Benchmarking Tabular Foundation Models for Total Volatile Fatty Acid Prediction in Anaerobic Digestion
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. Data Preparation and Feature Engineering
2.3. Machine Learning Models
3. Results
3.1. Model Performance Comparison
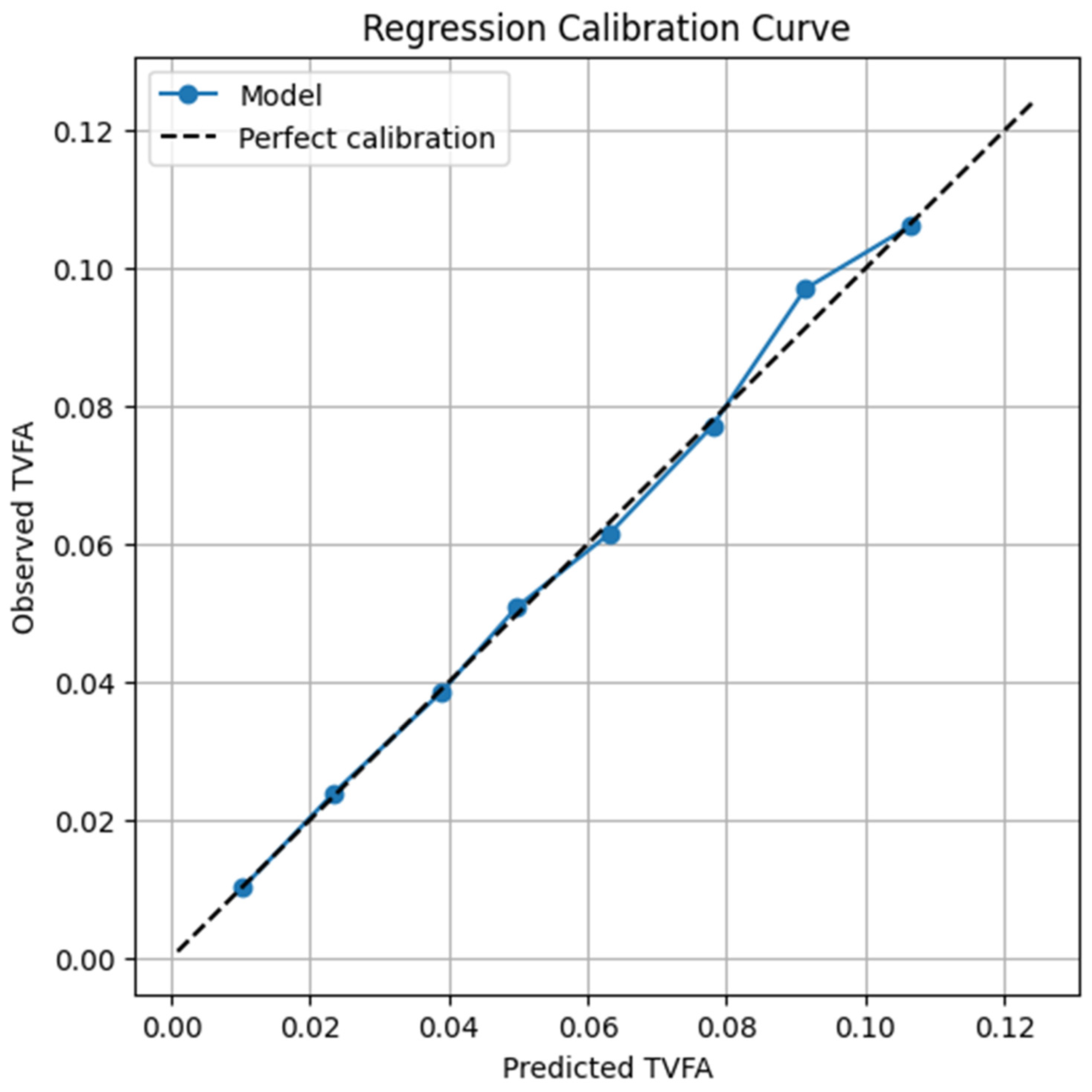
3.2. In-Depth Analysis of the Optimal (RealTabPFN-V2.5) Model
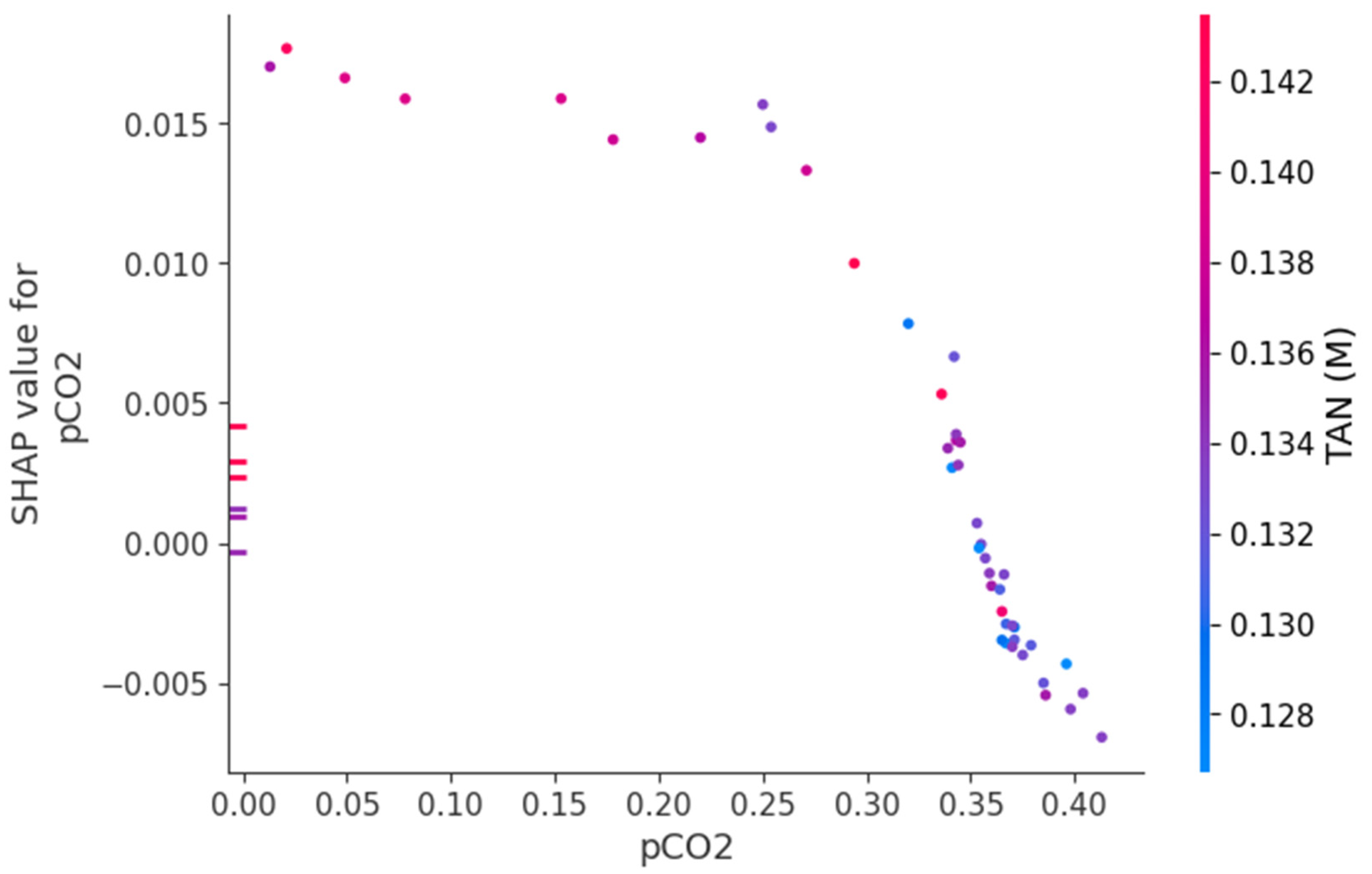
3.3. Feature Importance and Mechanistic Interpretation
3.4. Error Stratification and Reliability Analysis Across TVFA (M) Operating Regimes
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| TVFA (M) | Total Volatile Fatty Acids |
| AD | Anaerobic Digestion |
| pH | potential of Hydrogen |
| pCO2 | partial pressure of carbon dioxide |
| CO2 | carbon dioxide |
| TAN | Total Ammoniacal Nitrogen |
| SOTA | State-of-the-Art |
| RMSE | Root Mean Squared Error |
| MSE | Mean Squared Error |
| MAE | Mean Absolute Error |
| MAPE | Mean Absolute Percentage Error |
| R2 | coefficient of determination |
| SHAP | SHapley Additive exPlanations |
| XAI | eXplainable Artificial Intelligence |
| ANN | Artificial Neural Network(s) |
| SVM | Support Vector Machine(s) |
| ML | Machine Learning |
| IoT | Internet of Things |
| SCADA | Supervisory Control and Data Acquisition |
| TRL | Technology Readiness Level |
| MLR | Multiple Linear Regression |
| NH3 | ammonia |
| NH4+ | ammonium |
References
- Sevillano, C.A.; Pesantes, A.A.; Peña Carpio, E.; Martínez, E.J.; Gómez, X. Anaerobic digestion for producing renewable energy: The evolution of this technology in a new uncertain scenario. Entropy 2021, 23, 145. [Google Scholar] [CrossRef] [PubMed]
- Archana, K.; Visckram, A.; Kumar, P.S.; Manikandan, S.; Saravanan, A.; Natrayan, L. A review on recent technological breakthroughs in anaerobic digestion of organic biowaste for biogas generation: Challenges towards sustainable development goals. Fuel 2023, 358, 130298. [Google Scholar] [CrossRef]
- Wu, D.; Li, L.; Zhao, X.; Peng, Y.; Yang, P.; Peng, X. Anaerobic digestion: A review on process monitoring. Renew. Sustain. Energy Rev. 2019, 103, 1–12. [Google Scholar] [CrossRef]
- Rudakiya, D.; Narra, M. Microbial community dynamics in anaerobic digesters for biogas production. In Microbial Rejuvenation of Polluted Environment; Springer: Singapore, 2021; pp. 143–159. [Google Scholar] [CrossRef]
- Beschkov, V.N.; Angelov, I.K. Volatile Fatty Acid Production vs. Methane and Hydrogen in Anaerobic Digestion. Fermentation 2025, 11, 172. [Google Scholar] [CrossRef]
- Lee, D.J.; Teng, K.H.; Show, K.Y.; Chang, J.S. Effect of volatile fatty acid concentration on anaerobic degradation rate of food waste. J. Environ. Sci. Health A 2015, 50, 1253–1258. [Google Scholar] [CrossRef]
- Cruz, I.A.; Chomiak, M.; Cavaleiro, A.J.; Pereira, M.A.; Alves, M.M. An overview of process monitoring for anaerobic digestion. Bioresour. Technol. 2021, 337, 125402. [Google Scholar] [CrossRef]
- Kazemi, P.; Steyer, J.-P.; Bengoa, C.; Font, J.; Giralt, J. Robust data-driven soft sensors for online monitoring of volatile fatty acids in anaerobic digestion processes. Processes 2020, 8, 67. [Google Scholar] [CrossRef]
- Wang, X.; Rashid, I.; Zhao, Z.; Oladele, M.; Xiang, W.; Huang, Y.; Wazer, E.; McCutcheon, J.; Bollas, G.; Contreras, J.; et al. Machine learning algorithm integrated with real-time in situ sensors and physiochemical principle-driven soft sensors toward an anaerobic digestion data fusion framework. ACS ES&T Water 2023, 3, 1061–1072. [Google Scholar] [CrossRef]
