Evolution of Codon Usage Bias in Henipaviruses Is Governed by Natural Selection and Is Host-Specific
Abstract
1. Introduction
2. Materials and Methods
2.1. Sequence Data Analyzed
2.2. Phylogenetic Analysis
2.3. Nucleotide Composition Analysis
2.4. Relative Synonymous Codon Usage Analysis
2.5. Effective Number of Codons Analysis
2.6. ENc–GC3s Plot Analysis
2.7. Parity Rule 2 Analysis
2.8. Neutrality Plot Analysis
2.9. Codon Adaptation Index
2.10. tRNA Adaptation Index
2.11. Relative Codon Deoptimization Index
2.12. Similarity Index
2.13. Statistical Analysis
3. Results
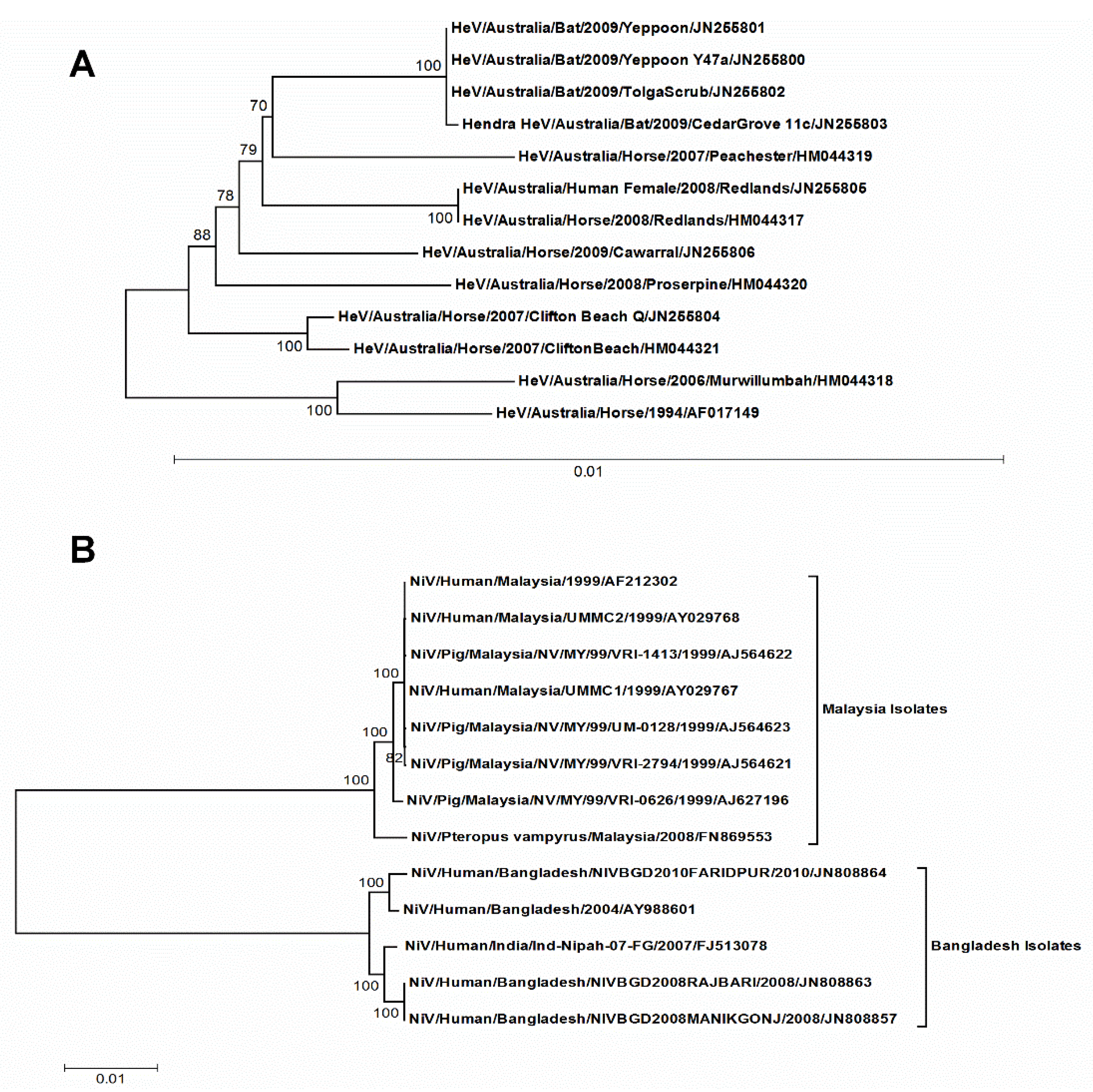
3.1. HeV and NiV Have Quite Distinct Evolutionary Patterns
3.2. Trends in Codon Usage Variations of Henipaviruses
3.3. Influence of Nucleotide Compositions on the Codon Usage Bias
3.4. Relative Synonymous Codon Usage (RSCU) Analysis
3.5. L Gene of NiV and N Gene of HeV Showed a Comparatively High Codon Bias
3.6. Mutation Bias Acts Differently on the Protein-Coding Genes of NiV and HeV
3.7. Natural Selection Prevails in Shaping the Codon Usage Patterns in Henipaviruses
3.8. HeV and NiV Showed Host-Specific Discrete Codon Adaptation Patterns
3.9. Henipaviruses Coding Sequences Showed Lowest Codon Usage Deoptimization for Pteropus alecto
3.10. Sus scrofa Had a High Similarity Index for Henipaviruses
3.11. Henipaviruses Are Better Adapted to tRNAs Pool of Homo sapiens
