Comparative Genomics and Characterization of the Late Promoter pR’ from Shiga Toxin Prophages in Escherichia coli
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Culture Conditions
2.2. Sequence Analysis and Phylogenetic Trees
2.3. Nomenclature of Promotor Constructs
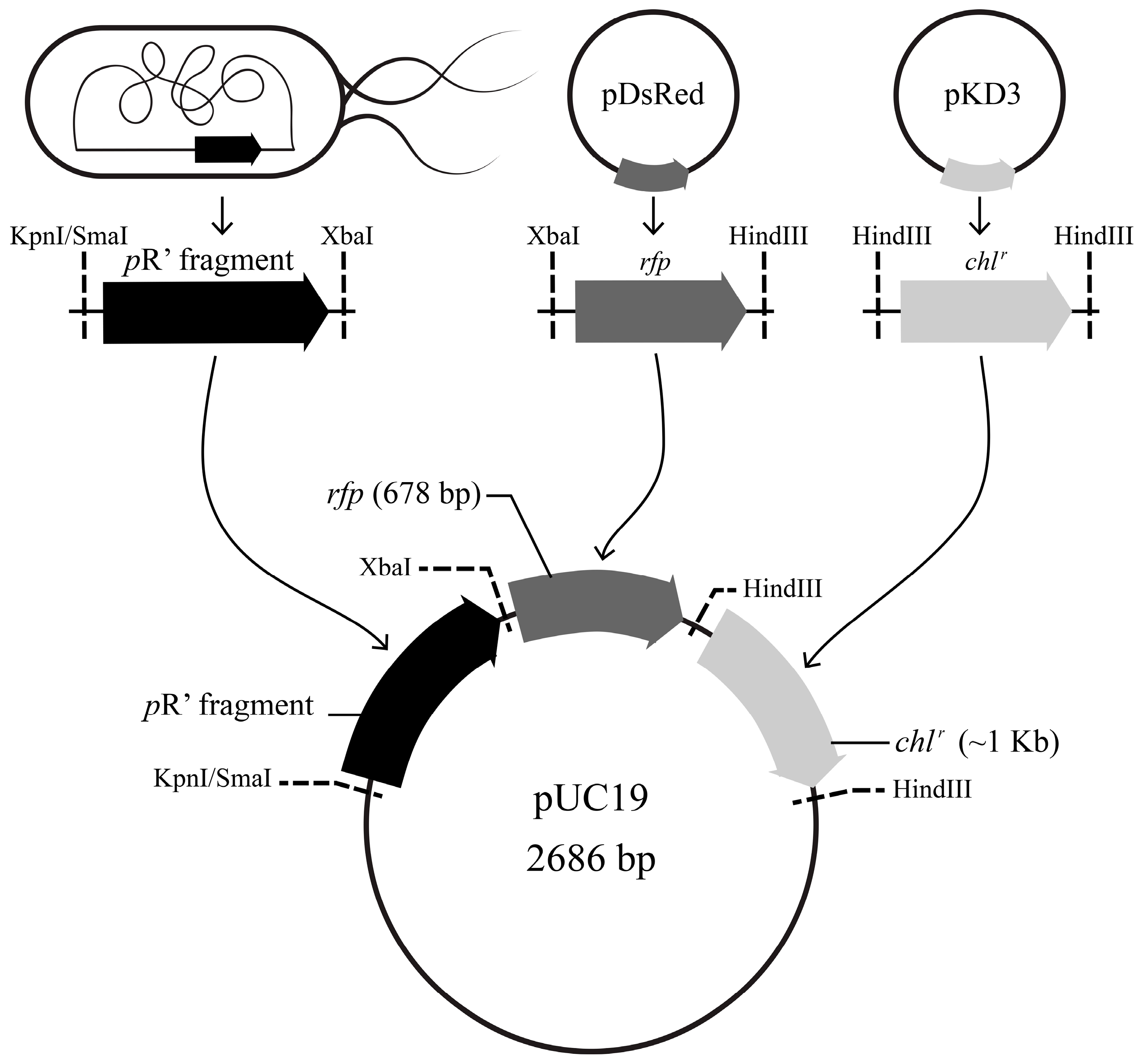
2.4. Construction and Validation of the pR’::rfp::chlr Transcriptional Fusion Reporter System
2.5. Determination of the Treatment Conditions for Flow Cytometry Detection
2.6. Flow Cytometry Detection of the Behavior of the pR’::rfp::chlr Constructs in Different Target Strains
2.7. Statistical Analysis
3. Results
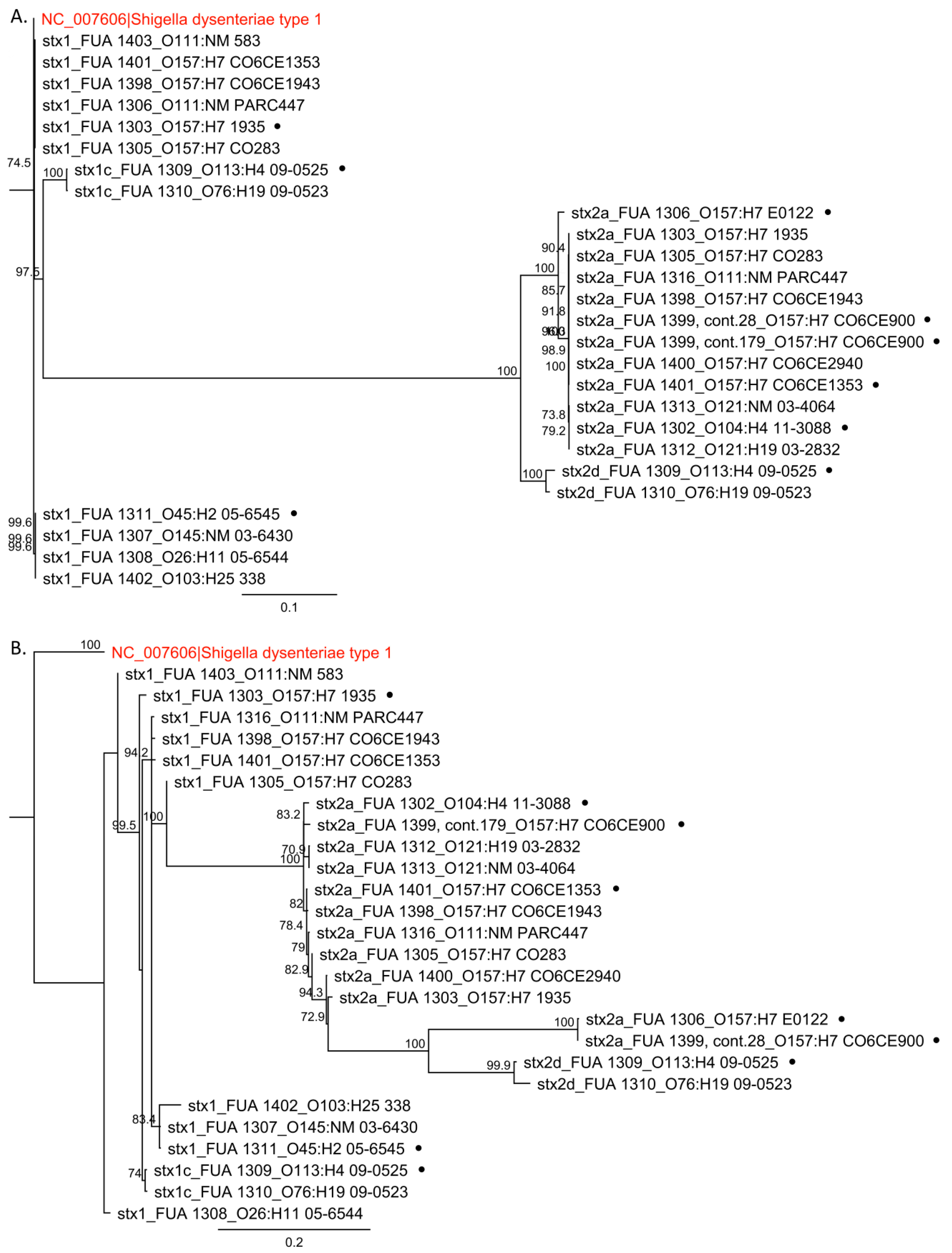
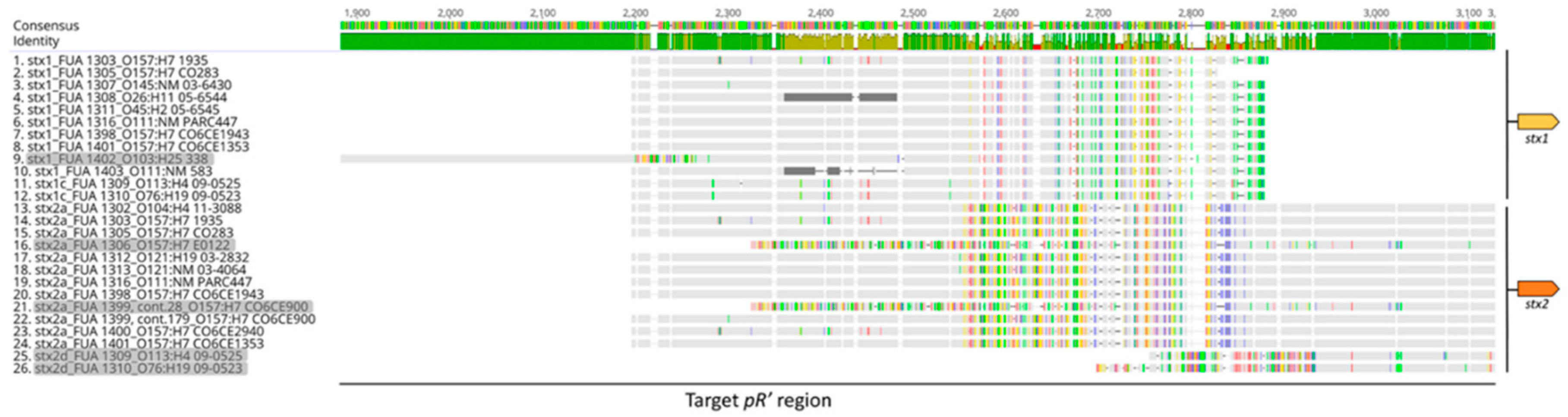
3.1. Sequence Alignment and Phylogenetic Analysis
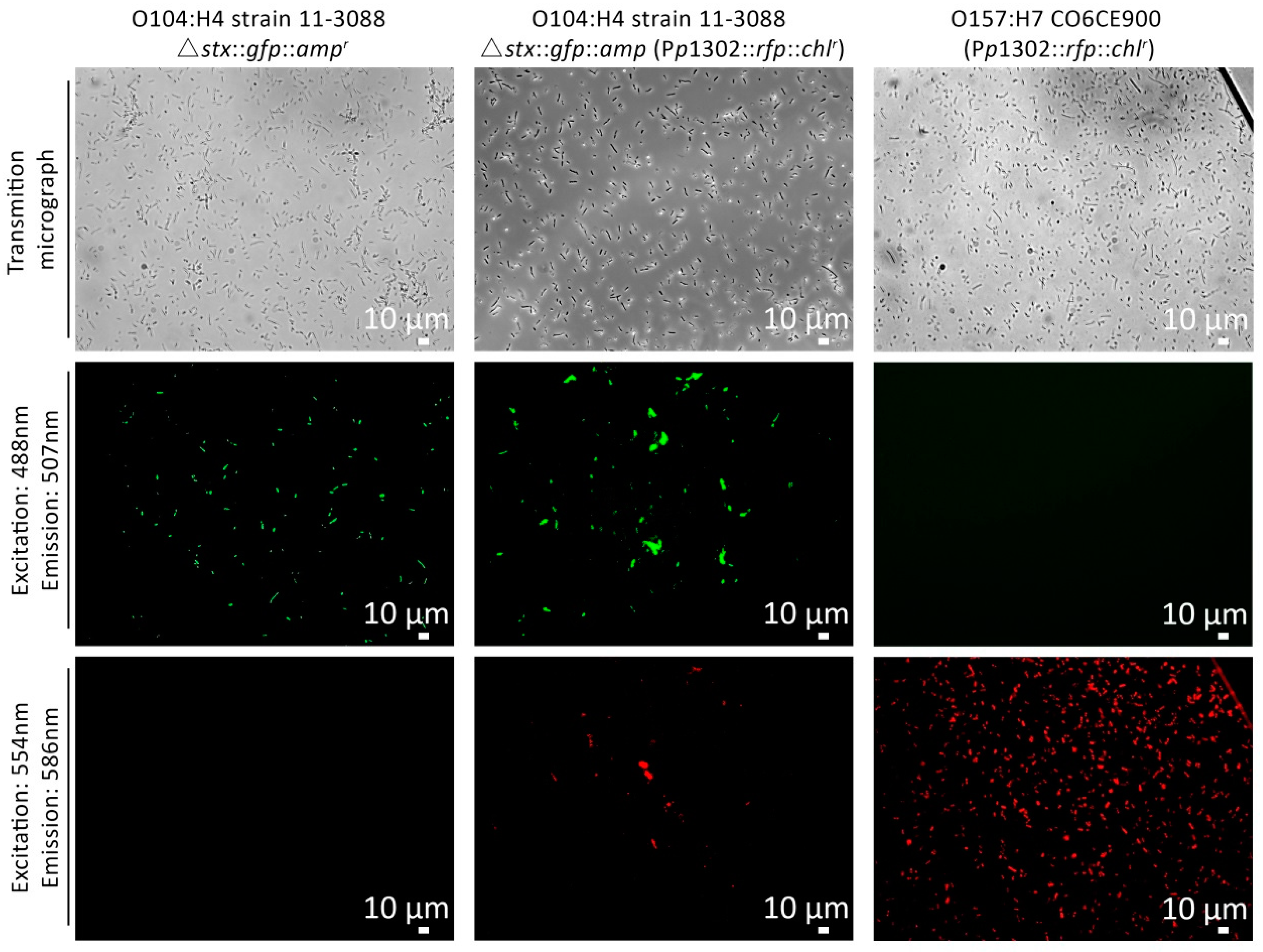
3.2. Construction and Validation of the pR’::rfp::chlr Transcriptional Fusion
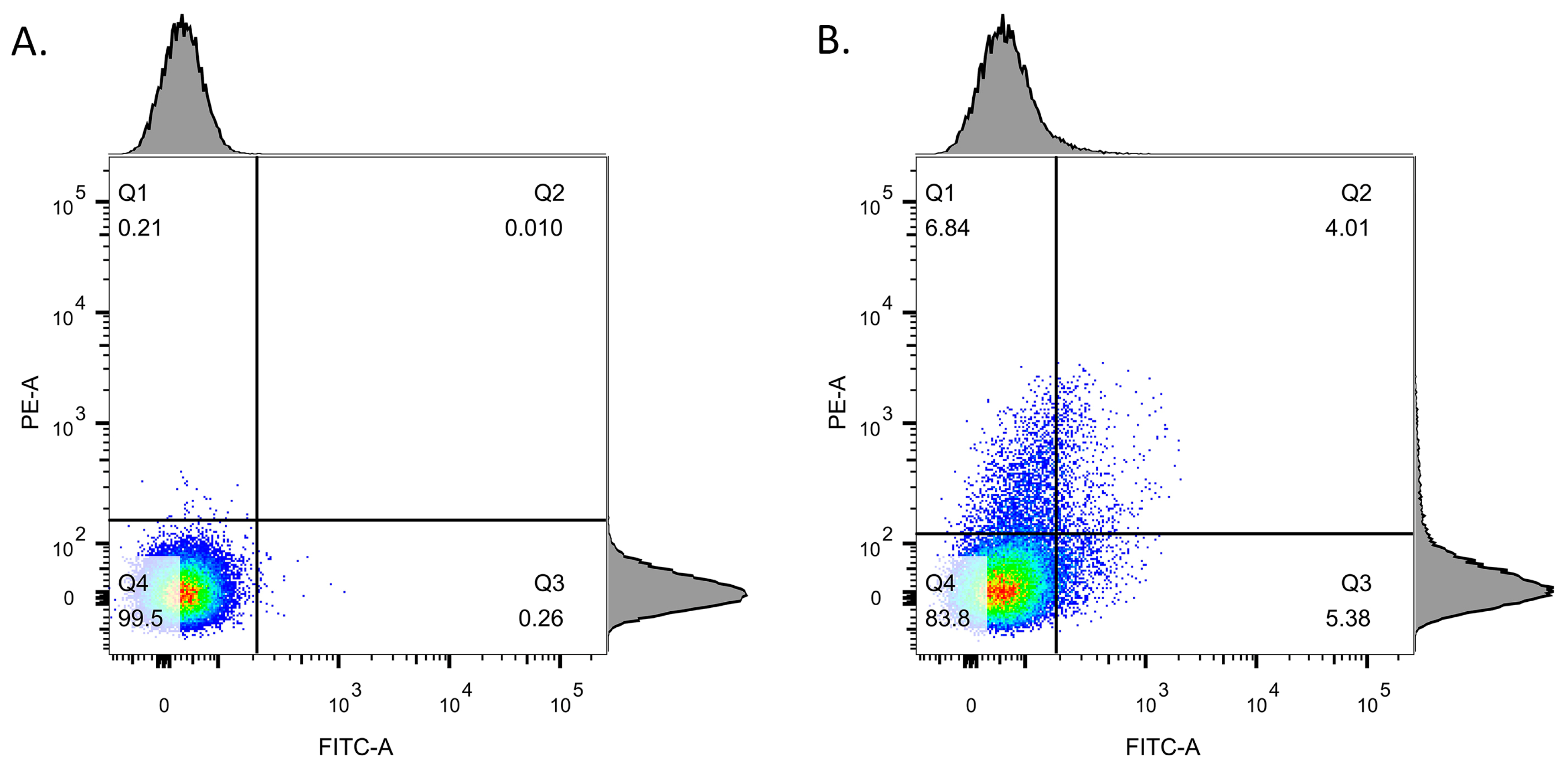
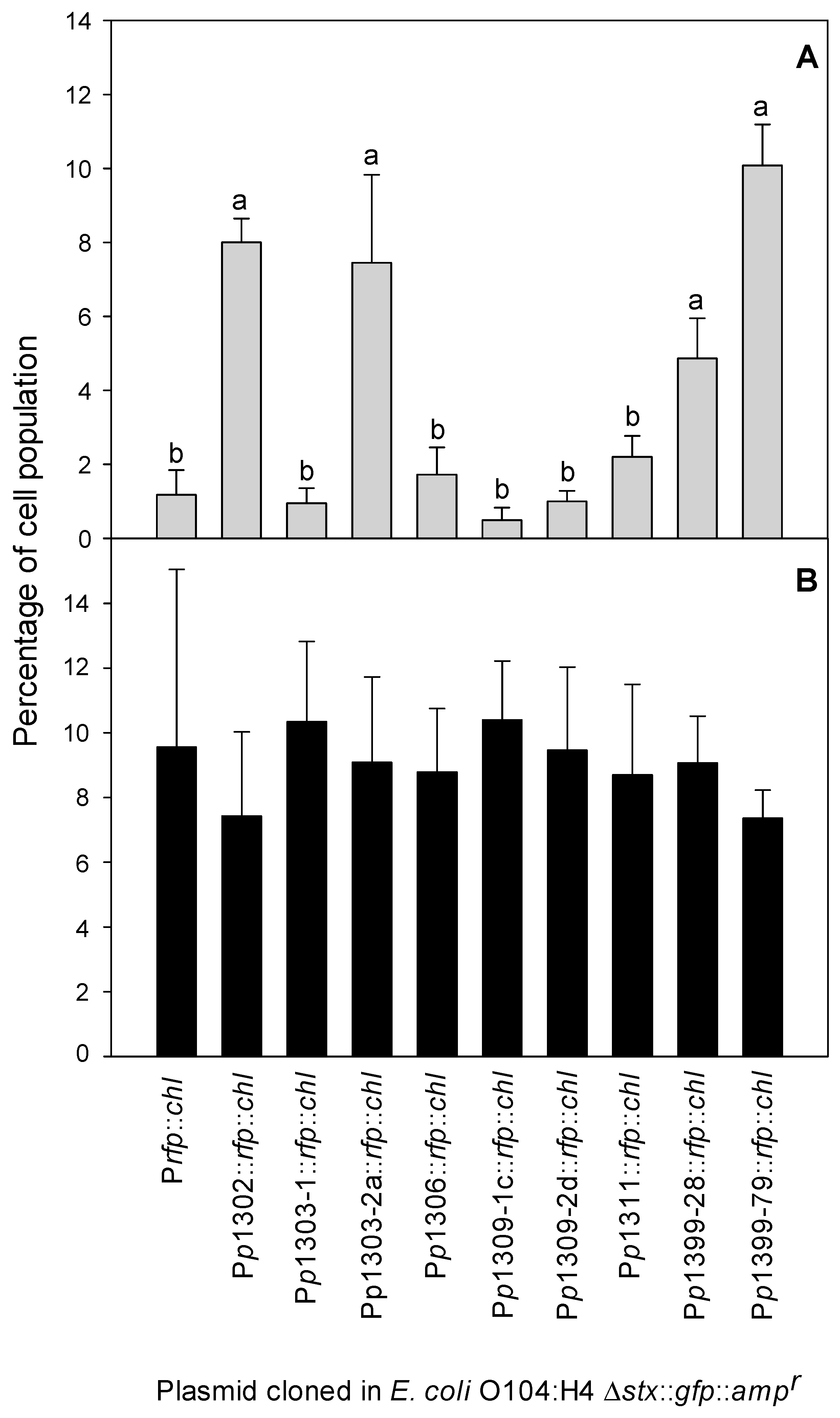
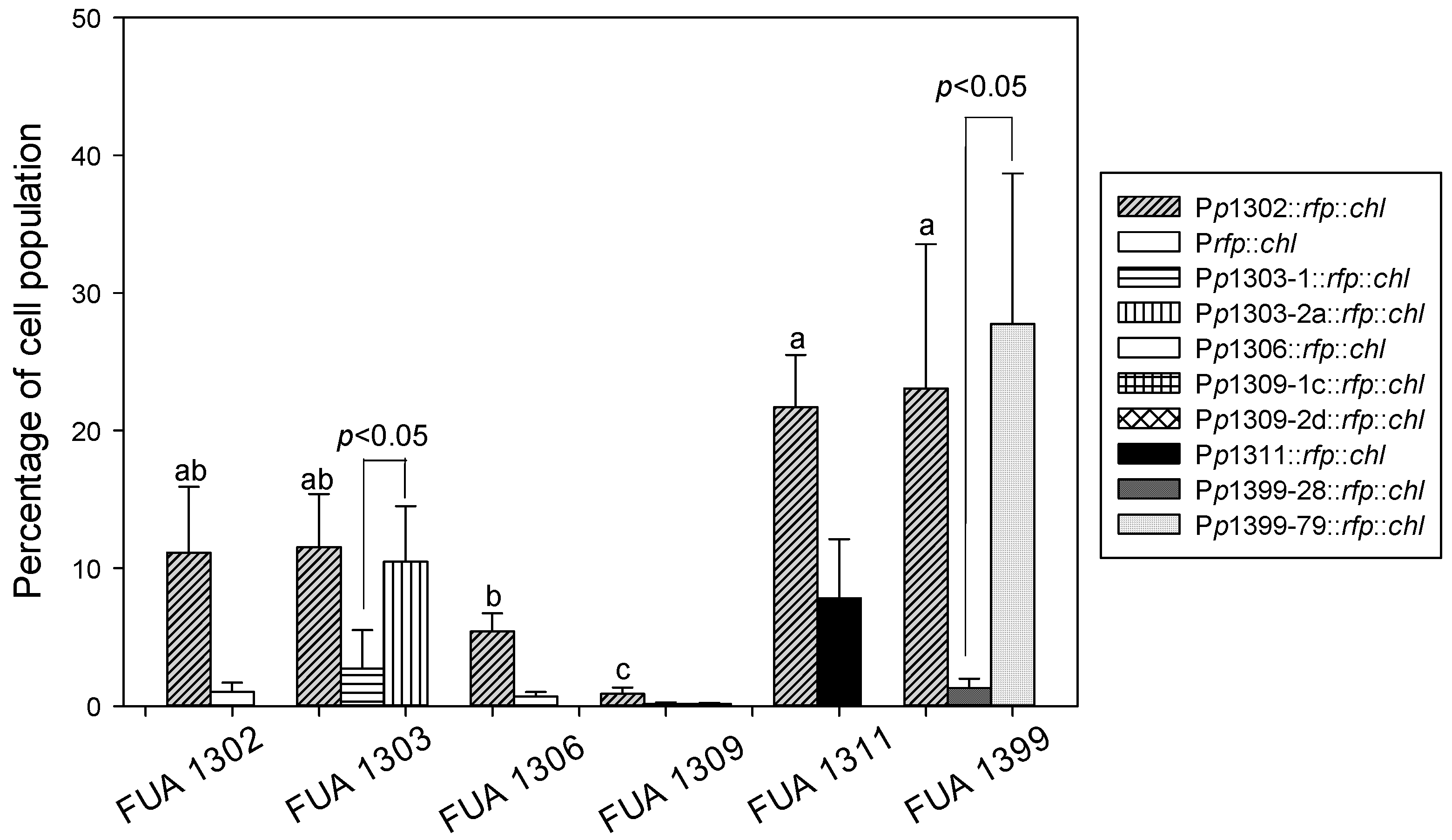
3.3. Detection of Stx Induction Levels in STEC Populations
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Van Elsas, J.D.; Semenov, A.V.; Costa, R.; Trevors, J.T. Survival of Escherichia coli in the environment: Fundamental and public health aspects. ISME J. 2011, 5, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Ochman, H.; Lawrence, J.G.; Groisman, E.A. Lateral gene transfer and the nature of bacterial innovation. Nature 2000, 405, 299–304. [Google Scholar] [CrossRef] [PubMed]
- Canchaya, C.; Fournous, G.; Chibani-Chennoufi, S.; Dillmann, M.-L.; Brüssow, H. Phage as agents of lateral gene transfer. Curr. Opin. Microbiol. 2003, 6, 417–424. [Google Scholar] [CrossRef]
- Blount, Z.D. The unexhausted potential of E. coli. eLife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- Fuhrman, J.A. Marine viruses and their biogeochemical and ecological effects. Nature 1999, 399, 541–548. [Google Scholar] [CrossRef] [PubMed]
- Erken, M.; Lutz, C.; McDougald, D. The rise of pathogens: predation as a factor driving the evolution of human pathogens in the environment. Microb. Ecol. 2013, 65, 860–868. [Google Scholar] [CrossRef] [PubMed]
- Łoś, J.M.; Łoś, M.; Węgrzyn, A.; Węgrzyn, G. Altruism of Shiga toxin-producing Escherichia coli: Recent hypothesis versus experimental results. Front. Cell. Infect. Microbiol. 2013, 2, 166. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.-S.; Koo, S.; Jeong, D.; Tesh, V.; Lee, M.-S.; Koo, S.; Jeong, D.G.; Tesh, V.L. Shiga toxins as multi-functional proteins: Induction of host cellular stress responses, role in pathogenesis and therapeutic applications. Toxins 2016, 8, 77. [Google Scholar] [CrossRef] [PubMed]
