdes, A.; Haren, L. Cell and molecular biology of spindle poles and numa. Int. Rev. Cytol. 2004, 238, 1–57. [Google Scholar]
- Chen, B.J.; Leser, G.P.; Jackson, D.; Lamb, R.A. The influenza virus m2 protein cytoplasmic tail interacts with the m1 protein and influences virus assembly at the site of virus budding. J. Virol. 2008, 82, 10059–10070. [Google Scholar] [CrossRef]
- Nayak, D.P.; Hui, E.K.; Barman, S. Assembly and budding of influenza virus. Virus Res 2004, 106, 147–165. [Google Scholar] [CrossRef]
- Rossman, J.S.; Lamb, R.A. Influenza virus assembly and budding. Virology 2011, 411, 229–236. [Google Scholar] [CrossRef]
- Dubois, J.; Terrier, O.; Rosa-Calatrava, M. Influenza viruses and mrna splicing: Doing more with less. MBio 2014, 5, e00070-14. [Google Scholar] [CrossRef]
- Lamb, R.A.; Choppin, P.W.; Chanock, R.M.; Lai, C.J. Mapping of the two overlapping genes for polypeptides ns1 and ns2 on rna segment 8 of influenza virus genome. Proc. Natl. Acad. Sci. USA 1980, 77, 1857–1861. [Google Scholar] [CrossRef]
- Lamb, R.A.; Choppin, P.W. Identification of a second protein (m2) encoded by rna segment 7 of influenza virus. Virology 1981, 112, 729–737. [Google Scholar] [CrossRef]
- Qiu, Y.; Nemeroff, M.; Krug, R.M. The influenza virus ns1 protein binds to a specific region in human u6 snrna and inhibits u6-u2 and u6-u4 snrna interactions during splicing. RNA 1995, 1, 304–316. [Google Scholar]
- Wang, W.; Krug, R.M. U6atac snrna, the highly divergent counterpart of u6 snrna, is the specific target that mediates inhibition of at-ac splicing by the influenza virus ns1 protein. RNA 1998, 4, 55–64. [Google Scholar]
- Park, Y.W.; Katze, M.G. Translational control by influenza virus. Identification of cis-acting sequences and trans-acting factors which may regulate selective viral mrna translation. J. Biol. Chem. 1995, 270, 28433–28439. [Google Scholar]
- Garfinkel, M.S.; Katze, M.G. Translational control by influenza-virus—Selective translation is mediated by sequences within the viral messenger-rna 5’-untranslated region. J. Biol. Chem. 1993, 268, 22223–22226. [Google Scholar]
- de la Luna, S.; Fortes, P.; Beloso, A.; Ortin, J. Influenza virus ns1 protein enhances the rate of translation initiation of viral mrnas. J. Virol. 1995, 69, 2427–2433. [Google Scholar]
- Aragon, T.; de la Luna, S.; Novoa, I.; Carrasco, L.; Ortin, J.; Nieto, A. Eukaryotic translation initiation factor 4gi is a cellular target for ns1 protein, a translational activator of influenza virus. Mol. Cell. Biol. 2000, 20, 6259–6268. [Google Scholar] [CrossRef]
- Marc, D. Influenza virus non-structural protein ns1: Interferon antagonism and beyond. J. Gen. Virol. 2014, 95, 2594–2611. [Google Scholar] [CrossRef]
- Satterly, N.; Tsai, P.L.; van Deursen, J.; Nussenzveig, D.R.; Wang, Y.M.; Faria, P.A.; Levay, A.; Levy, D.E.; Fontoura, B.M.A. Influenza virus targets the mrna export machinery and the nuclear pore complex. Proc. Natl. Acad. Sci. USA 2007, 104, 1853–1858. [Google Scholar] [CrossRef]
- Robb, N.C.; Chase, G.; Bier, K.; Vreede, F.T.; Shaw, P.C.; Naffakh, N.; Schwemmle, M.; Fodor, E. The influenza a virus ns1 protein interacts with the nucleoprotein of viral ribonucleoprotein complexes. J. Virol. 2011, 85, 5228–5231. [Google Scholar] [CrossRef]