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wang, L.F.; Mackenzie, J.S.; Broder, C.C. Henipaviruses. In Fields Virology; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2013; pp. 1070–1085. ISBN 9781451105636, 1451105630. [Google Scholar]
- Marsh, G.A.; de Jong, C.; Barr, J.A.; Tachedjian, M.; Smith, C.; Middleton, D.; Yu, M.; Todd, S.; Foord, A.J.; Haring, V.; et al. Cedar virus: A novel Henipavirus isolated from Australian bats. PLoS Pathog. 2012, 8, e1002836. [Google Scholar] [CrossRef] [PubMed]
- Murray, K.; Selleck, P.; Hooper, P.; Hyatt, A.; Gould, A.; Gleeson, L.; Westbury, H.; Hiley, L.; Selvey, L.; Rodwell, B.; et al. A morbillivirus that caused fatal disease in horses and humans. Science 1995, 268, 94–97. [Google Scholar] [CrossRef] [PubMed]
- Harcourt, B.H.; Tamin, A.; Ksiazek, T.G.; Rollin, P.E.; Anderson, L.J.; Bellini, W.J.; Rota, P.A. Molecular characterization of Nipah virus, a newly emergent paramyxovirus. Virology 2000, 271, 334–349. [Google Scholar] [CrossRef] [PubMed]
- Croser, E.L.; Marsh, G.A. The changing face of the henipaviruses. Vet. Microbiol. 2013, 167, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Hendra Virus. ProMED-Mail. Archive No. 20170819.5260496. 2017. Available online: http://www.promedmail.org/ (accessed on 20 November 2017).
- Nipah Virus. ProMED-Mail. Archive No. 20150407.3280088. 2015. Available online: http://www.promedmail.org/ (accessed on 20 November 2017).
- Chua, K.B.; Koh, C.L.; Hooi, P.S.; Wee, K.F.; Khong, J.H.; Chua, B.H.; Chan, Y.P.; Lim, M.E.; Lam, S.K. Isolation of Nipahvirus from Malaysian Island flying-foxes. Microbes Infect. 2002, 4, 145–151. [Google Scholar] [CrossRef]
- Halpin, K.; Young, P.L.; Field, H.E.; Mackenzie, J.S. Isolation of Hendra virus from pteropid bats: A natural reservoir of Hendra virus. J. Gen. Virol. 2000, 81, 1927–1932. [Google Scholar] [CrossRef] [PubMed]
- Rahman, S.A.; Hassan, S.S.; Olival, K.J.; Mohamed, M.; Chang, L.Y.; Hassan, L.; Saad, N.M.; Shohaimi, S.A.; Mamat, Z.C.; Naim, M.S.; et al. Characterization of Nipah virus from naturally infected Pteropus vampyrus bats, Malaysia. Emerg. Infect. Dis. 2010, 16, 1990–1993. [Google Scholar] [CrossRef] [PubMed]
- Peel, A.J.; Baker, K.S.; Crameri, G.; Barr, J.A.; Hayman, D.T.; Wright, E.; Broder, C.C.; Fernández-Loras, A.; Fooks, A.R.; Wang, L.F.; et al. Henipavirus neutralising antibodies in an isolated island population of African fruit bats. PLoS ONE. 2012, 7, e30346. [Google Scholar] [CrossRef] [PubMed]
- Hasebe, F.; Thuy, N.T.; Inoue, S.; Yu, F.; Kaku, Y.; Watanabe, S.; Akashi, H.; Dat, D.T.; Mai, L.T.Q.; Morita, K. Serologic evidence of Nipah virus infection in bats, Vietnam. Emerg. Infect. Dis. 2012, 18, 536–537. [Google Scholar] [CrossRef] [PubMed]
- Wacharapluesadee, S.; Lumlertdacha, B.; Boongird, K.; Wanghongsa, S.; Chanhome, L.; Rollin, P.; Stockton, P.; Rupprecht, C.E.; Ksiazek, T.G.; Hemachudha, T. Bat Nipah virus, Thailand. Emerg. Infect. Dis. 2005, 11, 1949–1951. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wang, J.; Hickey, A.C.; Zhang, Y.; Li, Y.; Wu, Y.; Zhang, H.; Yuan, J.; Han, Z.; McEachern, J.; et al. Antibodies to Nipah or Nipah-like viruses in bats, China. Emerg. Infect. Dis. 2008, 14, 1974–1976. [Google Scholar] [CrossRef] [PubMed]
- Grantham, R.; Gautier, C.; Gouy, M.; Mercier, R.; Pave, A. Codon catalog usage and the genome hypothesis. Nucleic Acids Res. 1980, 8, 49–62. [Google Scholar] [CrossRef]
- Sharp, P.M.; Averof, M.; Lloyd, A.T.; Matassi, G.; Peden, J.F. DNA sequence evolution: The sounds of silence. Philos. Trans. R. Soc. Lond. B Biol. Sci. 1995, 349, 241–247. [Google Scholar] [CrossRef] [PubMed]
- Sueoka, N. Directional mutation pressure and neutral molecular evolution. Proc. Natl. Acad. Sci. USA 1988, 85, 2653–2657. [Google Scholar] [CrossRef] [PubMed]
- Cooper, D.A.; Banerjee, S.; Chakrabarti, A.; García-Sastre, A.; Hesselberth, J.R.; Silberman, R.H.; Barton, D.J. RNase L targets distinct sites in influenza A virus RNAs. J. Virol. 2015, 89, 2764–2776. [Google Scholar] [CrossRef] [PubMed]
- Greenbaum, B.D.; Levine, A.J.; Bhanot, G.; Rabadan, R. Patterns of evolution and host gene mimicry in influenza and other RNA viruses. PLoS Pathog. 2008, 4, e1000079. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Liu, W.J.; Peng, S.W.; Sun, X.Y.; Frazer, I. Papillomavirus capsid protein expression level depends on the match between codon usage and tRNA availability. J. Virol. 1999, 73, 4972–4982. [Google Scholar] [PubMed]
- Belalov, I.S.; Lukashev, A.N. Causes and implications of codon usage bias in RNA viruses. PLoS ONE 2013, 8, e56642. [Google Scholar] [CrossRef] [PubMed]
- Bera, B.C.; Virmani, N.; Kumar, N.; Anand, T.; Pavulraj, S.; Rash, A.; Elton, D.; Rash, N.; Bhatia, S.; Sood, R.; et al. Genetic and codon usage bias analyses of polymerase genes of equine influenza virus and its relation to evolution. BMC Genom. 2017, 18, 652. [Google Scholar] [CrossRef] [PubMed]
- Simmonds, P.; Tuplin, A.; Evans, D.J. Detection of genome-scale ordered RNA structure (GORS) in genomes of positive-stranded RNA viruses: Implications for virus evolution and host persistence. RNA 2004, 10, 1337–1351. [Google Scholar] [CrossRef] [PubMed]
- Weill, L.; James, L.; Ulryck, N.; Chamond, N.; Herbreteau, C.H.; Ohlmann, T.; Sargueil, B. A new type of IRES within gag coding region recruits three initiation complexes on HIV-2 genomic RNA. Nucleic Acids Res. 2010, 38, 1367–1381. [Google Scholar] [CrossRef] [PubMed]
- Marsh, G.A.; Rabadan, R.; Levine, A.J.; Palese, P. Highly Conserved Regions of Influenza A Virus Polymerase Gene Segments Are Critical for Efficient Viral RNA Packaging. J. Virol. 2008, 82, 2295–2304. [Google Scholar] [CrossRef] [PubMed]
- Shackelton, L.A.; Parrish, C.R.; Holmes, E.C. Evolutionary basis of codon usage and nucleotide composition bias in vertebrate DNA viruses. J. Mol. Evol. 2006, 62, 551–563. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Sharp, P.M.; Li, W.H. Codon usage in regulatory genes in Escherichia coli does not reflect selection for ‘rare’ codons. Nucleic Acids Res. 1986, 14, 7737–7749. [Google Scholar] [CrossRef] [PubMed]
- Wright, F. The ‘effective number of codons’ used in a gene. Gene 1990, 87, 23–29. [Google Scholar] [CrossRef]
- Sueoka, N. Translation-coupled violation of Parity Rule 2 in human genes is not the cause of heterogeneity of the DNA G+C content of third codon position. Gene 1999, 238, 53–58. [Google Scholar] [CrossRef]
- Sharp, P.M.; Li, W.H. The codon Adaptation Index—A measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 1987, 15, 1281–1295. [Google Scholar] [CrossRef] [PubMed]
- Puigbo, P.; Bravo, I.G.; Garcia-Vallve, S. CAIcal: A combined set of tools to assess codon usage adaptation. Biol. Direct 2008, 3, 38. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, Y.; Gojobori, T.; Ikemura, T. Codon usage tabulated from international DNA sequence databases: Status for the year 2000. Nucleic Acids Res. 2000, 28, 292. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.P.; Ke, H.; Liang, Z.L.; Liu, Z.X.; Hao, L.; Ma, J.Y.; Li, Y.G. Multiple Evolutionary Selections Involved in Synonymous Codon Usages in the Streptococcus agalactiae Genome. Int. J. Mol. Sci. 2016, 17, 277. [Google Scholar] [CrossRef] [PubMed]
