1. Introduction
Medical imaging plays a crucial role in modern healthcare, enabling clinicians to perform preventative screening and monitor diseases with increasing precision. However, radiological departments face significant challenges in managing growing workloads while maintaining high diagnostic quality and consistency [
1]. The interpretation of radiographs remains highly variable between practitioners, creating potential disparities in patient care [
2].
The workflow within radiological departments encompasses multiple steps, from image acquisition to final diagnosis. Many of these steps, particularly those involving data retrieval, organisation, and preliminary analysis, are repetitive and time-consuming [
1]. These steps represent ideal candidates for automation through artificial intelligence (AI) solutions, which could significantly expedite workflows and enhance diagnostic accuracy.
Deep learning (DL), a subfield of AI, has demonstrated remarkable capabilities in image analysis tasks across numerous domains, particularly in the field of medical imaging [
3]. Within DL, transfer learning has emerged as a particularly promising approach for medical image applications. Transfer learning leverages knowledge gained from training on large general image datasets and applies it to more specialised tasks with limited data availability [
4]—a common constraint in medical imaging research.
Previous research addressed medical image modality classification using various convolutional neural network (CNN) architectures. Yu et al. [
5] applied deep transfer learning to modality classification but did not consider anatomical region prediction. Chiang et al. [
6] demonstrated combined modality–anatomy classification but limited the evaluation to three anatomical regions and two imaging modalities. Existing work predominantly treats modality and anatomy classification as independent tasks, evaluates a narrow range of modality–anatomy combinations, and lacks systematic comparison of modern architectures for joint classification.
This study addresses the aforementioned gaps through a two-component pipeline that separately optimises modality and anatomical region prediction across five imaging modalities and 17 anatomical regions. Both tasks are formulated as multi-class single-label classification, where each image receives exactly one modality label and one anatomical region label. The contributions of this study are as follows:
Systematic evaluation of modern CNN and vision transformer architectures across both classification tasks.
Construction of a comprehensive dataset integrating 11 publicly available datasets with standardised preprocessing, spanning five imaging modalities and 17 anatomical regions.
A two-stage pipeline design enabling targeted feature learning for each classification task, achieving 97.21% combined accuracy on unseen test data.
The remainder of this study is organised as follows:
Section 2 contains the background of the proposed models utilised in this study.
Section 3 presents the dataset utilised in this study as well as the specific models utilised.
Section 4 presents the results achieved by the various models on the dataset, with particular emphasis on the best-performing model.
Section 5 discusses the implications of the results and compares findings with the existing literature.
Section 6 concludes with the key findings of this study.
2. Background
This section details the various medical imaging modalities utilised in clinical settings and also provides a background of DL, specifically CNNs and transfer learning.
2.1. Medical Imaging
Medical imaging encompasses a variety of imaging technologies that allow for non-invasive visualisation of internal body structures. These technologies provide critical diagnostic insight across multiple medical modalities [
7]. The primary imaging modalities include the following:
X-ray radiography: X-ray radiography is the most common form of medical imaging due to affordability, speed, and availability [
8]. X-ray captures bone and soft tissue through the use of electromagnetic radiation.
Ultrasound: Ultrasound employs high-frequency sound waves to visualise internal organs and structures in real time.
Computed tomography: Computed tomography (CT) generates cross-sectional images through multiple X-ray projections.
Magnetic resonance imaging: Magnetic resonance imaging (MRI) utilises magnetic fields and radio waves to produce detailed anatomical representations.
Angiography: Angiography is utilised to image the flow of blood through blood vessels and organs of the body. A radiopaque substance is introduced to the blood system; X-ray images are then used to detect the radiopaque substance in the body.
2.2. Deep Learning
DL is a machine learning subdomain that emulates the neural structure of the human brain, comprising many hidden layers of trainable parameters to learn intricate patterns in data. The power of DL lies in its ability to process raw data with minimal manual feature extraction, making it particularly effective in complex domains [
9], including advanced image and speech recognition [
10,
11] for natural language processing [
12,
13] and predictive analytics [
14]. DL has revolutionised artificial intelligence by creating systems capable of learning and adapting with remarkable precision and sophistication.
2.3. Convolutional Neural Networks
CNNs are a specific type of DL architecture that are designed to process grid-like data such as images. CNNs consist of one or more convolutional layers responsible for feature extraction on the input image [
15]. Feature extraction is performed by applying a convolutional filter, which is systematically applied to the image in a sliding window fashion.
After the convolutional filter has been applied to the entire image, a feature map is produced. Pooling layers are then utilised to downscale the image and reduce the dimensionality of the feature maps. Finally, a feedforward network is applied to the final feature map to produce an output. These layers are repeated multiple times to increase the capacity of the network to recognise more complex shapes. The outputs of the feedforward network are passed to a softmax layer. This final layer outputs the predicted classes for the input image.
2.4. Residual Networks
Residual networks (ResNet), introduced by He et al. [
16], revolutionised CNN architecture design by addressing the vanishing gradient problem through residual connections. These skip connections allow information to bypass certain layers, enabling the training of much deeper networks, up to 152 layers in the original implementation. The key innovation of ResNet is the residual block, which learns residual mappings instead of direct mappings, making optimisation easier for deep networks.
2.5. Dense Networks
Dense convolutional networks (DenseNet), proposed by Huang et al. [
17], further expanded on connectivity patterns in CNNs by implementing dense connections, where each layer receives inputs from all preceding layers and passes its feature maps to all subsequent layers. This dense connectivity pattern creates short paths for gradient flow during backpropagation, alleviating the vanishing gradient problem [
18] while enhancing feature reuse. DenseNet requires fewer parameters than traditional architectures with comparable performance due to its efficient feature reuse mechanism.
2.6. Efficient Networks
EfficientNet, developed by Tan and Le [
19], introduced a systematic approach to CNN scaling through compound coefficient scaling that uniformly scales network width, depth, and resolution. Unlike previous architectures that scaled primarily in one dimension, the compound scaling method of EfficientNet improved accuracy while maintaining computational efficiency. The base architecture, EfficientNet-B0, was developed using neural architecture search optimised for both accuracy and efficiency, while subsequent models (B1 to B7) were derived through the compound scaling method.
2.7. Vision Transformers
Vision transformers (ViTs) [
20] adapted the transformer architecture, originally developed for natural language processing, to image classification by treating image patches as token sequences. While standard ViTs require large-scale pre-training data, hierarchical variants such as the Swin Transformer [
21] introduced shifted window-based self-attention to efficiently model local and global dependencies. Swin Transformers compute attention within non-overlapping windows and shift windows between layers to enable cross-window connections, achieving linear computational complexity with respect to image size. This design produces multi-scale feature maps similar to CNNs, making Swin Transformers well-suited for dense prediction tasks and competitive with CNNs on image classification benchmarks.
