The Effect of Aspergillus Thermomutatus Chrysovirus 1 on the Biology of Three Aspergillus Species
Abstract
:1. Introduction
2. Materials and Methods
2.1. Source of Aspergillus Isolates
2.2. Virus Purification and Detection
2.3. Protoplast Preparation and Virus Transfection
2.4. Quantifying the Biological Impacts of AthCV1 Infection on Three Aspergillus Species
2.5. AthCV1 Transmission Through Ascospores and Sclerotia
2.6. The Effects of AthCV1 on Gene Expression
3. Results
3.1. Transfection of Aspergillus Species
3.2. Impact of AthCV1 on Sporulation of Aspergillus Spp.
3.3. Effects of AthCV1 on Aspergillus Growth and Morphology
3.4. Transmission of AthCV1 Through Ascospores and Sclerotia
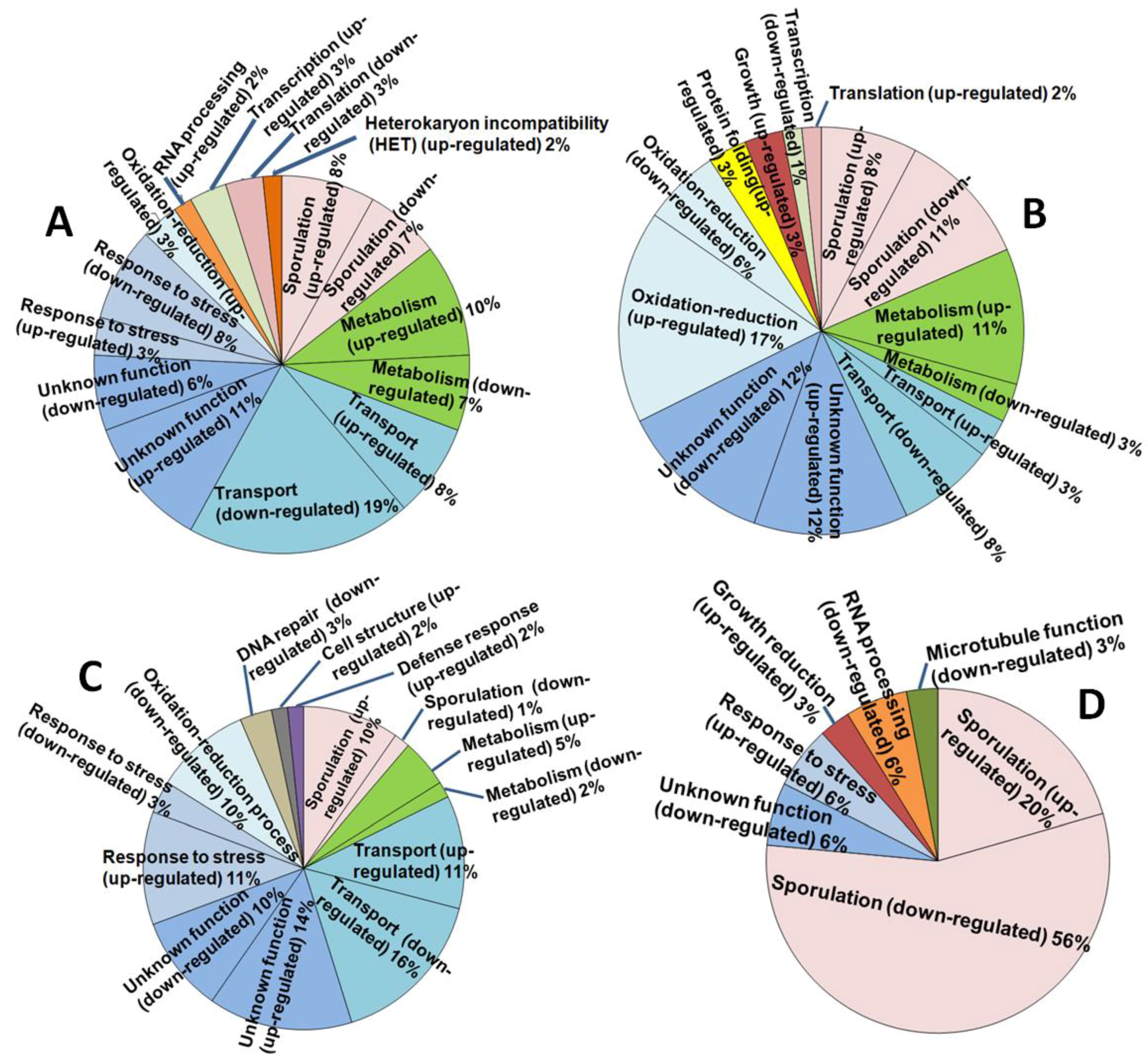
3.5. Changes in Gene Expression Associated with AthCV1 Infection
4. Discussion
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Son, M.; Yu, J.; Kim, K.H. Five Questions about Mycoviruses. PLoS Pathog. 2015, 11, e1005172. [Google Scholar] [CrossRef] [PubMed]
- Pearson, M.N.; Beever, R.E.; Boine, B.; Arthur, K. Mycoviruses of filamentous fungi and their relevance to plant pathology. Mol. Plant. Pathol. 2009, 10, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Boddy, L. Interactions between fungi and other microbes. In The Fungi; Watkinson, S.C., Money, N., Boddy, L., Eds.; Academic Press: Cambridge, MA, USA, 2016; pp. 337–360. [Google Scholar]
- Yu, X.; Li, B.; Fu, Y.; Xie, J.; Cheng, J.; Ghabrial, S.A.; Li, G.; Yi, X.; Jiang, D. Extracellular transmission of a DNA mycovirus and its use as a natural fungicide. Proc. Natl. Acad. Sci. USA 2013, 110, 1452–1457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marzano, S.L.; Nelson, B.D.; Ajayi-Oyetunde, O.; Bradley, C.A.; Hughes, T.J.; Hartman, G.L.; Eastburn, D.M.; Domier, L.L. Identification of Diverse Mycoviruses through Metatranscriptomics Characterization of the Viromes of Five Major Fungal Plant Pathogens. J. Virol. 2016, 90, 6846–6863. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.; Xie, J.; Cheng, J.; Li, B.; Chen, T.; Fu, Y.; Li, G.; Wang, M.; Jin, H.; Wan, H.; et al. Fungal DNA virus infects a mycophagous insect and utilizes it as a transmission vector. PNAS 2016, 113, 12803–12808. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boland, G.J.; Mould, M.J.R.; Robb, J. Ultrastructure of a hypovirulent isolate of Sclerotinia sclerotiorum containing double stranded RNA. Physiol. Mol. Plant. Pathol. 1993, 43, 21–32. [Google Scholar] [CrossRef]
- Hammar, S. Association of double-stranded RNA with low virulence in an isolate of Leucostoma persoonii. Phytopathology 1989, 79, 568–572. [Google Scholar] [CrossRef]
- Boland, G.J. Hypovirulence and double stranded RNA in Sclerotinia sclerotiorum. Can. J. Plant. Pathol. 1992, 14, 10–17. [Google Scholar] [CrossRef]
- Bottacin, A.M.; Levesque, C.A.; Punja, Z.K. Characterization of dsRNA in Chalara elegans and effects on growth and virulence. Phytopathology 1994, 84, 303–312. [Google Scholar] [CrossRef]
- Punja, Z.K. Influence of double-stranded RNAs on growth, sporulation, pathogenicity and survival of Chalara elegans. Can. J. Bot. 1995, 73, 1001–1009. [Google Scholar] [CrossRef]
- Newhouse, J.R.; Hoch, H.C.; Macdonald, W.L. The ultra structure of Endothia parasitica. Comparison of a virulent with a hypovirulent isolate. Can. J. Botany. 1983, 61, 389–399. [Google Scholar] [CrossRef]
- Chu, M.; Jean, J.; Yea, S.J.; Kim, Y.H.; Lee, Y.W.; Kim, K.H. Double-strand RNA mycoviruses from Fusarium graminearum. Appl. Environ. Microbiol. 2002, 68, 2529–2534. [Google Scholar] [CrossRef] [PubMed]