- Hatada, E.; Fukuda, R. Binding of influenza-a virus ns1 protein to dsrna in vitro. J. Gen. Virol. 1992, 73, 3325–3329. [Google Scholar] [CrossRef]
- Brydon, E.W.; Morris, S.J.; Sweet, C. Role of apoptosis and cytokines in influenza virus morbidity. FEMS Microbiol. Rev. 2005, 29, 837–850. [Google Scholar] [CrossRef]
- Hinshaw, V.S.; Olsen, C.W.; Dybdahl-Sissoko, N.; Evans, D. Apoptosis: A mechanism of cell killing by influenza a and b viruses. J. Virol. 1994, 68, 3667–3673. [Google Scholar]
- Takizawa, T.; Matsukawa, S.; Higuchi, Y.; Nakamura, S.; Nakanishi, Y.; Fukuda, R. Induction of programmed cell death (apoptosis) by influenza virus infection in tissue culture cells. J. Gen. Virol. 1993, 74, 2347–2355. [Google Scholar] [CrossRef]
- Radulescu, A.E.; Cleveland, D.W. Numa after 30 years: The matrix revisited. Trends Cell Biol. 2010, 20, 214–222. [Google Scholar] [CrossRef]
- Yamauchi, Y.; Kiriyama, K.; Kimura, H.; Nishiyama, Y. Herpes simplex virus induces extensive modification and dynamic relocalisation of the nuclear mitotic apparatus (numa) protein in interphase cells. J. Cell Sci. 2008, 121, 2087–2096. [Google Scholar] [CrossRef]
- Seldin, L.; Muroyama, A.; Lechler, T. Numa-microtubule interactions are critical for spindle orientation and the morphogenesis of diverse epidermal structures. Elife 2016, 5, e12504. [Google Scholar] [CrossRef]
- Haren, L.; Merdes, A. Direct binding of numa to tubulin is mediated by a novel sequence motif in the tail domain that bundles and stabilizes microtubules. J. Cell Sci. 2002, 115, 1815–1824. [Google Scholar]
- Merdes, A.; Ramyar, K.; Vechio, J.D.; Cleveland, D.W. A complex of numa and cytoplasmic dynein is essential for mitotic spindle assembly. Cell 1996, 87, 447–458. [Google Scholar] [CrossRef]
- Merdes, A.; Heald, R.; Samejima, K.; Earnshaw, W.C.; Cleveland, D.W. Formation of spindle poles by dynein/dynactin-dependent transport of numa. J. Cell Biol. 2000, 149, 851–862. [Google Scholar] [CrossRef]
- Bonifacino, J.S.; Lippincott-Schwartz, J. Coat proteins: Shaping membrane transport. Nat. Rev. Mol. Cell Biol. 2003, 4, 409–414. [Google Scholar] [CrossRef]
- Polishchuk, R.S.; Mironov, A.A.; Luini, A. Exocytic pathway. In Encyclopedic Reference of Genomics and Proteomics in Molecular Medicine; Springer: Berlin/Heidelberg, Germany, 2006; pp. 536–540. [Google Scholar]
- Boulo, S.; Akarsu, H.; Ruigrok, R.W.; Baudin, F. Nuclear traffic of influenza virus proteins and ribonucleoprotein complexes. Virus Res. 2007, 124, 12–21. [Google Scholar] [CrossRef]
- Bouvier, N.M.; Palese, P. The biology of influenza viruses. Vaccine 2008, 26 (Suppl. 4), D49–D53. [Google Scholar] [CrossRef]
- Calder, L.J.; Wasilewski, S.; Berriman, J.A.; Rosenthal, P.B. Structural organization of a filamentous influenza a virus. Proc. Natl. Acad. Sci. USA 2010, 107, 10685–10690. [Google Scholar] [CrossRef]
- Ali, A.; Avalos, R.T.; Ponimaskin, E.; Nayak, D.P. Influenza virus assembly: Effect of influenza virus glycoproteins on the membrane association of m1 protein. J. Virol. 2000, 74, 8709–8719. [Google Scholar] [CrossRef]
- Nayak, D.P.; Balogun, R.A.; Yamada, H.; Zhou, Z.H.; Barman, S. Influenza virus morphogenesis and budding. Virus Res. 2009, 143, 147–161. [Google Scholar] [CrossRef]