2.8. Transfer Learning
Transfer learning [
22] represents a pivotal advancement in DL, whereby CNNs are trained on a more general dataset before the models are fine-tuned to a more specific dataset. Transfer learning can significantly improve performance and reduce training time through the transfer of learned features and representations. Transfer learning helps to address the computational and data-intensive challenges of DL [
23].
ResNet, DenseNet, and EfficientNet demonstrate the effectiveness of transfer learning in deep learning applications. These architectures, when pre-trained on large-scale datasets such as ImageNet [
24], serve as powerful feature extractors that are fine-tuned for new tasks. Transfer learning leverages these pre-trained representations to accelerate training and improve model performance, particularly in domains with limited labelled data. This approach has proven especially valuable in medical imaging, where acquiring large annotated datasets is often challenging and expensive [
25].
3. Materials and Methods
This section details the data collection process and the methodology of this study. In particular, the datasets utilised in this study, the CNN architectures utilised, and the experimental framework employed for model training and evaluation are discussed.
3.1. Dataset Creation
The specific datasets utilised in this study and the specific preprocessing steps required to create a combined dataset with uniform images are discussed below.
3.1.1. National Institute of Health Chest X-Ray Dataset
The National Institute of Health chest X-ray (NIH-CXR) dataset is the largest publicly available collection of frontal chest X-rays from the NIH Clinical Centre’s picture archiving and communication system (PACS). The dataset features 14 thoracic pathology labels with over 90% accuracy via weak-supervised learning methods [
26]. For this study, 20,000 images were randomly sampled from the original 112,120 to avoid a skewed class representation. The original images were labelled as ‘xray’ (class 1) and ‘chest’ (class 2).
3.1.2. Curated Breast Imaging Subset of the Digital Database for Screening Mammography Dataset
The curated breast imaging subset of the digital database for screening mammography (CBIS-DDSM) dataset [
27] is a curated, standardised subset of the DDSM mammography database. It contains normal, benign, and malignant examples with verified pathology information, selected by a trained mammographer for improved segmentation accuracy. From the original 10,237 images (3520 cropped, 3567 masked, 3150 normal), the dataset was filtered to remove mask images (i.e., more than 200 black pixels) and retain normal images (i.e., more than 250,000 pixels in the 0 to 20 intensity range), with visual inspection confirmation. Patient IDs were used for train–test splitting to maintain patient separation across sets, with random shuffling before division. Images received random identifiers to prevent naming conflicts and were labelled as ‘xray’ (class 1) and ‘breast’ (class 2) in the constructed dataset.
3.1.3. Musculoskeletal Radiographs Dataset
The musculoskeletal radiographs (MURA-v1.1) [
28] dataset is a large collection of bone X-rays from the Stanford machine learning group, featuring seven bone categories, i.e., elbow, finger, forearm, hand, humerus, shoulder, and wrist. Board-certified Stanford Hospital radiologists manually labelled images as normal (uninjured bones) or abnormal (injuries or metal plates). Created for a Stanford competition on normal/abnormal image classification, the dataset was reorganised from its original training/validation split to match the desired ratio and include a test set. The images were labelled with class 1 ‘xray’ and class 2 specifying the bone type (i.e., ‘elbow’, ‘finger’, ‘forearm’, ‘hand’, ‘humerus’, ‘shoulder’, or ‘wrist’) in the constructed dataset.
3.1.4. COVID-19 Computed Tomography Dataset
The COVID-19 computed tomography (COVID-CT) dataset contains chest CT scans in three categories, i.e., COVID-19-infected patients, normal scans, and scans labelled as ‘other’. This dataset was used for an explainable COVID-19 detection model [
29]. The images in this dataset are organised by patient ID, with each patient having one or more CT slices. Patient IDs are assumed to be distinct across categories. The complete dataset of 4171 images was included in the constructed dataset. The images were labelled with class 1 ‘ct’ and class 2 ‘chest’ in the constructed dataset.
3.1.5. Moscow Medical Brain Computed Tomography Dataset
The Moscow medical brain computed tomography (MosMedData) dataset [
30] contains brain CT scans with and without intracranial haemorrhage. The dataset was collected from Moscow radiological departments between 2020 and 2023; it originally included 800 patients with 887 CT scans (patients with multiple scans were present).
Studies with quality issues were removed; in total, 65 patients were removed. Further processing issues during conversion to PNG eliminated 17 more patients, resulting in 718 patients. Random identifiers were assigned to track slices grouped per study. Due to computational constraints, only 10% of the images were utilised. The images were labelled with class 1 ‘ct’ and class 2 ‘brain’ in the constructed dataset.
3.1.6. Kidney Computed Tomography Dataset
The kidney computed tomography (Kidney-CT) dataset [
31] was collected from picture archiving and communication systems (PACSs) across multiple hospitals in Dhaka, Bangladesh. It contains four categories, i.e., normal kidneys and three pathological conditions, namely cyst, tumour, and stone. The dataset includes both coronal and axial abdominal cuts, as well as urograms. All patient data was anonymised and validated by a radiologist and medical technologist. Since unique patient identifiers were not provided, all images were assumed to belong to different patients. The images were labelled with class 1 ‘ct’ and class 2 ‘kidney’ in the constructed dataset.
3.1.7. Fetal Ultrasound Dataset
The fetal ultrasound (Fetal-US) dataset [
32] contains routine maternal–fetal ultrasound screening images from two hospitals across multiple machines, manually labelled by expert fetal clinicians. The dataset classifies images into six categories, i.e., four common fetal anatomical planes (abdomen, brain, femur, thorax), maternal cervix, and a general category for less common planes. Data is organised by patient ID, ultrasound machine, and operator.
From the original 12,400 images, images tagged as ‘Other’ or ‘Maternal cervix’ were filtered out—the latter because the images typically do not include fetal visualisation, and the former due to ambiguity. This filtering removed 5839 images, leaving 6561 images. The images were labelled with class 1 ‘ultrasound’ and class 2 ‘fetus’ in the constructed dataset.
3.1.8. Ultrasound-Neck-Nerve Dataset
This ultrasound-neck-nerve dataset [
33] originated from a Kaggle competition focused on pain management after surgery. Pain management catheters are thin tubes used to deliver medication, reducing narcotic dependence. Effective catheter placement requires accurate identification of nerve structures in the neck (brachial plexus).
The original dataset was divided into 5636 training instances and 5508 test instances. While the training set contained patient identifiers, the test set did not. The images were labelled with class 1 ‘ultrasound’ and class 2 ‘neck’ in the constructed dataset.