- Rigling, D.; van Alfen, N.K. Extra and intracellular laccase activity of the Chestnut blight fungus, Cryphonectria paraesitica. Appl. Environ. Microbial. 1993, 59, 3634–3639. [Google Scholar]
- Márquez, L.M.; Redman, R.S.; Rodriguez, R.J.; Roossinck, M.J. A Virus in a Fungus in a Plant: Three-Way Symbiosis Required for Thermal Tolerance. Science 2007, 315, 513–515. [Google Scholar] [CrossRef] [PubMed]
- Magliani, W.; Conti, S.; Gerloni, M.; Bertolotti, D.; Polonelli, L. Yeast killer systems. Clin. Microbiol. Rev. 1997, 10, 369–400. [Google Scholar] [PubMed]
- Park, C.M.; Banerjee, N.; Koltin, Y.; Bruenn, J.A. The Ustilago maydis virally encoded KP1 killer toxin. Mol. Microbiol. 1996, 20, 957–963. [Google Scholar] [CrossRef] [PubMed]
- Banks, G.T.; Buck, K.W.; Chain, E.B.; Darbyshire, J.E.; Himmelweit, F.; Ratti, G.; Sharpe, T.J.; Planterose, D.N. Antiviral activity of double stranded RNA from a virus isolated from Aspergillus foetidus. Nature 1970, 227, 505–507. [Google Scholar] [CrossRef] [PubMed]
- Ghabrial, S.A.; Caston, J.R. Family Chrysoviridae, genus Chrysovirus. In Virus Taxonomy: Classification and Nomenclature of Viruses; Ninth Report of the International Committee on Taxonomy of Viruses; King, A.M.Q., Adams, M.J., Carstens, E.B., Lefkowitz, E.J., Eds.; Elsevier: Amsterdam, The Netherlands; Academic Press: Cambridge, MA, USA, 2012; pp. 509–513. [Google Scholar]
- Wang, L.; Jiang, J.; Wang, Y.; Hong, N.; Zhang, F.; Xu, W.; Wang, G. Hypovirulence of the Phytopathogenic Fungus Botryosphaeria dothidea: Association with a Coinfecting Chrysovirus and a Partitivirus. J. Virol. 2014, 88, 7517–7527. [Google Scholar] [CrossRef] [PubMed]
- Urayama, S.; Kato, S.; Suzuki, Y.; Aoki, N.; Le, M.T.; Arie, T.; Teraoka, T.; Fukuhara, T.; Moriyama, H. Mycoviruses related to chrysovirus affect vegetative growth in the rice blast fungus Magnaporthe oryzae. J. Gen. Virol. 2010, 91, 3085–3094. [Google Scholar] [CrossRef] [PubMed]
- Urayama, S.; Sakoda, H.; Katoh, Y.; Takai, R.; Le, T.M.; Fukuhara, T.; Arie, T.; Teraoka, T.; Moriyama, H. A dsRNA mycovirus, Magnaporthe oryzae chrysovirus1-B, suppresses vegetative growth and development of the rice blast fungus. Virology 2014, 448, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.; Zhou, T.; Guo, L.Y. Characterization of a novel strain of Botryosphaeria dothidea chrysovirus 1 from the apple white rot pathogen Botryosphaeria dothidea. Arch. Virol. 2017, 162, 2097–2102. [Google Scholar] [CrossRef] [PubMed]
- Bhatti, M.F.; Jamal, A.; Petrou, M.A.; Cairns, T.C.; Bignell, E.M.; Coutts, R.H.A. The effects of dsRNA mycoviruses on growth and murine virulence of Aspergillus fumigatus. Fungal Genet. Biol. 2011, 48, 1071–1075. [Google Scholar] [CrossRef] [PubMed]
- Hammond, T.M.; Andrewski, M.D.; Roossinck, M.J.; Keller, N.P. Aspergillus mycoviruses are targets and suppressors of RNA silencing. Eukaryotic Cell 2008, 7, 350–357. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.M.; Cho, W.K.; Yu, J.; Son, M.; Choi, H.; Min, K.; Lee, Y.W.; Kim, K.H. A Comparison of Transcriptional Patterns and Mycological Phenotypes following Infection of Fusarium graminearum by Four Mycoviruses. PLoS ONE 2014, 9, e100989. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Nuss, D.L.; Suzuki, N. Synergism between a mycoreovirus and a hypovirus mediated by the papain-like protease p29 of the prototypic hypovirus CHV1-EP. J. Gen. Virol. 2006, 87, 3703–3714. [Google Scholar] [CrossRef] [PubMed]
- Ejmal, M.A.; Holland, D.J.; MacDiarmid, R.M.; Pearson, M.N. A novel chrysovirus from a clinical isolate of Aspergillus thermomutatus affects sporulation. PLoS ONE. under review.