- Karmali, M.A.; Petric, M.; Lim, C.; Fleming, P.C.; Arbus, G.S.; Lior, H. The association between idiopathic hemolytic uremic syndrome and infection by Verotoxin-producing Escherichia coli. J. Infect. Dis. 1985, 151, 775–782. [Google Scholar] [CrossRef] [PubMed]
- Kaper, J.B.; Nataro, J.P.; Mobley, H.L.T. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2004, 2, 123–140. [Google Scholar] [CrossRef] [PubMed]
- Beutin, L.; Hammerl, J.A.; Strauch, E.; Reetz, J.; Dieckmann, R.; Kelner-Burgos, Y.; Martin, A.; Miko, A.; Strockbine, N.A.; Lindstedt, B.A.; et al. Spread of a distinct Stx2-encoding phage prototype among Escherichia coli O104:H4 strains from outbreaks in Germany, Norway, and Georgia. J. Virol. 2012, 86, 10444–10455. [Google Scholar] [CrossRef] [PubMed]
- Karch, H.; Denamur, E.; Dobrindt, U.; Finlay, B.B.; Hengge, R.; Johannes, L.; Ron, E.Z.; Tønjum, T.; Sansonetti, P.J.; Vicente, M. The enemy within us: Lessons from the 2011 European Escherichia coli O104:H4 outbreak. EMBO Mol. Med. 2012, 4, 841–848. [Google Scholar] [CrossRef] [PubMed]
- Bielaszewska, M.; Mellmann, A.; Zhang, W.; Köck, R.; Fruth, A.; Bauwens, A.; Peters, G.; Karch, H. Characterisation of the Escherichia coli strain associated with an outbreak of haemolytic uraemic syndrome in Germany, 2011: A microbiological study. Lancet Infect. Dis. 2011, 11, 671–676. [Google Scholar] [CrossRef]
- Steyert, S.R.; Sahl, J.W.; Fraser, C.M.; Teel, L.D.; Scheutz, F.; Rasko, D.A. Comparative genomics and stx phage characterization of LEE-negative Shiga toxin-producing Escherichia coli. Front. Cell. Infect. Microbiol. 2012, 2, 133. [Google Scholar] [CrossRef] [PubMed]
- Croxen, M.A.; Law, R.J.; Scholz, R.; Keeney, K.M.; Wlodarska, M.; Finlay, B.B. Recent advances in understanding enteric pathogenic Escherichia coli. Clin. Microbiol. Rev. 2013, 26, 822–880. [Google Scholar] [CrossRef] [PubMed]
- Endo, Y.; Tsurugi, K.; Yutsudo, T.; Takeda, Y.; Ogasawara, T.; Igarashi, K. Site of action of a Vero toxin (VT2) from Escherichia coli O157:H7 and of Shiga toxin on eukaryotic ribosomes. RNA N-glycosidase activity of the toxins. Eur. J. Biochem. 1988, 171, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Saxena, S.K.; O’Brien, A.D.; Ackerman, E.J. Shiga toxin, Shiga-like toxin II variant, and ricin are all single-site RNA N-glycosidases of 28 S RNA when microinjected into Xenopus oocytes. J. Biol. Chem. 1989, 264, 596–601. [Google Scholar] [PubMed]
- Tesh, V.L. Induction of apoptosis by Shiga toxins. Futur. Microbiol. 2010, 5, 431–453. [Google Scholar] [CrossRef] [PubMed]
- Fuller, C.A.; Pellino, C.A.; Flagler, M.J.; Strasser, J.E.; Weiss, A.A. Shiga toxin subtypes display dramatic differences in potency. Infect. Immun. 2011, 79, 1329–1337. [Google Scholar] [CrossRef] [PubMed]
- Wagner, P.L.; Neely, M.N.; Zhang, X.; Acheson, D.W.K.; Waldor, M.K.; Friedman, D.I.; Acheson, W.K.; Waldor, M.K.; Friedman, D.I.; Acheson, D.W.K. Role for a phage promoter in Shiga toxin 2 expression from a pathogenic Escherichia coli strain. J. Bacteriol. 2001, 183, 2081–2085. [Google Scholar] [CrossRef] [PubMed]
- Yarnell, W.S.; Roberts, J.W. The phage λ gene Q transcription antiterminator binds DNA in the late gene promoter as it modifies RNA polymerase. Cell 1992, 69, 1181–1189. [Google Scholar] [CrossRef]
- Neely, M.N.; Friedman, D.I. Functional and genetic analysis of regulatory regions of coliphage H-19B: Location of shiga-like toxin and lysis genes suggest a role for phage functions in toxin release. Mol. Microbiol. 1998, 28, 1255–1267. [Google Scholar] [CrossRef] [PubMed]
- Plunkett, G.; Rose, D.J.; Durfee, T.J.; Blattner, F.R. Sequence of Shiga toxin 2 phage 933W from Escherichia coli O157:H7: Shiga toxin as a phage late-gene product. J. Bacteriol. 1999, 181, 1767–1778. [Google Scholar] [PubMed]
- Haugum, K.; Lindstedt, B.-A.; Løbersli, I.; Kapperud, G.; Brandal, L.T. Identification of the anti-terminator qO111:H− gene in Norwegian sorbitol-fermenting Escherichia coli O157:NM. FEMS Microbiol. Lett. 2012, 329, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Lejeune, J.T.; Abedon, S.T.; Takemura, K.; Christie, N.P.; Sreevatsan, S. Human Escherichia coli O157:H7 genetic marker in isolates of bovine origin. Emerg. Infect. Dis. 2004, 10, 1482–1485. [Google Scholar] [CrossRef] [PubMed]
- Olavesen, K.K.; Lindstedt, B.-A.; Løbersli, I.; Brandal, L.T. Expression of Shiga toxin 2 (Stx2) in highly virulent Stx-producing Escherichia coli (STEC) carrying different anti-terminator (q) genes. Microb. Pathog. 2016, 97, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Ogura, Y.; Mondal, S.I.; Islam, M.R.; Mako, T.; Arisawa, K.; Katsura, K.; Ooka, T.; Gotoh, Y.; Murase, K.; Ohnishi, M.; et al. The Shiga toxin 2 production level in enterohemorrhagic Escherichia coli O157:H7 is correlated with the subtypes of toxin-encoding phage. Sci. Rep. 2015, 5, 16663. [Google Scholar] [CrossRef] [PubMed]