3.1.9. COVID-19 Bedside Lung Ultrasound in Emergency Dataset
The COVID-19 Bedside Lung Ultrasound in Emergency (COVID-BLUES) [
34] dataset contains lung ultrasound videos using the BLUE (Bedside Lung Ultrasound in Emergency) protocol for rapid respiratory diagnosis. The dataset includes 362 MP4 videos from 63 patients (33 COVID-19-positive, 30 COVID-19-negative), with six distinct anatomical points captured per patient using a Philips Lumify ultrasound device. Each video was tagged with patient identifiers and specific BLUE point information.
For use in image classification, the videos were deconstructed into individual frames. The images were labelled with class 1 ‘ultrasound’ and class 2 ‘lungs’ in the constructed dataset.
3.1.10. Angiographic Stenosis Data Set
This dataset [
35] contains coronary angiographs showing heart blood vessels, specifically focusing on coronary stenosis. These images were captured using a General Electric Healthcare Innova image–surgery system. All patients had confirmed coronary artery disease, either through angiography or functional evaluation. Angiography represents a unique imaging modality in the constructed dataset, with data limitations resulting in only one angiography dataset being included.
Without patient ID data available, the dataset could not be split based on individual patients. The original dataset was pre-divided into 90% and 10% portions. For the constructed dataset, the original 10% was maintained as the test set, while the remaining 90% was further split into training and validation subsets. The images were labelled with class 1 ‘angiograph’ and class 2 ‘coronaryartery’ in the constructed dataset.
3.1.11. FastMRI
The fastMRI dataset comprises three main categories (i.e., knee, brain, and prostate MRI), each with training, validation, and test subsets, and further differentiated between single-coil and multi-coil MRIs. (Data used in the preparation of this article were obtained from the New York University (NYU) fastMRI Initiative database (
https://fastmri.med.nyu.edu/), accessed on 14 February 2025. As such, NYU fastMRI investigators provided data but did not participate in analysis or writing of this report. A listing of NYU fastMRI investigators, subject to updates, can be found at
https://fastmri.med.nyu.edu/. The primary goal of fastMRI is to test whether machine learning can aid in the reconstruction of medical images) [
36,
37]. Due to the considerable size of the dataset, which was over a terabyte large, only a subset was used for the experiments, i.e., knee single-coil and three batches of brain multi-coil training data.
The original dataset required additional processing from the hierarchical data format (H5), where each file contained MRI data with multiple slices that were extracted using Python 3.12.3, converted to tensors, processed via inverse Fourier transform to obtain complex images, converted to real images by calculating absolute pixel values, and finally normalised to create the final image arrays.
For single-coil knee MRIs, no further manipulation was needed. For multi-coil brain MRIs, an additional root-sum-of-squares (RSS) transform was applied to combine multiple coil regions into single images. All slices from the same MRI received a random globally unique identifier (GUID) prefix for identification. The images were labelled with class 1 ‘mri’ and class 2 specifying either ‘knee’ or ‘brain’.
3.1.12. Prostate National Cancer Institute Dataset
The Prostate National Cancer Institute (Prostate-NCI) dataset (
https://www.kaggle.com/datasets/sshikamaru/world-wide-covid-dataset/data), accessed on 14 February 2025, contains prostate MRI images from 26 subjects with biopsy-confirmed cancer. The images were acquired using a Philips Achieva scanner with an endorectal phased array surface coil. Data collection occurred at the National Cancer Institute in Bethesda, Maryland, between 2008 and 2010. The dataset was split based on patient ID. The images were labelled with class 1 ‘mri’ and class 2 ‘prostate’ in the constructed dataset.
3.1.13. Transformations
Various transformations were applied to the previously mentioned datasets to create a uniform constructed dataset. The following standard transformations were applied to all images in the constructed dataset to ensure consistency:
Resizing: Images of identical width and height were scaled down to 256 × 256 pixels.
Cropping: Images larger than the desired 256 × 256 image size were resized along the shortest dimension to 256 pixels. A centre crop of 256 × 256 pixels was then applied.
Padding: Images that were smaller than the target size of 256 × 256 were padded with black pixels on each side of the image that was undersized.
GUID: Several datasets stored images in different directories with the same filename. The constructed dataset combined these images into a single folder, which required the use of unique identifiers to avoid conflict.
Bit depth: All images in the constructed dataset were converted to 8-bit grayscale images. Images that were from data sources that had multiple channels were converted to a single channel.
Normalisation: All images were normalised to have pixel values in the range of 0–255.
3.2. Dataset Split and Class Distributions
To prevent data leakage, train/validation/test splits were performed at the patient or study level wherever patient identifiers were available, ensuring that all images from a given patient were assigned to the same split. This applied to the majority of the source datasets (fastMRI, Prostate-NCI, CBIS-DDSM, MosMedData, Fetal-US, COVID-BLUES, NIH-CXR, MURA, and COVID-CT). For the Kidney-CT dataset, unique patient identifiers were not provided; however, each image represents a distinct acquisition and was therefore treated as an independent sample. The ultrasound-neck-nerve dataset retained its original training partition (which was split by patient), and the remaining test portion was randomly divided into validation and test subsets. The angiographic stenosis dataset was pre-split by the source into 90% and 10% portions, which were used as the basis for the train/validation and test sets, respectively. All splits followed an approximate 75%/15%/10% train/validation/test ratio.
Figure 1 illustrates the modality class distribution of the constructed dataset, and
Figure 2 illustrates the anatomy class distribution.
Table 1 and
Table 2 present the exact number of images per class in each split.
The dataset was scaled down to manage computational constraints while preserving the clinical distribution patterns. Different scaling factors were applied to various medical image types, resulting in a representative subset of the original data. These scaling decisions were made strategically to maintain proportional representation similar to what would be encountered in clinical settings, ensuring that the resulting dataset remains practical for computational processing while preserving the essential characteristics of medical imaging distribution.
Table 3 presents the scaling factors utilised.
3.3. Proposed Pipeline
The proposed pipeline consists of two stages. The first stage classifies the modality of the input image, and the second stage classifies the anatomical region of the image. Each stage of the pipeline is assessed independently to ensure that the modality and anatomical region prediction are accurate and reliable. This two-stage approach allows for specialised processing based on imaging characteristics, because different modalities exhibit distinct visual features that require specific analysis techniques.
Table 4 provides an overview of the selected architectures, including model complexity and training efficiency measured on the same hardware. In addition to five CNN architectures, the Swin Transformer (Swin-T) [
21] was included to evaluate whether transformer-based architectures offer advantages over CNNs for this task. As shown in
Table 4, the Swin-T is the slowest architecture at 190.0 s per epoch (151.8 ms per batch), compared to 166.1 s (132.7 ms) for EfficientNet-B4 and 41.5 s (33.2 ms) for ResNet-18. Combined with the lower classification accuracy reported in
Section 4, these results indicate that CNNs remain the more practical choice for this classification task.
EfficientNet-B4 was employed for both the modality classification and anatomical region classification components because it achieved the best performance across all evaluation metrics (detailed results are presented in
Section 4).
