Description, Distribution, and Relevance of Viruses of the Forest Pathogen Gremmeniella abietina
Abstract
1. Taxonomy of G. abietina and Relevance in Forestry
2. Occurrence of Viruses in G. abietina: Description of Their Genome and Structure, and Phylogenetic Relationships
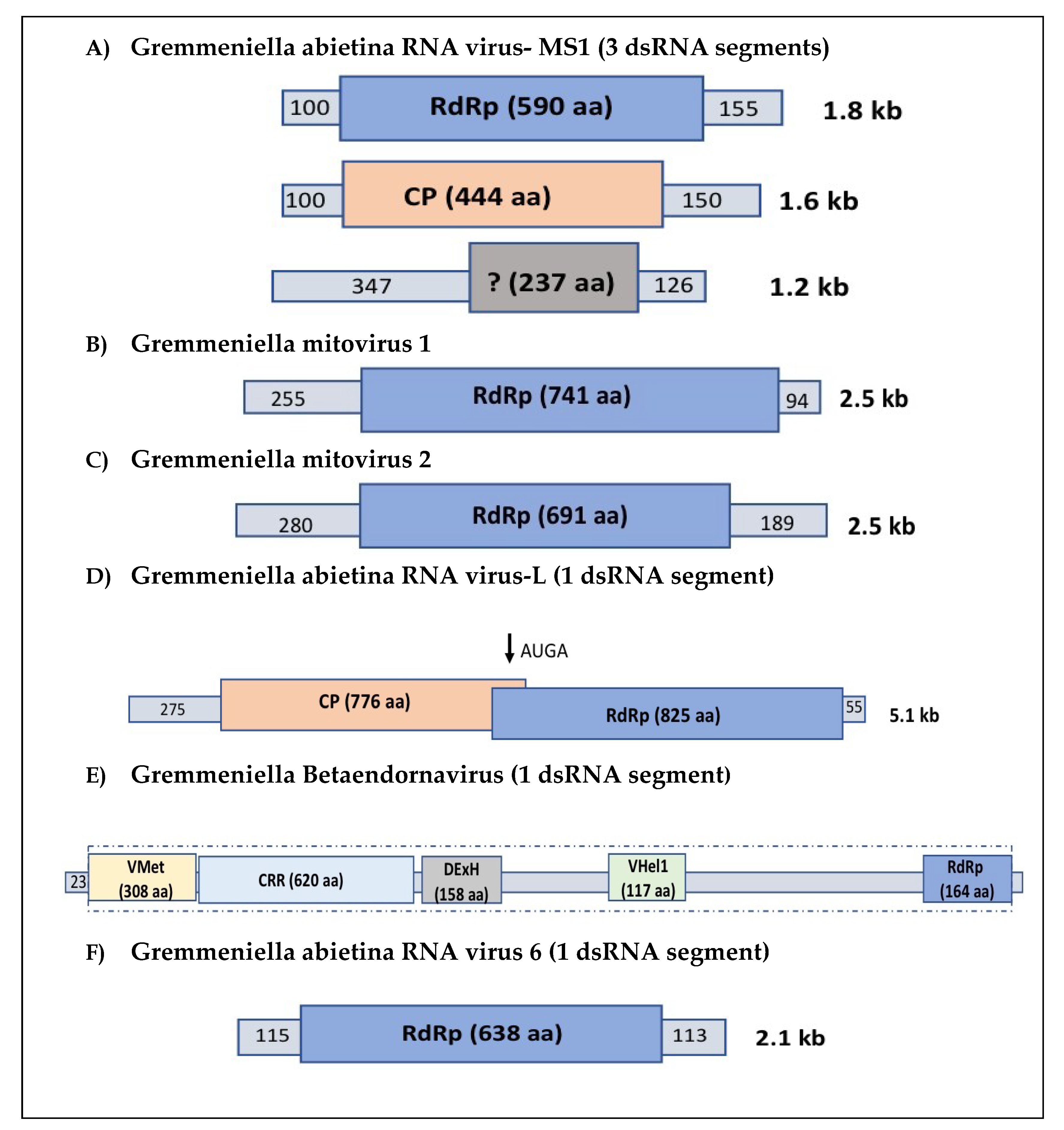
2.1. Partitiviruses
2.2. Narnaviruses
2.3. Totiviruses
2.4. Endornaviruses
2.5. Gremmeniella abietina RNA Virus 6
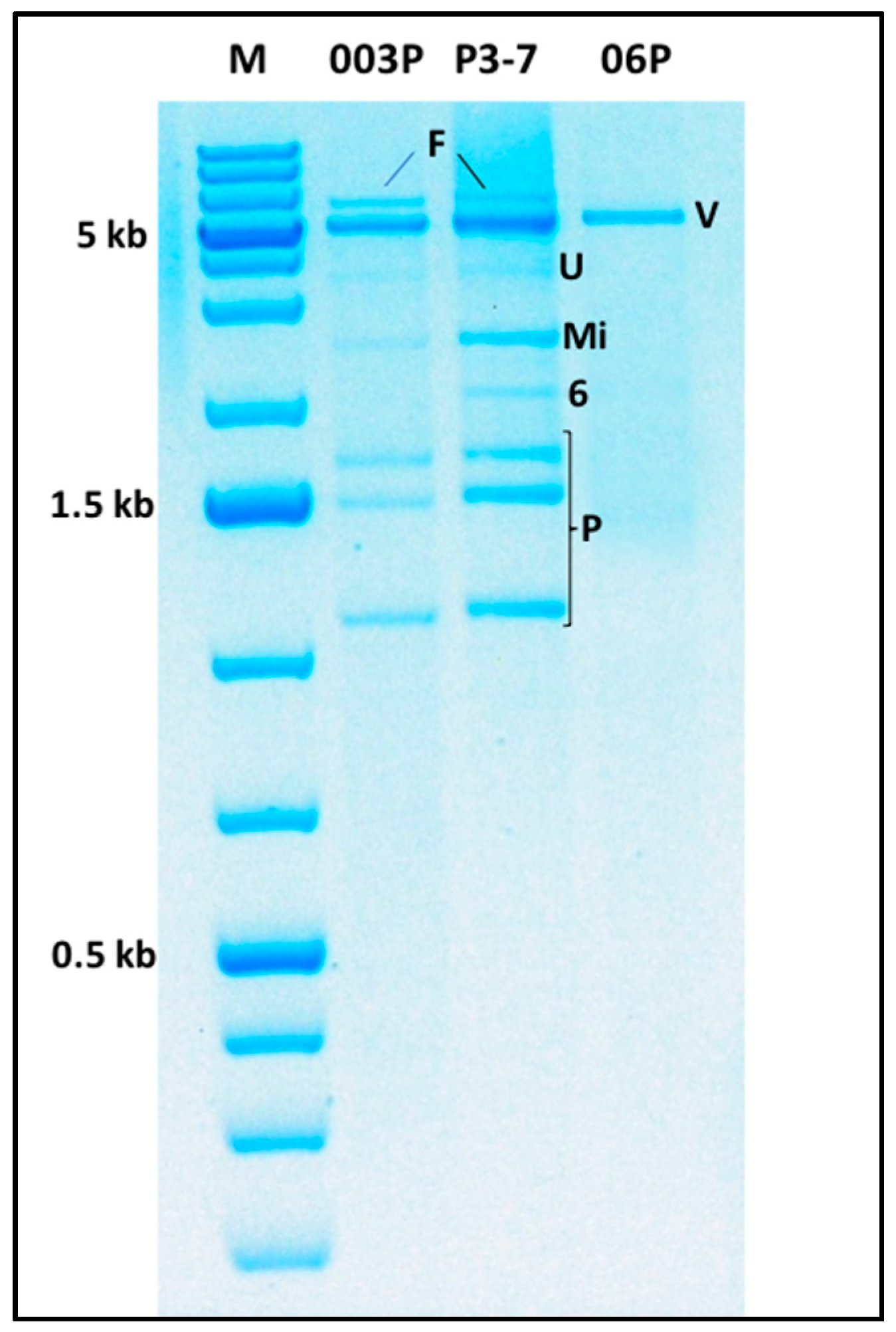
2.6. Multiple Virus Infections and Evidence of the Existence of Novel Viruses
2.6.1. The ca. 6 kb dsRNA Band
2.6.2. The ca. 5 kb dsRNA Band
3. Mycoviruses and Their Clarifying Message on the Complex Diversity of G. abietina
4. Consequences of Viral Infections on G. abietina
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Donaubauer, E. Distribution and hosts of Scleroderris lagerbergii in Europe and North America. For. Pathol. 1972, 2, 6–11. [Google Scholar] [CrossRef]
- Yokota, S.; Uozumi, T.; Matsuzaki, S. Scleroderris canker of Todo-Fir in Hokkaido, Northern Japan I. Present status of damage, and features of infected plantations. For. Pathol. 1974, 4, 65–74. [Google Scholar] [CrossRef]
- Morelet, M. La maladie à Brunchorstia. Eur. J. For. Pathol. 1980, 10, 268–277. [Google Scholar] [CrossRef]
- Barklund, P.; Rowe, J. Gremmeniella abietina (Scleroderris lagerbergii), a primary parasite in a Norway spruce die-back. For. Pathol. 1981, 11, 97–108. [Google Scholar] [CrossRef]
- Kaitera, J.; Seitamäki, L.; Jalkanen, R. Morphological and ecological variation of Gremmeniella abietina var. abietina in Pinus sylvestris, Pinus contorta and Picea abies sapling stands in northern Finland and the Kola Peninsula. Scand. J. For. Res. 2000, 15, 13–19. [Google Scholar] [CrossRef]
- Santamaria, O.; Alves-Santos, F.M.; Diez, J.J. Genetic characterization of Gremmeniella abietina var. abietina isolates from Spain. Plant Pathol. 2005, 54, 331–338. [Google Scholar] [CrossRef]
- Petrini, O.; Toti, L.; Petrini, L.E.; Heiniger, U. Gremmeniella abietina and G. laricina in Europe: characterization and identification of isolates and laboratory strains by soluble protein electrophoresis. Can. J. Bot. 1990, 68, 2629–2635. [Google Scholar] [CrossRef]
- Dorworth, C.E.; Krywienczyk, J. Comparisons among isolates of Gremmeniella abietina by means of growth rate, conidia measurement, and immunogenic reaction. Can. J. Bot. 1975, 53, 2506–2525. [Google Scholar] [CrossRef]
- Uotila, A.; Hantula, J.; Vaatanen, A.K.; Hamelin, R.C. Hybridization between two biotypes of Gremmeniella abietina var. abietina in artificial pairings. For. Pathol. 2000, 30, 211–219. [Google Scholar] [CrossRef]
