Complete Nucleotide Sequence of a Partitivirus from Rhizoctonia solani AG-1 IA Strain C24
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Strain
2.2. Isolation and Sequencing of Mycovirus dsRNA
2.3. Data Analysis
3. Results
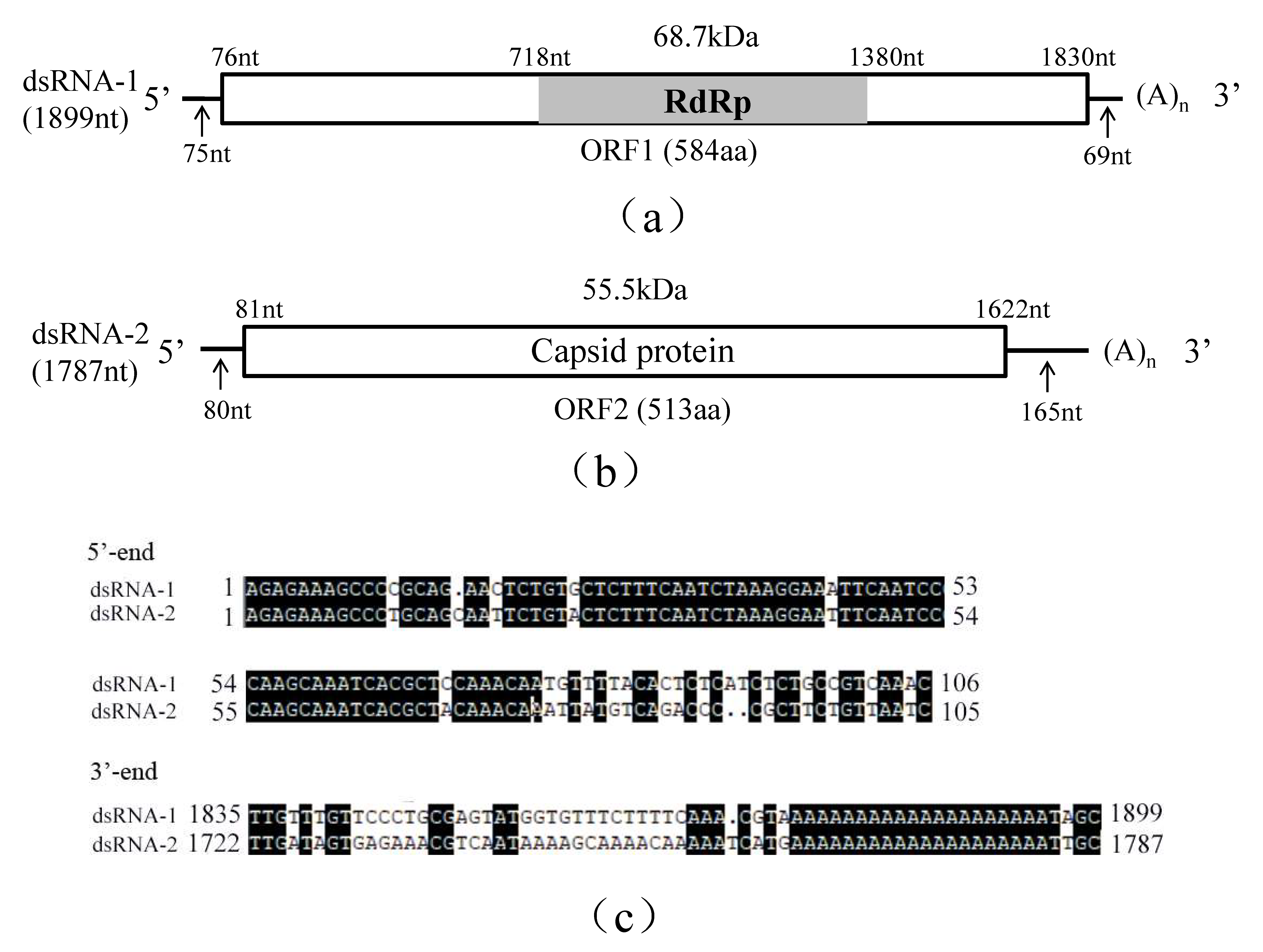
3.1. Genomic Structure Analysis
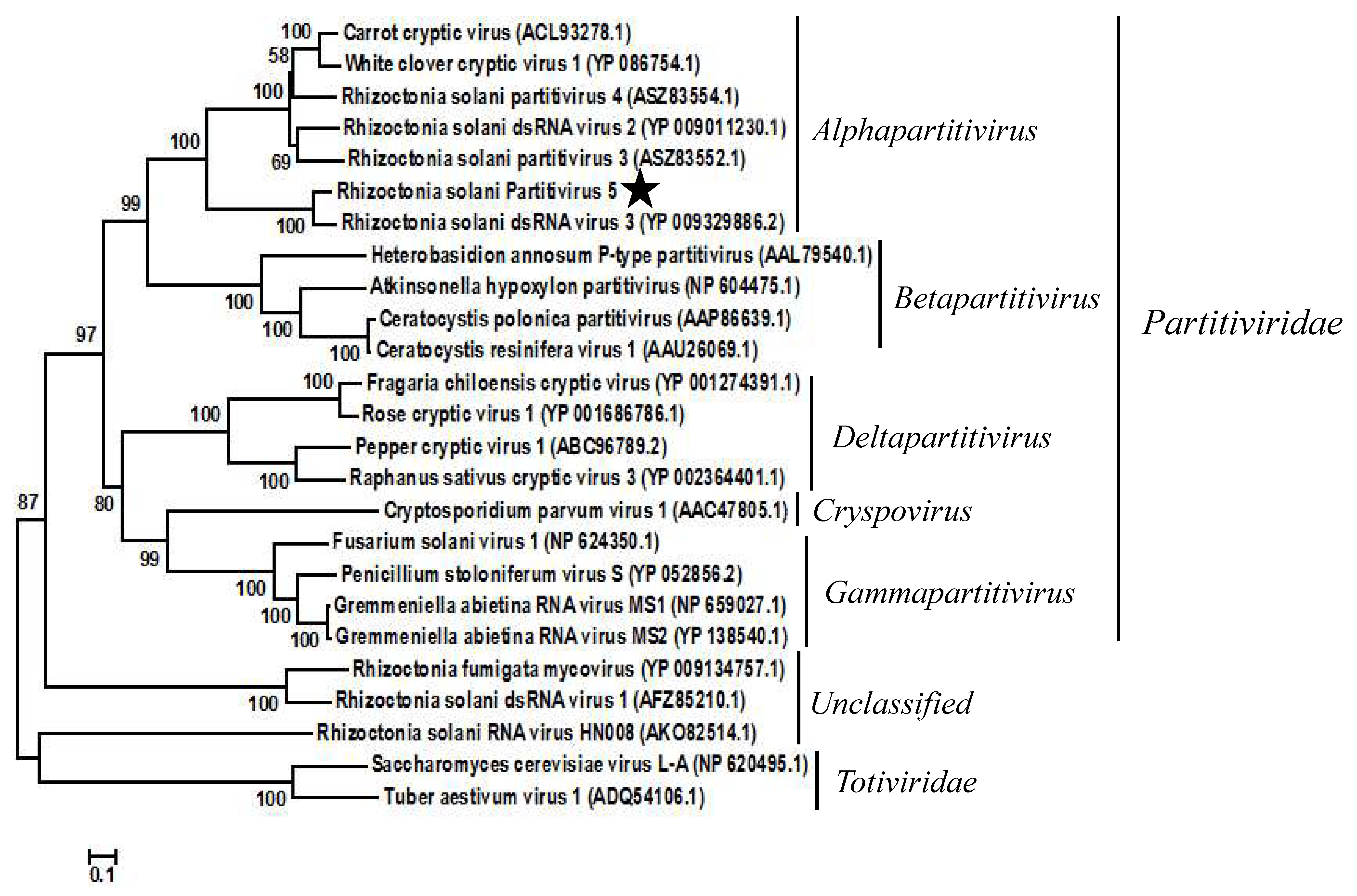
3.2. Phylogenetic Analysis
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Pearson, M.N.; Beever, R.E.; Boine, B.; Arthur, K. Mycoviruses of filamentous fungi and their relevance to plant pathology. Mol. Plant Path. 2009, 10, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Herrero, N.; Márquez, S.S.; Zabalgogeazcoa, I. Mycoviruses are common among different species of endophytic fungi of grasses. Arch. Virol. 2009, 154, 327–330. [Google Scholar] [CrossRef] [PubMed]
- Nuss, D.L. Hypovirulence: mycoviruses at the fungal-plant interface. Nat. Rev. Microbiol. 2005, 3, 632–642. [Google Scholar] [CrossRef] [PubMed]
- Grente, J.; Berthelay-Sauret, S. Biological control of chestnut blight in France. In Proceedings of the American Chestnut Symposium, Morgantown, WV, USA, 4–5 January 1978; pp. 30–34. [Google Scholar]
- Sasaki, A.; Miyanishi, M.; Ozaki, K.; Onoue, M.; Yoshida, K. Molecular characterization of a partitivirus from the plant pathogenic ascomycete Rosellinia necatrix. Arch. Virol. 2005, 150, 1069–1083. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Zhang, L.; Li, G.; Jiang, D.; Ghabrial, S.A. Genome characterization of a debilitation-associated mitovirus infecting the phytopathogenic fungus Botrytis cinerea. Virology 2010, 406, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Liu, H.; Zhang, M.; Cao, X.; Zhou, E. The complete genomic sequence of a novel mycovirus from Rhizoctonia solani AG-1 IA strain B275. Arch. Virol. 2013, 158, 1609–1612. [Google Scholar] [CrossRef] [PubMed]
- Ghabrial, S.A.; Castón, J.R.; Jiang, D.; Nibert, M.L.; Suzuki, N. 50-plus years of fungal viruses. Virology 2015, 479, 356–368. [Google Scholar] [CrossRef] [PubMed]
- Puchades, A.V.; Carpino, C.; Alfaro-Fernandez, A.; Font-San-Ambrosio, M.I.; Davin, S.; Guerri, J.; Rubio, L.; Galipienso, L. Detection of Southern tomato virus by molecular hybridisation. Ann. Appl. Biol. 2017, 171, 172–178. [Google Scholar] [CrossRef]
- Yu, X.; Li, B.; Fu, Y.; Jiang, D.; Ghabrial, S.A.; Li, G.; Peng, Y.; Xie, J.; Cheng, J.; Huang, J.; et al. A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus. Proc. Natl. Acad. Sci. USA 2010, 107, 8387–8392. [Google Scholar] [CrossRef] [PubMed]
- Cubeta, M.A.; Vilgalys, R. Population biology of the Rhizoctonia solani complex. Phytopathology 1997, 87, 480–484. [Google Scholar] [CrossRef]
- Abdoulaye, A.H.; Cheng, J.; Jiang, D.; Xie, J. Complete genome sequence of a novel mitovirus from the phytopathogenic fungus Rhizoctonia oryzae-sativae. Arch. Virol. 2017, 162, 1409–1412. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Zhang, M.; Chen, Q.; Zhu, M.; Zhou, E. A novel mycovirus closely related to viruses in the genus Alphapartitivirus confers hypovirulence in the phytopathogenic fungus Rhizoctonia solani. Virology 2014, 456, 220–226. [Google Scholar] [CrossRef] [PubMed]
- Castanho, B.; Butler, E.E.; Shepherd, R.J. Association of double-stranded-RNA with Rhizoctonia decline. Phytopathology 1978, 68, 1515–1519. [Google Scholar] [CrossRef]
- Bharathan, N.; Tavantzis, S.M. Genetic diversity of double-stranded RNA from Rhizoctonia solani. Phytopathology 1990, 80, 631–635. [Google Scholar] [CrossRef]
- Robinson, H.L.; Deacon, J.W. Double-stranded RNA elements in Rhizoctonia solani AG 3. Mycol. Res. 2002, 106, 12–22. [Google Scholar] [CrossRef]
- Bharathan, N.; Saso, H.; Gudipati, L.; Bharathan, S.; Whited, K.; Anthony, K. Double-stranded RNA: distribution and analysis among isolates of Rhizoctonia solani AG-2 to-13. Plant Pathol. 2005, 54, 196–203. [Google Scholar] [CrossRef]
- Zhang, M.; Zheng, L.; Liu, C.; Shu, C.W.; Zhou, E.X. Characterization of a novel dsRNA mycovirus isolated from strain A105 of Rhizoctonia solani AG-1 IA. Arch. Virol. 2018, 163, 427–430. [Google Scholar] [CrossRef] [PubMed]
- Lyu, R.L.; Zhang, Y.; Tang, Q.; Li, Y.; Cheng, J.; Fu, Y.; Chen, T.; Jiang, D.; Xie, J. Two alphapartitiviruses co-infecting a single isolate of the plant pathogenic fungus Rhizoctonia solani. Arch. Virol. 2018, 163, 515–520. [Google Scholar] [CrossRef]
- Strauss, E.E.; Lakshman, D.K.; Tavantzis, S.M. Molecular characterization of the genome of a partitivirus from the basidiomycete Rhizoctonia solani. J. General Virol. 2000, 81, 549–555. [Google Scholar] [CrossRef]
- Lakshman, D.K.; Jian, J.; Tavantzis, S.M. A double-stranded RNA element from a hypovirulent strain of Rhizoctonia solani occurs in DNA form and is genetically related to the pentafunctional AROM protein of the shikimate pathway. Proc. Natl. Acad. Sci. USA 1998, 95, 6425–6429. [Google Scholar] [CrossRef]
- Li, W.; Zhang, T.; Sun, H.; Deng, Y.; Zhang, A.; Chen, H.; Wang, K. Complete genome sequence of a novel endornavirus in the wheat sharp eyespot pathogen Rhizoctonia cerealis. Arch. Virol. 2014, 159, 1213–1216. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Falloon, R.E.; Stewart, A.; Pitman, A.R. Molecular characterisation of an endornavirus from Rhizoctonia solani AG-3PT infecting potato. Fungal Biol. 2014, 118, 924–934. [Google Scholar] [CrossRef] [PubMed]
- Zhong, J.; Chen, C.Y.; Gao, B.D. Genome sequence of a novel mycovirus of Rhizoctonia solani, a plant pathogenic fungus. Virus Genes 2015, 51, 167–170. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xu, P.; Zhang, L.; Xia, Z.; Qin, X.; Yang, G.; Mo, X. Molecular characterization of a novel mycovirus from Rhizoctonia fumigata AG-Ba isolate C-314 Baishi. Arch. Virol. 2015, 160, 2371–2374. [Google Scholar] [CrossRef] [PubMed]
- Morris, T.; Dodds, J. Isolation and analysis of double-stranded RNA from virus-infected plant and fungal tissue. Phytopathology 1979, 69, 854–858. [Google Scholar] [CrossRef]
- Potgieter, A.C.; Page, N.A.; Liebenberg, J.; Wright, I.M.; Landt, O.; van Dijk, A.A. Improved strategies for sequence-independent amplification and sequencing of viral dsRNA genomes. J. General Virol. 2009, 90, 1423–1432. [Google Scholar] [CrossRef] [PubMed]
- Lim, W.S.; Jeong, J.H.; Jeong, R.D.; Yoo, Y.B.; Yie, S.W.; Kim, K.H. Complete nucleotide sequence and genome organization of a dsRNA partitivirus infecting Pleurotus ostreatus. Virus Res. 2005, 108, 111–119. [Google Scholar] [CrossRef]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, C.; Zeng, M.; Zhang, M.; Shu, C.; Zhou, E. Complete Nucleotide Sequence of a Partitivirus from Rhizoctonia solani AG-1 IA Strain C24. Viruses 2018, 10, 703. https://doi.org/10.3390/v10120703
Liu C, Zeng M, Zhang M, Shu C, Zhou E. Complete Nucleotide Sequence of a Partitivirus from Rhizoctonia solani AG-1 IA Strain C24. Viruses. 2018; 10(12):703. https://doi.org/10.3390/v10120703
Chicago/Turabian StyleLiu, Chen, Miaolin Zeng, Meiling Zhang, Canwei Shu, and Erxun Zhou. 2018. "Complete Nucleotide Sequence of a Partitivirus from Rhizoctonia solani AG-1 IA Strain C24" Viruses 10, no. 12: 703. https://doi.org/10.3390/v10120703
APA StyleLiu, C., Zeng, M., Zhang, M., Shu, C., & Zhou, E. (2018). Complete Nucleotide Sequence of a Partitivirus from Rhizoctonia solani AG-1 IA Strain C24. Viruses, 10(12), 703. https://doi.org/10.3390/v10120703