Figure 3 illustrates the proposed pipeline.
3.4. Evaluation Metrics
Multiple evaluation metrics were employed to ensure that the proposed pipelines were effectively evaluated in this study. Each metric provides unique insight into a particular aspect of the performance of the model.
In the following equations, , , , and represent true positives, false positives, false negatives, and true negatives, respectively, for each class. The specific accuracy metrics employed in this study are described as follows:
Accuracy measures the proportion of correctly classified instances against the total number of instances. Accuracy is defined as
Precision quantifies the proportion of correctly identified positive instances against all instances predicted as positive. Precision is defined as
Recall measures the proportion of actual positive instances that were correctly identified. Recall is defined as
The
F1-score represents the harmonic mean of precision and recall, which is defined as
The macro-average of each metric was utilised as opposed to the micro-average in this study. A macro-average approach was chosen because the constructed dataset contains an unequal distribution of classes, which the macro-average takes into consideration by computing the average of each metric across each class. This is opposed to the micro-average, which computes the metric as a whole but does not account for classes of different sizes.
3.5. Model Architecture
Each model consists of a backbone network pre-trained on ImageNet [
24], with the original classification head replaced by a single linear layer mapping the backbone’s output features to the number of target classes. All layers of the backbone were fully fine-tuned during training (i.e., no layers were frozen). This approach allows the pre-trained feature representations to adapt to the medical imaging domain while benefiting from the learned low-level features.
3.6. Training Procedure and Hyperparameters
All architectures were trained with identical hyperparameters for fair comparison (refer to
Table 5). Images were converted to three-channel grayscale and resized to 224 × 224 pixels (380 × 380 for EfficientNet-B4) and then normalised using the standard mean and deviation values from the ImageNet dataset [
24]. Online data augmentation was applied during training to increase generalisability, which included horizontal flipping (50% probability) and random rotations of 15 degrees. Three different initial learning rates (i.e., 0.0001, 0.001, 0.01) were evaluated to determine optimal convergence for each architecture, spanning typical ranges used in CNN training. Each configuration was trained for three independent runs, and the model with the highest validation accuracy was selected. No learning rate scheduler or early stopping was employed. Training was performed on two NVIDIA RTX 3090 GPUs using PyTorch 2.8.0. Each model configuration required approximately 25 epochs, with per-epoch training times ranging from 41.5 s (ResNet-18) to 190.0 s (Swin-T), as detailed in
Table 4.
4. Results
This section presents the results achieved of the first and second components of the proposed pipeline, as well as the performance of the overall pipeline.
4.1. First Component Analysis
The performance of the first component in the proposed pipeline focused on medical imagery modality classification across the five modalities investigated. Each proposed model was trained for three independent trials, using different learning rates, namely 0.0001, 0.001, and 0.01.
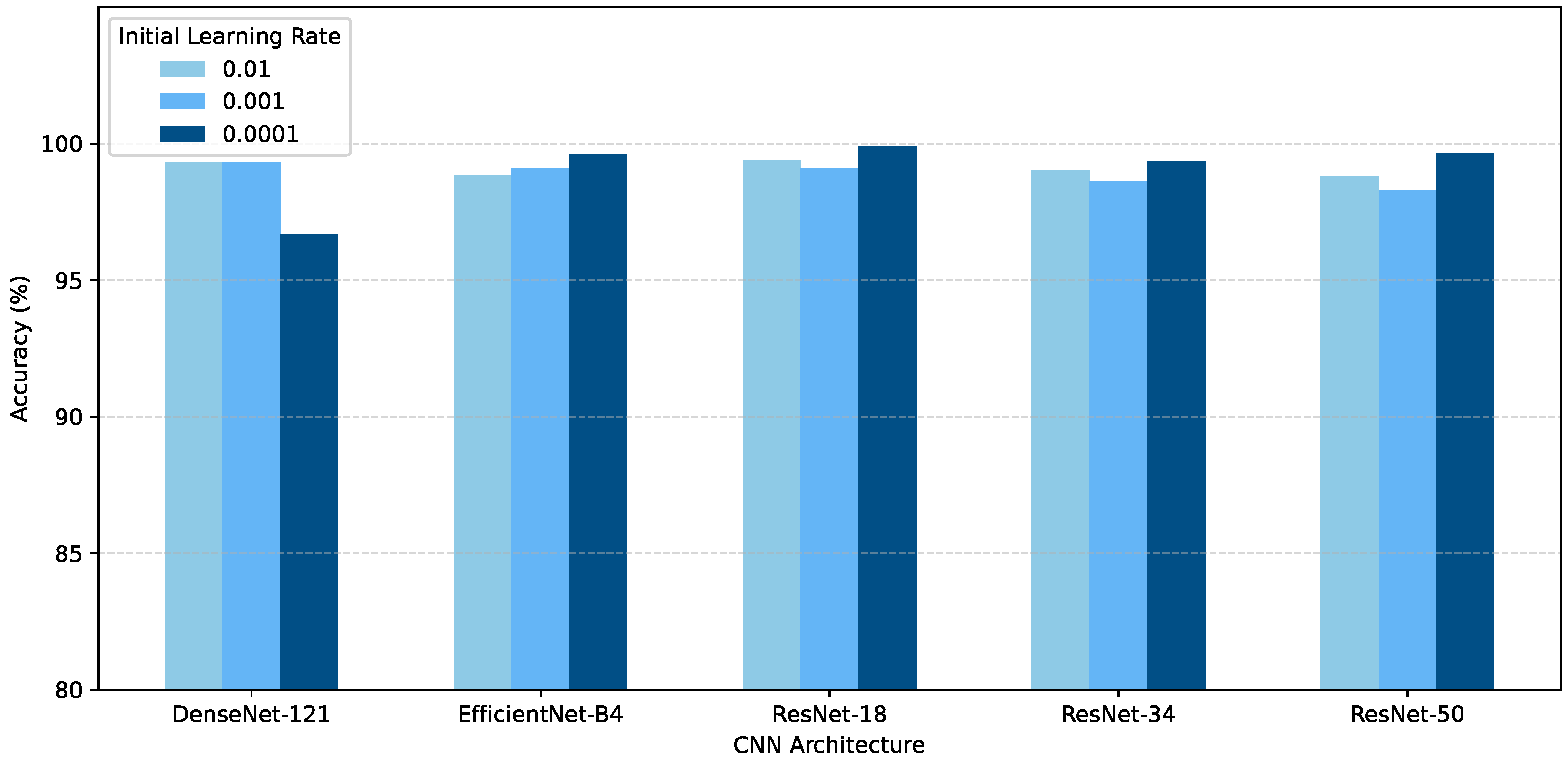
Figure 4 presents the performance of the best models of the three independent trials of the various architectures for component one on the test set. EfficientNet-B4 trained with a learning rate of 0.0001 performed the best, with an accuracy of 99.60%. The Swin-T achieved a best accuracy of 97.82%, which is competitive but below the best CNN results. DenseNet-121 performed the worst, with an accuracy of 96.69%.