- Aneja, K.R. Experiments in Microbiology, Plant Pathology and Biotechnology; New Age International (P) Ltd.: New Delhi, India, 2003. [Google Scholar]
- O’Gorman, C.M.; Fuller, H.T.; Dyer, P.S. Discovery of a sexual cycle in the opportunistic fungal pathogen Aspergillus fumigatus. Nature 2009, 457, 471–474. [Google Scholar] [CrossRef] [PubMed]
- Girardin, H.; Monod, M.; Latge, J. Molecular Characterization of the Food-Borne Fungus Neosartorya fischeri (Malloch and Cain). Appl. Environ. Microbiol. 1995, 61, 1378–1383. [Google Scholar] [PubMed]
- Utkhede, R.S.; Rahe, J.E. Wet-sieving floatation technique for isolation of sclerotia of Sclerotium cepivorum from muck soil. Phytopathology 1979, 69, 295–297. [Google Scholar] [CrossRef]
- Zhang, F.; Guo, Z.; Zhong, H.; Wang, S.; Yang, W.; Liu, Y.; Wang, S. RNA-Seq-based transcriptome analysis of aflatoxigenic Aspergillus flavus in response to water activity. Toxins 2014, 6, 3187–3207. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 12 December 2016).
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.M.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.; Pachter, L. Transcript assembly and abundance estimation from RNA-Seq reveals thousands of new transcripts and switching among isoforms. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haas, J.H.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dyer, P.S.; O’Gorman, C.M. Sexual development and cryptic sexuality in fungi: Insights from Aspergillus species. FEMS Microbiol. Rev. 2012, 36, 165–192. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, S.; Sarikaya Bayram, Ö.; Bayram, Ö.; Braus, G.H. ConF and conJ contribute to conidia germination and stress response in the filamentous fungus Aspergillus nidulans. Fungal Genet. Biol. 2013, 56, 42–53. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Sun, L.; Liang, Q.; Wang, J.; Mo, W.; Zhou, B. Yeast AMID homologue Ndi1p displays respiration-restricted apoptotic activity and is involved in chronological aging. Mol. Biol. Cell 2006, 17, 1802–1811. [Google Scholar] [CrossRef] [PubMed]
- Hyder, R.; Pennanen, T.; Hamberg, L.; Vainio, E.J.; Piri, T.; Hantula, J. Two viruses of Heterobasidion confer beneficial, cryptic or detrimental effects to their hosts in different situations. Fungal Ecology 2013, 6, 387–396. [Google Scholar] [CrossRef]
- Vainio, E.J.; Korhonen, K.; Tuomivirta, T.T.; Hantula, J. A novel putative partitivirus of the saprotrophic fungus Heterobasidion ecrustosum infects pathogenic species of the Heterobasidion annosum complex. Fungal Biology 2010, 114, 955–965. [Google Scholar] [CrossRef] [PubMed]
- Madhosingh, C. Production of intraspecific hybrids of Fusarium oxysporum f. sp. radicis-lycopersici and Fusarium oxysporum f. sp. lycopersici by protoplast fusions. J. Phytopathol. 1994, 142, 301–309. [Google Scholar] [CrossRef]
- Van Diepeningen, A.D.; Debets, A.J.; Hoekstra, R.F. Intra- and interspecies virus transfer in Aspergilli via protoplast fusion. Fungal Genet. Biol. 1998, 25, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Van Diepeningen, A.D.; Varga, J.; Hoekstra, R.F.; Debets, A.J. Mycoviruses in Aspergilli. In Aspergillus in the genomics era; Samson, R., Varga, J., Eds.; Wageningen Academic Publishers: Wageningen, The Netherlands, 2008; p. 133. ISBN 978-90-8686-065-4. [Google Scholar]
- Kanematsu, S.; Sasaki, A.; Onoue, M.; Oikawa, Y.; Ito, T. Extending the fungal host range of a partitivirus and a mycoreovirus from Rosellinia necatrix by Inoculation of protoplasts with virus particles. Phytopathology. 2010, 100, 922–930. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.M.; Yu, J.; Son, M.; Lee, Y.W.; Kim, K.H. Transmission of Fusarium boothii mycovirus via protoplast fusion causes hypovirulence in other phytopathogenic fungi. PLoS ONE 2011, 6, e21629. [Google Scholar] [CrossRef] [PubMed]
- Kanhayuwa, L.; Kotta-Loizou, I.; Özkan, S.; Gunning, A.P.; Coutts, R.H. A novel mycovirus from Aspergillus fumigatus contains four unique dsRNAs as its genome and is infectious as dsRNA. Proc. Natl. Acad. Sci. USA 2015, 112, 9100–9105. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Zhang, M.; Chen, Q.; Zhu, M.; Zhou, E. A novel mycovirus closely related to viruses in the genus Alphapartitivirus confers hypovirulence in the phytopathogenic fungus Rhizoctonia solani. Virology 2014, 456–457, 220–226. [Google Scholar] [CrossRef] [PubMed]
- Hillman, B.I.; Supyani, S.; Kondo, H.; Suzuki, N. A Reovirus of the fungus Cryphonectria parasitica that is infectious as particles and related to the Coltivirus genus of animal pathogens. J. Virol. 2004, 78, 892–898. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, A.; Kanematsu, S.; Onoue, M.; Oyama, Y.; Yoshida, K. Infection of Rosellinia necatrix with purified viral particles of a member of Partitiviridae (RnPV1-W8). Arch Virol. 2006, 151, 697–707. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, A.; Kanematsu, S.; Onoue, M.; Oikawa, Y.; Nakamura, H.; Yoshida, K. Artificial Infection of Rosellinia necatrix with Purified Viral Particles of a Member of the Genus Mycoreovirus Reveals Its Uneven Distribution in Single Colonies. Phytopathology 2007, 97, 278–286. [Google Scholar] [CrossRef] [PubMed]
- Nuss, D.L. Hypovirulence: Mycoviruses at the fungal-plant interface. Nat. Rev. Microbiol. 2005, 3, 632–642. [Google Scholar] [CrossRef] [PubMed]
- Ghabrial, S.A.; Suzuki, N. Viruses of Plant Pathogenic Fungi. Annu. Rev. Phytopathol. 2009, 47, 353–384. [Google Scholar] [CrossRef] [PubMed]
- Romeralo, C.; Bezos, D.; Martínez-Álvarez, P.; Diez, J.J. Vertical Transmission of Fusarium circinatum Mitoviruses FcMV1 and FcMV2-2 via Microconidia. Forests 2018, 9, 356. [Google Scholar] [CrossRef]
- Chu, Y.M.; Lim, W.S.; Yea, S.J.; Cho, J.D.; Lee, Y.W.; Kim, K.H. Complexity of dsRNA mycovirus isolated from Fusarium graminearum. Virus Genes 2004, 28, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Milgroom, M.G.; Cortesi, P. Biological control of chestnut blight with hypovirulence: A critical analysis. Annu. Rev. Phytopathol. 2004, 42, 3113–3118. [Google Scholar] [CrossRef] [PubMed]
- Van Diepeningen, A.D.; Debets, A.J.; Hoekstra, R.F. Heterokaryon incompatibility blocks virus transfer among natural isolates of black Aspergilli. Curr. Genet. 1997, 32, 2091–2097. [Google Scholar] [CrossRef]
- Refos, J.M.; Vonk, A.G.; Eadie, K.; Lo-Ten-Foe, J.R.; Verbrugh, H.A.; van Diepeningen, A.D.; van de Sande, W.W. Double-Stranded RNA Mycovirus Infection of Aspergillus fumigatus is Not Dependent on the Genetic Make-Up of the Host. PLoS ONE 2013, 8, e77381. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Coenen, A.; Kevei, F.; Hoekstra, R.F. Factors affecting the spread of double-stranded RNA viruses in Aspergillus nidulans. Genet. Res. 1997, 69, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.M.C.; Pearson, M.N.; Beever, R.E.; Parkes, S.L. Why Fungi Have Sex? In 14th International Botrytis Symposium abstract book; Cape Town, South Africa, 2007; p. 26. [Google Scholar]
- Brewer, B.J.; Fangman, W.L. Preferential inclusion of extrachromosomal genetic elements in yeast meiotic spores. Proc. Nat. Acad. Sci. USA 1980, 77, 5380–5384. [Google Scholar] [CrossRef]
- Bell, G. The Masterpiece of Nature: The Evolution and Genetics of Sexuality; University of California Press: Berkeley, CA, USA, 1982. [Google Scholar]
- Frisvad, J.C.; Petersen, L.M.; Lyhne, K.; Larsen, T.O. Formation of sclerotia and production of indoloterpenes by Aspergillus niger and other species in section Nigri. PLoS ONE 2014, 9, e94857. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rajendran, C.; Muthappa, B.N. Saitoa, a new genus of Plectomycetes. Proc. Plant Sci. 1980, 89, 185–191. [Google Scholar]
- Paoletti, M.; Seymour, F.A.; Alcocer, M.J.C.; Kaur, N.; Calvo, A.M.; Archer, D.B.; Dyer, P.S. Mating type and the genetic basis of self-fertility in the model fungus Aspergillus nidulans. Curr. Biol. 2007, 17, 1384–1389. [Google Scholar] [CrossRef] [PubMed]
- Baidya, S.; Cary, J.W.; Grayburn, W.S.; Calvo, A.M. Role of nitric oxide and flavohemoglobin homolog genes in Aspergillus nidulans sexual development and mycotoxin production. Appl. Environ. Microbiol. 2011, 77, 5524–5528. [Google Scholar] [CrossRef] [PubMed]
- De Souza, C.P.; Hashmi, S.B.; Osmani, A.H.; Andrews, P.; Ringelberg, C.S.; Dunlap, J.C.; Osmani, S.A. Functional Analysis of the Aspergillus nidulans Kinome. PLoS ONE 2013, 8, e58008. [Google Scholar] [CrossRef] [PubMed]
- Xiang, X.; Osmani, A.H.; Osmani, S.A.; Xin, M.; Morris, N.R. NudF, a nuclear migration gene in Aspergillus nidulans, is similar to the human LIS-1 gene required for neuronal migration. Mol. Biol. Cell 1995, 6, 297–310. [Google Scholar] [CrossRef] [PubMed]
- Bayram, O.; Biesemann, C.; Krappmann, S.; Galland, P.; Braus, G.H. More than a repair enzyme: Aspergillus nidulans photolyase-like CryA is a regulator of sexual development. Mol. Biol. Cell 2008, 19, 3254–3262. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, L.; Sánchez, O.; Shiozaki, K.; Aguirre, J. SakA MAP kinase is involved in stress signal transduction, sexual development and spore viability in Aspergillus nidulans. Mol. Microbiol. 2002, 45, 1153–1163. [Google Scholar] [CrossRef] [PubMed]
- Semighini, C.P.; Harris, S.D. Regulation of apical dominance in Aspergillus nidulans hyphae by reactive oxygen species. Genetics 2008, 179, 1919–1932. [Google Scholar] [CrossRef] [PubMed]
- Westfall, P.J.; Momany, M. Aspergillus nidulans septin AspB plays pre- and postmitotic roles in septum, branch, and conidiophore development. Mol. Biol. Cell 2002, 13, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Rodríguez, Y.; Hastings, S.; Momany, M. The septin AspB in Aspergillus nidulans forms bars and filaments and plays roles in growth emergence and conidiation. Eukaryot Cell 2012, 11, 3112–3113. [Google Scholar] [CrossRef] [PubMed]
- Busch, S.; Eckert, S.E.; Krappmann, S.; Braus, G.H. The COP9 signalosome is an essential regulator of development in the filamentous fungus Aspergillus nidulans. Mol. Microbiol. 2003, 49, 717–730. [Google Scholar] [CrossRef] [PubMed]
- Busch, S.; Schwier, E.U.; Nahlik, K.; Bayram, O.; Helmstaedt, K.; Draht, O.W.; Krappmann, S.; Valerius, O.; Lipscomb, W.N.; Braus, G.H. An eight-subunit COP9 signalosome with an intact JAMM motif is required for fungal fruit body formation. Proc. Natl. Acad. Sci. USA 2007, 104, 8089–8094. [Google Scholar] [CrossRef] [PubMed]
- Bok, J.W.; Keller, N.P. LaeA, a regulator of secondary metabolism in Aspergillus spp. Eukaryot Cell 2004, 3, 5273–5275. [Google Scholar] [CrossRef]
- Bayram, O.; Krappmann, S.; Ni, M.; Bok, J.W.; Helmstaedt, K.; Valerius, O.; Braus-Stromeyer, S.; Kwon, N.J.; Keller, N.P.; Yu, J.H.; et al. VelB/VeA/LaeA complex coordinates light signal with fungal development and secondary metabolism. Science 2008, 320, 1504–1506. [Google Scholar] [CrossRef] [PubMed]
- Sarikaya, B.O.; Bayram, O.; Valerius, O.; Park, H.S.; Irniger, S.; Gerke, J.; Ni, M.; Han, K.H.; Yu, J.H.; Braus, G.H. LaeA control of velvet family regulatory proteins for light-dependent development and fungal cell-type specificity. PLoS Genet. 2010, 6, e1001226. [Google Scholar] [CrossRef] [PubMed]
- Bayram, O.; Braus, G.H. Coordination of secondary metabolism and development in fungi: The velvet family of regulatory proteins. FEMS Microbiol. Rev. 2012, 36, 12–14. [Google Scholar] [CrossRef] [PubMed]
- Patananan, A.N.; Palmer, J.M.; Garvey, G.S.; Keller, N.P.; Clarke, S.G. A novel automethylation reaction in the Aspergillus nidulans LaeA protein generates S-methylmethionine. J. Biol. Chem. 2013, 288, 14032–14045. [Google Scholar] [CrossRef] [PubMed]
- Nitsche, B.M.; Jørgensen, T.R.; Akeroyd, M.; Meyer, V.; Ram, A.F. The carbon starvation response of Aspergillus niger during submerged cultivation: Insights from the transcriptome and secretome. BMC Genomics 2012, 13, 380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jørgensen, T.R.; Nitsche, B.M.; Lamers, G.E.; Arentshorst, M.; van den Hondel, C.A.; Ram, A.F. Transcriptomic insights into the physiology of Aspergillus niger approaching a specific growth rate of zero. Appl. Environ. Microbiol. 2010, 76, 5344–5355. [Google Scholar] [CrossRef] [PubMed]
- Van Leeuwen, M.R.; Krijgsheld, P.; Bleichrodt, R.; Menke, H.; Stam, H.; Stark, J.; Wösten, H.A.; Dijksterhuis, J. Germination of conidia of Aspergillus niger is accompanied by major changes in RNA profiles. Stud. Mycol. 2013, 74, 59–70. [Google Scholar] [CrossRef] [PubMed]
- Fu, C.; Iyer, P.; Herkal, A.; Abdullah, J.; Stout, A.; Free, S.J. Identification and characterization of genes required for cell-to-cell fusion in Neurospora crassa. Eukaryot Cell. 2011, 10, 1100–1109. [Google Scholar] [CrossRef] [PubMed]
- Wada, R.; Jin, F.J.; Koyama, Y.; Maruyama, J.; Kitamoto, K. Efficient formation of heterokaryotic sclerotia in the filamentous fungus Aspergillus oryzae. Appl. Microbiol. Biotechnol. 2014, 98, 3253–3254. [Google Scholar] [CrossRef] [PubMed]
- Carlile, M.J. The Success of the Hypha and Mycelium. In The Growing Fungus; Gow, N.A.R., Gadd, G.M., Eds.; Chapman and Hall: London, UK, 1995; pp. 3–19. [Google Scholar]
- Teutschbein, J.; Albrecht, D.; Pötsch, M.; Guthke, R.; Aimanianda, V.; Clavaud, C.; Latgé, J.P.; Brakhage, A.A.; Kniemeyer, O. Proteome profiling and functional classification of intracellular proteins from conidia of the human-pathogenic mold Aspergillus fumigatus. J. Proteome Res. 2010, 9, 3427–3442. [Google Scholar] [CrossRef] [PubMed]
- Albrecht, D.; Guthke, R.; Brakhage, A.A.; Kniemeyer, O. Integrative analysis of the heat shock response in Aspergillus fumigatus. BMC Genom. 2010, 11, 32. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsitsigiannis, D.I.; Zarnowski, R.; Keller, N.P. The lipid body protein, PpoA, coordinates sexual and asexual sporulation in Aspergillus nidulans. J. Biol. Chem. 2004, 279, 11344–11353. [Google Scholar] [CrossRef] [PubMed]
- Duarte, M.; Peters, M.; Schulte, U.; Videira, A. The internal alternative NADH dehydrogenase of Neurospora crassa mitochondria. Biochem. J. 2003, 371, 1005–1011. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.F.; Wheeler, M.H.; Chang, Y.C.; Kwon-Chung, K.J. A Developmentally Regulated Gene Cluster Involved in Conidial Pigment Biosynthesis in Aspergillus fumigatus. J. Bacteriol. 1999, 181, 6469–6477. [Google Scholar] [PubMed]
- Tsai, H.F.; Washburn, R.G.; Chang, Y.C.; Kwon-Chung, K.J. Aspergillus fumigatus arp1 modulates conidial pigmentation and complement deposition. Mol. Microbiol. 1997, 26, 175–183. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.F.; Fujii, I.; Watanabe, A.; Wheeler, M.H.; Chang, Y.C.; Yasuoka, Y.; Ebizuka, Y.; Kwon-Chung, K.J. Pentaketide melanin biosynthesis in Aspergillus fumigatus requires chain-length shortening of a heptaketide precursor. J. Biol. Chem. 2001, 276, 29292–29298. [Google Scholar] [CrossRef] [PubMed]
- Langfelder, K.; Jahn, B.; Gehringer, H.; Schmidt, A.; Wanner, G.; Brakhage, A.A. Identification of a polyketide synthase gene (pksP) of Aspergillus fumigatus involved in conidial pigment biosynthesis and virulence. Med. Microbiol. Immunol. 1998, 187, 798–799. [Google Scholar] [CrossRef]
- Jackson, J.C.; Higgins, L.A.; Lin, X. Conidiation color mutants of Aspergillus fumigatus are highly pathogenic to the heterologous insect host Galleria mellonella. PLoS ONE 2009, 4, e4224. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Adalia, E.J.; Fernández, M.M.; Diez, J.J. The use of mycoviruses in the control of forest diseases. Biocontrol. Sci. Technol. 2016, 26, 577–604. [Google Scholar] [CrossRef]

| A. thermomutatus | A. fumigatus | A. niger | A. nidulans | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Property | Growth Temp | Virus-Free | AthCV1 Infected | Virus-Free | AthCV1 Infected | Virus-Free | AthCV1 Infected | Virus-Free | AthCV1 Infected |
| Conidia production (per plate) | 37 °C | 2.1 (± 0.36) × 105 | 2.0 (± 0.10) × 106 * | 3.8 (± 0.42) × 108 | 1.6 (± 0.12) × 107 * | 1.8 (± 0.12) × 108 | 2.0 (± 0.14) × 107 * | 2.2 (± 0.62) × 108 | 9.5 (± 0.79) × 107 * |
| 20 °C | 2.4 (± 0.36) × 106 | 2.1 (± 0.24) × 105 * | 1.5 (± 0.21) × 108 | 3.6 (± 0.31) × 107 * | 2.2 (± 0.26) × 108 | 1.9 (± 0.24) × 108 | 1.4 (± 0.16) × 108 | 1.4 (± 0.12) × 108 | |
| Ascospore production (per plate) | 37 °C | 0 | 7.2 (± 0.73) × 105 * | 0 | 0 | 0 | 0 | 1.1 (± 0.10) × 106 | 2.1 (± 0.42) × 106 * |
| 20 °C | 0 | 3.6 (± 0.37) × 105 * | 0 | 0 | 0 | 0 | 2.9 (± 0.22) × 105 | 8.0 (± 0.55) × 105 * | |
| Sclerotia production (per plate) | 37 °C | 0 | 0 | 0 | 0 | 0 | 54 (± 4.2) * | 0 | 0 |
| 20 °C | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Growth (mm) 1 | 37 °C | 74 (± 0.32) | 75 (± 1.00) | 72 (± 0.37) | 73 (± 1.00) | 77 (± 0.20) | 67 (± 0.68) * | 76 (± 0.49) | 75 (± 0.71) |
| 20 °C | 76 (± 0.32) | 75 (± 0.32) | 67 (± 0.32) | 72 (± 0.20) * | 77 (± 0.60) | 67 (± 0.74) | 33 (± 0.45) | 30 (± 0.66) * | |
| Fungal Biomass 2 (g dry wt) | 37 °C | 1.25 (± 0.012) | 1.28 (± 0.009) | 1.30 (± 0.008) | 1.23 (± 0.008) * | 1.25 (± 0.023) | 1.30 (± 0.007) | 1.36 (± 0.021) | 1.12 (± 0.005) * |
| 20 °C | 1.13 (± 0.011) | 1.13 (± 0.009) | 1.15 (± 0.011) | 1.13 (± 0.007) | 1.09 (± 0.004) | 1.12 (± 0.003) * | 1.12 (± 0.016) | 1.12 (± 0.010) | |
| Sector formation | 37 °C | Creamy, rough ascospore-rich sectors | Clear, elongated sectors in the grey mycelium | Conidia-free sectors with no pigmentation | Ascospore-rich sectors in the green mycelium | ||||
| 20 °C | Creamy, rough ascospore-rich sectors | Clear, elongated sectors in the grey mycelium | Conidia-free sectors with no pigmentation | No sector formation | |||||
| Pigment change | 37 °C | Creamy sectors | Sectors lack pigmentation | Sectors lack pigmentation | No change | ||||
| 20 °C | Creamy sectors | Sectors lack pigmentation | Sectors lack pigmentation | No change | |||||
| Gene ID | Verified and Predicted Gene and Function ◊ retrieved from www.aspergillusgenome.org, www.uniprot.org and fungi.ensembl.org/index.html | Fold Change |
|---|---|---|
| A. thermomutatus∆ | ||
| NFIA_070550 | csnB; orthologue(s) subunit2 of the COP9 signalosome; required for formation of cleistothecia | 7 ↑ |
| NFIA_010750 | laeA; orthologue (laeA/AN0807, A. nidulans) coordinates asexual development in response to light, involved in the regulation of secondary metabolism and required for the formation Hulle cells. | 1913 ↓ |
| NFIA_030070 | fhbA; flavohemoprotein, regulates sexual sporulation and sterigmatocystin production | 100 ↓ |
| NFIA_071100 | MAT1; positive regulation of mating type specific transcription, DNA-templated | 35 ↓ |
| A. fumigatus | ||
| Afu1g14520 | Pyridine nucleotide-disulphide oxidoreductase; NADPH dehydrogenase activity, spore germination | 25 ↑ |
| Afu2g17560 | Hydroxynaphthalene reductase arp2; involved in conidial pigment biosynthesis; conidia-enriched protein | 16 ↑ |
| Afu2g17600 | Conidial pigment polyketide synthase PksP/Alb1; conidial pigment biosynthesis; conidia wall assembly | 13 ↑ |
| Afu2g17550 | Heptaketide hydrolase ayg1; conidial pigment; polyketide shortening; conidia-enriched protein | 13 ↑ |
| Afu2g17580 | Scytalone dehydratase arp1; conidial pigment biosynthesis; conidia formation and sporulation | 9 ↑ |
| Afu1g09750 | Aldehyde reductase (AKR1), putative; conidia-enriched protein | 38 ↓ |
| Afu4g10770 | ppoA; response to oxidative stress; orthologue (AN1967/ppoA) negative regulation of sexual sporulation and positive regulation asexual sporulation. | 7 ↓ |
| A. niger | ||
| An18g06650 | Orthologue (Afu3g14540, A. fumigatus), 30-kilodalton heat shock protein; conidia-enriched protein | 157 ↑ |
| An01g12490 | Orthologue (Neurospora crassa) has NADPH dehydrogenase activity, role in spore germination | 18 ↑ |
| An14g02460 | fhbA; orthologue in A. nidulans regulation of sexual sporulation and sterigmatocystin biosynthetic process | 12 ↑ |
| An04g07400 | Putative C6 zinc finger transcription factor; orthologue (AN1848/nosA, A. nidulans) positive regulation of sexual development; orthologue (adv-1) in N. crassa has predicted role in conidium formation and hyphal growth. | 326 ↓ |
| An12g00710 | Orthologue (AN9121/esdC, A. nidulans); negative regulation of conidiation and positive regulation of sexual sporulation | 200 ↓ |
| An02g05420 | flbC; putative C2H2 transcription factor; predicted role in conidiation; expressed in germinating conidia | 31 ↓ |
| An04g05880 | ppoA; response to oxidative stress; orthologue (AN1967/ppoA) negative regulation of sexual sporulation and positive regulation of asexual sporulation. | 20 ↓ |
| An01g04830 | flbD; Myb-like DNA-binding protein; positive regulation of conidiation; expressed in germinating conidia | 12 ↓ |
| An17g01580 | Orthologue (AN2290/steA, A. nidulans) has negative regulation of transcription by RNA polymerase II promoter; regulation of secondary metabolite biosynthetic process; sporocarp development involved in sexual reproduction | 5 ↓ |
| An05g00480 | stuA; positive regulation of conidium formation and conidiophore development | 5 ↓ |
| A. nidulans | ||
| AN7795 | gprK; positive regulation of sexual sporulation [40] | 39 ↑ |
| AN3387 | gprD; Putative G-protein coupled receptor; Deletion of gprD resulted in delayed conidial germination and enhanced sexual development | 16 ↑ |
| AN3695 | Putative anthranilate synthase with a predicted role in aromatic amino acid biosynthesis and cleistothecium development | 13 ↑ |
| AN7169 | fhbA; flavohemoprotein; sterigmatocystin biosynthetic process and regulates sexual development | 12 ↑ |
| AN5844 | Controls conidia germination and adjusts cellular substances which protect conidia against dryness [41] | 7 ↑ |
| AN5046 | anisin-1; asexual sporulation, response to oxidative stress and defense response | 7 ↑ |
| AN5156 | pho80; overexpression decreases conidiation and increases formation of cleistothecia | 6 ↑ |
| AN3148 | PH domain protein; have role in ascospore wall assembly, ascospore-type prospore membrane assembly | 44 ↓ |
| AN0387 | cryA; negative regulation of cleistothecium development | 37 ↓ |
| AN8640 | conF; Conidiation protein Con-6, putative; contributes in conidia germination and desiccation resistance | 21 ↓ |
| AN1848 | nosA; Zinc(II)2Cys6 putative transcription factor involved in the regulation of sexual development | 20 ↓ |
| AN2755 | MAT1; regulator of sexual development; acts with Mat2 HMG domain protein | 16 ↓ |
| AN6046 | noxR; P67phox regulatory subunit homolog; required for normal sexual and asexual development | 13 ↓ |
| AN6688 | aspB; conidiophore development and hyphal growth | 12 ↓ |
| AN4163 | cpcB; required for sexual development; positive regulation of cleistothecium development | 11 ↓ |
| AN4351 | palA; pH-response regulator protein palA; orthologue in Saccharomyces cerevisiae (RIM20) role in sporulation resulting in formation of a cellular spore | 6 ↓ |
| AN1017 | hOGA; Putative mitogen-activated protein kinase; required for sexual development and sporulation | 6 ↓ |
| AN7553 | devR; Basic helix-loop-helix transcription factor required for normal conidiophore development | 5 ↓ |
| AN10306 | candA-N; role in sexual development, secondary metabolism and light control of asexual development | 5 ↓ |