- Sabi, R.; Volvovitch Daniel, R.; Tuller, T. stAIcalc: TRNA adaptation index calculator based on species-specific weights. Bioinformatics 2017, 33, 589–591. [Google Scholar] [CrossRef] [PubMed]
- dos Reis, M.; Savva, R.; Wernisch, L. Solving the riddle of codon usage preferences: A test for translational selection. Nucleic Acids Res. 2004, 32, 5036–5044. [Google Scholar] [CrossRef] [PubMed]
- Chan, P.P.; Lowe, T.M. GtRNAdb: A database of transfer RNA genes detected in genomic sequence. Nucleic Acids Res. 2009, 37, D93–D97. [Google Scholar] [CrossRef] [PubMed]
- Puigbò, P.; Aragonès, L.; Garcia-Vallvé, S. RCDI/eRCDI: A web-server to estimate codon usage deoptimization. BMC Res. Notes 2010, 3, 87. [Google Scholar] [CrossRef] [PubMed]
- Mueller, S.; Papamichail, D.; Coleman, J.R.; Skiena, S.; Wimmer, E. Reduction of the rate of poliovirus protein synthesis through large-scale codon deoptimization causes attenuation of viral virulence by lowering specific infectivity. J. Virol. 2006, 80, 9687–9696. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.H.; Zhang, J.; Sun, D.J.; Ma, Q.; Chen, H.T.; Ma, L.N.; Ding, Y.Z.; Liu, Y.S. The distribution of synonymous codon choice in the translation initiation region of dengue virus. PLoS ONE 2013, 8, e77239. [Google Scholar] [CrossRef] [PubMed]
- Greenacre, M. Theory and Applications of Correspondence Analysis; Academic Press: London, UK, 1984; ISBN 0122990501. [Google Scholar]
- Weis, M.; Behner, L.; Hoffmann, M.; Krüger, N.; Herrler, G.; Drosten, C.; Drexler, J.F.; Dietzel, E.; Maisner, A. Characterization of African bat henipavirus GH-M74a glycoproteins. J. Gen. Virol. 2014, 95, 539–548. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Yang, L.; Yang, F.; Ren, X.; Jiang, J.; Dong, J.; Sun, L.; Zhu, Y.; Zhou, H.; Jin, Q. Novel Henipa-like virus, Mojiang Paramyxovirus, in rats, China, 2012. Emerg. Infect. Dis. 2014, 20, 1064–1066. [Google Scholar] [CrossRef] [PubMed]
- Lo Presti, A.; Cella, E.; Giovanetti, M.; Lai, A.; Angeletti, S.; Zehender, G.; Ciccozzi, M. Origin and evolution of Nipah virus. J. Med. Virol. 2016, 88, 380–388. [Google Scholar] [CrossRef] [PubMed]
- Marsh, G.A.; Todd, S.; Foord, A.; Hansson, E.; Davies, K.; Wright, L.; Morrissy, C.; Halpin, K.; Middleton, D.; Field, H.E.; et al. Genome sequence conservation of Hendra virus isolates during spillover to horses, Australia. Emerg. Infect. Dis. 2010, 16, 1767–1769. [Google Scholar] [CrossRef] [PubMed]
- Di Giallonardo, F.; Schlub, T.E.; Shi, M.; Holmes, E.C. Dinucleotide Composition in Animal RNA Viruses Is Shaped More by Virus Family than by Host Species. J. Virol. 2017, 91, e02381-16. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, G.; Bosch, A.; Pinto, R.M. Genome variability and capsid structural constraints of hepatitis A virus. J. Virol. 2003, 77, 452–459. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.S.; Wang, Q.Q.; Zhang, J.; Chen, H.T.; Xu, Z.W.; Zhu, L.; Ding, Y.Z.; Ma, L.N.; Xu, K.; Gu, Y.X.; et al. The characteristic of codon usage pattern and its evolution of hepatitis C virus. Infect. Genet. Evol. 2011, 11, 2098–2102. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Liu, S.; Zhang, B.; Wei, W. Analysis of Synonymous Codon Usage Bias of Zika Virus and Its Adaption to the Hosts. PLoS ONE 2016, 11, e0166260. [Google Scholar] [CrossRef] [PubMed]
- Cristina, J.; Moreno, P.; Moratorio, G.; Musto, H. Genome-wide analysis of codon usage bias in Ebolavirus. Virus Res. 2014, 196, 87–93. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Bera, B.C.; Greenbaum, B.D.; Bhatia, S.; Sood, R.; Selvaraj, P.; Anand, T.; Tripathi, B.N.; Virmani, N. Revelation of influencing factors in overall codon usage bias of Equine Influenza Viruses. PLoS ONE 2016, 11, e0154376. [Google Scholar] [CrossRef] [PubMed]
- Carbone, A.; Zinovyev, A.; Kepes, F. Codon adaptation index as a measure of dominating codon bias. Bioinformatics 2003, 19, 2005–2015. [Google Scholar] [CrossRef] [PubMed]
- Yob, J.M.; Field, H.; Rashdi, A.M.; Morrissy, C.; van der Heide, B.; Rota, P.; bin Adzhar, A.; White, J.; Daniels, P.; Jamaluddin, A.; et al. Nipah virus infection in bats (order Chiroptera) in peninsular Malaysia. Emerg. Infect. Dis. 2001, 7, 439–441. [Google Scholar] [CrossRef] [PubMed]
- Clayton, B.A. Nipah virus: Transmission of a zoonotic paramyxovirus. Curr. Opin. Virol. 2017, 22, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Chong, H.T.; Hossain, M.J.; Chong, T.T. Differences in epidemiologic and clinical features of Nipah virus encephalitis between the Malaysian and Bangladesh outbreaks. Neurol. Asia 2008, 13, 23–26. [Google Scholar]
- Rahman, M.A.; Hossain, M.J.; Sultana, S.; Homaira, N.; Khan, S.U.; Rahman, M.; Gurley, E.S.; Rollin, P.E.; Lo, M.K.; Comer, J.A.; et al. Date palm sap linked to Nipah virus outbreak in Bangladesh, 2008. Vector Borne Zoonotic Dis. 2012, 12, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Escaffre, O.; Borisevich, V.; Vergara, L.A.; Wen, J.W.; Long, D.; Rockx, B. Characterization of Nipah virus infection in a model of human airway epithelial cells cultured at an air–liquid interface. J. Gen. Virol. 2016, 97, 1077–1086. [Google Scholar] [CrossRef] [PubMed]