The model configuration that achieved the highest validation accuracy across the three tested learning rates was selected as the final model for each CNN architecture.
Figure 5 illustrates the loss and accuracy curves on the training and validation sets, respectively. The accuracy and loss curves show consistent decreases in loss and increases in accuracy on both the training and validation sets. Notably, the final model achieved 100% validation accuracy after only three epochs of training.
Figure 6 presents the confusion matrix for the modality prediction component, which illustrates the strong discriminative capability of the EfficientNet-B4 model across the five modalities. The matrix reveals nearly perfect classification performance, with all modalities showing high true positive rates along the diagonal. Notably, angiography and CT images achieved 100% correct classification. The only observed misclassifications were that 20 MRI images were incorrectly classified as ultrasound and one image as X-ray.
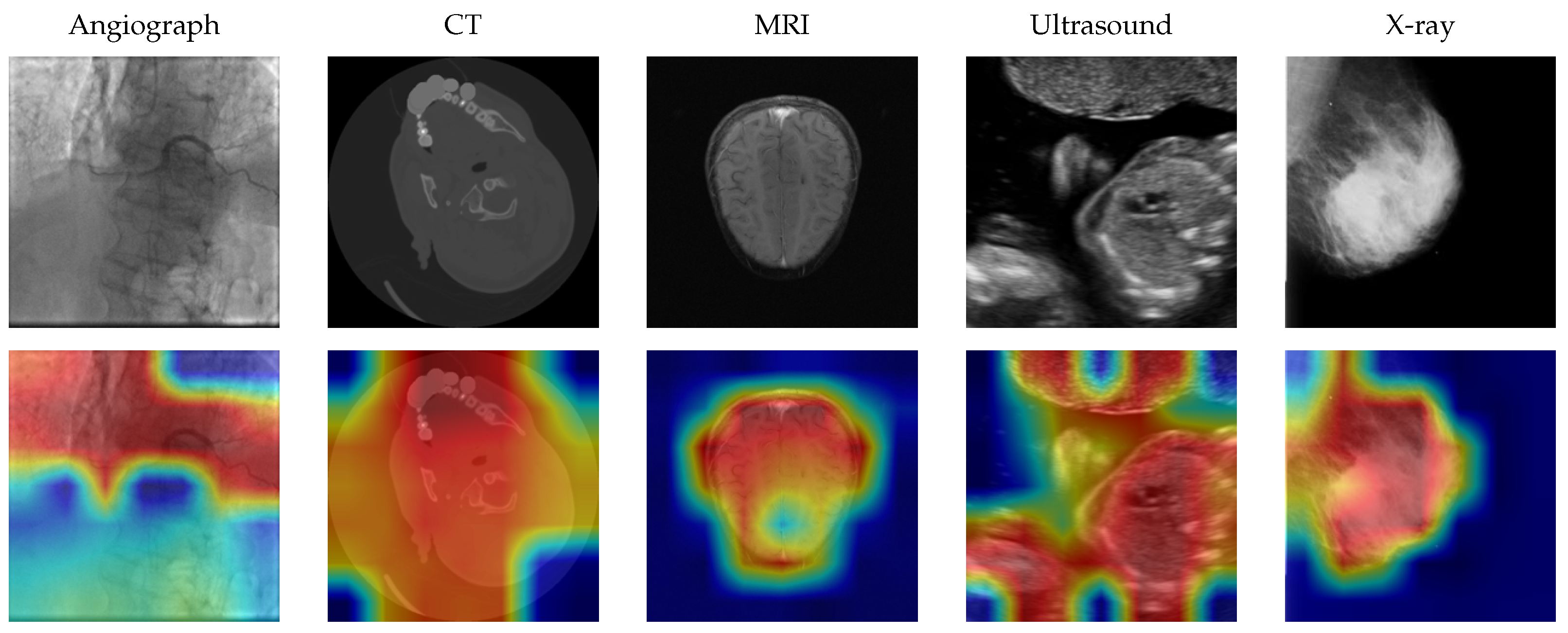
To verify that the model attends to clinically relevant image regions, Grad-CAM [
39] visualisations were generated for representative samples from each modality class.
Figure 7 illustrates the resulting heatmaps overlaid on the original images. The activations consistently highlight anatomically relevant structures—for example, cardiac vessels in angiographs, brain tissue in CT scans, and bone structures in X-rays—rather than image borders or acquisition artifacts.
Table 6 presents the per-class precision, recall, and F1-score for the best modality classifier, together with macro-averaged and weighted-averaged summaries. The weighted average accounts for class size, whereas the macro-average weights all classes equally. All five modalities achieve near-perfect classification, with angiograph and CT achieving perfect scores. The only notable confusion involves MRI, where 20 images were misclassified as ultrasound and one as X-ray.