| AN2458 | candA-C; role in sexual development, secondary metabolism and light control of asexual development | 5 ↓ |
| Fold Change in AthCV1-infected Samples | Gene and Function From www.uniport.org & www.aspergillusgenome.org | ||||
|---|---|---|---|---|---|
| Gene ID A. nidulans | A. thermomutatus | A. fumigatus | A. nidulans | A. niger | |
| AN10660 | - | 25 | - | 18 | ndiA; Pyridine nucleotide-disulphide oxidoreductase, putative; NADPH dehydrogenase activity and spore germination. |
| AN2755 | −35 | - | −16 | - | MAT1: Alpha-domain mating-type protein; regulator of sexual development; acts with Mat2 HMG domain protein. |
| AN1652 | - | −5 | - | - | sebA; C2H2 transcription factor; required for virulence; response to oxidative stress and heat shock. |
| AN5781 | - | 31 | 6 | 157 | Heat shock protein Hsp30, transcript increase during the unfolded-protein response; palA-dependent expression independent of pH |
| AN2458 | 3 | - | −5 | - | canadA-C; orthologue(s) N-terminal subunit of Cand1; sexual development, secondary metabolism and light control of asexual development. |
| AN10306 | 5 | - | −5 | 3 | candA-N; role in sexual development and secondary metabolism. |
| AN10049 | - | 9 | −4 | - | mdpB; Probable scytalone dehydratase; involved in conidial pigment biosynthesis, conidium formation and sporulation. |
| AN5836 | - | - | −4 | −5 | stuA; Positive regulation of asexual sporulation and conidiophore development. |
| AN1848 | - | −2 | −20 | −326 | nosA; Zinc(II)2Cys6 putative transcription factor; positive regulation of sexual development. |
| AN5156 | −2 | - | 6 | - | Pho80-like cyclin; overexpression decreases conidiation and increases cleistothecia. |
| AN7169 | −100 | 4 | 12 | 12 | fhbA; Flavohemoglobin, negative regulation of sexual sporulation and positive regulation of sterigmatocystin biosynthesis. |
| AN0807 | −1913 | −2 | - | −4 | laeA; Methyltransferase-domain protein; self-methylates; coordinates asexual development in response to light; regulates secondary metabolism and is required for Hulle cell formation. |
| AN0387 | - | 2 | −37 | - | cryA: Negative regulation of cleistothecium development. |
| AN4783 | 7 | - | −2 | - | csnB; COP9 signalosome subunit 2 (CsnB) formation of cleistothecia. |
| AN2421 | 4 | −3 | - | −31 | flbC: regulation of conidium formation and spore germination. |
| AN7553 | - | −3 | −5 | - | devR; Basic helix-loop-helix transcription factor; conidiophore development. |
| AN5893 | −5 | 2 | - | - | flbA; Developmental regulator FlbA, conidiophore development and asexual sporulation. |
| AN0082 | −6 | −2 | −2 | −2 | phnA; Phosducin, putative; regulates sporulation |
| AN3387 | - | 2 | 16 | −2 | gprD: Deletion of gprD resulted in delayed conidial germination and enhanced sexual development. |
| AN0279 | - | −2 | 2 | −12 | flbD: Myb-like DNA-binding protein; positive regulation of conidiation. |
| AN2290 | - | - | - | −5 | steA; Sexual development transcription factor SteA; Required for cleistothecial development and ascosporogenesis. Not required for conidiation. |
| AN0170 | - | - | - | −24 | trxA; Thioredoxin, required for conidiation; expression upregulated after exposure to farnesol. |
| AN9121 | 2 | - | - | −200 | esdC; sexual development protein, involved in early sexual development and regulated by VeA and FlbA. |
| AN4163 | - | 2 | −11 | - | cpcB: G-protein complex beta subunit CpcB; positive regulation of cleistothecium development. |
| AN1017 | 2 | −2 | −6 | 2 | hogA; Putative mitogen-activated protein kinase; required for sexual development and sporulation. |
| AN7795 | - | - | 39 | - | gprK; positive regulation of sexual sporulation [40]. |
| Up-regulated | Down-regulated | ||||
| Colour | Fold change | Colour | Fold change | ||
| <5 | <5 | ||||
| 5 to 20 | 5 to 20 | ||||
| 21 to 100 | 21 to 100 | ||||
| 101 to 500 | 101 to 500 | ||||
| > 500 | > 500 | ||||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ejmal, M.A.; Holland, D.J.; MacDiarmid, R.M.; Pearson, M.N. The Effect of Aspergillus Thermomutatus Chrysovirus 1 on the Biology of Three Aspergillus Species. Viruses 2018, 10, 539. https://doi.org/10.3390/v10100539
Ejmal MA, Holland DJ, MacDiarmid RM, Pearson MN. The Effect of Aspergillus Thermomutatus Chrysovirus 1 on the Biology of Three Aspergillus Species. Viruses. 2018; 10(10):539. https://doi.org/10.3390/v10100539
Chicago/Turabian StyleEjmal, Mahjoub A., David J. Holland, Robin M. MacDiarmid, and Michael N. Pearson. 2018. "The Effect of Aspergillus Thermomutatus Chrysovirus 1 on the Biology of Three Aspergillus Species" Viruses 10, no. 10: 539. https://doi.org/10.3390/v10100539