- Avalos, R.T.; Yu, Z.; Nayak, D.P. Association of influenza virus np and m1 proteins with cellular cytoskeletal elements in influenza virus-infected cells. J. Virol. 1997, 71, 2947–2958. [Google Scholar]
- Amorim, M.J.; Bruce, E.A.; Read, E.K.C.; Foeglein, A.; Mahen, R.; Stuart, A.D.; Digard, P. A rab11-and microtubule-dependent mechanism for cytoplasmic transport of influenza a virus viral RNA. J. Virol. 2011, 85, 4143–4156. [Google Scholar] [CrossRef]

| Uniprot | Gene Symbol | Protein | Peptides (95%) | Unused Score * | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Bio1 | Bio2 | Bio3 Tech1 | Bio3 Tech2 | Bio1 | Bio2 | Bio3 Tech1 | Bio3 Tech2 | |||
| P55265 | ADAR | Double-stranded RNA-specific adenosine deaminase | 4 | 4 | 20 | 20 | 2.86 | 7.54 | 37.77 | 35.31 |
| Q14692 | BMS1 | Ribosome biogenesis protein BMS1 homolog | 3 | 9 | 7 | 5 | 4.29 | 17.6 | 16.75 | 10.34 |
| Q14137 | BOP1 | Ribosome biogenesis protein BOP1 | 4 | 9 | 7 | 9 | 8 | 17.8 | 14.19 | 15.1 |
| Q8TDN6 | BRIX1 | Ribosome biogenesis protein BRX1 homolog | 2 | 5 | 8 | 10 | 3.54 | 11.06 | 15.72 | 16.79 |
| Q9Y224 | C14orf166# | UPF0568 protein C14orf166 | 3 | - | 8 | 9 | 6 | - | 13.56 | 14.89 |
| Q1ED39 | C16orf88 | Protein C16orf88 | - | 4 | 6 | 8 | - | 8 | 13.53 | 16.01 |
| Q9Y3I0 | C22orf28 | UPF0027 protein C22orf28 | 6 | - | 12 | 10 | 9.47 | - | 20.52 | 21.07 |
| Q7Z7K6 | CENPV | Centromere protein V | 4 | 2 | 3 | 3 | 7.76 | 2.96 | 5.84 | 5.67 |
| Q969X6 | CIRH1A | Cirhin OS | - | 10 | 6 | 11 | - | 18.31 | 13.36 | 19.31 |
| Q13206 | DDX10 | Probable ATP-dependent RNA helicase DDX10 | - | 2 | 4 | 3 | - | 2.29 | 8.05 | 5.44 |
| Q92841 | DDX17# | Probable ATP-dependent RNA helicase DDX17 | 5 | 2 | 18 | 16 | 7.4 | 4.03 | 29.5 | 28.76 |
| Q9NVP1 | DDX18 | ATP-dependent RNA helicase DDX18 | 9 | 14 | 12 | 13 | 17.53 | 20.77 | 22.19 | 20.23 |
| Q9NR30 | DDX21# | Nucleolar RNA helicase 2 | 14 | 5 | 14 | 16 | 27.57 | 10.05 | 25.03 | 30.43 |
| Q9GZR7 | DDX24 | ATP-dependent RNA helicase DDX24 | 4 | 4 | 11 | 8 | 8.15 | 7.07 | 20.93 | 15.19 |
| Q96GQ7 | DDX27 | Probable ATP-dependent RNA helicase DDX27 | - | 4 | 5 | 5 | - | 9.11 | 10.71 | 10.45 |
| O00571 | DDX3X# | ATP-dependent RNA helicase DDX3X | 4 | - | 9 | 12 | 8 | - | 14.93 | 18.22 |
| P17844 | DDX5 | Probable ATP-dependent RNA helicase DDX5 | 3 | - | - | 7 | 5.88 | - | - | 11.15 |
| Q8TDD1 | DDX54 | ATP-dependent RNA helicase DDX54 | 2 | 6 | 2 | 2 | 4 | 11.81 | 3.78 | 3 |
| Q9NY93 | DDX56# | Probable ATP-dependent RNA helicase DDX56 | - | 4 | - | 2 | - | 7.22 | - | 4.45 |
| O43143 | DHX15# | Putative pre-mRNA-splicing factor ATP-dependent RNA helicase DHX15 | 5 | - | 28 | 22 | 8.66 | - | 46.27 | 40.65 |
| Q7L2E3 | DHX30# | Putative ATP-dependent RNA helicase DHX30 | 2 | 9 | 53 | 59 | 3.05 | 17.57 | 87.85 | 100.67 |
| O60832 | DKC1 | H/ACA ribonucleoprotein complex subunit 4 | 3 | 3 | 5 | 3 | 4.52 | 6.14 | 10.8 | 6.18 |
| Q5QJE6 | DNTTIP2 | Deoxynucleotidyltransferase terminal-interacting protein 2 | - | 8 | 5 | 4 | - | 14.42 | 10.49 | 8.82 |
| Q99848 | EBNA1BP2 | Probable rRNA-processing protein EBP2 | 2 | 7 | 7 | 7 | 3.5 | 11.2 | 12.59 | 13.09 |
| P19525 | EIF2AK2 | Interferon-induced, double-stranded RNA-activated protein kinase | - | 3 | 6 | 6 | - | 6.6 | 12.89 | 12.21 |
| P56537 | EIF6 | Eukaryotic translation initiation factor 6 | 2 | 6 | 4 | 5 | 2.6 | 9.54 | 4.01 | 5.14 |