4.2. Second Component Analysis
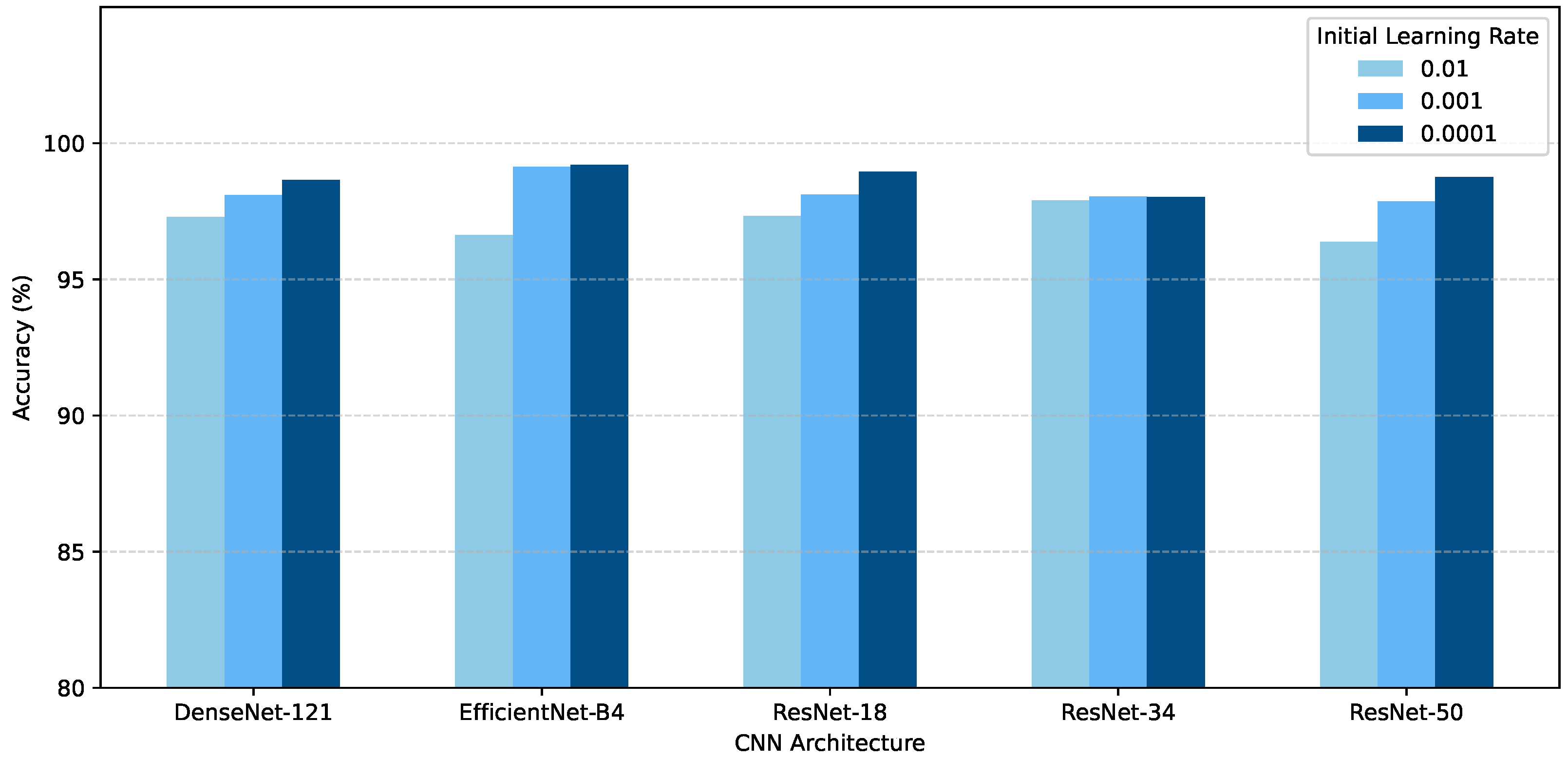
Figure 8 compares the performance of the various architectures for anatomical region prediction. Each architecture was again trained for three independent trials on a range of learning rates, namely 0.0001, 0.001, and 0.01. EfficientNet-B4 performed the best when trained with a learning rate of 0.0001, which achieved an accuracy of 99.21% on the test set. The Swin-T achieved a best accuracy of 98.88%, again competitive but slightly below that of the best CNN. ResNet-50 performed the worst out of all of the models when trained with a learning rate of 0.01, with an accuracy of 97.32%.
Figure 9 showcases the accuracy and loss curves of the EfficientNet-B4 model during training. The training loss rapidly decreases in the first few epochs before gradually stabilising at near-zero, while the validation loss stabilises around 0.05. The accuracy curves display similar behaviour, with training accuracy quickly rising to nearly 100% and consistently remaining above validation accuracy, which stabilises around 99.2%.
Figure 10 illustrates the confusion matrix obtained by the best model in component two, i.e., EfficientNet-B4. The confusion matrix shows that the model achieves high classification accuracy across all anatomical regions, with particularly strong performance in the kidney, brain, and coronary-artery regions. The most notable confusion patterns occur among upper limb categories, where anatomical proximity results in some misclassifications between elbow, forearm, hand, and wrist images.
Grad-CAM visualisations were also generated for the anatomical region classifier.
Figure 11 shows representative examples, confirming that the model focuses on the relevant anatomical structures for each class.
Table 7 presents the per-class metrics for the anatomical region classifier. Non-X-ray anatomical regions (prostate, coronary artery, neck, kidney, fetus, breast, lungs) all achieve perfect or near-perfect F1-scores. The primary source of confusion is among upper limb X-ray categories, where anatomical similarity between adjacent body parts (e.g., forearm and wrist, elbow and humerus) leads to occasional misclassifications.
4.3. Proposed Pipeline Results
The best-performing models of component one and component two were combined to construct the proposed pipeline. EfficientNet-B4 achieved the best results with regard to both modality detection and anatomy detection. The combined prediction metrics were calculated in such a way that only when both predictions were correct was the pipeline considered to provide a correct prediction. If one or both of the models had an incorrect prediction, the data instance was labelled to have been incorrectly predicted.
Table 8 presents the metrics of the combined pipeline on the test set. The overall performance demonstrates excellent results across all evaluation metrics, with an accuracy of 97.21%, precision of 96.19%, recall of 94.57%, and F1-score of 94.88%. These consistently high metrics indicate robust performance across the diverse range of modality–anatomy combinations, with the slight variation between precision and recall suggesting balanced classification performance.
5. Discussion
This study presented a comprehensive approach to medical image classification using transfer learning techniques. The proposed two-component pipeline demonstrated high performance in classifying both imaging modality and anatomical region, achieving an overall accuracy of 97.21% on unseen test data. This performance supports the viability of automated classification systems in clinical radiological workflows.
The accuracy and loss curves indicate that the selected model demonstrates good generalisation ability, with excellent model convergence and minimal overfitting. The model generalises well to unseen data while maintaining high performance across all anatomical regions. The EfficientNet-B4 architecture consistently outperformed other models in both pipeline components, suggesting its effectiveness for medical imaging tasks.
The analysis of confusion patterns revealed that classification errors primarily occurred among anatomically similar structures, particularly in upper limb X-rays, indicating areas for future improvement. These misclassifications likely reflect genuine anatomical similarity that challenges even experienced radiologists.
The strong results demonstrate the effectiveness of the two-component pipeline approach for automated medical image classification. This approach offers the potential for streamlined radiological workflows whilst maintaining diagnostic quality.
The inclusion of the Swin-T (97.82% modality, 98.88% anatomical) demonstrates that performance on this task is not architecture-dependent. The Swin-T achieves competitive results despite having more parameters (27.5 M) than EfficientNet-B4 (17.6 M), which confirms that the high classification accuracy reflects the visual distinctiveness of the modality and anatomy classes rather than overfitting to a particular architecture.
Regarding image resolution, all models were trained at 224 × 224 pixels (380 × 380 for EfficientNet-B4). Modality and anatomy classification relies on global structural features (e.g., tissue contrast, imaging geometry, and overall organ shape) rather than fine-grained details such as lesion boundaries. The consistently high accuracy across all architectures at these resolutions supports the conclusion that higher resolutions are unlikely to yield meaningful improvements for this classification task.
To assess whether the models rely on clinically relevant features rather than dataset-specific artifacts, Grad-CAM [
39] visualisations were generated for both classification components (see
Section 4). The resulting heatmaps confirm that the models attend to anatomically relevant regions of each image rather than background borders or acquisition artifacts, providing evidence that the learned representations are clinically meaningful.