- Sauerhering, L.; Zickler, M.; Elvert, M.; Behner, L.; Matrosovich, T.; Erbar, S.; Matrosovich, M.; Maisner, A. Species-specific and individual differences in Nipah virus replication in porcine and human airway epithelial cells. J. Gen. Virol. 2016, 97, 1511–1519. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, S.; Volchkova, V.; Basler, C.F.; Palese, P.; Volchkov, V.E.; Shaw, M.L. Nipah virus edits its P gene at high frequency to express the V and W proteins. J. Virol. 2009, 83, 3982–3987. [Google Scholar] [CrossRef] [PubMed]


| Amino Acid | Codons | HeV | NiV | Bat * | Human | Horse | Pig | Dog | Amino Acid | Codons | HeV | NiV | Bat * | Human | Horse | Pig | Dog |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Phe | UUU | 1.08 | 1.14 | 0.95 | 0.93 | 0.83 | 0.79 | 0.82 | Ser | UCA | 1.78 | 1.79 | 1.02 | 0.90 | 0.80 | 0.73 | 0.81 |
| UUC | 0.92 | 0.86 | 1.05 | 1.07 | 1.17 | 1.21 | 1.16 | UCG | 0.30 | 0.27 | 0.37 | 0.33 | 0.34 | 0.39 | 0.38 | ||
| Leu | UUA | 0.95 | 0.96 | 0.78 | 0.46 | 0.33 | 0.32 | 0.35 | AGU | 1.23 | 1.18 | 0.82 | 0.90 | 0.86 | 0.77 | 0.89 | |
| UUG | 1.05 | 1.04 | 1.22 | 0.77 | 0.72 | 0.67 | 0.68 | AGC | 0.77 | 0.83 | 1.18 | 1.44 | 1.48 | 1.62 | 1.56 | ||
| CUU | 1.14 | 1.16 | 0.71 | 0.79 | 0.73 | 0.65 | 0.67 | Arg | AGA | 1.36 | 1.38 | 1.03 | 1.29 | 1.30 | 1.12 | 1.20 | |
| CUC | 0.94 | 0.78 | 0.96 | 1.17 | 1.32 | 1.35 | 1.25 | AGG | 0.64 | 0.62 | 0.97 | 1.27 | 1.32 | 1.23 | 1.32 | ||
| CUA | 0.99 | 1.06 | 0.39 | 0.43 | 0.34 | 0.33 | 0.37 | CGU | 1.06 | 1.42 | 0.59 | 0.48 | 0.55 | 0.44 | 0.46 | ||
| CUG | 0.94 | 1.01 | 1.93 | 2.37 | 2.56 | 2.68 | 2.45 | CGC | 0.76 | 0.25 | 1.18 | 1.10 | 1.15 | 1.31 | 1.26 | ||
| Ile | AUU | 1.02 | 1.03 | 1.11 | 1.08 | 0.92 | 0.91 | 0.96 | CGA | 1.79 | 1.82 | 0.82 | 0.65 | 0.61 | 0.60 | 0.67 | |
| AUC | 1.00 | 0.94 | 1.34 | 1.41 | 1.66 | 1.67 | 1.61 | CGG | 0.39 | 0.52 | 1.40 | 1.21 | 1.08 | 1.29 | 1.31 | ||
| AUA | 0.99 | 1.03 | 0.55 | 0.51 | 0.42 | 0.42 | 0.45 | Cys | UGU | 1.23 | 1.31 | 0.96 | 0.91 | 0.89 | 0.79 | 0.85 | |
| Val | GUU | 1.14 | 1.47 | 0.75 | 0.73 | 0.60 | 0.57 | 0.58 | UGC | 0.77 | 0.69 | 1.04 | 1.09 | 1.11 | 1.21 | 1.10 | |
| GUC | 1.01 | 0.75 | 0.95 | 0.95 | 1.08 | 1.07 | 1.10 | His | CAU | 1.43 | 1.19 | 0.86 | 0.84 | 0.81 | 0.70 | 0.78 | |
| GUA | 0.78 | 1.03 | 0.52 | 0.47 | 0.35 | 0.34 | 0.42 | CAC | 0.57 | 0.81 | 1.14 | 1.16 | 1.19 | 1.30 | 1.22 | ||
| GUG | 1.07 | 0.75 | 1.77 | 1.85 | 1.97 | 2.03 | 1.98 | Gln | CAA | 1.17 | 1.26 | 0.55 | 0.53 | 0.52 | 0.44 | 0.50 | |
| Pro | CCU | 1.44 | 1.68 | 1.18 | 1.15 | 1.19 | 1.05 | 1.08 | CAG | 0.83 | 0.74 | 1.46 | 1.47 | 1.48 | 1.56 | 1.46 | |
| CCC | 0.67 | 0.68 | 1.24 | 1.29 | 1.38 | 1.46 | 1.47 | Asn | AAU | 1.34 | 1.29 | 0.98 | 0.94 | 0.84 | 0.79 | 0.87 | |