| Q15717 | ELAVL1# | ELAV-like protein 1 | 5 | 9 | 15 | 16 | 7.14 | 19.31 | 24.84 | 23.5 |
| Q8IY81 | FTSJ3 | Putative rRNA methyltransferase 3 | 5 | 16 | 10 | 8 | 9.87 | 30.25 | 21.22 | 15.88 |
| Q96I24 | FUBP3 | Far upstream element-binding protein 3 | 3 | 3 | 11 | 11 | 4.21 | 6.85 | 21.46 | 20.94 |
| P35637 | FUS# | RNA-binding protein FUS | 4 | - | 3 | 3 | 7.38 | - | 4 | 5.4 |
| P51114 | FXR1# | Fragile X mental retardation syndrome-related protein 1 | 2 | - | 3 | 2 | 4.01 | - | 2.9 | 2.68 |
| Q13823 | GNL2 | Nucleolar GTP-binding protein 2 | - | 3 | 2 | 2 | - | 4.88 | 2.57 | 3.41 |
| Q9BVP2 | GNL3 | Guanine nucleotide-binding protein-like 3 | - | 4 | 4 | 4 | - | 9.03 | 8.98 | 8.39 |
| Q9BZE4 | GTPBP4 | Nucleolar GTP-binding protein 1 | 5 | 9 | 14 | 14 | 9.64 | 18.5 | 26.81 | 25.87 |
| P07305 | H1F0# | Histone H1.0 | - | 3 | 5 | 3 | - | 4.23 | 6.25 | 4.09 |
| Q92522 | H1FX | Histone H1x | - | 2 | 2 | 2 | - | 4.36 | 3.34 | 2.82 |
| Q9BTM1 | H2AFJ | Histone H2A.J | - | 2 | 2 | - | - | 2.8 | 4.4 | - |
| Q71UI9 | H2AFV | Histone H2A.V | - | 2 | 5 | 4 | - | 3.35 | 6.96 | 6.22 |
| O75367 | H2AFY | Core histone macro-H2A.1 | 2 | 3 | 5 | 8 | 5.1 | 4.84 | 9.07 | 14 |
| Q9H583 | HEATR1 | HEAT repeat-containing protein 1 | 7 | 14 | 14 | 9 | 12.41 | 27.31 | 26.58 | 18.09 |
| P16401 | HIST1H1B# | Histone H1.5 | 4 | 6 | 10 | 8 | 6.05 | 9.63 | 17.07 | 14 |
| Q93077 | HIST1H2AC | Histone H2A type 1-C | - | 4 | 2 | - | - | 6.87 | 4.27 | - |
| Q8IUE6 | HIST2H2AB | Histone H2A type 2-B | - | 4 | 11 | 11 | - | 8.13 | 10.21 | 12.76 |
| Q13151 | HNRNPA0 | Heterogeneous nuclear ribonucleoprotein A | 4 | 3 | 5 | 5 | 5.07 | 4.36 | 9.15 | 8.94 |
| P22626 | HNRNPA2B1 | Heterogeneous nuclear ribonucleoproteins A2/B1 | 3 | 3 | 11 | 15 | 6.66 | 5.21 | 18.47 | 23.97 |
| P51991 | HNRNPA3# | Heterogeneous nuclear ribonucleoprotein A3 | 4 | - | 8 | 37 | 8.06 | - | 17.75 | 58.02 |
| Q99729 | HNRNPAB# | Heterogeneous nuclear ribonucleoprotein A/B | 3 | - | 3 | 3 | 3.2 | - | 3.05 | 2.57 |
| Q14103 | HNRNPD | Heterogeneous nuclear ribonucleoprotein D0 | - | 2 | 3 | 4 | - | 4 | 5.38 | 7.08 |
| P52597 | HNRNPF# | Heterogeneous nuclear ribonucleoprotein F | 3 | - | 16 | 14 | 5.21 | - | 18.27 | 19.02 |
| P31943 | HNRNPH1 | Heterogeneous nuclear ribonucleoprotein H | 6 | 4 | - | - | 7.82 | 6.59 | - | - |
| P55795 | HNRNPH2# | Heterogeneous nuclear ribonucleoprotein H2 | 2 | - | 10 | - | 4 | - | 12.4 | - |
| P31942 | HNRNPH3 | Heterogeneous nuclear ribonucleoprotein H3 | 2 | - | 3 | 2 | 2.6 | - | 5.07 | 4.08 |
| P14866 | HNRNPL# | Heterogeneous nuclear ribonucleoprotein L | - | 30 | 33 | 36 | - | 32.72 | 28.44 | 32.15 |
| P52272 | HNRNPM# | Heterogeneous nuclear ribonucleoprotein M | 12 | 6 | 26 | 26 | 16.03 | 11.55 | 37.44 | 32.3 |
| O43390 | HNRNPR# | Heterogeneous nuclear ribonucleoprotein R | 13 | 12 | 27 | 27 | 19.81 | 21.8 | 43.09 | 37.93 |
| Q9BUJ2 | HNRNPUL1# | Heterogeneous nuclear ribonucleoprotein U-like protein 1 | 7 | 5 | 10 | 9 | 13.72 | 10.45 | 13.58 | 17.97 |
| Q1KMD3 | HNRNPUL2# | Heterogeneous nuclear ribonucleoprotein U-like protein 2 | 5 | 7 | 19 | 20 | 8.38 | 12.28 | 26.64 | 24.43 |
| Q5SSJ5 | HP1BP3# | Heterochromatin protein 1-binding protein 3 | 2 | 6 | 13 | 15 | 3.12 | 10.71 | 25.87 | 27.64 |
| Q58FF8 | HSP90AB2P | Putative heat shock protein HSP 90-beta 2 | 6 | - | - | 2 | 8.86 | - | - | 3.89 |
| P08107 | HSPA1A# | Heat shock 70 kDa protein 1A/1B | 3 | - | 5 | 5 | 6.23 | - | 10.38 | 9.08 |
| P54652 | HSPA2# | Heat shock-related 70 kDa protein 2 | 3 | - | 7 | 9 | 5.8 | - | 13.36 | 17.7 |
| P04792 | HSPB1# | Heat shock protein beta-1 | 2 | 2 | 2 | 2 | 4.25 | 2.77 | 4 | 4 |
| Q9NZI8 | IGF2BP1 | Insulin-like growth factor 2 mRNA-binding protein 1 | 4 | 5 | 12 | 15 | 57 | 7.42 | 20.61 | 15.59 |
| O00425 | IGF2BP3 | Insulin-like growth factor 2 mRNA-binding protein 3 | 3 | 3 | 9 | 8 | 6.03 | 5.03 | 17.24 | 15.92 |
| Q12905 | ILF2# | Interleukin enhancer-binding factor 2 | 9 | - | 18 | 27 | 18.29 | - | 26.79 | 40.47 |
| Q12906 | ILF3# | Interleukin enhancer-binding factor 3 | 23 | 19 | 48 | 53 | 42.2 | 35.75 | 60.93 | 74.73 |
| Q96G21 | IMP4 | U3 small nucleolar ribonucleoprotein protein IMP4 | - | 3 | 2 | 4 | - | 6.11 | 2.35 | 7.13 |
| Q07666 | KHDRBS1 | KH domain-containing, RNA-binding, signal transduction-associated protein 1 | 3 | - | 5 | 4 | 4.51 | - | 7.85 | 6.39 |
| P48668 | KRT6C | Keratin, type II cytoskeletal 6C | - | 6 | - | 5 | - | 10.19 | - | 8.84 |
| P83111 | LACTB | Serine beta-lactamase-like protein LACTB, mitochondrial | 2 | 3 | 2 | - | 3.56 | 4.77 | 2.4 | - |
| P02545 | LMNA | Lamin-A/C | 3 | - | - | 5 | 4.97 | - | - | 9.82 |
| Q9BXY0 | MAK16 | Protein MAK16 homolog | - | 4 | 5 | 4 | - | 6.29 | 9.39 | 8.01 |
| P43243 | MATR3# | Matrin-3 | 4 | 15 | 22 | 25 | 7.12 | 29.4 | 36.94 | 40.98 |
| Q9BYG3 | MKI67IP | MKI67 FHA domain-interacting nucleolar phosphoprotein | 7 | 5 | 6 | 5 | 12.02 | 7.21 | 11.84 | 7.51 |
| O00566 | MPHOSPH10 | U3 small nucleolar ribonucleoprotein protein MPP10 | - | 6 | 4 | 5 | - | 11.27 | 3.88 | 10.01 |
| Q9BQG0 | MYBBP1A# | Myb-binding protein 1A | 5 | 11 | 5 | 14 | 10.67 | 20.55 | 11.75 | 27.46 |
| O00159 | MYO1C# | Myosin-Ic | - | 2 | 3 | 4 | - | 4.43 | 5.67 | 7.7 |
| Q9H0A0 | NAT10# | N-acetyltransferase 10 | 4 | 3 | 4 | 7 | 8.49 | 6.54 | 7.26 | 13.43 |
| P19338 | NCL | Nucleolin | 5 | - | 7 | 9 | 8.15 | - | 12.97 | 18.66 |
| Q9Y221 | NIP7 | 60S ribosome subunit biogenesis protein NIP7 homolog | - | 2 | 4 | - | - | 3.74 | 5.8 | - |
| O15226 | NKRF# | NF-kappa-B-repressing factor | - | 8 | 11 | 12 | - | 12.94 | 22.84 | 22.1 |
| Q9H8H0 | NOL11 | Nucleolar protein 11 | - | 9 | 7 | 6 | - | 15.16 | 12.65 | 12.92 |
| Q9H6R4 | NOL6 | Nucleolar protein 6 | 2 | 5 | 7 | 8 | 2.27 | 7.92 | 13.17 | 13.42 |
| Q9UMY1 | NOL7 | Nucleolar protein 7 | - | 5 | 3 | 3 | - | 8.47 | 6 | 5.82 |
| Q9Y3C1 | NOP16 | Nucleolar protein 16 | 2 | 3 | 3 | 4 | 4 | 6.58 | 3.74 | 8.07 |
| P46087 | NOP2 | Putative ribosomal RNA methyltransferase NOP2 | 11 | 12 | 22 | 23 | 19.36 | 25.03 | 37.05 | 40.73 |
| O00567 | NOP56# | Nucleolar protein 56 | 13 | 15 | 24 | 27 | 23.87 | 22.4 | 43.65 | 44.13 |
| Q9Y2X3 | NOP58# | Nucleolar protein 58 | 9 | - | 17 | 19 | 12.35 | - | 29.8 | 31.99 |
| Q14980 | NUMA1 | Nuclear mitotic apparatus protein 1 | 6 | 4 | 9 | 12 | 11.12 | 8.69 | 17.94 | 23.27 |
| Q13310 | PABPC4# | Polyadenylate-binding protein 4 | 4 | 6 | 7 | 10 | 7.81 | 6.46 | 14.6 | 15.16 |
| Q9NWT1 | PAK1IP1 | p21-activated protein kinase-interacting protein 1 | 3 | 2 | 2 | 3 | 6 | 3.47 | 2.92 | 5.89 |
| Q15365 | PCBP1# | Poly(rC)-binding protein 1 | 2 | - | 2 | - | 3.8 | - | 4.01 | - |
| Q14690 | PDCD11 | Protein RRP5 homolog | 13 | 19 | 27 | 28 | 24.61 | 37.82 | 56.56 | 54.56 |
| O00541 | PES1 | Pescadillo homolog | 3 | 4 | 7 | 5 | 6.72 | 9.09 | 12.5 | 9.42 |
| Q96HS1 | PGAM5 | Serine/threonine-protein phosphatase PGAM5, mitochondrial | 3 | - | 4 | 5 | 4.34 | - | 7.22 | 8.01 |
| P78527 | PRKDC | DNA-dependent protein kinase catalytic subunit | 2 | - | 3 | - | 3.27 | - | 3.73 | - |
| Q9UMS4 | PRPF19# | Pre-mRNA-processing factor 19 | - | 4 | 10 | 10 | - | 8.67 | 16.95 | 18.41 |
| Q13523 | PRPF4B | Serine/threonine-protein kinase PRP4 homolog | 2 | - | 8 | 5 | 3.42 | - | 15.43 | 10.27 |
| P26599 | PTBP1# | Polypyrimidine tract-binding protein 1 | 10 | 9 | 14 | 15 | 14.9 | 18.56 | 24.28 | 23.42 |
| Q15269 | PWP2 | Periodic tryptophan protein 2 homolog | 2 | 6 | 6 | 6 | 2.73 | 11.3 | 11.52 | 11.13 |
| Q9P0K7 | RAI14 | Ankycorbin | 7 | - | - | 2 | 11.95 | - | - | 4 |
| Q9UKM9 | RALY# | RNA-binding protein Raly | 3 | 11 | 20 | 23 | 6.36 | 17.52 | 26.01 | 28.69 |
| Q96PK6 | RBM14 | RNA-binding protein 14 | - | 5 | 7 | 9 | - | 8.46 | 13.26 | 17.49 |
| Q96T37 | RBM15 | Putative RNA-binding protein 15 | 3 | - | 6 | 8 | 3.57 | - | 12.18 | 15.66 |
| P49756 | RBM25 | RNA-binding protein 25 | 2 | - | - | 5 | 3.6 | - | - | 9.28 |
| Q9NW13 | RBM28 | RNA-binding protein 28 | 2 | 6 | 10 | 8 | 4.01 | 12.02 | 18.29 | 14.38 |
| P42696 | RBM34 | RNA-binding protein 34 | - | 4 | 2 | 3 | - | 8 | 4.45 | 6.33 |
| Q9BWF3 | RBM4 | RNA-binding protein 4 | - | 2 | 4 | 3 | - | 4.22 | 7.82 | 4.92 |
| P38159 | RBMX# | Heterogeneous nuclear ribonucleoprotein G | 5 | 15 | 12 | - | 7.4 | 25.26 | 21.86 | |
| O95758 | ROD1 | Regulator of differentiation 1 | - | 2 | 5 | 4 | - | 4.42 | 8.43 | 6.59 |
| Q9H9Y2 | RPF1 | Ribosome production factor 1 | - | 4 | 2 | 3 | - | 7.3 | 3.59 | 6 |
| Q9H7B2 | RPF2 | Ribosome production factor 2 homolog | 3 | 7 | 8 | 8 | 6.15 | 12.33 | 13.61 | 14 |
| P62913 | RPL11# | 60S ribosomal protein L11 | 2 | - | 2 | 4 | 3.16 | - | 4 | 7.32 |
| P30050 | RPL12 | 60S ribosomal protein L12 | 2 | - | 2 | 2 | 4 | - | 4 | 2.79 |
| P26373 | RPL13# | 60S ribosomal protein L13 | 2 | - | - | 4 | 3.46 | - | - | 8.4 |
| P50914 | RPL14# | 60S ribosomal protein L14 | 2 | - | 2 | 3 | 3.59 | - | 4.01 | 3.06 |
| P61313 | RPL15# | 60S ribosomal protein L15 | 2 | - | 3 | 3 | 4.03 | - | 4.97 | 6.04 |
| P62829 | RPL23# | 60S ribosomal protein L23 | 3 | - | 5 | 3 | 2 | - | 8.8 | 4.02 |
| P62750 | RPL23A# | 60S ribosomal protein L23a | 2 | - | 2 | - | 3.57 | - | 4.39 | - |
| P61353 | RPL27 | 60S ribosomal protein L27 | 2 | - | 6 | 5 | 4 | - | 10.64 | 8.68 |
| P39023 | RPL3# | 60S ribosomal protein L3 | 6 | - | 9 | 8 | 9.09 | - | 18.72 | 15.38 |
| Q9Y3U8 | RPL36# | 60S ribosomal protein L36 | 2 | - | 2 | 3 | 3.54 | - | 4.38 | 4.94 |
| P46777 | RPL5 | 60S ribosomal protein L5 | - | 4 | 3 | - | - | 6.66 | 5.48 | - |
| Q02878 | RPL6# | 60S ribosomal protein L6 | 3 | 7 | 11 | 13 | 6.41 | 14.44 | 17.38 | 21.66 |
| P18124 | RPL7# | 60S ribosomal protein L7 | 4 | - | 3 | 6 | - | 5.31 | 6.6 | 11.46 |
| P62424 | RPL7A# | 60S ribosomal protein L7a | 4 | - | 8 | 11 | 7.11 | - | 15.34 | 18.93 |
| Q6DKI1 | RPL7L1 | 60S ribosomal protein L7-like 1 | - | 3 | 3 | 4 | - | 6.44 | 5.02 | 8.34 |
| P62263 | RPS14# | 40S ribosomal protein S14 | 3 | - | - | 2 | 6 | - | - | 2.26 |
| P23396 | RPS3# | 40S ribosomal protein S3 | 3 | - | 2 | 4 | 4.51 | - | 2.55 | 7.39 |
| P62241 | RPS8# | 40S ribosomal protein S8 | 2 | - | 3 | 4 | 4 | - | 4.82 | 8 |
| Q9P2E9 | RRBP1 | Ribosome-binding protein 1 | 7 | - | 18 | 19 | 15.03 | - | 36.88 | 38.71 |
| P56182 | RRP1 | Ribosomal RNA processing protein 1 homolog A | 2 | 3 | 6 | 4 | 4.19 | 6.66 | 9.28 | 8.38 |
| Q5JTH9 | RRP12 | RRP12-like protein | 2 | 3 | 4 | 2 | 2.49 | 6.02 | 5.4 | 3.86 |
| Q14684 | RRP1B | Ribosomal RNA processing protein 1 homolog B | 4 | 4 | 11 | 12 | 7.29 | 7.06 | 18.98 | 20.86 |
| Q9Y3A4 | RRP7A | Ribosomal RNA-processing protein 7 homolog A | - | 2 | - | 3 | - | 4.13 | - | 4.17 |
| O43159 | RRP8 | Ribosomal RNA-processing protein 8 | - | 2 | 4 | 3 | - | 3.32 | 8.02 | 6 |
| O43818 | RRP9 | U3 small nucleolar RNA-interacting protein 2 | 3 | 4 | 5 | 4 | 6 | 7.01 | 11.04 | 8.16 |
| Q15050 | RRS1 | Ribosome biogenesis regulatory protein homolog | 4 | 11 | 7 | 9 | 7.01 | 18.42 | 9.17 | 17.81 |
| O76021 | RSL1D1 | Ribosomal L1 domain-containing protein 1 | 9 | - | 20 | 17 | 17.26 | - | 37.44 | 34.24 |
| P60903 | S100A10 | Protein S100-A10 | 3 | - | - | 2 | 6 | - | - | 2.49 |
| Q14151 | SAFB2# | Scaffold attachment factor B2 | 6 | 2 | 10 | 12 | 12.26 | 2.64 | 20.4 | 22.78 |
| Q9H7N4 | SCAF1 | Splicing factor, arginine/serine-rich 19 | - | 2 | 3 | - | - | 4 | 5.92 | - |
| Q13435 | SF3B2 | Splicing factor 3B subunit 2 | - | 3 | 9 | 8 | - | 3.83 | 17.43 | 15.05 |
| Q15393 | SF3B3 | Splicing factor 3B subunit 3 | 3 | - | 7 | 7 | 5.85 | - | 13.05 | 9.58 |
| Q08170 | SFRS4 | Splicing factor, arginine/serine-rich 4 | 3 | - | - | 2 | 4 | - | - | 3.59 |
| Q13247 | SFRS6 | Splicing factor, arginine/serine-rich 6 | 3 | - | 3 | 3 | 4 | - | 5.48 | 3.59 |
| Q16629 | SFRS7 | Splicing factor, arginine/serine-rich 7 | 2 | - | 3 | - | 3.28 | - | 5.57 | - |
| Q9NWH9 | SLTM | SAFB-like transcription modulator | 3 | 3 | 11 | 10 | 5.82 | 3.09 | 20.01 | 18.56 |
| O60264 | SMARCA5 | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 | 2 | - | 3 | 3 | 3.35 | - | 6.1 | 5.79 |
| O75643 | SNRNP200 | U5 small nuclear ribonucleoprotein 200 kDa helicase | - | 10 | 23 | 20 | - | 19.16 | 48.66 | 37.55 |
| P08621 | SNRNP70 | U1 small nuclear ribonucleoprotein 70 kDa | 2 | - | 6 | 7 | 3.25 | - | 11.58 | 13.86 |
| P18583-5 | SON# | Isoform D of Protein SON | - | 5 | 4 | 3 | - | 9.39 | 8.16 | 6.11 |
| Q13501 | SQSTM1 | Sequestosome-1 | 2 | - | 2 | 6 | 3.89 | - | 4.37 | 10 |
| Q8IYB3 | SRRM1 | Serine/arginine repetitive matrix protein 1 | - | 2 | 2 | 4 | - | 2.67 | 4.12 | 7.68 |
| O95793 | STAU1# | Double-stranded RNA-binding protein Staufen homolog 1 | - | 2 | 5 | 8 | - | 3.7 | 9.86 | 16.03 |
| O75683 | SURF6 | Surfeit locus protein 6 | 2 | - | - | 3 | 3.1 | - | - | 6.24 |
| O60506 | SYNCRIP# | Heterogeneous nuclear ribonucleoprotein Q | 8 | - | 11 | 10 | 14.4 | - | 17.7 | 19.41 |
| Q92804 | TAF15 | TATA-binding protein-associated factor 2N | 2 | - | - | 3 | 3.06 | - | - | 4 |
| Q13148 | TARDBP | TAR DNA-binding protein 43 | 2 | - | 3 | 2 | 4 | - | 5.96 | 3.26 |
| Q12788 | TBL3 | Transducin beta-like protein 3 | 3 | 8 | 5 | 9 | 6.01 | 16.72 | 10.19 | 18.46 |
| Q13428 | TCOF1 | Treacle protein | 3 | - | 6 | 7 | 6 | - | 12.4 | 13.24 |
| Q9NXF1 | TEX10 | Testis-expressed sequence 10 protein | - | 4 | 3 | 3 | - | 6.75 | 4.79 | 6 |
| P42166 | TMPO | Lamina-associated polypeptide 2, isoform alpha | 2 | - | - | 6 | 4 | - | - | 12.22 |
| Q13595 | TRA2A | Transformer-2 protein homolog alpha | 2 | - | 5 | 6 | 2.04 | - | 8.55 | 9.86 |
| Q14258 | TRIM25# | E3 ubiquitin/ISG15 ligase TRIM25 | 2 | - | 2 | 2 | 2.66 | - | 2.62 | 4 |
| Q14669 | TRIP12 | Probable E3 ubiquitin-protein ligase TRIP12 | - | 2 | 10 | 7 | - | 4.39 | 21.08 | 12.7 |
| Q9BVJ6 | UTP14A | U3 small nucleolar RNA-associated protein 14 homolog A | - | 3 | 6 | 5 | - | 6.01 | 11.86 | 8.75 |
| Q8TED0 | UTP15 | U3 small nucleolar RNA-associated protein 15 homolog | 3 | 8 | 6 | 9 | 4.65 | 13.57 | 6.6 | 17.59 |
| Q9Y5J1 | UTP18 | U3 small nucleolar RNA-associated protein 18 homolog | 4 | 12 | 10 | 5 | 8.99 | 17.71 | 15.09 | 10.54 |
| Q9NYH9 | UTP6 | U3 small nucleolar RNA-associated protein 6 homolog | 3 | 3 | 7 | 7 | 6 | 6.4 | 13.09 | 12.81 |
| P21796 | VDAC1 | Voltage-dependent anion-selective channel protein 1 | 2 | - | - | 2 | 2.46 | - | - | 4.01 |
| Q9Y277 | VDAC3 | Voltage-dependent anion-selective channel protein 3 | 2 | - | 3 | 2 | 3.02 | - | 6.01 | 4.12 |
| Q9GZL7 | WDR12 | Ribosome biogenesis protein WDR12 | 2 | 7 | 3 | 2 | 3.38 | 14.26 | 4.27 | 4.26 |
| Q9UNX4 | WDR3 | WD repeat-containing protein 3 | 3 | 10 | 9 | 8 | 6 | 20.94 | 18.64 | 15.57 |
| Q8NI36 | WDR36 | WD repeat-containing protein 36 | - | 8 | 12 | 17 | - | 19.87 | 25.61 | 33.36 |
| Q15061 | WDR43 | WD repeat-containing protein 43 | - | 4 | 6 | 7 | - | 6.52 | 8.33 | 12 |
| O15213 | WDR46 | WD repeat-containing protein 46 | 3 | 6 | 9 | 9 | 6 | 10.03 | 16.71 | 17.11 |
| Q6RFH5 | WDR74 | WD repeat-containing protein 74 | 2 | 6 | - | - | 4.14 | 10.11 | - | - |
| Q8IWA0 | WDR75 | WD repeat-containing protein 75 | 4 | 10 | 5 | 13 | 6.17 | 17.11 | 12.14 | 21.8 |
| Q9H0D6 | XRN2 | 5’-3’ exoribonuclease 2 | 2 | 6 | 14 | 12 | 2.71 | 12.14 | 28.48 | 23.17 |
| Q7Z2W4 | ZC3HAV1 | Zinc finger CCCH-type antiviral protein 1 | 5 | 6 | 8 | 6 | 9.06 | 9.94 | 16.91 | 11.3 |
| Q96KR1 | ZFR | Zinc finger RNA-binding protein | 10 | 20 | 34 | 33 | 15.16 | 35.58 | 42.02 | 48.56 |
| Q5BKZ1 | ZNF326 | Zinc finger protein 326 | 3 | - | 5 | 7 | 6 | - | 8.37 | 10.49 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rahim, M.N.; Klewes, L.; Zahedi-Amiri, A.; Mai, S.; Coombs, K.M. Global Interactomics Connect Nuclear Mitotic Apparatus Protein NUMA1 to Influenza Virus Maturation. Viruses 2018, 10, 731. https://doi.org/10.3390/v10120731
Rahim MN, Klewes L, Zahedi-Amiri A, Mai S, Coombs KM. Global Interactomics Connect Nuclear Mitotic Apparatus Protein NUMA1 to Influenza Virus Maturation. Viruses. 2018; 10(12):731. https://doi.org/10.3390/v10120731
Chicago/Turabian StyleRahim, Md Niaz, Ludger Klewes, Ali Zahedi-Amiri, Sabine Mai, and Kevin M. Coombs. 2018. "Global Interactomics Connect Nuclear Mitotic Apparatus Protein NUMA1 to Influenza Virus Maturation" Viruses 10, no. 12: 731. https://doi.org/10.3390/v10120731
APA StyleRahim, M. N., Klewes, L., Zahedi-Amiri, A., Mai, S., & Coombs, K. M. (2018). Global Interactomics Connect Nuclear Mitotic Apparatus Protein NUMA1 to Influenza Virus Maturation. Viruses, 10(12), 731. https://doi.org/10.3390/v10120731