Limitations
The constructed dataset exhibits inherent heterogeneity, as the 11 source datasets originate from different institutions, scanners, and acquisition protocols. While heterogeneity introduces variation in image appearance, the variation also strengthens the evaluation by exposing models to realistic clinical diversity. Labelling noise in the source datasets cannot be entirely excluded, particularly for datasets where labels were derived through weak-supervision methods, such as the NIH-CXR dataset [
26]. The consistently high classification performance across all classes suggests that labelling noise did not materially affect the results. Class imbalance is present in the constructed dataset, with anatomical region class sizes ranging from 1061 (forearm) to 6866 (knee), as detailed in
Table 2. The use of macro-averaged metrics addresses class imbalance by weighting each class equally regardless of sample size.
Angiography and fetal ultrasound are each represented by a single source dataset. A leave-one-dataset-out evaluation is therefore not feasible without removing entire classes. Dataset-specific visual signatures, such as acquisition device characteristics and image borders, could contribute to classification performance alongside genuine modality and anatomical features. The Grad-CAM analysis partially mitigates the concern by confirming that models attend to anatomically relevant regions rather than artifacts. The dataset was further limited to five imaging modalities and 17 anatomical regions; generalisation to a broader range of clinical imaging scenarios remains to be validated.
The performance differences between architectures are substantive. EfficientNet-B4 achieved 99.60% modality accuracy compared to 96.69% for DenseNet-121, a difference corresponding to hundreds of additional correctly classified images. EfficientNet-B4 consistently achieved the highest performance across both classification tasks and all evaluation metrics, reinforcing the reliability of the observed architecture ranking.
Direct comparison with prior work is constrained by the absence of studies operating at a comparable scale. Yu et al. [
5] addressed modality classification alone, and Chiang et al. [
6] evaluated two modalities and three anatomical regions. The present study spans five modalities and 17 anatomical regions using a different dataset composition. The contribution of this work lies in demonstrating that joint modality–anatomy classification is tractable at a significantly larger scale than previously attempted.
Future research directions include expansion to additional medical modalities and finer anatomical region granularity, multi-task learning for simultaneous modality, anatomy, and pathology prediction, attention mechanisms for improved discrimination between anatomically similar body regions, and clinical workflow integration with explainability features.
6. Conclusions
This study demonstrated that transfer learning techniques applied to convolutional neural networks (CNNs) and vision transformers (ViTs) effectively automate medical image classification across multiple modalities and anatomical regions. The two-component pipeline approach enabled specialised feature learning for each classification task, with the efficient network B4 (EfficientNet-B4) architecture achieving superior performance in both modality prediction (99.60% accuracy) and anatomical region prediction (99.21%). The combined pipeline achieved an accuracy of 97.21% on unseen test data, indicating strong generalisability for practical implementation in radiological departments.
Author Contributions
Conceptualisation, J.d.S. and A.E.; methodology, J.d.S. and A.E.; software, J.d.S.; validation, J.d.S.; formal analysis, J.d.S.; investigation, J.d.S. and A.E.; resources, J.d.S.; data curation, J.d.S.; writing—original draft preparation, J.d.S.; writing—review and editing, A.E. and K.A.; visualisation, J.d.S.; supervision, A.E.; project administration, J.d.S., A.E. and K.A. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Informed Consent Statement
Not applicable.
Data Availability Statement
The datasets generated during and/or analysed during the current study are available from the corresponding author upon reasonable request.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| AI | Artificial Intelligence |
| ANN | Artificial Neural Network |
| BLUE | Bedside Lung Ultrasound in Emergency |
| CBIS-DDSM | Curated Breast Imaging Subset of the Digital Database for Screening Mammography |
| CNN | Convolutional Neural Network |
| COVID-BLUES | COVID-19 Bedside Lung Ultrasound in Emergency |
| COVID-CT | COVID-19 Computed Tomography |
| CT | Computed Tomography |
| DenseNet | Dense Convolutional Network |
| DL | Deep Learning |
| EfficientNet | Efficient Network |
| Fetal-US | Fetal Ultrasound |
| GUID | Globally Unique Identifier |
| Kidney-CT | Kidney Computed Tomography |
| MosMedData | Moscow Medical Brain Computed Tomography |
| MRI | Magnetic Resonance Imaging |
| MURA | Musculoskeletal Radiographs |
| NIH-CXR | National Institute of Health Chest X-ray |
| PACS | Picture Archiving and Communication System |
| Prostate-NCI | Prostate National Cancer Institute |
| ResNet | Residual Network |
| RSS | Root-Sum-of-Squares |
| Swin-T | Swin Transformer-Tiny |
| ViT | Vision Transformer |
References
- Wendler, T.; Loef, C. Workflow management-integration technology for efficient radiology. Medicamundi 2001, 45, 41–49. [Google Scholar]
- Ganesan, K.; Acharya, U.R.; Chua, C.K.; Min, L.C.; Abraham, K.T.; Ng, K.H. Computer-aided breast cancer detection using mammograms: A review. IEEE Rev. Biomed. Eng. 2012, 6, 77–98. [Google Scholar] [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Pan, S.J.; Yang, Q. A Survey on Transfer Learning. IEEE Trans. Knowl. Data Eng. 2010, 22, 1345–1359. [Google Scholar] [CrossRef]
- Yu, Y.; Lin, H.; Meng, J.; Wei, X.; Guo, H.; Zhao, Z. Deep transfer learning for modality classification of medical images. Information 2017, 8, 91. [Google Scholar] [CrossRef]
- Chiang, C.H.; Weng, C.L.; Chiu, H.W. Automatic classification of medical image modality and anatomical location using convolutional neural network. PLoS ONE 2021, 16, e0253205. [Google Scholar]
- Hussain, S.; Mubeen, I.; Ullah, N.; Shah, S.S.U.D.; Khan, B.A.; Zahoor, M.; Ullah, R.; Khan, F.A.; Sultan, M.A. Modern diagnostic imaging technique applications and risk factors in the medical field: A review. BioMed Res. Int. 2022, 2022, 5164970. [Google Scholar] [CrossRef]
