Pinus sylvestris Breeding for Resistance against Natural Infection of the Fungus Heterobasidion annosum
Abstract
1. Introduction
2. Materials and Methods
2.1. Studied Trial and Measurements
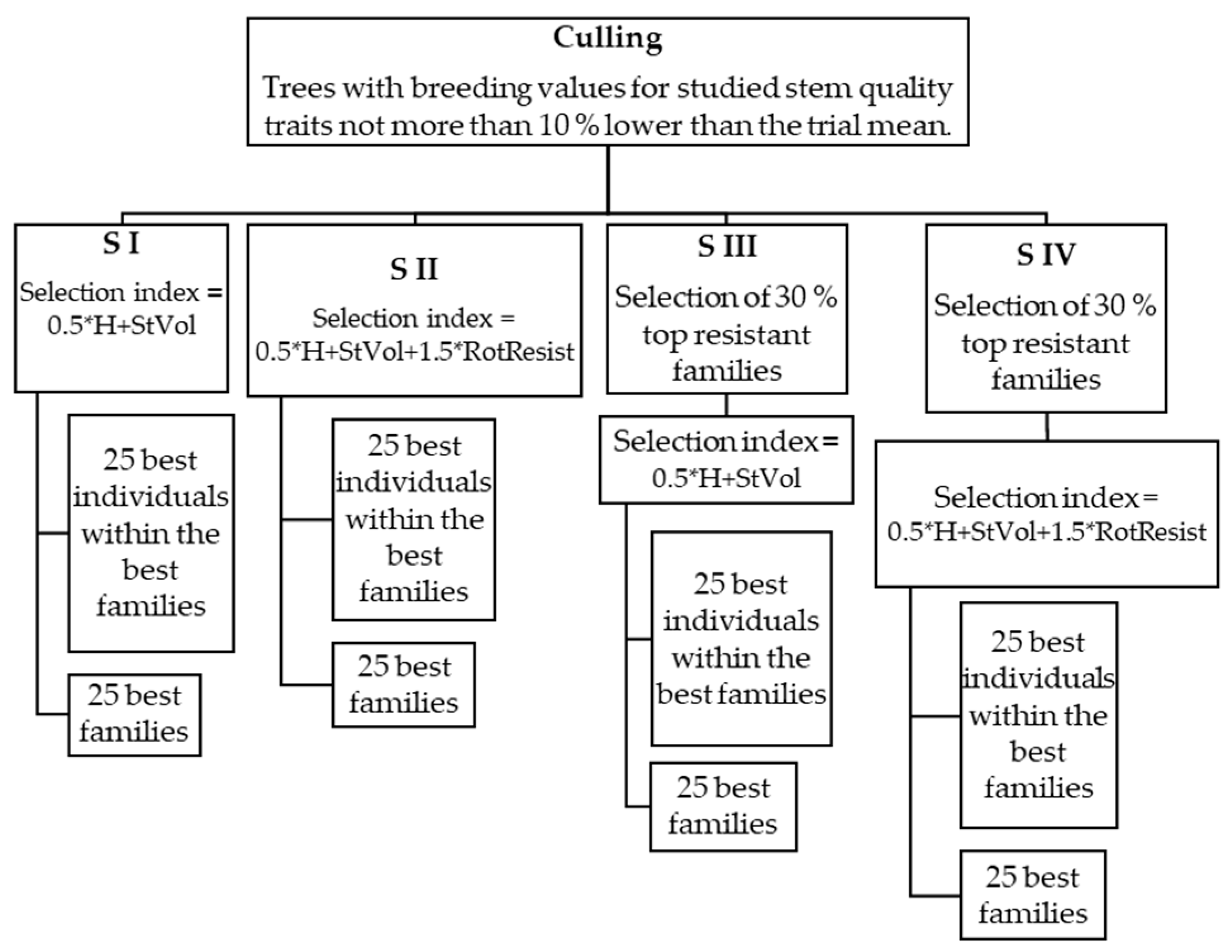
2.2. Data Analysis
3. Results
4. Discussion
4.1. Genetic Parameters
4.2. Selection Methods
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jansson, G.; Hansen, J.K.; Haapanen, M.; Kvaalen, H.; Steffenrem, A. The genetic and economic gains from forest tree breeding programmes in Scandinavia and Finland. Scand. J. For. Res. 2017, 32, 273–286. [Google Scholar] [CrossRef]
- Persson, T. Genotype by environment interaction for survival, growth and Cronartium resistance in northern Scots pine. In Genetic Aspects of Adaptation and Mitigation: Forest Health, Wood Quality and Biomass Production, Proceedings of the Book of Abstracts of International Scientific Conference, Riga, Latvia, 3–5 October 2012; Jansons, Ā., Konstantinova, I., Eds.; LSFRI Silava: Salaspils, Latvia, 2012; p. 150. [Google Scholar]
- Krakau, U.-K.; Liesebach, M.; Aronen, T.; Lelu-Walter, M.-A.; Schneck, V. Scots Pine (Pinus sylvestris L.); Springer: Dordrecht, Germany, 2013; pp. 267–323. [Google Scholar]
- Neimane, U.; Polmanis, K.; Zaluma, A.; Klavina, D.; Gaitnieks, T.; Jansons, A. Damage caused by Lophodermium needle cast in open-pollinated and control-crossed progeny trials of Scots pine (Pinus sylvestris L.). For. Chron. 2018, 94, 155–161. [Google Scholar] [CrossRef]
- Gonthier, P.; Thor, M. Annosus root and butt rots. In Infectious Forest Diseases; Gonthier, P., Nicolotti, G., Eds.; CAB International: Wallingford, UK, 2013; pp. 128–158. [Google Scholar]
- Garbelotto, M.; Gonthier, P. Biology, Epidemiology, and Control of Heterobasidion Species Worldwide. Annu. Rev. Phytopathol. 2013, 51, 39–59. [Google Scholar] [CrossRef] [PubMed]
- Asiegbu, F.O.; Adomas, A.; Stenlid, J. Conifer root and butt rot caused by Heterobasidion annosum (Fr.) Bref. s.l. Mol. Plant Pathol. 2005, 6, 395–409. [Google Scholar] [CrossRef] [PubMed]
- Swedjemark, G.; Stenlid, J.; Karlsson, B. Genetic variation among clones of Picea abies in resistance to growth of Heterobasidion annosum. Silvae Genet. 1998, 46, 369–374. [Google Scholar]
- Chen, Z.-Q.; Lundén, K.; Karlsson, B.; Vos, I.; Olson, Å.; Lundqvist, S.-O.; Stenlid, J.; Wu, H.X.; García Gil, M.R.; Elfstrand, M. Early selection for resistance to Heterobasidion parviporum in Norway spruce is not likely to adversely affect growth and wood quality traits in late-age performance. Eur. J. For. Res. 2018, 137, 517–525. [Google Scholar] [CrossRef]
- Korshikov, I.I.; Demkovich, A.E. Genetic features of root fungus-resistant scotch pine trees in artificial stands in the steppe zone of Ukraine. Cytol. Genet. 2008, 42, 323–328. [Google Scholar] [CrossRef]
- Vasiliauskas, A. Šakninės pinties (Heterobasidion annosum (Fr.) Bref.) paplitimas pušynuose, įveistuose žemės ūkio naudmenuose, kovos su ja priemonės ir kovos rezultatai [Root rot caused by Heterobasidion annosum in Pinus sylvestris plantations on former agricultural land. Miskininkyste 2001, 2, 42–59. [Google Scholar]
- Zaluma, A.; Gailis, A.; Burņeviča, N.; Korhonen, K.; Gaitnieks, T. Susceptibility of Picea abies and Pinus sylvestris seedlings of various origins to Heterobasidion annosum and H. parviporum. Proc. Latv. Acad. Sci. 2016, 70, 29–33. [Google Scholar]
- Mukrimin, M.; Kovalchuk, A.; Ghimire, R.P.; Kivimäenpää, M.; Sun, H.; Holopainen, J.K.; Asiegbu, F.O. Evaluation of potential genetic and chemical markers for Scots pine tolerance against Heterobasidion annosum infection. Planta 2019, 250, 1881–1895. [Google Scholar] [CrossRef]
- Šņepste, I.; Šķipars, V.; Krivmane, B.; Brūna, L.; Ruņģis, D. Characterization of a Pinus sylvestris thaumatin-like protein gene and determination of antimicrobial activity of the in vitro expressed protein. Tree Genet. Genomes 2018, 14, 58. [Google Scholar] [CrossRef]
- Skrøppa, T.; Solheim, H.; Steffenrem, A. Genetic variation, inheritance patterns and parent–offspring relationships after artificial inoculations with Heterobasidion parviporum and Ceratosystis polonica in Norway spruce seed orchards and progeny tests. Silva Fenn. 2015, 49, 1191. [Google Scholar] [CrossRef]
- Marčiulynas, A.; Sirgedaitė-Šėžienė, V.; Žemaitis, P.; Baliuckas, V. The Resistance of Scots Pine (Pinus sylvestris L.) Half-sib Families to Heterobasidion annosum. Forests 2019, 10, 287. [Google Scholar] [CrossRef]
- Zaļuma, A.; Muižnieks, I.; Gaitnieks, T.; Burņeviča, N.; Jansons, Ā.; Jansons, J.; Stenlid, J.; Vasaitis, R. Infection and spread of root rot caused by Heterobasidion spp. in Pinus contorta plantations in Northern Europe: Three case studies. Can. J. For. Res. 2019, 49, 969–977. [Google Scholar] [CrossRef]
- White, T.L.; Adams, W.T.; Neale, D.B. Forest Genetics; CABI Publishing: Wallingford, UK, 2007; ISBN 9780851990835. [Google Scholar]
- Falconer, D.S.; Mackay, T.F. Introduction to Quantitative Genetics, 4th ed.; Longman Group Ltd.: London, UK, 1996. [Google Scholar]
- Hazel, L.N. The Genetic Basis for Constructing Selection Indexes. Genetics 1943, 28, 476–490. [Google Scholar]
- Laiviņš, M.; Melecis, V. Bio-geographical interpretation of climate data in Latvia: Multidimensional analysis. Acta Univ. Latv. 2003, 654, 7–22. [Google Scholar]
- Harris, I.; Jones, P.D.; Osborn, T.J.; Lister, D.H. Updated high-resolution grids of monthly climatic observations—The CRU TS3.10 Dataset. Int. J. Climatol. 2014, 34, 623–642. [Google Scholar] [CrossRef]
- Bušs, K. Meža Ekoloģija un Tipoloģija [Forest ecology and typology]; Zinātne: Rīga, Latvia, 1981. [Google Scholar]
- Liepa, I. Pieauguma Mācība [Increment Science]; LLU: Jelgava, Latvia, 1996. [Google Scholar]
- SAS. SAS/IML 9.3 User’s Guide; Sas Institute: Cary, NC, USA, 2011; ISBN 1607649136. [Google Scholar]
- Oliver, M.A.; Webster, R. Kriging: A method of interpolation for geographical information systems. Int. J. Geogr. Inf. Syst. 1990, 4, 313–332. [Google Scholar] [CrossRef]
- Littell, R.C.; Milliken, G.A.; Stroup, W.W.; Wolfinger, R.D. SAS System for Mixed Models; Sas Institute: Cary, NC, USA, 1996. [Google Scholar]
- Littel, R.C.; Milliken, G.A.; Stroup, W.W.; Wolfinger, R.D.; Schabenberger, O. SAS for Mixed Models, 2nd ed.; Sas Institute: Cary, NC, USA, 2006; Volume 816. [Google Scholar]
- Dickerson, G.E. Techniques for research in quantitative animal genetics. In Techniques and Procedures in Animal Science Research; American Society of Animal Production: Albany, NY, USA, 1969; pp. 36–79. [Google Scholar]
- Baumanis, I.; Jansons, A.; Neimane, U. Priede: Selekcija, Ģenētika un Sēklkopība Latvijā [Scots Pine: Breeding, Genetics and Seed Orchard Management]; Daugavpils Universitātes Akadēmiskais apgāds “Saule”: Daugavpils, Latvia, 2014. [Google Scholar]
- Evans, J.D. Straightforward Statistics for the Behavioral Sciences; Thomson Brooks/Cole Publishing Co.: Belmont, CA, USA, 1996. [Google Scholar]
- Swedjemark, G.; Karlsson, B. Genotypic variation in susceptibility following artificial Heterobasidion annosum inoculation of Picea abies clones in a 17-year-old field test. Scand. J. For. Res. 2004, 19, 103–111. [Google Scholar] [CrossRef]
- Swedjemark, G.; Karlsson, B. Variation in incidence and genetic impact on natural infection of Heterobasidion annosum in Picea abies (L.) Karst. in genetic trials in south Sweden. For. Ecol. Manag. 2004, 203, 135–145. [Google Scholar] [CrossRef]
- Wellendorf, H.; Thomsen, I.M. Genetic variation in resistance against Heterobasidion annosum (Fr.) bref. in Picea abies (L.) karst. expressed after inoculation of neighboring stumps. Silvae Genet. 2008, 57, 312–324. [Google Scholar] [CrossRef]
- Arnerup, J.; Swedjemark, G.; Elfstrand, M.; Karlsson, B.; Stenlid, J. Variation in growth of Heterobasidion parviporum in a full-sib family of Picea abies. Scand. J. For. Res. 2010, 25, 106–110. [Google Scholar] [CrossRef]
- Skrøppa, T.; Solheim, H.; Hietala, A. Variation in phloem resistance of Norway spruce clones and families to Heterobasidion parviporum and Ceratocystis polonica and its relationship to phenology and growth traits. Scand. J. For. Res. 2015, 30, 103–111. [Google Scholar] [CrossRef]
- Steffenrem, A.; Solheim, H.; Skrøppa, T. Genetic parameters for wood quality traits and resistance to the pathogens Heterobasidion parviporum and Endoconidiophora polonica in a Norway spruce breeding population. Eur. J. For. Res. 2016, 135, 815–825. [Google Scholar] [CrossRef]
- Perry, A.; Wachowiak, W.; Brown, A.V.; Ennos, R.A.; Cottrell, J.E.; Cavers, S. Substantial heritable variation for susceptibility to Dothistroma septosporum within populations of native British Scots pine (Pinus sylvestris). Plant Pathol. 2016, 65, 987–996. [Google Scholar] [CrossRef]
- Wang, L.; Zhang, J.; Drobyshev, I.; Cleary, M.; Rönnberg, J. Incidence and impact of root infection by Heterobasidion spp., and the justification for preventative silvicultural measures on Scots pine trees: A case study in southern Sweden. For. Ecol. Manag. 2014, 315, 153–159. [Google Scholar] [CrossRef]
- Rönnberg, J.; Petrylaite, E.; Nilsson, G.; Pratt, J. Two studies to assess the risk to Pinus sylvestris from Heterobasidion spp. in southern Sweden. Scand. J. For. Res. 2006, 21, 405–413. [Google Scholar] [CrossRef]
- Allikmäe, E.; Laarmann, D.; Korjus, H. Vitality assessment of visually healthy trees in Estonia. Forests 2017, 8, 223. [Google Scholar] [CrossRef]
| Trait | Mean ± Standard Deviation | Min | Max | Narrow-Sense Heritability h2 ± standard Error |
|---|---|---|---|---|
| Height (m) | 9.7 ± 2.42 | 1.9 | 16.1 | 0.45 ± 0.094 |
| Diameter at breast height (cm) | 10.3 ± 4.06 | 1.0 | 23.0 | 0.25 ± 0.069 |
| Stem volume (dm3) | 55.6 ± 43.39 | 0.3 | 274.9 | 0.28 ± 0.072 |
| Branch diameter (cm) | 1.5 ± 0.43 | 0.3 | 4.5 | 0.13 ± 0.053 |
| Branchiness (score) | 1.4 | 1.0 | 3.0 | 0.10 ± 0.048 |
| Spike knots (% of trees) | 21.8 | - | - | 0.03 ± 0.077 |
| Stem straightness (score) | 1.2 | 1.0 | 3.0 | 0.02 ± 0.042 |
| Root rot resistance index | 0.78 ± 1.34 | −1.9 | 8.6 | 0.37 ± 0.220 |
| H | DBH | StVol | BrD | Branch | StStr | SpKn | RotResist | |
|---|---|---|---|---|---|---|---|---|
| H | 1 | 0.89 | 0.88 | 0.51 | −0.10 | 0.39 | −0.27 | 0.15 |
| DBH | <0.01 | 1 | 0.98 | 0.65 | −0.34 | 0.31 | −0.25 | 0.16 |
| Vol | <0.01 | <0.01 | 1 | 0.62 | −0.34 | 0.26 | −0.25 | 0.19 |
| BrD | <0.01 | <0.01 | <0.01 | 1 | −0.58 | 0.04 | −0.13 | 0.28 |
| Branch | 0.19 | <0.01 | <0.01 | <0.01 | 1 | 0.15 | 0.02 | −0.15 |
| StStr | <0.01 | <0.01 | <0.01 | 0.59 | 0.06 | 1 | −0.24 | −0.03 |
| SpKn | <0.01 | <0.01 | <0.01 | 0.11 | 0.76 | <0.01 | 1 | 0.07 |
| RotResist | 0.06 | 0.05 | 0.02 | <0.01 | 0.06 | 0.71 | 0.38 | 1 |
| Selection Criterion | Genetic Gain (%) | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Height | Diameter at Breast Height | Branch Diameter | Branchiness | Stem Straightness | Spike Knots | Stem Volume | Root Rot Resistance Index | ||
| S I (0.5 × H + StVol) | 25 best individuals | 16.5 | 13.2 | 3.0 | −0.6 | 0.1 | −1.1 | 44.3 | 2.3 |
| 25 best families | 14.5 | 17.8 | 6.3 | −2.4 | 0.2 | −15.4 | 50.6 | 4.9 | |
| S II (0.5 × H + StVol + 1.5*RotResist) | 25 best individuals | 6.5 | 4.7 | 1.6 | 0.3 | 0.1 | −1.6 | 10.9 | 30.0 |
| 25 best families | 10.4 | 13.5 | 6.0 | −2.4 | 0.0 | −0.5 | 38.1 | 33.7 | |
| S III (Two-stage selection/0.5 × H + StVol) | 25 best individuals | 16.4 | 12.5 | 2.8 | −0.2 | 0.0 | −1.0 | 41.0 | 15.4 |
| 25 best families | 11.5 | 14.3 | 4.8 | −2.4 | 0.1 | −2.3 | 39.7 | 27.0 | |
| S IV (Two-stage selection/0.5 × H + StVol + 1.5 × RotResist) | 25 best individuals | 6.5 | 4.6 | 1.7 | 0.2 | 0.1 | −1.4 | 10.7 | 27.7 |
| 25 best families | 9.5 | 12.8 | 5.5 | −2.3 | 0.0 | 0.6 | 35.7 | 35.2 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rieksts-Riekstiņš, R.; Zeltiņš, P.; Baliuckas, V.; Brūna, L.; Zaļuma, A.; Kāpostiņš, R. Pinus sylvestris Breeding for Resistance against Natural Infection of the Fungus Heterobasidion annosum. Forests 2020, 11, 23. https://doi.org/10.3390/f11010023
Rieksts-Riekstiņš R, Zeltiņš P, Baliuckas V, Brūna L, Zaļuma A, Kāpostiņš R. Pinus sylvestris Breeding for Resistance against Natural Infection of the Fungus Heterobasidion annosum. Forests. 2020; 11(1):23. https://doi.org/10.3390/f11010023
Chicago/Turabian StyleRieksts-Riekstiņš, Raitis, Pauls Zeltiņš, Virgilijus Baliuckas, Lauma Brūna, Astra Zaļuma, and Rolands Kāpostiņš. 2020. "Pinus sylvestris Breeding for Resistance against Natural Infection of the Fungus Heterobasidion annosum" Forests 11, no. 1: 23. https://doi.org/10.3390/f11010023
APA StyleRieksts-Riekstiņš, R., Zeltiņš, P., Baliuckas, V., Brūna, L., Zaļuma, A., & Kāpostiņš, R. (2020). Pinus sylvestris Breeding for Resistance against Natural Infection of the Fungus Heterobasidion annosum. Forests, 11(1), 23. https://doi.org/10.3390/f11010023