- Hamelin, R.C.; Rail, J. Phylogeny of Gremmeniella spp. based on sequences of the 5.8S rDNA and internal transcribed spacer region. Can. J. Bot. 1997, 75, 693–698. [Google Scholar] [CrossRef]
- Uotila, A. Physiological and morphological variation among Finnish Gremmeniella abietina isolates. Commun. Inst. For. Fenn. 1983, 119, 12. [Google Scholar]
- Hamelin, R.C.; Lecours, N.; Hansson, P.; Hellgren, M.; Laflamme, G. Genetic differentiation within the European race of Gremmeniella abietina. Mycol. Res. 1996, 100, 49–56. [Google Scholar] [CrossRef]
- Hellgren, M.; Högberg, N. Ecotypic variation of Gremmeniella abietina in northern Europe: Disease patterns reflected by DNA variation. Can. J. Bot. 1995, 73, 1531–1539. [Google Scholar] [CrossRef]
- Hantula, J.; Muller, M. Variation within Gremmeniella abietina in Finland and other countries as determined by Random Amplified Microsatellites (RAMS). Mycol. Res. 1997, 101, 169–175. [Google Scholar] [CrossRef]
- Santamaria, O.; Pajares, J.A.; Diez, J.J. First report of Gremmeniella abietina on Pinus halepensis in Spain. Plant Pathol. 2003, 52, 425. [Google Scholar] [CrossRef]
- Botella, L.; Tuomivirta, T.T.; Kaitera, J.; Carrasco Navarro, V.; Diez, J.J.; Hantula, J. Spanish population of Gremmeniella abietina is genetically unique but related to type A in Europe. Fungal Biol. 2010, 114, 778–789. [Google Scholar] [CrossRef] [PubMed]
- Ghabrial, S.A.; Suzuki, N. Viruses of Plant Pathogenic Fungi. Annu. Rev. Phytopathol. 2009, 47, 353–384. [Google Scholar] [CrossRef] [PubMed]
- Schoebel, C.N.; Botella, L.; Lygis, V.; Rigling, D. Population genetic analysis of a parasitic mycovirus to infer the invasion history of its fungal host. Mol. Ecol. 2017, 26, 2482–2497. [Google Scholar] [CrossRef] [PubMed]
- Bryner, S.F.; Rigling, D.; Brunner, P.C. Invasion history and demographic pattern of Cryphonectria hypovirus 1 across European populations of the chestnut blight fungus. Ecol. Evol. 2012, 2, 3227–3241. [Google Scholar] [CrossRef] [PubMed]
- Vainio, E.J.; Hyder, R.; Aday, G.; Hansen, E.; Piri, T.; Doğmuş-Lehtijärvi, T.; Lehtijärvi, A.; Korhonen, K.; Hantula, J. Population structure of a novel putative mycovirus infecting the conifer root-rot fungus Heterobasidion annosum sensu lato. Virology 2012, 422, 366–376. [Google Scholar] [CrossRef] [PubMed]
- Milgroom, M.G.; Lipari, S.E.; Ennos, R.A.; Liu, Y.-C. Estimation of the outcrossing rate in the chestnut blight fungus, Cryphonectria parasitica. Heredity 1993, 70, 385–892. [Google Scholar] [CrossRef]
- Liu, Y.-C.; Linder-Basso, D.; Hillman, B.I.; Kaneko, S.; Milgroom, M.G. Evidence for interspecies transmission of viruses in natural populations of filamentous fungi in the genus Cryphonectria. Mol. Ecol. 2003, 12, 1619–1628. [Google Scholar] [CrossRef] [PubMed]
- Deng, F.; Xu, R.; Boland, G.J. Hypovirulence-associated double-stranded RNA from Sclerotinia homoeocarpa is conspecific with Ophiostoma novo-ulmi Mitovirus 3a-Ld. Phytopathology 2003, 93, 1407–1414. [Google Scholar] [CrossRef] [PubMed]
- Buck, K.W.; Brasier, C.M.; Paoletti, M.; Crawford, L.J. Virus transmission and gene flow between two species of the Dutch elm disease fungi, Ophiostoma ulmi and O. novo-ulmi: Deleterious viruses as selective. Br. Ecol. Soc. 2003, 15, 26–45. [Google Scholar]
- Vainio, E.J.; Hakanpää, J.; Dai, Y.-C.; Hansen, E.; Korhonen, K.; Hantula, J. Species of Heterobasidion host a diverse pool of partitiviruses with global distribution and interspecies transmission. Fungal Biol. 2011, 115, 1234–1243. [Google Scholar] [CrossRef] [PubMed]
- Kashif, M.; Hyder, R.; De Vega Perez, D.; Hantula, J.; Vainio, E.J. Heterobasidion wood decay fungi host diverse and globally distributed viruses related to Helicobasidium mompa partitivirus V70. Virus Res. 2015, 195, 119–123. [Google Scholar] [CrossRef] [PubMed]
- Melzer, M.S.; Ikeda, S.S.; Boland, G.J. Interspecific transmission of double-stranded RNA and hypovirulence from Sclerotinia sclerotiorum to S. minor. Phytopathology 2002, 92, 780–784. [Google Scholar] [CrossRef] [PubMed]
- Boland, G.J. Fungal viruses, hypovirulence, and biological control of Sclerotinia species. Can. J. Plant Pathol. 2004, 26, 6–18. [Google Scholar] [CrossRef]
- Charlton, N.D.; Carbone, I.; Tavantzis, S.M.; Cubeta, M.A. Phylogenetic relatedness of the M2 double-stranded RNA in Rhizoctonia fungi. Mycologia 2008, 100, 555–564. [Google Scholar] [CrossRef] [PubMed]
- Tuomivirta, T.T.; Uotila, A.; Hantula, J. Two independent double-stranded RNA patterns occur in the Finnish Gremmeniella abietina var. abietina type A. For. Pathol. 2002, 32, 197–205. [Google Scholar] [CrossRef]
- Tuomivirta, T.T.; Hantula, J. Two unrelated double-stranded RNA molecule patterns in Gremmeniella abietina type A code for putative viruses of the families Totiviridae and Partitiviridae. Arch. Virol. 2003, 148, 2293–2305. [Google Scholar] [CrossRef] [PubMed]
- Tuomivirta, T.T.; Hantula, J. Three unrelated viruses occur in a single isolate of Gremmeniella abietina var. abietina type A. Virus Res. 2005, 110, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Tuomivirta, T.T.; Hantula, J. Gremmeniella abietina mitochondrial RNA virus S1 is phylogenetically related to the members of the genus Mitovirus. Arch. Virol. 2003, 148, 2429–2436. [Google Scholar] [CrossRef] [PubMed]