- Krüger, A.; Lucchesi, P.M.A. Shiga toxins and stx phages: Highly diverse entities. Microbiology 2015, 161, 451–462. [Google Scholar] [CrossRef] [PubMed]
- Yin, S.; Rusconi, B.; Sanjar, F.; Goswami, K.; Xiaoli, L.; Eppinger, M.; Dudley, E.G. Escherichia coli O157:H7 strains harbor at least three distinct sequence types of Shiga toxin 2a-converting phages. BMC Genom. 2015, 16, 733. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.L.; Rooks, D.J.; Fogg, P.C.M.; Darby, A.C.; Thomson, N.R.; McCarthy, A.J.; Allison, H.E. Comparative genomics of Shiga toxin encoding bacteriophages. BMC Genom. 2012, 13, 311. [Google Scholar] [CrossRef] [PubMed]
- Livny, J.; Friedman, D.I. Characterizing spontaneous induction of Stx encoding phages using a selectable reporter system. Mol. Microbiol. 2004, 51, 1691–1704. [Google Scholar] [CrossRef] [PubMed]
- Mercer, R.G.; Zheng, J.; Garcia-Hernandez, R.; Ruan, L.; Gänzle, M.G.; McMullen, L.M.; Allard, M.W.; Gänzle, M.G.; McMullen, L.M. Genetic determinants of heat resistance in Escherichia coli. Front. Microbiol. 2015, 6, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, H.M. Microbial analysis at the single-cell level: Tasks and techniques. J. Microbiol. Methods 2000, 42, 3–16. [Google Scholar] [CrossRef]
- Fang, Y.; Mercer, R.G.; McMullen, L.M.; Gänzle, M.G. Induction of Shiga toxin-encoding prophage by abiotic environmental stress in food. Appl. Environ. Microbiol. 2017, 83, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Datsenko, K.A.; Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 2000, 97, 6640–6645. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Gill, A.; McmMullen, L.; Gänzle, M.G. Variation in heat and pressure resistance of Verotoxigenic and nontoxigenic Escherichia coli. J. Food Prot. 2015, 78, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.L.; Wareing, B.M.; Fogg, P.C.M.M.; Riley, L.M.; Spencer, M.; Cox, M.J.; Saunders, J.R.; McCarthy, A.J.; Allison, H.E. Multilocus characterization scheme for Shiga toxin-encoding bacteriophages. Appl. Environ. Microbiol. 2007, 73. [Google Scholar] [CrossRef] [PubMed]
- Casjens, S.R.; Hendrix, R.W. Bacteriophage lambda: Early pioneer and still relevant. Virology 2015, 479–480, 310–330. [Google Scholar] [CrossRef] [PubMed]
- Manning, S.D.; Motiwala, A.S.; Springman, A.C.; Qi, W.; Lacher, D.W.; Ouellette, L.M.; Mladonicky, J.M.; Somsel, P.; Rudrik, J.T.; Dietrich, S.E.; et al. Variation in virulence among clades of Escherichia coli O157:H7 associated with disease outbreaks. Proc. Natl. Acad. Sci. USA 2008, 105, 4868–4873. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, K.; Makino, K.; Kubota, Y.; Watanabe, M.; Kimura, S.; Yutsudo, C.H.; Kurokawa, K.; Ishii, K.; Hattori, M.; Tatsuno, I.; et al. Complete nucleotide sequence of the prophage VT1-Sakai carrying the Shiga toxin 1 genes of the enterohemorrhagic Escherichia coli O157:H7 strain derived from the Sakai outbreak. Gene 2000, 258, 127–139. [Google Scholar] [CrossRef]
- Wegrzyn, G.; Licznerska, K.; Wegrzyn, A. Phage λ-New Insights into Regulatory Circuits. Adv. Virus Res. 2012, 82, 155–178. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, M.; Terajima, J.; Kurokawa, K.; Nakayama, K.; Murata, T.; Tamura, K.; Ogura, Y.; Watanabe, H.; Hayashi, T. Genomic diversity of enterohemorrhagic Escherichia coli O157 revealed by whole genome PCR scanning. Proc. Natl. Acad. Sci. USA 2002, 99, 17043–17048. [Google Scholar] [CrossRef] [PubMed]
- Muniesa, M.; Blanco, J.E.; de Simón, M.; Serra-Moreno, R.; Blanch, A.R.; Jofre, J. Diversity of stx2 converting bacteriophages induced from Shiga-toxin-producing Escherichia coli strains isolated from cattle. Microbiology 2004, 150, 2959–2971. [Google Scholar] [CrossRef] [PubMed]
- Boyd, E.F.; Brüssow, H. Common themes among bacteriophage-encoded virulence factors and diversity among the bacteriophages involved. Trends Microbiol. 2002, 10, 521–529. [Google Scholar] [CrossRef]
- Miyamoto, H.; Nakai, W.; Yajima, N.; Fujibayashi, A.; Higuchi, T.; Sato, K.; Matsushiro, A. Sequence analysis of Stx2-converting phage VT2-Sa shows a great divergence in early regulation and replication regions. DNA Res. 1999, 6, 235–240. [Google Scholar] [CrossRef] [PubMed]
- Wagner, P.L.; Acheson, D.W.; Waldor, M.K. Isogenic lysogens of diverse shiga toxin 2-encoding bacteriophages produce markedly different amounts of shiga toxin. Infect. Immun. 1999, 67, 6710–6714. [Google Scholar] [PubMed]
- Tyler, J.S.; Mills, M.J.; Friedman, D.I. The operator and early promoter region of the Shiga toxin type 2-encoding bacteriophage 933W and control of toxin expression. J. Bacteriol. 2004, 186. [Google Scholar] [CrossRef] [PubMed]
- Muniesa, M.; de Simon, M.; Prats, G.; Ferrer, D.; Pañella, H.; Jofre, J. Shiga toxin 2-converting bacteriophages associated with clonal variability in Escherichia coli O157:H7 strains of human origin isolated from a single outbreak. Infect. Immun. 2003, 71, 4554–4562. [Google Scholar] [CrossRef] [PubMed]
- Phage, T.; Gamage, S.D.; Patton, A.K.; Hanson, J.F.; Weiss, A.A. Diversity and host range of Shiga diversity and host range of Shiga toxin-encoding phage. Society 2004, 72, 7131–7139. [Google Scholar] [CrossRef]
- Hao, W.; Allen, V.G.; Jamieson, F.B.; Low, D.E.; Alexander, D.C. Phylogenetic incongruence in E. coli O104: understanding the evolutionary relationships of emerging pathogens in the face of homologous recombination. PLoS ONE 2012, 7, e33971. [Google Scholar] [CrossRef] [PubMed]
- Łoś, J.M.; Łoś, M.; Węgrzyn, G.; Węgrzyn, A.; Joanna, M.; Alicja, W.; Łoś, J.M.; Łoś, M.; Węgrzyn, G.; Węgrzyn, A. Differential efficiency of induction of various lambdoid prophages responsible for production of Shiga toxins in response to different induction agents. Microb. Pathog. 2009, 47, 289–298. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, T.; Ohta, Y.; Tsutsuki, H.; Noda, M. Construction of a novel bioluminescent reporter system for investigating Shiga toxin expression of enterohemorrhagic Escherichia coli. Gene 2011, 478, 1–10. [Google Scholar] [CrossRef] [PubMed]
- King, O.D.; Masel, J. The evolution of bet-hedging adaptations to rare scenarios. Theor. Popul. Biol. 2007, 72, 560–575. [Google Scholar] [CrossRef] [PubMed]
- Koitabashi, T.; Vuddhakul, V.; Radu, S.; Morigaki, T.; Asai, N.; Nakaguchi, Y.; Nishibuchi, M. Genetic characterization of Escherichia coli O157:H7/- strains carrying the stx2 gene but not producing Shiga toxin 2. Microbiol. Immunol. 2006, 50, 135–148. [Google Scholar] [CrossRef] [PubMed]
- Vorobiev, S.M.; Gensler, Y.; Vahedian-Movahed, H.; Seetharaman, J.; Su, M.; Huang, J.Y.; Xiao, R.; Kornhaber, G.; Montelione, G.T.; Tong, L.; et al. Structure of the DNA-binding and RNA-polymerase-binding region of transcription antitermination factor λQ. Structure 2014, 22, 488–495. [Google Scholar] [CrossRef] [PubMed]
- Fogg, P.C.M.; Saunders, J.R.; McCarthy, A.J.; Allison, H.E. Cumulative effect of prophage burden on Shiga toxin production in Escherichia coli. Microbiology 2012, 158, 488–497. [Google Scholar] [CrossRef] [PubMed]