- Olczak, J.; Emilson, F.; Razavian, A.; Antonsson, T.; Stark, A.; Gordon, M. Ankle fracture classification using deep learning: Automating detailed AO Foundation/Orthopedic Trauma Association (AO/OTA) 2018 malleolar fracture identification reaches a high degree of correct classification. Acta Orthop. 2020, 92, 102–108. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef]
- Mehrish, A.; Majumder, N.; Bharadwaj, R.; Mihalcea, R.; Poria, S. A review of deep learning techniques for speech processing. Inf. Fusion 2023, 99, 101869. [Google Scholar] [CrossRef]
- Li, Y. Research and application of deep learning in image recognition. In Proceedings of the 2022 IEEE 2nd International Conference on Power, Electronics and Computer Applications; IEEE: Piscataway, NJ, USA, 2022; pp. 994–999. [Google Scholar]
- Lauriola, I.; Lavelli, A.; Aiolli, F. An introduction to deep learning in natural language processing: Models, techniques, and tools. Neurocomputing 2022, 470, 443–456. [Google Scholar] [CrossRef]
- Khurana, D.; Koli, A.; Khatter, K.; Singh, S. Natural language processing: State of the art, current trends and challenges. Multimed. Tools Appl. 2023, 82, 3713–3744. [Google Scholar] [CrossRef]
- Saeed, M.K.; Al Mazroa, A.; Alghamdi, B.M.; Alallah, F.S.; Alshareef, A.; Mahmud, A. Predictive analytics of complex healthcare systems using deep learning based disease diagnosis model. Sci. Rep. 2024, 14, 27497. [Google Scholar] [CrossRef]
- Yamashita, R.; Nishio, M.; Do, R.K.G.; Togashi, K. Convolutional neural networks: An overview and application in radiology. Insights Imaging 2018, 9, 611–629. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. [Google Scholar]
- Hochreiter, S. The vanishing gradient problem during learning recurrent neural nets and problem solutions. Int. J. Uncertainty Fuzziness Knowl.-Based Syst. 1998, 6, 107–116. [Google Scholar] [CrossRef]
- Tan, M.; Le, Q. Efficientnet: Rethinking model scaling for convolutional neural networks. In Proceedings of the International Conference on Machine Learning PMLR, Long Beach, CA USA, 9–15 June 2019; pp. 6105–6114. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An image is worth 16x16 words: Transformers for image recognition at scale. In Proceedings of the International Conference on Learning Representations, Virtual Event, 3–7 May 2021. [Google Scholar]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin transformer: Hierarchical vision transformer using shifted windows. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, QC, Canada, 11–17 October 2021; pp. 10012–10022. [Google Scholar]
- Yosinski, J.; Clune, J.; Bengio, Y.; Lipson, H. How transferable are features in deep neural networks? In Advances in Neural Information Processing Systems 27; NIPS Foundation: Lake Tahoe, CA, Canada, 2014. [Google Scholar]
- Weiss, K.; Khoshgoftaar, T.M.; Wang, D. A survey of transfer learning. J. Big Data 2016, 3, 9. [Google Scholar] [CrossRef]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Fei-Fei, L. Imagenet: A large-scale hierarchical image database. In Proceedings of the 2009 IEEE Conference on Computer Vision and Pattern Recognition; IEEE: Piscataway, NJ, USA, 2009; pp. 248–255. [Google Scholar]
- Zhuang, F.; Qi, Z.; Duan, K.; Xi, D.; Zhu, Y.; Zhu, H.; Xiong, H.; He, Q. A comprehensive survey on transfer learning. Proc. IEEE 2020, 109, 43–76. [Google Scholar] [CrossRef]
- Wang, X.; Peng, Y.; Lu, L.; Lu, Z.; Bagheri, M.; Summers, R.M. Chestx-ray8: Hospital-scale chest X-ray database and benchmarks on weakly-supervised classification and localization of common thorax diseases. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2097–2106. [Google Scholar]
- Lee, R.S.; Gimenez, F.; Hoogi, A.; Miyake, K.K.; Gorovoy, M.; Rubin, D.L. A curated mammography data set for use in computer-aided detection and diagnosis research. Sci. Data 2017, 4, 170177. [Google Scholar] [CrossRef]
- Raj, R.; Mathew, J.; Kannath, S.K.; Rajan, J. Crossover based technique for data augmentation. Comput. Methods Programs Biomed. 2022, 218, 106716. [Google Scholar] [CrossRef]
- Soares, E.; Angelov, P.; Zhang, Z. An explainable approach to deep learning from CT-scans for COVID identification. Evol. Syst. 2024, 15, 2159–2168. [Google Scholar] [CrossRef]
- Khoruzhaya, A.N.; Bobrovskaya, T.M.; Kozlov, D.V.; Kuligovskiy, D.; Novik, V.P.; Arzamasov, K.M.; Kremneva, E.I. Expanded Brain CT Dataset for the Development of AI Systems for Intracranial Hemorrhage Detection and Classification. Data 2024, 9, 30. [Google Scholar] [CrossRef]
- Islam, M.N.; Hasan, M.; Hossain, M.K.; Alam, M.G.R.; Uddin, M.Z.; Soylu, A. Vision transformer and explainable transfer learning models for auto detection of kidney cyst, stone and tumor from CT-radiography. Sci. Rep. 2022, 12, 11440. [Google Scholar] [CrossRef] [PubMed]
- Burgos-Artizzu, X.P.; Coronado-Gutiérrez, D.; Valenzuela-Alcaraz, B.; Bonet-Carne, E.; Eixarch, E.; Crispi, F.; Gratacós, E. Evaluation of deep convolutional neural networks for automatic classification of common maternal fetal ultrasound planes. Sci. Rep. 2020, 10, 10200. [Google Scholar] [CrossRef]
- Montoya, A.; Hasnin; kaggle446; shirzad; Cukierski, W.; yffud. Ultrasound Nerve Segmentation. Kaggle. 2016. Available online: https://kaggle.com/competitions/ultrasound-nerve-segmentation (accessed on 15 February 2025).
- Born, J.; Wiedemann, N.; Cossio, M.; Buhre, C.; Brändle, G.; Leidermann, K.; Goulet, J.; Aujayeb, A.; Moor, M.; Rieck, B.; et al. Accelerating detection of lung pathologies with explainable ultrasound image analysis. Appl. Sci. 2021, 11, 672. [Google Scholar] [CrossRef]
- Danilov, V.; Klyshnikov, K.; Kutikhin, A.; Gerget, O.; Frangi, A.; Ovcharenko, E. Angiographic dataset for stenosis detection. Mendeley Data 2021, 2. [Google Scholar] [CrossRef]
- Knoll, F.; Zbontar, J.; Sriram, A.; Muckley, M.J.; Bruno, M.; Defazio, A.; Parente, M.; Geras, K.J.; Katsnelson, J.; Chandarana, H.; et al. fastMRI: A publicly available raw k-space and DICOM dataset of knee images for accelerated MR image reconstruction using machine learning. Radiol. Artif. Intell. 2020, 2, e190007. [Google Scholar] [CrossRef]
- Zbontar, J.; Knoll, F.; Sriram, A.; Murrell, T.; Huang, Z.; Muckley, M.; Defazio, A.; Stern, R.; Johnson, P.; Bruno, M.; et al. fastMRI: An open dataset and benchmarks for accelerated MRI. arXiv 2018, arXiv:1811.08839. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. In Proceedings of the 3rd International Conference on Learning Representations, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual explanations from deep networks via gradient-based localization. In Proceedings of the IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 618–626. [Google Scholar]
| Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |