Genetic Diversity of Paeonia rockii (Flare Tree Peony) Germplasm Accessions Revealed by Phenotypic Traits, EST-SSR Markers and Chloroplast DNA Sequences
Abstract
1. Introduction
2. Materials and Methods
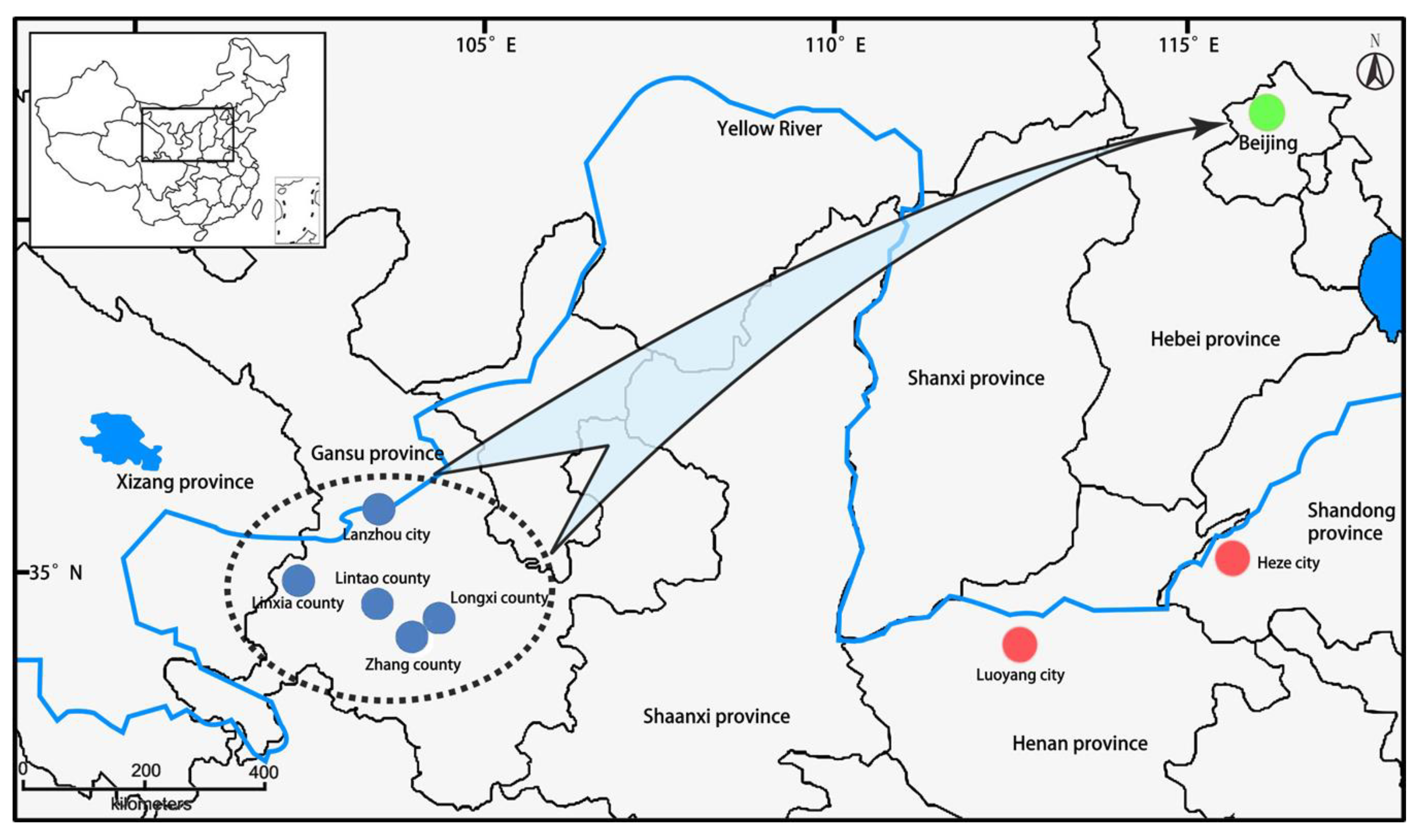
2.1. Plant Materials and DNA Extraction
2.2. Measurement of Phenotypic Traits
2.3. Microsatellites Markers
2.4. Chloroplast DNA Sequences
2.5. Statistical Analyses
2.5.1. Analysis of Phenotypic Data
2.5.2. Analysis of EST-SSR Data
2.5.3. Analysis of Chloroplast DNA Sequences
3. Results
3.1. Genetic Diversity Based on Phenotypic Traits
3.1.1. Phenotypic Traits Variation
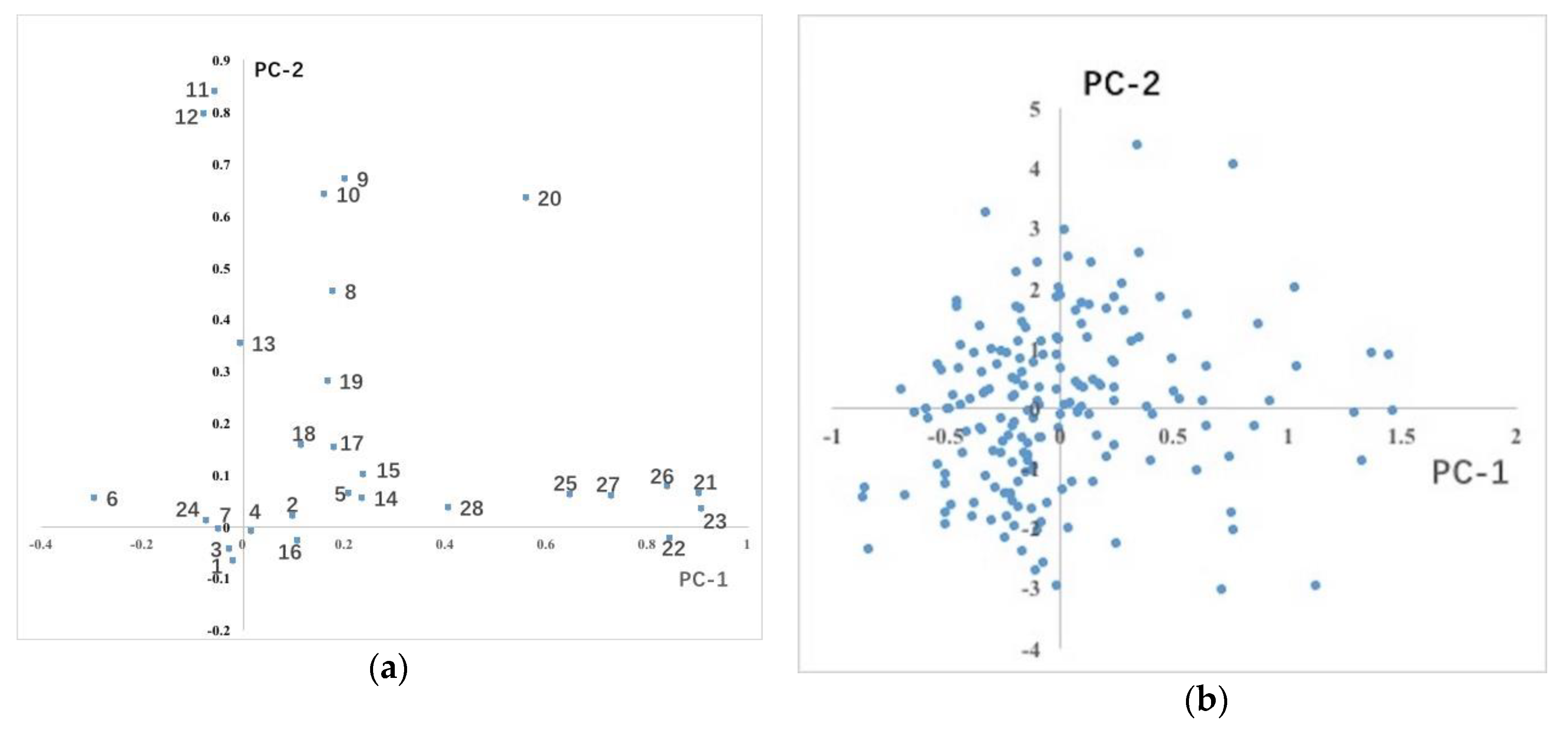
3.1.2. Distribution of Phenotypic Traits Variation in the Population
3.1.3. Correlation Analysis of Quantitative Traits
3.2. Genetic Diversity Based on the EST-SSR
3.2.1. EST-SSR Polymorphism
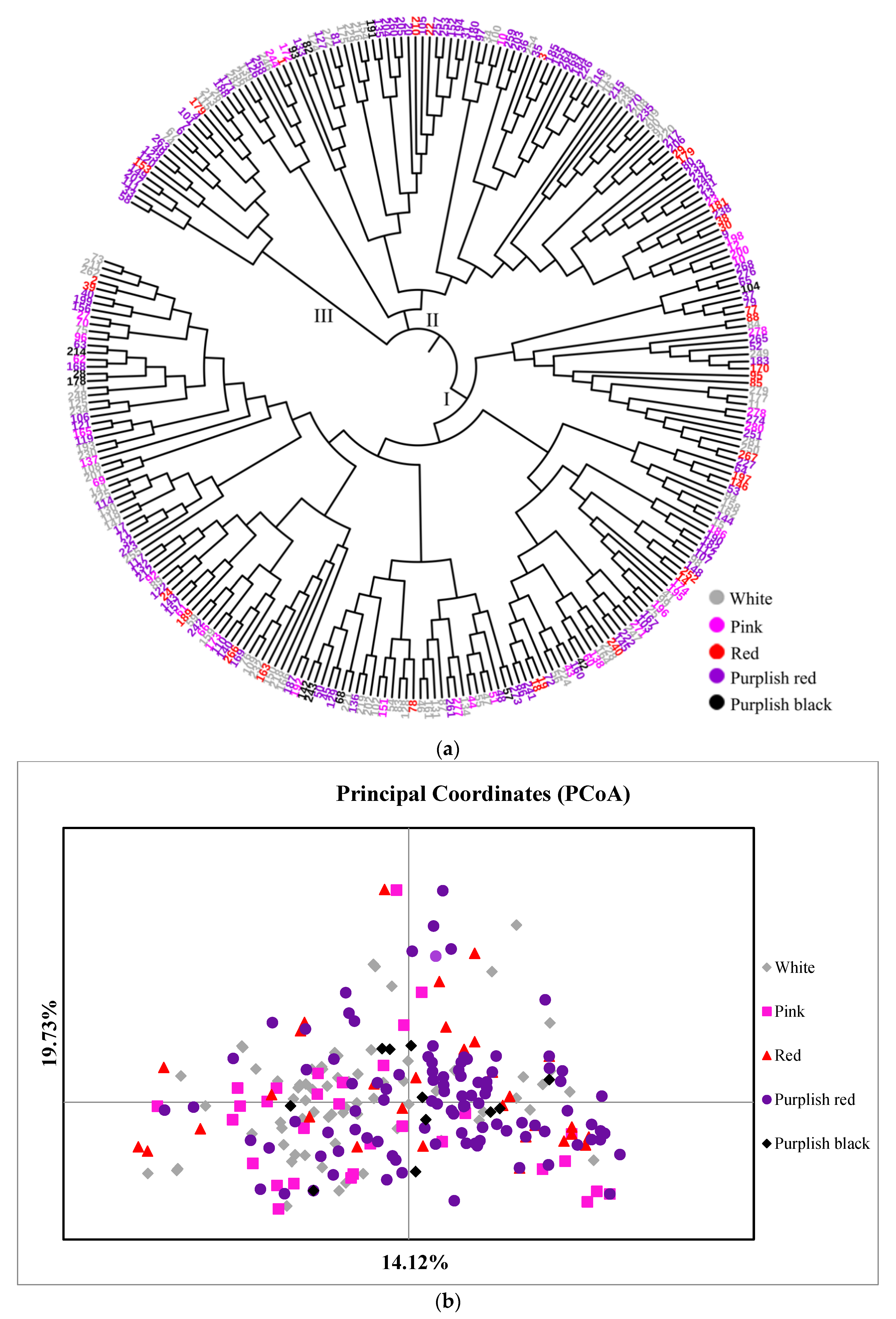
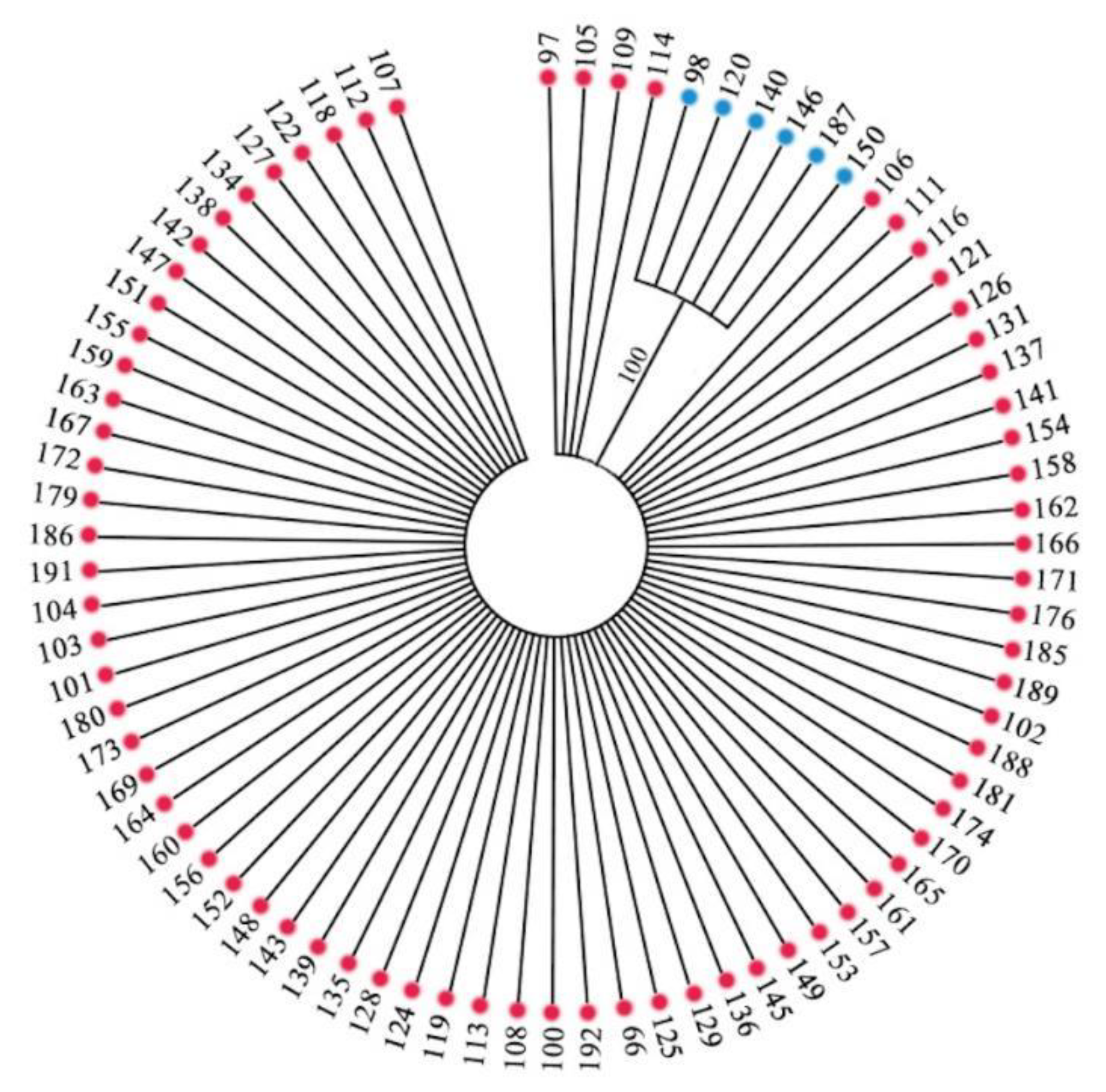
3.2.2. Genetic Relationships among 282 Accessions
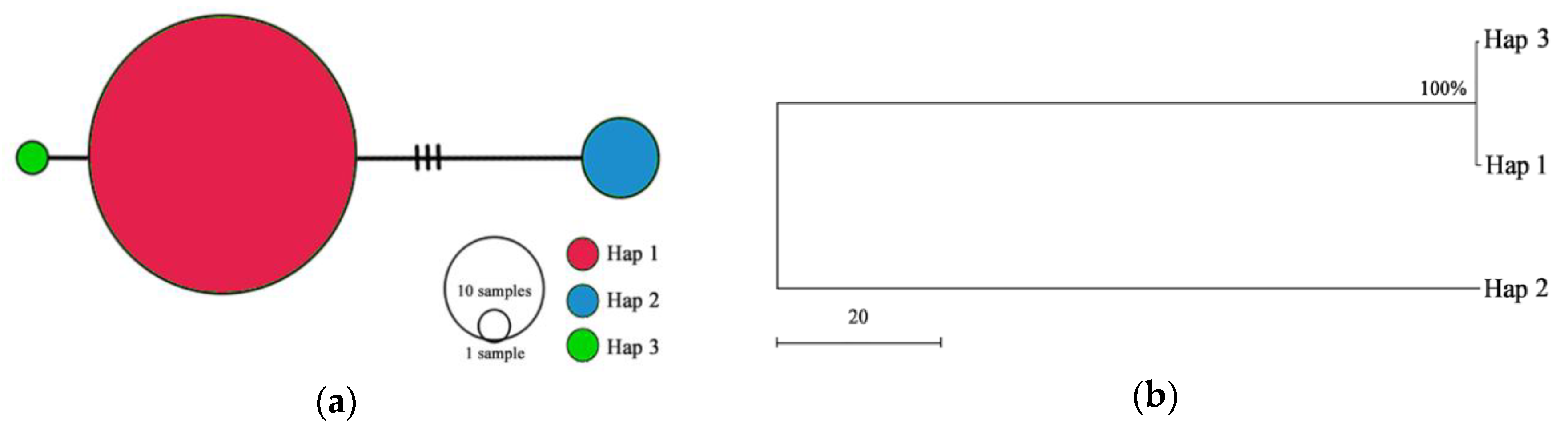
3.3. Genetic Diversity Based on the cpDNA
4. Discussion
4.1. Diversity of Phenotypic Traits
4.2. Genetic Diversity Based on SSR Markers
4.3. Genetic Diversity Based on the cpDNA
4.4. Conversation and Utilization of FTP Germplasm Accessions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cheng, F.Y.; Li, J.J.; Chen, D.Z.; Zhang, Z.S. Chinese Paeonia rockii; Chinese Forestry Publishing House: Beijing, China, 2005; pp. 20–30. [Google Scholar]