- Botella, L.; Tuomivirta, T.T.; Vervuurt, S.; Diez, J.J.; Hantula, J. Occurrence of two different species of mitoviruses in the European race of Gremmeniella abietina var. abietina, both hosted by the genetically unique Spanish population. Fungal Biol. 2012, 116, 872–882. [Google Scholar] [CrossRef] [PubMed]
- Tuomivirta, T.T.; Kaitera, J.; Hantula, J. A novel putative virus of Gremmeniella abietina type B (Ascomycota: Helotiaceae) has a composite genome with endornavirus affinities. J. Gen. Virol. 2009, 90, 2299–2305. [Google Scholar] [CrossRef] [PubMed]
- Botella, L.; Vainio, E.J.; Hantula, J.; Diez, J.J.; Jankovsky, L. Description and prevalence of a putative novel mycovirus within the conifer pathogen Gremmeniella abietina. Arch. Virol. 2015, 160, 1967–1975. [Google Scholar] [CrossRef] [PubMed]
- Nibert, M.L.; Ghabrial, S.A.; Maiss, E.; Lesker, T.; Vainio, E.J.; Jiang, D.; Suzuki, N. Taxonomic reorganization of family Partitiviridae and other recent progress in partitivirus research. Virus Res. 2014, 188, 128–141. [Google Scholar] [CrossRef] [PubMed]
- Botella, L.; Tuomivirta, T.T.; Hantula, J.; Diez, J.J.; Jankovsky, L. The European race of Gremmeniella abietina hosts a single species of gammapartitivirus showing a global distribution and possible recombinant events in its history. Fungal Biol. 2015, 119, 125–135. [Google Scholar] [CrossRef] [PubMed]
- Kuraku, S.; Zmasek, C.M.; Nishimura, O.; Katoh, K. aLeaves facilitates on-demand exploration of metazoan gene family trees on MAFFT sequence alignment server with enhanced interactivity. Nucleic Acids Res. 2013, 41, W22–W28. [Google Scholar] [CrossRef] [PubMed]
- Thapa, V.; Turner, G.G.; Hafenstein, S.; Overton, B.E.; Vanderwolf, K.J.; Roossinck, M.J. Using a novel partitivirus in Pseudogymnoascus destructans to understand the epidemiology of White-Nose Syndrome. PLoS Pathog. 2016, 12, e1006076. [Google Scholar] [CrossRef] [PubMed]
- Marzano, S.-Y.L.; Domier, L.L. Novel mycoviruses discovered from metatranscriptomics survey of soybean phyllosphere phytobiomes. Virus Res. 2016, 213, 332–342. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, K.; Katayama, Y.; Omatsu, T.; Mizutani, T.; Fukuhara, T.; Kodama, M.; Arie, T.; Teraoka, T.; Moriyama, H. Genome sequence of a novel mitovirus identified in the phytopathogenic fungus Alternaria arborescens. Arch. Virol. 2016, 161, 2627–2631. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Shang, H.H.; Yang, H.Q.; Da Gao, B.; Zhong, J. A mitovirus isolated from the phytopathogenic fungus Alternaria brassicicola. Arch. Virol. 2017, 162, 2869–2874. [Google Scholar] [CrossRef] [PubMed]
- Bruenn, J.A. A closely related group of RNA-dependent RNA polymerases from double-stranded RNA viruses. Nucleic Acids Res. 1993, 21, 5667–5669. [Google Scholar] [CrossRef] [PubMed]
- Nerva, L.; Ciuffo, M.; Vallino, M.; Margaria, P.; Varese, G.C.; Gnavi, G.; Turina, M. Multiple approaches for the detection and characterization of viral and plasmid symbionts from a collection of marine fungi. Virus Res. 2016, 219, 22–38. [Google Scholar] [CrossRef] [PubMed]
- Niu, Y.; Zhang, T.; Zhu, Y.; Yuan, Y.; Wang, S.; Liu, J.; Liu, D. Isolation and characterization of a novel mycovirus from Penicillium digitatum. Virology 2016, 494, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Hammond, T.M.; Andrewski, M.D.; Roossinck, M.J.; Keller, N.P. Aspergillus mycoviruses are targets and suppressors of RNA silencing. Eukaryot. Cell 2008, 7, 350–357. [Google Scholar] [CrossRef] [PubMed]
- Rong, R.; Rao, S.; Scott, S.W.; Carner, G.R.; Tainter, F.H. Complete sequence of the genome of two dsRNA viruses from Discula destructiva. Virus Res. 2002, 90, 217–224. [Google Scholar] [CrossRef]
- Khalifa, M.E.; Pearson, M.N. Molecular characterisation of an endornavirus infecting the phytopathogen Sclerotinia sclerotiorum. Virus Res. 2014, 189, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Kwon, S.-J.; Lee, K.-M.; Son, M.; Kim, K.-H. Complete nucleotide sequence of double-stranded RNA viruses from Fusarium graminearum strain DK3. Arch. Virol. 2009, 154, 1855–1858. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Liu, H.; Zhang, M.; Cao, X.; Zhou, E. The complete genomic sequence of a novel mycovirus from Rhizoctonia solani AG-1 IA strain B275. Arch. Virol. 2013, 158, 1609–1612. [Google Scholar] [CrossRef] [PubMed]
- Marquez, L.M.; Redman, R.S.; Rodriguez, R.; Stout, R.G.; Rodriguez, R.J.; Roossinck, M. A Virus in a fungus in a plant: Three-way symbiosis required for thermal tolerance. Science 2007, 315, 513–515. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, S.; Yang, X.; Zeng, H.; Qiu, D.; Guo, L. The complete genome sequence of a double-stranded RNA mycovirus from Fusarium graminearum strain HN1. Arch. Virol. 2017, 162, 2119–2124. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Zhang, T.; Luo, C.; Jiang, D.; Li, G.; Li, Q.; Hsiang, T.; Huang, J. Prevalence and diversity of mycoviruses infecting the plant pathogen Ustilaginoidea virens. Virus Res. 2015, 195, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Botella, L.; Dvořák, M.; Capretti, P.; Luchi, N. Effect of temperature on GaRV6 accumulation and its fungal host, the conifer pathogen Gremmeniella abietina. For. Pathol. 2017, 47, e12291. [Google Scholar] [CrossRef]
