Fetal Brain Infection Is Not a Unique Characteristic of Brazilian Zika Viruses
Abstract
:1. Introduction
2. Materials and Methods
2.1. Generation of ZIKV-MY and Chimeric Virus Using Circular Polymerase Extension Reaction (CPER)
2.2. Growth Kinetics
2.3. Mice and ZIKV Infection
2.4. Ethics Statement
2.5. ZIKV CCID50 Assays
2.6. Statistics
3. Results
3.1. De Novo Generation and Characterization of ZIKV-MY
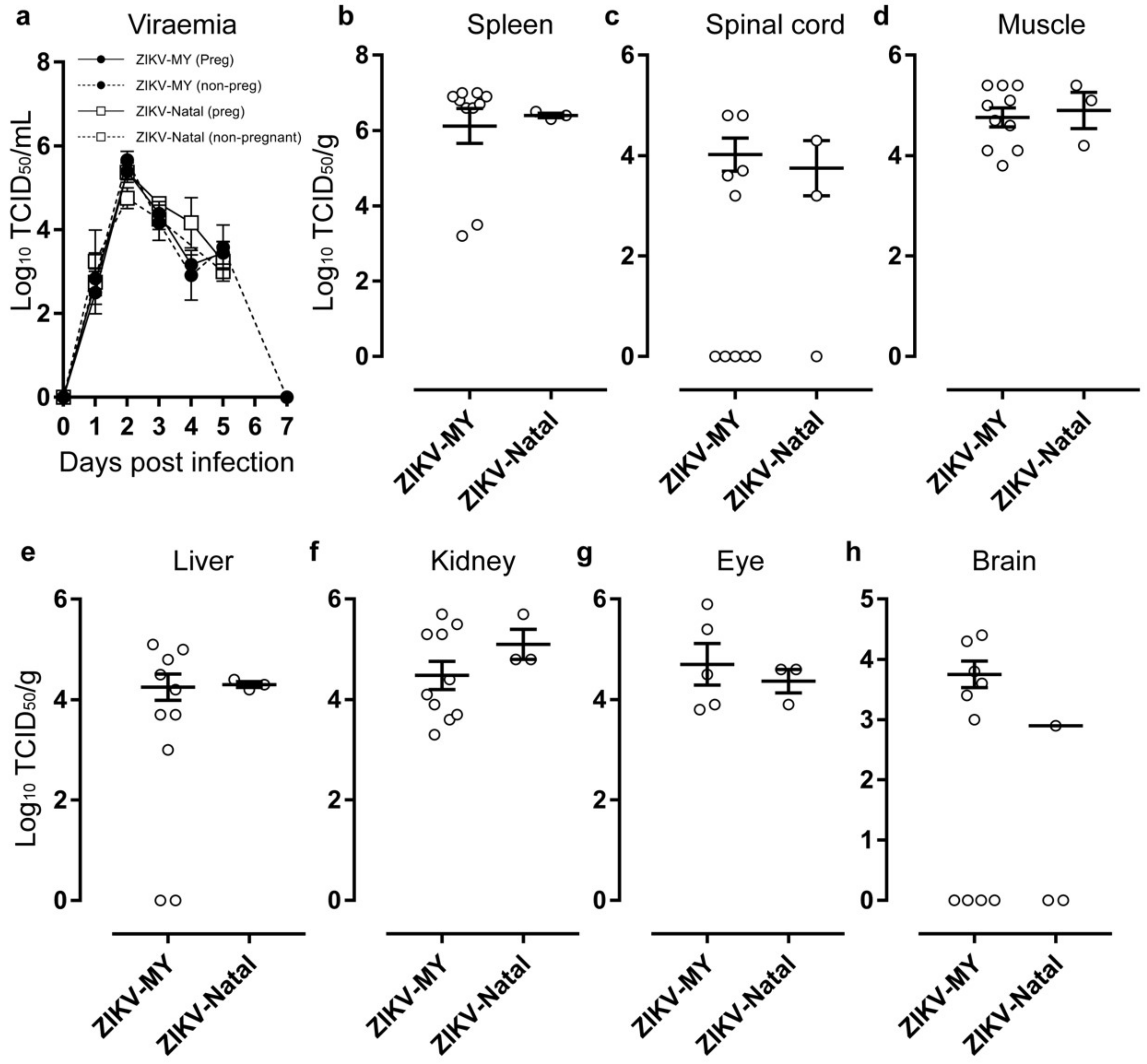
3.2. ZIKV-MY and ZIKV-Natal Both Infect IFNAR1−/− Mice and Produce Similar Viraemia and Viral Titres in Different Organs
3.3. ZIKV-MY and ZIKV-Natal Infection of Placenta and Foetal Heads in Pregnant IFNAR1−/− Mice
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kuno, G.; Chang, G.J. Full-length sequencing and genomic characterization of Bagaza, Kedougou, and Zika viruses. Arch. Virol. 2007, 152, 687–696. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wikan, N.; Smith, D.R. Zika virus: History of a newly emerging arbovirus. Lancet Infect. Dis. 2016, 16, e119–e126. [Google Scholar] [CrossRef]
- Roby, J.A.; Setoh, Y.X.; Hall, R.A.; Khromykh, A.A. Post-translational regulation and modifications of flavivirus structural proteins. J. Gen. Virol. 2015, 96, 1551–1569. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lanciotti, R.S.; Kosoy, O.L.; Laven, J.J.; Velez, J.O.; Lambert, A.J.; Johnson, A.J.; Stanfield, S.M.; Duffy, M.R. Genetic and serologic properties of Zika virus associated with an epidemic, yap state, Micronesia, 2007. Emerg. Infect. Dis. 2008, 14, 1232–1239. [Google Scholar] [CrossRef] [PubMed]
- Faye, O.; Freire, C.C.; Iamarino, A.; Faye, O.; de Oliveira, J.V.; Diallo, M.; Zanotto, P.M.; Sall, A.A. Molecular evolution of Zika virus during its emergence in the 20(th) century. PLoS Negl. Trop. Dis. 2014, 8, e2636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gatherer, D.; Kohl, A. Zika virus: A previously slow pandemic spreads rapidly through the Americas. J. Gen. Virol. 2016, 97, 269–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Faria, N.R.; Azevedo Rdo, S.; Kraemer, M.U.; Souza, R.; Cunha, M.S.; Hill, S.C.; Theze, J.; Bonsall, M.B.; Bowden, T.A.; Rissanen, I.; et al. Zika virus in the Americas: Early epidemiological and genetic findings. Science 2016, 352, 345–349. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Valderramos, S.G.; Wu, A.; Ouyang, S.; Li, C.; Brasil, P.; Bonaldo, M.; Coates, T.; Nielsen-Saines, K.; Jiang, T.; et al. From mosquitos to humans: Genetic evolution of Zika virus. Cell Host Microbe 2016, 19, 561–565. [Google Scholar] [CrossRef] [PubMed]
- Pettersson, J.H.; Eldholm, V.; Seligman, S.J.; Lundkvist, A.; Falconar, A.K.; Gaunt, M.W.; Musso, D.; Nougairede, A.; Charrel, R.; Gould, E.A.; et al. How did Zika virus emerge in the pacific islands and Latin America? MBio 2016, 7, e01239-16. [Google Scholar] [CrossRef] [PubMed]
- Weaver, S.C. Emergence of epidemic Zika virus transmission and congenital Zika syndrome: Are recently evolved traits to blame? MBio 2017, 8, e02063-16. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, S.; Starmer, W.T. Possible roles of new mutations shared by Asian and American Zika viruses. Mol. Biol. Evol. 2017, 34, 525–534. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, J.; Du, S.; Shan, C.; Nie, K.; Zhang, R.; Li, X.F.; Zhang, R.; Wang, T.; Qin, C.F.; et al. Evolutionary enhancement of Zika virus infectivity in Aedes aegypti mosquitoes. Nature 2017, 545, 482–486. [Google Scholar] [CrossRef] [PubMed]
- Seferovic, M.; Martin, C.S.; Tardif, S.D.; Rutherford, J.; Castro, E.C.C.; Li, T.; Hodara, V.L.; Parodi, L.M.; Giavedoni, L.; Layne-Colon, D.; et al. Experimental Zika virus infection in the pregnant common marmoset induces spontaneous fetal loss and neurodevelopmental abnormalities. Sci. Rep. 2018, 8, 6851. [Google Scholar] [CrossRef] [PubMed]
- Mlakar, J.; Korva, M.; Tul, N.; Popovic, M.; Poljsak-Prijatelj, M.; Mraz, J.; Kolenc, M.; Resman Rus, K.; Vesnaver Vipotnik, T.; Fabjan Vodusek, V.; et al. Zika virus associated with microcephaly. N. Engl. J. Med. 2016, 374, 951–958. [Google Scholar] [CrossRef] [PubMed]
- Ventura, C.V.; Maia, M.; Bravo-Filho, V.; Gois, A.L.; Belfort, R., Jr. Zika virus in Brazil and macular atrophy in a child with microcephaly. Lancet 2016, 387, 228. [Google Scholar] [CrossRef]
- Cauchemez, S.; Besnard, M.; Bompard, P.; Dub, T.; Guillemette-Artur, P.; Eyrolle-Guignot, D.; Salje, H.; Van Kerkhove, M.D.; Abadie, V.; Garel, C.; et al. Association between zika virus and microcephaly in French polynesia, 2013–15: A retrospective study. Lancet 2016, 387, 2125–2132. [Google Scholar] [CrossRef]
- Simonin, Y.; van Riel, D.; Van de Perre, P.; Rockx, B.; Salinas, S. Differential virulence between Asian and African lineages of Zika virus. PLoS Negl. Trop. Dis. 2017, 11, e0005821. [Google Scholar] [CrossRef] [PubMed]
- Cugola, F.R.; Fernandes, I.R.; Russo, F.B.; Freitas, B.C.; Dias, J.L.; Guimaraes, K.P.; Benazzato, C.; Almeida, N.; Pignatari, G.C.; Romero, S.; et al. The Brazilian Zika virus strain causes birth defects in experimental models. Nature 2016, 534, 267–271. [Google Scholar] [CrossRef] [PubMed]
- Xia, H.; Luo, H.; Shan, C.; Muruato, A.E.; Nunes, B.T.D.; Medeiros, D.B.A.; Zou, J.; Xie, X.; Giraldo, M.I.; Vasconcelos, P.F.C.; et al. An evolutionary ns1 mutation enhances Zika virus evasion of host interferon induction. Nat. Commun. 2018, 9, 414. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.R.; Sprague, T.R.; Hollidge, B.S.; Valdez, S.M.; Padilla, S.L.; Bellanca, S.A.; Golden, J.W.; Coyne, S.R.; Kulesh, D.A.; Miller, L.J.; et al. African and Asian Zika virus isolates display phenotypic differences both in vitro and in vivo. Am. J. Trop. Med. Hyg. 2018, 98, 432–444. [Google Scholar] [CrossRef] [PubMed]
- Rosenfeld, A.B.; Doobin, D.J.; Warren, A.L.; Racaniello, V.R.; Vallee, R.B. Replication of early and recent Zika virus isolates throughout mouse brain development. Proc. Natl. Acad. Sci. USA 2017, 114, 12273–12278. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nutt, C.; Adams, P. Zika in Africa-the invisible epidemic? Lancet 2017, 389, 1595–1596. [Google Scholar] [CrossRef]
- Setoh, Y.X.; Prow, N.A.; Peng, N.; Hugo, L.E.; Devine, G.; Hazlewood, J.E.; Suhrbier, A.; Khromykh, A.A. De novo generation and characterization of new Zika virus isolate using sequence data from a microcephaly case. mSphere 2017, 2, e00190-17. [Google Scholar] [CrossRef] [PubMed]
- Prow, N.A.; Liu, L.; Nakayama, E.; Cooper, T.H.; Yan, K.; Eldi, P.; Hazlewood, J.E.; Tang, B.; Le, T.T.; Setoh, Y.X.; et al. A vaccinia-based single vector construct multi-pathogen vaccine protects against both Zika and Chikungunya viruses. Nat. Commun. 2018, 9, 1230. [Google Scholar] [CrossRef] [PubMed]
- Marchette, N.J.; Garcia, R.; Rudnick, A. Isolation of Zika virus from Aedes aegypti mosquitoes in Malaysia. Am. J. Trop. Med. Hyg. 1969, 18, 411–415. [Google Scholar] [CrossRef] [PubMed]
- Swann, J.B.; Hayakawa, Y.; Zerafa, N.; Sheehan, K.C.; Scott, B.; Schreiber, R.D.; Hertzog, P.; Smyth, M.J. Type I IFN contributes to NK cell homeostasis, activation, and antitumor function. J. Immunol. 2007, 178, 7540–7549. [Google Scholar] [CrossRef] [PubMed]
- Gardner, J.; Anraku, I.; Le, T.T.; Larcher, T.; Major, L.; Roques, P.; Schroder, W.A.; Higgs, S.; Suhrbier, A. Chikungunya virus arthritis in adult wild-type mice. J. Virol. 2010, 84, 8021–8032. [Google Scholar] [CrossRef] [PubMed]
- Hugo, L.E.; Prow, N.A.; Tang, B.; Devine, G.; Suhrbier, A. Chikungunya virus transmission between Aedes albopictus and laboratory mice. Parasit. Vectors 2016, 9, 555. [Google Scholar] [CrossRef] [PubMed]
- Grant, A.; Ponia, S.S.; Tripathi, S.; Balasubramaniam, V.; Miorin, L.; Sourisseau, M.; Schwarz, M.C.; Sanchez-Seco, M.P.; Evans, M.J.; Best, S.M.; et al. Zika virus targets human stat2 to inhibit type I interferon signaling. Cell Host Microbe 2016, 19, 882–890. [Google Scholar] [CrossRef] [PubMed]