- Krüger, A.; Lucchesi, P.M.A.; Parma, A.E. Verotoxins in bovine and meat verotoxin-producing Escherichia coli isolates: Type, number of variants, and relationship to cytotoxicity. Appl. Environ. Microbiol. 2011, 77, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Eppinger, M.; Mammel, M.K.; Leclerc, J.E.; Ravel, J.; Cebula, T.A. Genomic anatomy of Escherichia coli O157:H7 outbreaks. Proc. Natl. Acad. Sci. USA 2011, 108, 20142–20147. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, A.D.; Newland, J.W.; Miller, S.F.; Holmes, R.K.; Smith, H.W.; Formal, S.B. Shiga-like toxin-converting phages from Escherichia coli strains that cause hemorrhagic colitis or infantile diarrhea. Science 1984, 226, 694–696. [Google Scholar] [CrossRef]
- Chang, D.-E.; Smalley, D.J.; Tucker, D.L.; Leatham, M.P.; Norris, W.E.; Stevenson, S.J.; Anderson, A.B.; Grissom, J.E.; Laux, D.C.; Cohen, P.S.; et al. Carbon nutrition of Escherichia coli in the mouse intestine. Proc. Natl. Acad. Sci. USA 2004, 101, 7427–7432. [Google Scholar] [CrossRef] [PubMed]
- Yatsuyanagi, J.; Saito, S.; Ito, I. A case of hemolytic-uremic syndrome associated with shiga toxin 2-producing Escherichia coli O121 infection caused by drinking water contaminated with bovine feces. Jpn. J. Infect. Dis. 2002, 55, 174–176. [Google Scholar] [PubMed]
- Eurosurveillance editorial team. The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2011 has been published. Eurosurveillance 2013, 18, 20449. [Google Scholar] [CrossRef]
- Newbold, C.J.; de la Fuente, G.; Belanche, A.; Ramos-Morales, E.; McEwan, N.R. The role of ciliate protozoa in the rumen. Front. Microbiol. 2015, 6, 1313. [Google Scholar] [CrossRef] [PubMed]
| Accession Numbers | Strains and Serotype | FUA Number Used for Plasmid Nomenclature | Description | Ref. |
|---|---|---|---|---|
| LDYN00000000 | E. coli O26:H11 05-6544 | 1308 | stx1 | [32] |
| LDZZ00000000 | E. coli O121:H19 03-2832 | 1312 | stx2a | [32] |
| LEAA00000000 | E. coli O121:NM 03-4064 | 1313 | stx2a | [32] |
| LEAB00000000 | E. coli O145:NM 03-6430 | 1307 | stx1 | [32] |
| LEAD00000000 | E. coli O157:H7 1935 | 1303 | stx1 stx2a | [32] |
| LEAE00000000 | E. coli O157:H7 CO6CE900 | 1399 | stx2a | [32] |
| LEAF00000000 | E. coli O157:H7 CO6CE1353 | 1401 | stx1 stx2a | [32] |
| LEAG00000000 | E. coli O157:H7 CO6CE1943 | 1398 | stx1 stx2a | [32] |