- Leticia Botella, L.; Tuomivirta, T.T.; Hantula, J.; Diez, J.J. Presence of viral dsRNA molecules in the Spanish population of Gremmeniella abietina. J. Agric. Ext. Rural Dev. 2012, 4, 211–213. [Google Scholar] [CrossRef]
- Göker, M.; Scheuner, C.; Klenk, H.-P.; Stielow, J.B.; Menzel, W. Codivergence of mycoviruses with their hosts. PLoS ONE 2011, 6, e22252. [Google Scholar] [CrossRef] [PubMed]
- Pearson, M.N.; Beever, R.E.; Boine, B.; Arthur, K. Mycoviruses of filamentous fungi and their relevance to plant pathology. Mol. Plant Pathol. 2009, 10, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Varga, J.; Vágvölgyi, C.; Tóth, B. Recent advances in mycovirus research. Acta Microbiol. Immunol. Hung. 2003, 50, 77–94. [Google Scholar] [CrossRef] [PubMed]
- Linder-Basso, D.; Dynek, J.N.; Hillman, B.I. Genome analysis of Cryphonectria hypovirus 4, the most common hypovirus species in North America. Virology 2005, 337, 192–203. [Google Scholar] [CrossRef] [PubMed]
- Dorworth, C.E.; Krywienczyk, J.; Skilling, D.D. New York isolates of Gremmeniella abietina (Scleroderris lagerbergii) identical in immunogenic reaction to European isolates [Pinus]. Plant Dis. Rep. 1977, 61, 887–890. [Google Scholar]
- Marosy, M.; Patton, R.F.; Upper, C.D. A conductive day concept to explain the effect of low temperature on the development of Scleroderris shoot blight. Ecol. Epidemiol. 1989, 79, 1293–1301. [Google Scholar]
- Yaegashi, H.; Kanematsu, S. Natural infection of the soil-borne fungus Rosellinia necatrix with novel mycoviruses under greenhouse conditions. Virus Res. 2016, 219, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Yaegashi, H.; Nakamura, H.; Sawahata, T.; Sasaki, A.; Iwanami, Y.; Ito, T.; Kanematsu, S. Appearance of mycovirus-like double-stranded RNAs in the white root rot fungus, Rosellinia necatrix, in an apple orchard. FEMS Microbiol. Ecol. 2013, 83, 49–62. [Google Scholar] [CrossRef] [PubMed]
- Vainio, E.J.; Pennanen, T.; Rajala, T.; Hantula, J. Occurrence of similar mycoviruses in pathogenic, saprotrophic and mycorrhizal fungi inhabiting the same forest stand. FEMS Microbiol. Ecol. 2017, 93, fix003. [Google Scholar] [CrossRef] [PubMed]
- Romeralo Tapia, C.; Botella, L.; Santamaría, O.; Diez, J. Effect of putative mitoviruses on in vitro growth of Gremmeniella abietina isolates under different laboratory conditions. For. Syst. 2012, 21, 515. [Google Scholar] [CrossRef]
| Virus Name | Virus Abbreviation | Full-Length Virus Strains | GenBank Accessions * | Virus Genus | Genome | Fungal Population | Population Studies | Countries Detected | References |
|---|---|---|---|---|---|---|---|---|---|
| Gremmeniella abietina RNA virus multi segmented 1 | GaRV-MS1 | 3 | KJ786411-KJ786413, AY089993-5, AY615211-13, | Gammapartitivirus | dsRNA | A, B, SP | YES | Canada, USA, Finland, Spain, Montenegro, Italy, Turkey | [31,32,38] |
| Gremmeniella abietina mitovirus 1 | GMV1 | 3 | HE586988, AF534641, AY615209 | Mitovirus | (+) ssRNA | A, SP | YES | Finland, Spain | [32,33,34] |
| Gremmeniella abietina mitovirus 2 | GMV2 | 1 | JN654496 | Mitovirus | (+) ssRNA | B, SP | YES | Finland, Spain | [34] |
| Gremmeniella abietina RNA virus lone | GaRV-L | 2 | AF337175, AY615210 | Totivirus | dsRNA | A | NO | Finland | [31,32] |
| Gremmeniella abietina RNA virus XL | GaBRV-XL | 2 | DQ399289-90 | Betaendornavirus | dsRNA | B | NO | Finland | [35] |
| Gremmeniella abietina RNA virus 6 | GaRV6 | 1 | KJ742567.1 | Unassigned † | dsRNA | A, SP | YES | Finland, Canada, Italy, Spain | [36] |
| GenBank BLASTX Database | Sequence ID | Family | Identities | Matching Region 1 |
|---|---|---|---|---|
| Macrophomina phaseolina single-stranded RNA virus 1 | ALD89094.1 | Fusariviridae * | 217/335 (65%) | 512 to 845 |
| Neofusicoccum luteum fusarivirus 1 | ARO52688.1 | Fusariviridae * | 205/333 (62%) | 527 to 856 |
| Penicillium aurantiogriseum fusarivirus 1 | YP_009182154.1 | Fusariviridae * | 198/328 (60%) | 511 to 838 |
| Penicillium roqueforti ssRNA mycovirus 1 | YP_009052456.1 | Fusariviridae * | 177/313 (57%) | 500 to 812 |
| Fusarium graminearum dsRNA mycovirus-1 | YP_223920.2 | Fusariviridae * | 179/334 (54%) | 530 to 863 |
| Sodiomyces alkalinus fusarivirus 1 | ATP75827.1 | Fusariviridae * | 183/326 (56%) | 445 to 770 |
| Rosellinia necatrix fusarivirus 1 | YP_009047147.1 | Fusariviridae * | 80/329 (55%) | 522 to 850 |
| Pleospora typhicola fusarivirus 1 | YP_009182158.1 | Fusariviridae * | 178/329 (54%) | 521 to 849 |
| Fusarium poae fusarivirus 1 | YP_009272906.1 | Fusariviridae * | 172/326 (53%) | 496 to 818 |
| Agaricus bisporus virus 11 | AQM49938.1 | Fusariviridae * | 151/314 (48%) | 592 to 901 |
| GenBank BLASTX database | Sequence ID | Family | Identities | Matching Region 1 |
|---|---|---|---|---|
| Sclerotinia nivalis victorivirus 1 | YP_009259368.1 | Totiviridae | 456/587 (78%) | 247 to 833 |
| Aspergillus foetidus slow virus 1 | YP_009508249.1 | Totiviridae | 306/584 (52%) | 254 to 837 |
| Beauveria bassiana victorivirus 1 | AMQ11131.1 | Totiviridae | 302/585 (52%) | 258 to 841 |
| Sphaeropsis sapinea RNA virus | NP_047558.1 | Totiviridae | 314/587 (53%) | 254 to 837 |