- Han, P.; Ruan, C.J.; Ding, J.; Wu, B.; Zhang, W.C.; Ruan, D.; Xiong, C.W.; Liu, W.H.; Wang, G.H. Polygene regulation of high accumulation of carbon 18 unsaturated fatty acids in seeds of Paeonia suffruticosa. Mol. Plant Breed. 2019, 17, 2101–2108. (In Chinese) [Google Scholar]
- Cheng, F.Y. Advances in the breeding of tree peonies and a cultivar system for the cultivar group. Int. J. Plant Breed. 2007, 1, 90–104. [Google Scholar]
- Cheng, F.Y.; Yu, X.N. Flare tree peonies (Paeonia rockii hybrids) and the origin of the cultivar group. Acta Horticulturae 2008, 766, 375–382. [Google Scholar] [CrossRef]
- Li, J.J. Chinese Tree Peony (Xibei, Xinan, Jiangnan Volume); Chinese Forestry Publishing House: Beijing, China, 2005; pp. 73–82. [Google Scholar]
- Zhang, Q.Y.; Niu, L.X.; Yu, R.; Zhang, X.X.; Bai, Z.Z.; Duan, K.; Gao, Q.H.; Zhang, Y.L. Cloning, characterization, and expression analysis of a gene encoding a putative lysophosphatidic acid acyltransferase from seeds of Paeonia rockii. Appl. Biochem. Biotech. 2017, 182, 721–741. [Google Scholar] [CrossRef]
- Shi, Q.Q.; Li, L.; Zhang, X.X.; Luo, J.R.; Li, X.; Zhai, L.J.; He, L.X.; Zhang, Y.L. Biochemical and Comparative Transcriptomic Analyses Identify Candidate Genes Related to Variegation Formation in Paeonia rockii. Molecules 2017, 22, 1364. [Google Scholar] [CrossRef]
- Zhang, Q.Y.; Yu, R.; Sun, D.Y.; Bai, Z.Z.; Li, H.; Xue, L.; Zhang, Y.L.; Niu, L.X. PrLPAAT4, a putative lysophosphatidic acid acyltransferase from Paeonia rockii, plays an important role in seed fatty acid biosynthesis. Molecules 2017, 22, 1694. [Google Scholar] [CrossRef]
- Cui, H.L.; Chen, C.R.; Huang, N.Z.; Cheng, F.Y. Association analysis of yield, oil and fatty acid content, and main phenotypic traits in Paeonia rockii as an oil crop. J. Hortic. Sci. Biotechnol. 2017, 93, 425–432. [Google Scholar] [CrossRef]
- Shi, Q.Q.; Zhang, X.X.; Li, X.; Zhai, L.J.; Luo, X.N.; Luo, J.R.; He, L.X.; Zhang, Y.L.; Li, L. Identification of microRNAs and their targets involved in Paeonia rockii petal variegation using high-throughput sequencing. J. Am. Soc. Hortic. Sci. 2019, 144, 118–129. [Google Scholar] [CrossRef]
- Liu, N.; Cheng, F.Y.; Zhong, Y.; Guo, X. Comparative transcriptome and coexpression network analysis of carpel quantitative variation in Paeonia rockii. BMC Genomics 2019, 20, 683. [Google Scholar] [CrossRef]
- Soltis, P.S.; Soltis, D.E. Genetic variation in endemic and widespread plant species. Aliso J. Syst. Evolut. Bot. 1991, 13, 215–223. [Google Scholar] [CrossRef]
- Wang, X.Q. Studies on Genetic Diversity of Paeonia delavayi in Shangri-la. Ph.D. Thesis, Beijing Forestry University, Beijing, China, 2009. Available online: https://kns.cnki.net/KCMS/detail/detail.aspx?dbcode=CDFD&dbname=CDFD0911&filename=2009134629.nh&v=MTA1MDExMjdGN0s3R3RmT3BwRWJQSVI4ZVgxTHV4WVM3RGgxVDNxVHJXTTFGckNVUjdxZlplUnFGQ25sVzd2S1Y= (accessed on 12 June 2020). (In Chinese).
- Wang, J. Genetic Diversity of Paeonia ostii and Germplasm Resources of Tree Peony Cultivars from Chinese Jiangnan Area. Ph.D. Thesis, Beijing Forestry University, Beijing, China, 2009. (In Chinese). [Google Scholar]
- Peng, L.P.; Cheng, F.Y.; Zhong, Y.; Xu, X.X.; Xian, H.L. Phenotypic variation in cultivar populations of Paeonia ostii. Plant Sci. J. 2018, 36, 170–180. (In Chinese) [Google Scholar]
- Li, Z.Y.; Zhang, H.Y. Morphological variation and diversity in populations of Paeonia lutea. J. Northwest. For. Univ. 2011, 26, 117–122. (In Chinese) [Google Scholar]
- Li, B.Y. Studies on Genetic Diversity and Construction of Core Collection of the Tree Peony Cultivars from Chinese Central Plains. Ph.D. Thesis, Beijing Forestry University, Beijing, China, 2007. (In Chinese). [Google Scholar]
- Yuan, T.; Wang, L.Y. Discussion on the origination of Chinese tree-peony cultivars according to pollen grain morphology. J. Beijing Fore. Univ. 2002, 24, 5–11. (In Chinese) [Google Scholar]
- Zhou, B.; Jiang, H.D.; Zhang, X.X.; Xue, J.Q.; Shi, Y.T. Morphological diversity of some introduced tree peony cultivars. Biodiv. Sci. 2011, 19, 543–550. (In Chinese) [Google Scholar] [CrossRef]
- Hosoki, T.; Kimura, D.; Hasegawa, R.; Nagasako, T.; Nishimoto, K.; Ohta, K.; Sugiyama, M.; Haruki, K. Comparative study of Chinese tree peony cultivars by random amplified polymorphic DNA (RAPD) analysis. Sci. Hortic. 1997, 70, 67–72. [Google Scholar] [CrossRef]
- Yu, H.P.; Cheng, F.Y.; Zhong, Y.; Cai, C.F.; Wu, J.; Cui, H.L. Development of simple sequence repeat (SSR) markers from Paeonia ostii to study the genetic relationships among tree peonies (Paeoniaceae). Sci. Hortic. 2013, 164, 58–64. [Google Scholar] [CrossRef]
- Wang, X.W.; Fan, H.M.; Li, Y.Y.; Sun, X.; Sun, X.Z.; Wang, W.L.; Zheng, C.S. Analysis of genetic relationships in tree peony of different colors using conserved DNA-derived polymorphism markers. Sci. Hortic. 2014, 175, 68–73. [Google Scholar] [CrossRef]
- Suo, Z.L.; Li, W.Y.; Yao, J.; Zhang, H.J.; Zhang, Z.M.; Zhao, D.X. Applicability of leaf morphology and intersimple sequence repeat markers in classification of tree peony (Paeoniaceae) cultivars. Hortscience 2005, 40, 329–334. [Google Scholar] [CrossRef]
- Zhang, J.J.; Shu, Q.Y.; Liu, Z.A.; Ren, H.X.; Wang, L.S.; De Keyser, E. Two EST-derived marker systems for cultivar identification in tree peony. Plant Cell Rep. 2012, 31, 299–310. [Google Scholar] [CrossRef]
- Duan, Y.B.; Guo, D.L.; Guo, L.L.; Wei, D.F.; Hou, X.G. Genetic diversity analysis of tree peony germplasm using iPBS markers. Genet. Mol. Res. 2015, 14, 7556–7566. [Google Scholar] [CrossRef]
- Suo, Z.L.; Zhang, C.H.; Zheng, Y.Q.; He, L.X.; Jin, X.B.; Hou, B.X.; Li, J.J. Revealing genetic diversity of tree peonies at micro-evolution level with hyper-variable chloroplast markers and floral traits. Plant Cell Rep. 2012, 31, 2199–2213. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.H.; Cheng, F.Y.; Zhou, S.L. Hybrid origin of Paeonia × yananensis revealed by microsatellite markers, chloroplast gene sequences, and morphological characteristics. Int. J. Plant Sci. 2010, 171, 409–420. [Google Scholar] [CrossRef]
- Yuan, J.H.; Cheng, F.Y.; Zhou, S.L. The phylogeographic structure and conservation genetics of the endangered tree peony, Paeonia rockii (Paeoniaceae), inferred from chloroplast gene sequences. Conserv. Genet. 2011, 12, 1539–1549. [Google Scholar] [CrossRef]
- Yuan, J.H.; Cheng, F.Y.; Zhou, S.L. Genetic structure of the tree peony (Paeonia rockii) and the Qinling Mountains as a geographic barrier driving the fragmentation of a large population. PLoS ONE 2012, 7, e34955. [Google Scholar] [CrossRef] [PubMed]
- Ren, X.X.; Zhang, Y.; Xue, J.Q.; Zhu, F.Y.; Shi, F.R.; Wang, S.L.; Zhang, X.X. Genetic diversity analysis of natural populations in Paeonia delavayi. J. Plant Gen. Res. 2015, 16, 772–780. (In Chinese) [Google Scholar]
- Tang, Q.; Zeng, X.L.; Liao, M.A.; Pan, G.T.; Zha, X.; Gong, J.H.; Ci Ren, Z.G. SRAP analysis of genetic diversity of Paeonia ludlowii in Tibet. Sci. Silva. Sin. 2012, 48, 70–76. (In Chinese) [Google Scholar]
- Zhang, J.M.; Liu, J.; Sun, H.L.; Yu, J.; Wang, J.X.; Zhou, S.L. Nuclear and chloroplast SSR markers in Paeonia delavayi (Paeoniaceae) and cross-species amplification in P. ludlowii. Am. J. Bot. 2011, 98, e346–e348. [Google Scholar] [CrossRef]
- Zhang, J.M.; López-Pujol, J.; Gong, X.; Wang, H.F.; Vilatersana, R.; Zhou, S.L. Population genetic dynamics of Himalayan-Hengduan tree peonies, Paeonia subsect. Delavayanae. Mol. Phylogenet. Evol. 2018, 125, 62–77. [Google Scholar] [CrossRef]
- Tong, F.; Xie, D.F.; Zeng, X.M.; He, X.J. Genetic diversity of Paeonia decomposita and Paeonia decomposita subsp. rotundiloba detected by ISSR markers. Acta Bot. Boreal.-Occident. Sin. 2016, 36, 1968–1976. (In Chinese) [Google Scholar]
- Xu, X.X.; Cheng, F.Y.; Xian, H.L.; Peng, L.P. Genetic diversity and population structure of endangered endemic Paeonia jishanensis in China and conservation implications. Biochem. Syst. Ecol. 2016, 66, 319–325. [Google Scholar] [CrossRef]
- Peng, L.P.; Cai, C.F.; Zhong, Y.; Xu, X.X.; Xian, H.L.; Cheng, F.Y.; Mao, J.F. Genetic analyses reveal independent domestication origins of the emerging oil crop Paeonia ostii, a tree peony with a long-term cultivation history. Sci. Rep. 2017, 7, 5340–5352. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.X.; Cheng, F.Y.; Peng, L.P.; Sun, Y.Q.; Hu, X.G.; Li, S.Y.; Xian, H.L.; Jia, K.H.; Abbott, R.J.; Mao, J.F. Late Pleistocene speciation of three closely related tree peonies endemic to the Qinling-Daba Mountains, a major glacial refugium in Central China. Ecol. Evol. 2019, 9, 7528–7548. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Cheng, F.Y.; Cai, C.F.; Zhong, Y.; Jie, X. Association mapping for floral traits in cultivated Paeonia rockii based on SSR markers. Mol. Genet. Genom. 2016, 292, 187–200. [Google Scholar] [CrossRef] [PubMed]
- Weber, J.L.; May, P.E. Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Am. J. Hum. Genet. 1989, 44, 388–396. [Google Scholar] [PubMed]
- Dumolin, S.; Demesure, B.; Petit, R.J. Inheritance of chloroplast and mitochondrial genomes in pedunculate oak investigated with an efficient PCR method. Theor. Appl. Genet. 1995, 91, 1253–1256. [Google Scholar] [CrossRef]
- Comes, H.P.; Kadereit, J.W. The effect of Quaternary climatic changes on plant distribution and evolution. Trends Plant Sci. 1998, 3, 1360–1385. [Google Scholar] [CrossRef]
- Qiu, Y.X.; Fu, C.X.; Comes, H.P. Plant molecular phylogeography in China and adjacent regions: Tracing the genetic imprints of Quaternary climate and environmental change in the world’s most diverse temperate flora. Mol. Phylogenet. Evol. 2011, 59, 225–244. [Google Scholar] [CrossRef]
- Cheng, Y.P.; Hwang, S.Y.; Lin, T.P. Potential refugia in Taiwan revealed by the phylogeographical study of Castanopsis carlesii Hayata (Fagaceae). Mol. Ecol. 2005, 14, 2075–2085. [Google Scholar] [CrossRef]
- Cai, C.F. High-Density Genetic Linkage Map Construction and QTLs Analyses for Phenotypic Traits in Tree Peony. Ph.D. Thesis, Beijing Forestry University, Beijing, China, 2015. (In Chinese). [Google Scholar]
- Wu, J.; Cai, C.F.; Cheng, F.Y.; Cui, H.L.; Zhou, H. Characterisation and development of EST-SSR markers in tree peony using transcriptome sequences. Mol. Breed. 2014, 34, 1853–1866. [Google Scholar] [CrossRef]
- Bousquet, J.; Strauss, S.H.; Doerksen, A.H.; Price, R.A. Extensive variation in evolutionary rate of rbcL gene sequences among seed plants. Proc. Natl. Acad. Sci. USA 1992, 89, 7844–7848. [Google Scholar] [CrossRef]
- Gaut, B.S.; Muse, S.V.; Clark, Z.W.D.; Clegg, M.T. Relative rates of nucleotide substitution at the rbcl locus of monocotyledonous plants. J. Mol. Evol. 1992, 35, 292–303. [Google Scholar] [CrossRef]
- Baker, W.J.; Hedderson, T.A.; Dransfield, J. Molecular phylogenetics of subfamily Calamoideae (Palmae) based on nrDNA ITS and cpDNA rps16 intron sequence data. Mol. Phylogenet. Evol. 2000, 14, 195–217. [Google Scholar] [CrossRef] [PubMed]
- Grivet, D.; Heinze, B.; Vendramin, G.G.; Petit, R.J. Genome walking with consensus primers application to the large single copy region of chloroplast DNA. Mol. Ecol. Notes 2001, 1, 345–349. [Google Scholar] [CrossRef]
- Davis, J.W. Handbook of univariate and multivariate data analysis and interpretation with SPSS; CRC Press: Boca Raton, FL, USA, 2008. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Rousset, F. Genepop’007: A complete re-implementation of the genepop software for Windows and Linux. Mol. Ecol. Resour. 2008, 8, 103–106. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K.; Battistuzzi, F.U. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.-i.; Miyata, T. MAFFT a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef]
- Vaidya, G.; Lohman, D.J.; Meier, R. SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2011, 27, 171–180. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Leigh, J.W.; Bryant, D.; Nakagawa, S. Popart: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Swofford, D.L. Paup*: Phylogenetic Analysis Using Parsimony (*and other Methods); Version 4; Sinauer: Sunderland, MA, USA, 2003; pp. 233–234. [Google Scholar]
- Pang, L.Z.; Cheng, F.Y.; Zhong, Y.; Cai, C.F.; Cui, H.L. Phenotypic analysis of association population for flare tree peony. J. Beijing Fore. Univ. 2012, 34, 115–120. (In Chinese) [Google Scholar]
- Wu, J. Dissection of Allelic Variation Underling Imoortant Traits in Paeonia rockii by Using Association Mapping. Ph.D. Thesis, Beijing Forestry University, Beijing, China, 2016. (In Chinese). [Google Scholar]
- Warschefsky, E.; Penmetsa, R.V.; Cook, D.R.; Von Wettberg, E.J.B. Back to the wilds: Tapping evolutionary adaptations for resilient crops through systematic hybridization with crop wild relatives. Am. J. Bot. 2014, 101, 1791–1800. [Google Scholar] [CrossRef]
- Bernard, A.; Lheureux, F.; Dirlewanger, E. Walnut: Past and future of genetic improvement. Tree Genet. Genomes 2017, 14, 1–28. [Google Scholar] [CrossRef]
- Yuan, J.H.; Cornille, A.; Giraud, T.; Cheng, F.Y.; Hu, Y.H. Independent domestications of cultivated tree peonies from different wild peony species. Mol. Ecol. 2014, 23, 82–95. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.L.; Hou, X.G.; Zhang, J. Sequence-related amplified polymorphism analysis of tree peony (Paeonia suffruticosa Andrews) cultivars with different flower colours. J. Hortic. Sci. Biotechnol. 2015, 84, 131–136. [Google Scholar] [CrossRef]
- Nielsen, R. Molecular signatures of natural selection. Annu. Rev. Genet. 2005, 39, 197–218. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.J.; Gross, B.L. From forest to field: Perennial fruit crop domestication. Am. J. Bot. 2011, 98, 1389–1414. [Google Scholar] [CrossRef] [PubMed]
- Margules, C.R.; Pressey, R.L. Systematic conservation planning. Nature 2000, 405, 243–253. [Google Scholar] [CrossRef]
- Rodrigues, L.; van den Berg, C.; Póvoa, O.; Monteiro, A. Low genetic diversity and significant structuring in the endangered Mentha cervina populations and its implications for conservation. Biochem. Syst. Ecol. 2013, 50, 51–61. [Google Scholar] [CrossRef]
- Migicovsky, Z.; Warschefsky, E.; Klein, L.L.; Miller, A.J. Using living germplasm collections to characterize, improve, and conserve woody perennials. Crop Sci. 2019, 59, 2365–2380. [Google Scholar] [CrossRef]

| Type | Code | Trait | Measurement Standard |
|---|---|---|---|
| Floral trait | 1 | Petal length | The length of the outer petal (cm) |
| 2 | Petal width | The width of the outer petal (cm) | |
| 3 | Flare length | The length of flare (cm) | |
| 4 | Flare width | The width of flare (cm) | |
| 5 | Flower diameter | The maximum width of flower at full bloom (cm) | |
| 6 | Petal number | The number of petals in the whole flower (score) | |
| 7 | Carpel number | The number of carpels in the whole flower (score) | |
| Branch and leaf trait | 8 | Plant height | The aboveground height of the whole plant (cm) |
| 9 | East-west crown breadth | The width of a plant in the east-west direction (cm) | |
| 10 | North-south crown breadth | The width of a plant in the north-south direction (cm) | |
| 11 | Fruit number | The number of follicles per plant (score) | |
| 12 | Flower number | The number of flowers per plant (score) | |
| 13 | Tiller number | The number of branches at the base of a plant (score) | |
| 14 | Compound leaf length | The length from tip of compound leaf to petiole (cm) | |
| 15 | Compound leaf width | The length at widest part of compound leaf (cm) | |
| 16 | Petiole length | The length of petiole (cm) | |
| 17 | Leaflet number | The number of leaflets in a compound leaf (score) | |
| 18 | Terminal leaflet length | The length of terminal leaflet (cm) | |
| 19 | Terminal leaflet width | The width of terminal leaflet (cm) | |
| Fruit trait | 20 | Fruit weight per plant | The weight of fruits per plant (g) |
| 21 | Individual fruit weight | The weight of one fruit (g) | |
| 22 | Number of seeds per fruit | The number of seeds per fruit (score) | |
| 23 | Seed weight per fruit | The weight of seeds per fruit (g) | |
| 24 | Effective carpel number | The number of carpels that produce seeds (score) | |
| 25 | Fruit length | The length of a single carpel of mature follicles (mm) | |
| 26 | Fruit width | The width of a single carpel of mature follicles (mm) | |
| 27 | Fruit height | The height of a single carpel of mature follicles (mm) | |
| 28 | Pericarp thickness | The thickness of pericarp of mature follicles (mm) |
| Locus | Primer Sequence | Tm (°C) | Expected Size (bp) | Modified Type | Na | Ne | I | Ho | He | FIS | PIC | p |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PS004 | F: GTGCTTAGCCTCTAATCTG R: CTTTGCTCCAAGTCTGTC | 50.5 | 274 | 5-FAM | 7 | 2.859 | 1.330 | 0.688 | 0.650 | −0.058 | 0.611 | 0.000 *** |
| PS026 | F: TTCCCTCCATTCTAACAC R: ACCCTAGCCTCTGACATT | 54 | 187 | 5-FAM | 5 | 1.761 | 0.715 | 0.475 | 0.432 | −0.100 | 0.358 | 0.000 *** |
| PS030 | F: ACCCTCCACCACCATCTT R: TACTCCATCTCGTGACCC | 57 | 237 | 5-HEX | 3 | 1.290 | 0.414 | 0.216 | 0.225 | 0.038 | 0.203 | 0.000 *** |
| PS047 | F: AGACGACGAGCAAAGATAT R: AAAGGGCAAGATTGGAAAT | 54 | 126 | 5-FAM | 4 | 1.643 | 0.735 | 0.397 | 0.391 | −0.015 | 0.356 | 0.343 ns |
| PS061 | F: CTCCTCCAACATTGACCC R: CACCCTCCCAAACATCTC | 57 | 154 | 5-FAM | 7 | 2.178 | 0.901 | 0.532 | 0.541 | 0.017 | 0.436 | 0.000 *** |
| PS068 | F: CTTTGGCATTCTCATTCA R: GGTGGTATTGGGCTTCTT | 52.5 | 174 | 5-FAM | 6 | 3.231 | 1.335 | 0.738 | 0.690 | −0.068 | 0.641 | 0.000 *** |
| PS073 | F: GTCGGTGAATGAAGGGTT R: ATTTCTGGTCAATGTGGC | 53.5 | 269 | 5-HEX | 5 | 2.066 | 0.780 | 0.574 | 0.516 | −0.113 | 0.400 | 0.000 *** |
| PS074 | F: TGCCTTGCTCCTCCTTGT R: CGGTTAGCCATGAATCCC | 57 | 236 | 5-HEX | 13 | 2.947 | 1.457 | 0.702 | 0.661 | −0.063 | 0.621 | 0.000 *** |
| PS095 | F: TCCCAAGACCTCAAACAAC R: CCATCAATACGAGCCAAC | 55 | 394 | 5-TAMRA | 9 | 5.394 | 1.857 | 0.819 | 0.815 | −0.006 | 0.792 | 0.000 *** |
| PS119 | F: GCAAAGACAACAGCCTCG R: CTCACCATCCAATCCCAC | 57 | 289 | 5-HEX | 4 | 3.627 | 1.331 | 0.745 | 0.724 | −0.028 | 0.673 | 0.000 *** |
| PS139 | F: CAACAATTTAACACGCAGAG R: GCCTTAGACGGAGACCAG | 56.5 | 482 | 5-FAM | 7 | 1.765 | 0.853 | 0.447 | 0.433 | −0.031 | 0.391 | 0.000 *** |
| PS144 | F: CAACCTACAATCCGACAATG R: CGACTTCCCTTCAATACA | 54.5 | 317 | 5-TAMRA | 6 | 1.378 | 0.553 | 0.273 | 0.274 | 0.005 | 0.252 | 0.000 *** |
| PS149 | F: AGTCGCCTCCTACACCTC R: TCCGTAAAGCCCACAATAC | 55.5 | 173 | 5-FAM | 5 | 1.777 | 0.798 | 0.426 | 0.437 | 0.027 | 0.391 | 0.741 ns |
| PS157 | F: CTCCCTGAACTCCCTACC R: CTTTCTAAACAGCCAACG | 56 | 322 | 5-TAMRA | 4 | 2.565 | 1.091 | 0.677 | 0.610 | −0.110 | 0.551 | 0.000 *** |
| PS158 | F: TTTCCCTGCTTCTTCTGAC R: CACCTCCTTCCTTTCTTACT | 55 | 423 | 5-ROX | 7 | 2.976 | 1.298 | 0.638 | 0.664 | 0.039 | 0.608 | 0.000 *** |
| PS159 | F: CCTCCATTCATTCCTGTC R: GCAATAAATAGCCGTCCT | 52 | 124 | 5-FAM | 11 | 2.384 | 1.134 | 0.567 | 0.581 | 0.023 | 0.506 | 0.000 *** |
| PS166 | F: TTCAGTGGGCAAGACCTAC R: TAGCCAATACAGAACAAACC | 55 | 337 | 5-TAMRA | 10 | 3.698 | 1.455 | 0.805 | 0.730 | −0.103 | 0.683 | 0.000 *** |
| PS180 | F: CCCCGAAATGGAGGAGTC R: AGGGCAGTAGCAGAAGAAAGTC | 60 | 188 | 5-FAM | 6 | 1.331 | 0.485 | 0.238 | 0.248 | 0.044 | 0.226 | 0.992 ns |
| PS187 | F: AAGCGGCGTCCATCATAC R: TCACAAGCCCAACCCAGA | 57 | 233 | 5-HEX | 4 | 2.399 | 1.000 | 0.784 | 0.583 | −0.344 | 0.498 | 0.000 *** |
| PS221 | F: GATACAAGGCGGAAAGTG R: AGAGTTGGGAACCAGACC | 56 | 301 | 5-TAMRA | 5 | 2.488 | 1.007 | 0.823 | 0.598 | −0.375 | 0.514 | 0.000 *** |
| PS260 | F: ATTCACGCCAGTATCAAAG R: TGTAAATGCCCATGTCTAG | 53 | 349 | 5-TAMRA | 2 | 1.595 | 0.560 | 0.376 | 0.373 | −0.007 | 0.304 | 0.905 ns |
| PS265 | F: TTTTATGGGTCCTGTTGC R: GAAGAGTAAGCCTTTGTCG | 54 | 290 | 5-TAMRA | 4 | 1.897 | 0.759 | 0.564 | 0.473 | −0.192 | 0.386 | 0.008 ** |
| PS271 | F: AGAATCCACCTCCTGTCAC R: AACCCTGCCCTAAACTAAAC | 56.5 | 406 | 5-ROX | 4 | 3.516 | 1.317 | 0.716 | 0.716 | −0.001 | 0.664 | 0.903 ns |
| PS276 | F: CTGTATCCTATCGGTTCTT R: CCTCATCTGCCTTTATCT | 52.5 | 447 | 5-ROX | 4 | 2.029 | 0.749 | 0.436 | 0.507 | 0.140 | 0.389 | 0.000 ns |
| PS296 | F: CTCTTTCGCTGCCACAAC R: CTCTGCTCTTCCCGTCTT | 57.5 | 419 | 5-ROX | 4 | 1.985 | 0.937 | 0.521 | 0.496 | −0.051 | 0.456 | 0.002 ** |
| PS309 | F: AAGCAAAGCCGTGGAGAT R: GTGCGTGAAAAGGAGACAGAAC | 55 | 257 | 5-HEX | 7 | 1.321 | 0.536 | 0.255 | 0.243 | −0.051 | 0.230 | 0.000 *** |
| PS311 | F: AACGCCACCATCACCTTT R: CACCTGAACTCACCCTCC | 60 | 277 | 5-HEX | 2 | 1.988 | 0.690 | 0.915 | 0.497 | −0.841 | 0.374 | 0.000 *** |
| PS323 | F: CTCACCCGTTCTAAAGTCA R: CCTCCTCCCTGTTCTTCT | 53 | 466 | 5-FAM | 2 | 1.004 | 0.013 | 0.004 | 0.004 | −0.002 | 0.004 | 0.976 ns |
| PS335 | F: TAATCACCCAATGAGCCA R: CGTCGTCGCCGAATACTT | 50 | 395 | 5-ROX | 5 | 1.172 | 0.340 | 0.096 | 0.147 | 0.347 | 0.141 | 0.000 *** |
| PS337 | F: ATCCTCTTCACGGCAATC R: CGTCCACTCTTCCTCCTC | 53.5 | 448 | 5-ROX | 5 | 1.638 | 0.695 | 0.461 | 0.390 | −0.183 | 0.338 | 0.013 * |
| PS339 | F: TGAGGCAGCCAAAGAATT R: GGCAGGTGTAGGGTATGTT | 50 | 175 | 5-FAM | 3 | 1.025 | 0.074 | 0.021 | 0.025 | 0.134 | 0.024 | 0.000 *** |
| PS345 | F: TGAAGTGAATCGAAGCAT R: CAACAGGCAGAAGAAAGG | 48 | 366 | 5-ROX | 4 | 2.078 | 0.944 | 0.589 | 0.519 | −0.135 | 0.469 | 0.000 *** |
| PS356 | F: TCAAGCCCAAGGTCATTC R: ACTTGCTCACCTCGCTCT | 53 | 354 | 5-TAMRA | 7 | 3.647 | 1.476 | 0.730 | 0.726 | −0.006 | 0.683 | 0.000 *** |
| PS367 | F: AGACGGACGGAAATAGGG R: ACGAGCGATCTCAACCAT | 53.5 | 265 | 5-HEX | 4 | 3.238 | 1.267 | 0.993 | 0.691 | −0.437 | 0.636 | 0.000 *** |
| Mean | 5.441 | 2.291 | 0.908 | 0.537 | 0.489 | −0.074 | 0.611 | |||||
| SE | 0.429 | 0.166 | 0.072 | 0.043 | 0.035 | 0.034 | 0.358 |
| Number | Loci | Primer Pairs | Primer Sequences (5′-3′) | Tm (°C) |
|---|---|---|---|---|
| 1 | petB-petD | petB | CTATCGTCCRACCGTTACWGAGGCT | 54 |
| petD | CAAAYGGATAYGCAGGTTCACC | |||
| 2 | accD-psaI | accD-psaI-21F | AACATTGAATAAGACAGTACCTGAG | 52 |
| accD-psaI-747R | GTAAGTTAAGAGTTGTCATAGGATGG | |||
| 3 | psbE-petL | psbE-petL-356F | CCTTCTTCTGACACAGCAATG | 52 |
| psbE-petL-1219R | TTACCATTATAGACAGCACTAACAA |
| Trait | Mean | Standard Deviation | Minimum | Maximum | F-Value | p-Value | Variable Coefficient |
|---|---|---|---|---|---|---|---|
| 1 | 7.54 | 0.93 | 4.80 | 10.07 | 15.96 | 0.00 | 12.33 |
| 2 | 7.41 | 1.19 | 2.73 | 9.80 | 18.11 | 0.00 | 16.06 |
| 3 | 2.71 | 0.63 | 1.57 | 4.37 | 16.09 | 0.00 | 23.25 |
| 4 | 1.97 | 0.58 | 0.67 | 3.77 | 18.43 | 0.00 | 29.44 |
| 5 | 15.41 | 1.90 | 9.67 | 19.63 | 6.97 | 0.00 | 12.33 |
| 6 | 32.55 | 35.39 | 9.00 | 238.33 | 22.72 | 0.00 | 108.73 |
| 7 | 5.28 | 1.05 | 0.00 | 10.33 | 8.08 | 0.00 | 19.89 |
| 8 | 125.92 | 22.56 | 71.33 | 206.67 | 7.42 | 0.00 | 17.92 |
| 9 | 149.02 | 32.42 | 78.67 | 276.00 | 4.15 | 0.00 | 21.76 |
| 10 | 163.84 | 34.41 | 75.33 | 277.00 | 5.37 | 0.00 | 21.00 |
| 11 | 28.24 | 18.88 | 0.00 | 97.33 | 2.99 | 0.00 | 66.86 |
| 12 | 32.04 | 18.40 | 4.33 | 86.00 | 1.56 | 0.08 | 57.43 |
| 13 | 21.02 | 19.03 | 1.00 | 169.33 | 7.71 | 0.00 | 90.53 |
| 14 | 36.98 | 6.24 | 20.67 | 55.84 | 9.46 | 0.00 | 16.87 |
| 15 | 23.13 | 5.57 | 11.67 | 45.07 | 7.26 | 0.00 | 24.08 |
| 16 | 12.00 | 2.52 | 6.97 | 21.57 | 9.96 | 0.00 | 21.00 |
| 17 | 15.15 | 3.41 | 9.00 | 31.00 | 14.87 | 0.00 | 22.51 |
| 18 | 6.95 | 1.46 | 3.53 | 11.13 | 10.99 | 0.00 | 21.01 |
| 19 | 7.06 | 2.07 | 3.13 | 15.30 | 5.21 | 0.00 | 29.32 |
| 20 | 478.26 | 408.66 | 25.33 | 2661.64 | 4.95 | 0.00 | 85.45 |
| 21 | 62.26 | 31.57 | 16.89 | 237.01 | 3.75 | 0.00 | 50.71 |
| 22 | 33.66 | 31.74 | 0.00 | 151.33 | 4.15 | 0.00 | 94.30 |
| 23 | 15.01 | 15.00 | 0.00 | 79.67 | 3.32 | 0.00 | 99.93 |
| 24 | 5.49 | 1.20 | 2.33 | 12.33 | 19.08 | 0.00 | 21.86 |
| 25 | 40.75 | 7.27 | 23.74 | 75.64 | 11.29 | 0.00 | 17.84 |
| 26 | 15.43 | 3.27 | 7.60 | 25.24 | 10.12 | 0.00 | 21.19 |
| 27 | 14.79 | 2.91 | 6.97 | 29.07 | 10.73 | 0.00 | 19.68 |
| 28 | 2.51 | 0.64 | 0.79 | 4.52 | 12.65 | 0.00 | 25.50 |
| Trait | Principal Component | |||||||
|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | |
| 1 | −0.019 | −0.068 | 0.831 | 0.150 | 0.076 | −0.065 | −0.064 | −0.017 |
| 2 | 0.100 | 0.019 | 0.776 | 0.043 | 0.134 | 0.153 | −0.277 | 0.148 |
| 3 | −0.025 | −0.044 | 0.669 | −0.164 | −0.204 | 0.238 | 0.165 | −0.273 |
| 4 | 0.019 | −0.010 | 0.543 | −0.182 | −0.030 | 0.525 | 0.125 | −0.138 |
| 5 | 0.211 | 0.063 | 0.688 | 0.146 | 0.040 | −0.211 | −0.133 | 0.163 |
| 6 | −0.293 | 0.053 | −0.196 | 0.065 | 0.027 | −0.077 | 0.633 | 0.286 |
| 7 | −0.047 | −0.007 | 0.167 | −0.069 | −0.053 | 0.001 | −0.629 | 0.413 |
| 8 | 0.180 | 0.453 | 0.129 | 0.377 | 0.068 | 0.379 | 0.203 | 0.222 |
| 9 | 0.203 | 0.669 | 0.184 | 0.083 | 0.084 | 0.006 | 0.398 | 0.176 |
| 10 | 0.163 | 0.641 | 0.342 | 0.144 | 0.103 | 0.030 | 0.308 | 0.092 |
| 11 | −0.053 | 0.839 | −0.153 | −0.001 | −0.057 | −0.160 | −0.074 | −0.136 |
| 12 | −0.076 | 0.796 | −0.144 | −0.073 | 0.054 | −0.017 | −0.126 | 0.042 |
| 13 | −0.005 | 0.352 | 0.077 | 0.026 | −0.144 | −0.684 | 0.140 | −0.099 |
| 14 | 0.237 | 0.053 | 0.059 | 0.866 | 0.120 | 0.006 | −0.012 | −0.036 |
| 15 | 0.240 | 0.098 | 0.160 | 0.569 | −0.029 | 0.466 | −0.042 | −0.060 |
| 16 | 0.110 | −0.029 | −0.018 | 0.842 | 0.043 | −0.175 | 0.129 | −0.038 |
| 17 | 0.181 | 0.153 | 0.071 | 0.206 | −0.675 | 0.193 | −0.137 | 0.073 |
| 18 | 0.116 | 0.156 | 0.079 | 0.438 | 0.716 | 0.179 | 0.083 | −0.049 |
| 19 | 0.169 | 0.279 | −0.015 | 0.315 | 0.494 | 0.251 | −0.070 | −0.165 |
| 20 | 0.562 | 0.633 | −0.031 | 0.133 | −0.018 | −0.193 | −0.102 | −0.144 |
| 21 | 0.904 | 0.063 | 0.074 | 0.249 | 0.115 | 0.032 | −0.050 | 0.017 |
| 22 | 0.846 | −0.024 | −0.032 | 0.047 | −0.247 | −0.033 | −0.196 | −0.066 |
| 23 | 0.909 | 0.033 | −0.003 | 0.105 | −0.130 | −0.031 | −0.132 | −0.022 |
| 24 | −0.070 | 0.011 | 0.001 | −0.077 | −0.078 | 0.030 | 0.009 | 0.795 |
| 25 | 0.648 | 0.060 | 0.160 | 0.287 | 0.311 | 0.107 | 0.210 | −0.128 |
| 26 | 0.842 | 0.076 | 0.077 | 0.015 | 0.117 | 0.086 | 0.072 | 0.034 |
| 27 | 0.730 | 0.059 | 0.088 | 0.136 | 0.297 | 0.085 | 0.080 | −0.032 |
| 28 | 0.408 | 0.036 | 0.144 | 0.062 | 0.566 | 0.105 | −0.106 | 0.091 |
| Total variance/% | 17.722 | 11.066 | 10.356 | 9.328 | 7.026 | 5.387 | 5.207 | 4.423 |
| Cumulative total variance/% | 17.722 | 28.789 | 39.144 | 48.472 | 55.499 | 60.886 | 66.093 | 70.516 |
| Trait | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 | 25 | 26 | 27 | 28 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | ||||||||||||||||||||||||||||
| 2 | 0.657 ** | |||||||||||||||||||||||||||
| 3 | 0.505 ** | 0.294 ** | ||||||||||||||||||||||||||
| 4 | 0.354 ** | 0.469 ** | 0.655 ** | |||||||||||||||||||||||||
| 5 | 0.519 ** | 0.539 ** | 0.309 ** | 0.272 ** | ||||||||||||||||||||||||
| 6 | −0.167 * | −0.287 ** | −0.081 | −0.122 | −0.170 ** | |||||||||||||||||||||||
| 7 | 0.102 | 0.220 ** | 0.163 * | 0.115 | 0.148 ** | −0.111 * | ||||||||||||||||||||||
| 8 | 0.135 * | 0.232 ** | 0.034 | 0.180 ** | 0.146 ** | 0.016 | 0.048 | |||||||||||||||||||||
| 9 | 0.160 * | 0.126 | 0.062 | 0.052 | 0.117 * | 0.049 | −0.013 | 0.577 ** | ||||||||||||||||||||
| 10 | 0.192 ** | 0.146 * | 0.096 | 0.095 | 0.165 ** | −0.031 | 0.050 | 0.652 ** | 0.708 ** | |||||||||||||||||||
| 11 | −0.119 | −0.041 | −0.123 | −0.058 | 0.010 | −0.018 | −0.046 | 0.176 ** | 0.291 ** | 0.281 ** | ||||||||||||||||||
| 12 | −0.134 | −0.021 | −0.171 | −0.087 | 0.008 | 0.056 | −0.013 | 0.284 ** | 0.317 ** | 0.357 ** | 0.581 ** | |||||||||||||||||
| 13 | −0.037 | −0.118 | −0.108 | −0.191 ** | −0.044 | 0.001 | −0.105 | −0.106 | 0.203 ** | 0.124 * | 0.167 ** | 0.199 | ||||||||||||||||
| 14 | 0.167 ** | 0.115 ** | −0.008 | 0.039 | 0.160 ** | 0.027 | −0.014 | 0.211 ** | 0.150 ** | 0.212 ** | −0.025 | −0.067 | −0.005 | |||||||||||||||
| 15 | 0.164 ** | 0.191 ** | 0.097 * | 0.160 ** | 0.143 ** | 0.030 | −0.015 | 0.345 ** | 0.241 ** | 0.265 ** | −0.022 | −0.034 | −0.065 | 0.570 ** | ||||||||||||||
| 16 | 0.102 ** | −0.032 | −0.046 | −0.084* | 0.044 | 0.027 | −0.091 | 0.056 | 0.049 | 0.081 | −0.021 | −0.046 | 0.089 | 0.723 ** | 0.310 ** | |||||||||||||
| 17 | 0.048 | 0.050 | −0.017 | −0.024 | 0.081 | −0.162 ** | 0.066 | 0.165 ** | 0.154 ** | 0.144 ** | 0.106 | 0.057 | 0.108 | 0.043 | 0.054 * | −0.056 * | ||||||||||||
| 18 | 0.114 ** | 0.110 ** | 0.040 | 0.131 ** | 0.084 | 0.145 * | −0.036 | 0.218 * * | 0.143 ** | 0.181 ** | −0.001 | 0.078 | −0.086 | 0.532 ** | 0.389 ** | 0.310 ** | −0.327 ** | |||||||||||
| 19 | 0.054 | 0.121 ** | 0.028 | 0.11 2** | 0.133 * | 0.026 | −0.032 | 0.222 ** | 0.197 ** | 0.143 ** | 0.130 * | 0.162 | −0.038 | 0.357 ** | 0.298 ** | 0.185 ** | −0.164 ** | 0.538 ** | ||||||||||
| 20 | 0.029 | 0.109 | −0.067 | 0.007 | 0.164 ** | −0.181 ** | −0.076 | 0.229 ** | 0.345 ** | 0.310 ** | 0.570 ** | 0.381 ** | 0.059 | 0.237 ** | 0.163 ** | 0.144 ** | 0.099 | 0.107 * | 0.162 ** | |||||||||
| 21 | 0.147 * | 0.207 ** | 0.021 | 0.111 | 0.266 ** | −0.206 ** | −0.085 | 0.242 ** | 0.181 ** | 0.198 ** | 0.057 | 0.017 | −0.082 | 0.402 ** | 0.333 ** | 0.224 ** | 0.044 | 0.239 ** | 0.199 ** | 0.615 ** | ||||||||
| 22 | 0.023 | 0.040 | −0.034 | 0.008 | 0.155 ** | −0.211 ** | −0.096 | 0.005 | 0.006 | −0.038 | 0.081 | 0.000 | 0.038 | 0.174 ** | 0.191 ** | 0.106 * | 0.133 * | −0.048 | 0.019 | 0.467 ** | 0.696 ** | |||||||
| 23 | 0.061 | 0.094 | −0.031 | 0.027 | 0.185 ** | −0.199 ** | −0.101 | 0.070 | 0.064 | 0.031 | 0.074 | 0.036 | 0.019 | 0.262 ** | 0.231 ** | 0.148 ** | 0.093 | 0.059 | 0.085 | 0.541 ** | 0.817 ** | 0.952 ** | ||||||
| 24 | 0.085 * | 0.115 ** | 0.114 * | 0.082 | 0.029 | 0.167 ** | 0.397** | 0.053 | 0.025 | −0.014 | −0.042 | 0.088 | −0.069 | 0.003 | 0.014 | −0.027 | −0.017 | 0.029 | 0.018 | −0.007 | −0.003 | −0.094 * | −0.079 | |||||
| 25 | 0.197 ** | 0.182 ** | 0.059 | 0.154 ** | 0.186 ** | −0.073 | −0.163 ** | 0.254 ** | 0.217 ** | 0.232 ** | 0.011 | −0.020 | −0.066 | 0.316 ** | 0.279 ** | 0.152 ** | −0.009 | 0.258 ** | 0.171 ** | 0.359 ** | 0.675 ** | 0.367 ** | 0.467 ** | −0.160 ** | ||||
| 26 | 0.161 ** | 0.204 ** | −0.012 | 0.115 ** | 0.165 ** | −0.185 * * | −0.086 | 0.182 ** | 0.137 ** | 0.176 ** | 0.064 | −0.016 | −0.127* | 0.248 ** | 0.207 ** | 0.115 ** | 0.082** | 0.122 ** | 0.135 ** | 0.453 ** | 0.684 ** | 0.463 ** | 0.534 ** | −0.108 ** | 0.652 ** | |||
| 27 | 0.178 ** | 0.212 ** | −0.007 | 0.124 ** | 0.184 ** | −0.171 ** | −0.115 | 0.184 ** | 0.119 * | 0.173 ** | −0.010 | 0.006 | −0.189 ** | 0.251 ** | 0.201 ** | 0.121 ** | 0.052 | 0.153 ** | 0.147 ** | 0.394 ** | 0.674 ** | 0.358 ** | 0.463 ** | −0.151 ** | 0.625 ** | 0.718 ** | ||
| 28 | 0.220 ** | 0.286 ** | 0.042 | 0.197 ** | 0.193 ** | −0.149 ** | 0.009 | 0.136 ** | 0.130 ** | 0.183 ** | −0.063 | −0.025 | −0.159 ** | 0.263 ** | 0.189 ** | 0.081 ** | −0.031 | 0.231** | 0.178 ** | 0.285 ** | 0.502 ** | 0.087 * | 0.199 ** | 0.001 | 0.450 ** | 0.518 ** | 0.515 ** | 1 |
| Domain | Variation Site | Hap 1 | Hap 2 | Hap 3 |
|---|---|---|---|---|
| accD-psaI | 1 | A | C | C |
| psbE-petL | 2 | G | A | A |
| petB-petD | 3 | T | C | C |
| 4 | G | G | A |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, X.; Cheng, F.; Zhong, Y. Genetic Diversity of Paeonia rockii (Flare Tree Peony) Germplasm Accessions Revealed by Phenotypic Traits, EST-SSR Markers and Chloroplast DNA Sequences. Forests 2020, 11, 672. https://doi.org/10.3390/f11060672
Guo X, Cheng F, Zhong Y. Genetic Diversity of Paeonia rockii (Flare Tree Peony) Germplasm Accessions Revealed by Phenotypic Traits, EST-SSR Markers and Chloroplast DNA Sequences. Forests. 2020; 11(6):672. https://doi.org/10.3390/f11060672
Chicago/Turabian StyleGuo, Xin, Fangyun Cheng, and Yuan Zhong. 2020. "Genetic Diversity of Paeonia rockii (Flare Tree Peony) Germplasm Accessions Revealed by Phenotypic Traits, EST-SSR Markers and Chloroplast DNA Sequences" Forests 11, no. 6: 672. https://doi.org/10.3390/f11060672
APA StyleGuo, X., Cheng, F., & Zhong, Y. (2020). Genetic Diversity of Paeonia rockii (Flare Tree Peony) Germplasm Accessions Revealed by Phenotypic Traits, EST-SSR Markers and Chloroplast DNA Sequences. Forests, 11(6), 672. https://doi.org/10.3390/f11060672