| LEAH00000000 | E. coli O157:H7 CO6CE2940 | 1400 | stx2a | [32] |
| LEAI00000000 | E. coli O157:H7 CO283 | 1305 | stx1 stx2a | [32] |
| LEAJ00000000 | E. coli O157:H7 E0122 | 1306 | stx2a | [32] |
| LECF00000000 | E. coli O103:H25 338 | 1402 | stx1 | [32] |
| LECH00000000 | E. coli O104:H4 11-3088 | 1302 | stx2a | [32] |
| LECI00000000 | E. coli O111:NM 583 | 1403 | stx1 | [32] |
| LECJ00000000 | E. coli O111:NM PARC447 | 1316 | stx1 stx2 | [32] |
| LECK00000000 | E. coli O113:H4 09-0525 | 1309 | stx1c stx2d | [32] |
| LECM00000000 | E. coli O45:H2 05-6545 | 1311 | stx1 | [32] |
| LECN00000000 | E. coli O76:H19 09-0523 | 1310 | stx1c stx2d | [32] |
| E. coli DH5α | ||||
| E. coli Top10 | pUC19 | |||
| E. coli Top10 | pRFP | |||
| E. coli O104:H4 11-3088 Δstx::gfp::amp | stx gene replaced with gfp | [34] |
| Primer | Sequence (5′-3′) a) | Restriction Site |
|---|---|---|
| LP F1-1 | 5′-CGGGAAGGTACCACCTCTGTATTTTATCAG-3′ | KpnI |
| LP R1-3 | 5′-GGGCCGTCTAGAAAAGAAAAAAGTTAGCAC-3′ | XbaI |
| LP F2-2 | 5′-ATTAGTCCCGGGCTTGGATTTATTGATGGT-3′ | SmaI |
| LP R3-2 | 5′-ATAACGTCTAGATAACAGGCACAGTACCCA-3′ | XbaI |
| LP F3-2 | 5′-AGCGGTACCAAAAACCGGAAACGTGTA-3′ | KpnI |
| LP F4-1 | 5′-TGCGTAGGTACCAGCGTCTATAATTGTATG-3′ | KpnI |
| LP R4-2 | 5′-GCATTATCTAGACAACAGGCACAGTATCCA-3′ | XbaI |
| RFP F-2 | 5′-CTGATATCTAGAATGGCCTCCTCCGAG-3′ | XbaI |
| RFP R-5 | 5′-ATCTGTAAGCTTCTACAGGAACAGGTGGT-3′ | HindIII |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, L.X.; Simpson, D.J.; McMullen, L.M.; Gänzle, M.G. Comparative Genomics and Characterization of the Late Promoter pR’ from Shiga Toxin Prophages in Escherichia coli. Viruses 2018, 10, 595. https://doi.org/10.3390/v10110595
Zhang LX, Simpson DJ, McMullen LM, Gänzle MG. Comparative Genomics and Characterization of the Late Promoter pR’ from Shiga Toxin Prophages in Escherichia coli. Viruses. 2018; 10(11):595. https://doi.org/10.3390/v10110595
Chicago/Turabian StyleZhang, Ling Xiao, David J. Simpson, Lynn M. McMullen, and Michael G. Gänzle. 2018. "Comparative Genomics and Characterization of the Late Promoter pR’ from Shiga Toxin Prophages in Escherichia coli" Viruses 10, no. 11: 595. https://doi.org/10.3390/v10110595
APA StyleZhang, L. X., Simpson, D. J., McMullen, L. M., & Gänzle, M. G. (2018). Comparative Genomics and Characterization of the Late Promoter pR’ from Shiga Toxin Prophages in Escherichia coli. Viruses, 10(11), 595. https://doi.org/10.3390/v10110595