| Rosellinia necatrix victorivirus 1 | BAM36400.1 | Totiviridae | 397/587 (67% | 107 to 692 |
| Beauveria bassiana victorivirus NZL/1980 | YP_009032633.1 | Totiviridae | 301/585 (51%) | 258 to 841 |
| Soybean-associated double-stranded RNA virus 1 | ALM62239.1 | Totiviridae | 311/587 (53%) | 255 to 840 |
| Ustilaginoidea virens RNA virus 1 | AGO04407.1 | Totiviridae | 302/584 (52%) | 243 to 825 |
| Bipolaris maydis victorivirus 1 | AXB26764.1 | Totiviridae | 289/588 (49%) | 253 to 835 |
| Botryotinia fuckeliana totivirus 1 | YP_001109580.1 | Totiviridae | 291/581 (50%) | 266 to 836 |
| GenBank BLASTX Database | Sequence ID | Family | Identities | Matching Region 1 |
|---|---|---|---|---|
| Sclerotinia nivalis victorivirus 1 | YP_009259367.1 | Totiviridae | 293/363 (81%) | 1 to 363 |
| Tolypocladium cylindrosporum virus 1 | YP_004089629.1 | Totiviridae | 247/362 (68%) | 1 to 362 |
| Beauveria bassiana victorivirus NZL/1980 | YP_009032632.1 | Totiviridae | 237/361 (66%) | 1 to 361 |
| Helminthosporium victoriae virus 190S | NP_619669.2 | Totiviridae | 240/363 (66%) | 1 to 363 |
| Bipolaris maydis victorivirus 1 | AXB26763.1 | Totiviridae | 240/363 (66%) | 1 to 363 |
| Botryotinia fuckeliana totivirus 1 | YP_001109579.1 | Totiviridae | 227/363 (63%) | 1 to 363 |
| Aspergillus foetidus slow virus 1 | YP_009508248.1 | Totiviridae | 221/360 (61%) | 1 to 360 |
| Fusarium asiaticum victorivirus 1 | AYD49681.1 | Totiviridae | 225/364 (62%) | 1 to 363 |
| Ustilaginoidea virens RNA virus L | YP_009094184.1 | Totiviridae | 223/363 (61%) | 1 to 363 |
| Rosellinia necatrix victorivirus 1 | BAM36399.1 | Totiviridae | 227/362 (63%) | 1 to 361 |
| Sclerotinia sclerotiorum victorivirus 1 (partial) | AWY10937.1 | Totiviridae | 208/263 (79%) | 1 to 233 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Botella, L.; Hantula, J. Description, Distribution, and Relevance of Viruses of the Forest Pathogen Gremmeniella abietina. Viruses 2018, 10, 654. https://doi.org/10.3390/v10110654
Botella L, Hantula J. Description, Distribution, and Relevance of Viruses of the Forest Pathogen Gremmeniella abietina. Viruses. 2018; 10(11):654. https://doi.org/10.3390/v10110654
Chicago/Turabian StyleBotella, Leticia, and Jarkko Hantula. 2018. "Description, Distribution, and Relevance of Viruses of the Forest Pathogen Gremmeniella abietina" Viruses 10, no. 11: 654. https://doi.org/10.3390/v10110654
APA StyleBotella, L., & Hantula, J. (2018). Description, Distribution, and Relevance of Viruses of the Forest Pathogen Gremmeniella abietina. Viruses, 10(11), 654. https://doi.org/10.3390/v10110654