| CCA | 1.42 | 1.16 | 1.13 | 1.11 | 0.97 | 0.94 | 1.05 | AAC | 0.66 | 0.71 | 1.02 | 1.06 | 1.16 | 1.21 | 1.12 | ||
| CCG | 0.47 | 0.48 | 0.45 | 0.45 | 0.45 | 0.56 | 0.51 | Lys | AAA | 1.04 | 1.17 | 0.91 | 0.87 | 0.79 | 0.76 | 0.79 | |
| Thr | ACU | 1.32 | 1.42 | 1.04 | 0.99 | 0.94 | 0.83 | 0.89 | AAG | 0.96 | 0.83 | 1.09 | 1.13 | 1.21 | 1.24 | 1.13 | |
| ACC | 0.69 | 0.71 | 1.33 | 1.42 | 1.58 | 1.68 | 1.58 | Asp | GAU | 1.34 | 1.25 | 0.95 | 0.93 | 0.83 | 0.80 | 0.86 | |
| ACA | 1.71 | 1.66 | 1.16 | 1.14 | 0.96 | 0.92 | 1.05 | GAC | 0.66 | 0.75 | 1.05 | 1.07 | 1.17 | 1.20 | 1.09 | ||
| ACG | 0.28 | 0.22 | 0.48 | 0.46 | 0.52 | 0.57 | 0.53 | Glu | GAA | 1.12 | 1.12 | 0.90 | 0.84 | 0.76 | 0.72 | 0.79 | |
| Ala | GCU | 1.53 | 1.51 | 1.09 | 1.06 | 1.05 | 0.96 | 1.00 | GAG | 0.88 | 0.89 | 1.10 | 1.16 | 1.24 | 1.28 | 1.23 | |
| GCC | 0.57 | 0.63 | 1.57 | 1.60 | 1.72 | 1.80 | 1.78 | Gly | GGU | 1.05 | 0.92 | 0.67 | 0.65 | 0.65 | 0.57 | 0.65 | |
| GCA | 1.65 | 1.59 | 0.94 | 0.91 | 0.77 | 0.74 | 0.81 | GGC | 0.49 | 0.53 | 1.31 | 1.35 | 1.43 | 1.46 | 1.45 | ||
| GCG | 0.25 | 0.28 | 0.40 | 0.42 | 0.45 | 0.50 | 0.47 | GGA | 1.36 | 1.51 | 1.04 | 1.00 | 0.95 | 0.91 | 1.02 | ||
| Tyr | UAU | 1.14 | 1.14 | 0.92 | 0.89 | 0.75 | 0.73 | 0.79 | GGG | 1.10 | 1.05 | 0.98 | 1.00 | 0.97 | 1.05 | 1.05 | |
| UAC | 0.86 | 0.86 | 1.08 | 1.11 | 1.25 | 1.27 | 1.15 | Trp | TGG | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | |
| Ser | UCU | 1.29 | 1.37 | 1.27 | 1.13 | 1.09 | 0.99 | 1.09 | Met | ATG | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| UCC | 0.63 | 0.56 | 1.34 | 1.31 | 1.43 | 1.50 | 1.52 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kumar, N.; Kulkarni, D.D.; Lee, B.; Kaushik, R.; Bhatia, S.; Sood, R.; Pateriya, A.K.; Bhat, S.; Singh, V.P. Evolution of Codon Usage Bias in Henipaviruses Is Governed by Natural Selection and Is Host-Specific. Viruses 2018, 10, 604. https://doi.org/10.3390/v10110604
Kumar N, Kulkarni DD, Lee B, Kaushik R, Bhatia S, Sood R, Pateriya AK, Bhat S, Singh VP. Evolution of Codon Usage Bias in Henipaviruses Is Governed by Natural Selection and Is Host-Specific. Viruses. 2018; 10(11):604. https://doi.org/10.3390/v10110604
Chicago/Turabian StyleKumar, Naveen, Diwakar D. Kulkarni, Benhur Lee, Rahul Kaushik, Sandeep Bhatia, Richa Sood, Atul Kumar Pateriya, Sushant Bhat, and Vijendra Pal Singh. 2018. "Evolution of Codon Usage Bias in Henipaviruses Is Governed by Natural Selection and Is Host-Specific" Viruses 10, no. 11: 604. https://doi.org/10.3390/v10110604
APA StyleKumar, N., Kulkarni, D. D., Lee, B., Kaushik, R., Bhatia, S., Sood, R., Pateriya, A. K., Bhat, S., & Singh, V. P. (2018). Evolution of Codon Usage Bias in Henipaviruses Is Governed by Natural Selection and Is Host-Specific. Viruses, 10(11), 604. https://doi.org/10.3390/v10110604