- Cui, L.; Zou, P.; Chen, E.; Yao, H.; Zheng, H.; Wang, Q.; Zhu, J.N.; Jiang, S.; Lu, L.; Zhang, J. Visual and motor deficits in grown-up mice with congenital Zika virus infection. EBioMedicine 2017, 20, 193–201. [Google Scholar] [CrossRef] [PubMed]
- Vermillion, M.S.; Lei, J.; Shabi, Y.; Baxter, V.K.; Crilly, N.P.; McLane, M.; Griffin, D.E.; Pekosz, A.; Klein, S.L.; Burd, I. Intrauterine Zika virus infection of pregnant immunocompetent mice models transplacental transmission and adverse perinatal outcomes. Nat. Commun. 2017, 8, 14575. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yockey, L.J.; Varela, L.; Rakib, T.; Khoury-Hanold, W.; Fink, S.L.; Stutz, B.; Szigeti-Buck, K.; Van den Pol, A.; Lindenbach, B.D.; Horvath, T.L.; et al. Vaginal exposure to Zika virus during pregnancy leads to fetal brain infection. Cell 2016, 166, 1247–1256. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.W.; Young, M.P.; Mamidi, A.; Regla-Nava, J.A.; Kim, K.; Shresta, S. A mouse model of Zika virus sexual transmission and vaginal viral replication. Cell Rep. 2016, 17, 3091–3098. [Google Scholar] [CrossRef] [PubMed]
- Miner, J.J.; Cao, B.; Govero, J.; Smith, A.M.; Fernandez, E.; Cabrera, O.H.; Garber, C.; Noll, M.; Klein, R.S.; Noguchi, K.K.; et al. Zika virus infection during pregnancy in mice causes placental damage and fetal demise. Cell 2016, 165, 1081–1091. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, S.; Balasubramaniam, V.R.; Brown, J.A.; Mena, I.; Grant, A.; Bardina, S.V.; Maringer, K.; Schwarz, M.C.; Maestre, A.M.; Sourisseau, M.; et al. A novel Zika virus mouse model reveals strain specific differences in virus pathogenesis and host inflammatory immune responses. PLoS Pathog. 2017, 13, e1006258. [Google Scholar] [CrossRef] [PubMed]
| WT ZIKV-MY Primers | Primer Sequence (5′ to 3′) |
|---|---|
| Fragment 1 Forward | AGTTGTTGATCTGTGTGAATCAGAC |
| Fragment 1 Reverse | GTGAACGCCGCGGTACATAAGGAGTATG |
| Fragment 2 Forward | CATACTCCTTATGTACCGCGGCGTTCAC |
| Fragment 2 Reverse | GATGAAAGAGACCAGCAACGCGGG |
| Fragment 3 Forward | CCCGCGTTGCTGGTCTCTTTCATC |
| Fragment 3 Reverse | CAGCAGCGACAACCCTGGTTGGAG |
| Fragment 4 Forward | CTCCAACCAGGGTTGTCGCTGCTG |
| Fragment 4 Reverse | GTACATGTAGTGCGCCACGAGCAGAATG |
| Fragment 5 Forward | CATTCTGCTCGTGGCGCACTACATGTAC |
| Fragment 5 Reverse | CTCCTGGTGTGCGGCTCATTTCTTC |
| Fragment 6 Forward | GAAGAAATGAGCCGCACACCAGGAG |
| Fragment 6 Reverse | AGACCCATGGATTTCCCCACACCG |
| Chimeric primers | Primer sequence (5′ to 3′) |
| Natal 5′C (MY1966) R | CTCCCACGTCTGGTGACCTCCACTGCCATAGCTGTGGTCAGCAG |
| MY1966prM_F | GTGGAGGTCACCAGACGTG |
| MY1966E_R | AGCAGAGACGGCTGTAGATAGG |
| Natal NS1-junc2 (MY1966) F | CTTCCTATCTACAGCCGTCTCTGCTGATGTGGGGTGCTCGGTG |
| Protein | Amino Acid Position (Within Protein) | African, MR766 1947 (146× SM, 1× C6/36, 1× Vero) | African, Nigeria 1968 (21× SM, 1× Vero) | Malaysian ZIKV-MY 1966 (6× SM, 1× BHK, 1× C6/36, 8× Vero) | African, DakAr41525 1985 (5× Vero, 1× C6/36, 2× Vero) | Brazilian ZIKV-Natal 2015 (No Passaging) | Brazilian, Suriname 2015 (4× Vero) |
|---|---|---|---|---|---|---|---|
| prM | 1 | A | A | V | A | A | A |
| 17 | S | S | S | S | N | N | |
| 21 | K | K | K | K | E | E | |
| 31 | V | V | V | V | M | M | |
| E | 154–156 | NDT | Deleted | NDI | NDT | NDT | NDT |
| 393 | D | D | D | D | E | E | |
| 473 | V | V | V | V | M | M | |
| 487 | T | T | T | T | M | M | |
| NS1 | 146 | E | K | K | K | E | K |
| 188 | V | V | A | V | V | V | |
| 233 | T | T | T | T | A | T | |
| 264 | V | V | V | V | M | M | |
| 349 | M | M | M | M | V | M | |
| NS2A | 117 | A | A | V | A | A | A |
| 143 | A | A | A | A | V | V | |
| NS3 | 400 | N | N | N | N | H | H |
| 472 | M | M | M | M | L | L | |
| 584 | Y | Y | Y | Y | H | H | |
| NS4B | 14 | A | A | G | A | S | S |
| 26 | M | V | M | M | I | I | |
| 49 | L | L | L | L | F | F | |
| 98 | M | M | M | M | I | I | |
| 180 | I | A | I | V | V | V | |
| 184 | V | V | V | V | I | I | |
| 186 | L | L | L | L | S | S | |
| 240 | T | T | T | T | I | T | |
| NS5 | 115 | M | M | T | M | V | V |
| 140 | P | P | S | P | P | P | |
| 230 | I | I | I | I | T | T | |
| 268 | A | A | A | A | V | V | |
| 276 | M | M | L | M | M | M | |
| 283 | I | I | V | I | I | I | |
| 377 | N | N | N | N | S | S | |
| 527 | A | A | T | A | I | I | |
| 531 | K | K | K | K | R | R | |
| 588 | G | G | R | G | K | K | |
| 643 | P | P | P | P | S | S | |
| 648 | R | R | S | R | N | N | |
| 704 | S | S | S | S | D | D | |
| 714 | H | H | Y | H | H | H | |
| 720 | Y | H | Y | H | H | H | |
| 868 | D | D | D | D | N | N |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Setoh, Y.X.; Peng, N.Y.; Nakayama, E.; Amarilla, A.A.; Prow, N.A.; Suhrbier, A.; Khromykh, A.A. Fetal Brain Infection Is Not a Unique Characteristic of Brazilian Zika Viruses. Viruses 2018, 10, 541. https://doi.org/10.3390/v10100541
Setoh YX, Peng NY, Nakayama E, Amarilla AA, Prow NA, Suhrbier A, Khromykh AA. Fetal Brain Infection Is Not a Unique Characteristic of Brazilian Zika Viruses. Viruses. 2018; 10(10):541. https://doi.org/10.3390/v10100541
Chicago/Turabian StyleSetoh, Yin Xiang, Nias Y. Peng, Eri Nakayama, Alberto A. Amarilla, Natalie A. Prow, Andreas Suhrbier, and Alexander A. Khromykh. 2018. "Fetal Brain Infection Is Not a Unique Characteristic of Brazilian Zika Viruses" Viruses 10, no. 10: 541. https://doi.org/10.3390/v10100541