- Choi, S.; Kim, S.I.; Yulisa, A.; Aghasa, A.; Hwang, S. Proactive prediction of total volatile fatty acids concentration in multiple full-scale food waste anaerobic digestion systems using substrate characteristics with machine learning and feature analysis. Waste Biomass Valor. 2023, 14, 593–608. [Google Scholar] [CrossRef]
- Mahmoodi-Eshkaftaki, M.; Mockaitis, G.; Rafiee, M.R. Dynamic optimization of volatile fatty acids to enrich biohydrogen production using a deep learning neural network. Biomass Conv. Bioref. 2024, 14, 8003–8014. [Google Scholar] [CrossRef]
- Kim, H.G.; Yu, S.I.; Shin, S.G.; Cho, K.H. Graph-based deep learning for predictions on changes in microbiomes and biogas production in anaerobic digestion systems. Water Res. 2025, 274, 123144. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Lü, F.; Chen, L.; Zhang, H.; He, P. Machine learning for enhancing prediction of biogas production and building a VFA/ALK soft sensor in full-scale dry anaerobic digestion of kitchen food waste. J. Environ. Manag. 2024, 371, 123190. [Google Scholar] [CrossRef] [PubMed]
- Rutland, H.; You, J.; Liu, H.; Bull, L.; Reynolds, D. A Systematic Review of Machine-Learning Solutions in Anaerobic Digestion. Bioengineering 2023, 10, 1410. [Google Scholar] [CrossRef] [PubMed]
- Behera, S.R.; Balasundaram, G. Artificial intelligence in anaerobic digestion: A review of sensors, soft sensors, and machine learning applications. Bioresour. Technol. 2025, 425, 131850. [Google Scholar]
- He, L.; Niu, M.; Tiwari, P.; Marttinen, P.; Su, R.; Jiang, J.; Guo, C.; Wang, H.; Ding, S.; Wang, Z.; et al. Deep learning for depression recognition with audiovisual cues: A review. Inf. Fusion 2022, 80, 56–86. [Google Scholar] [CrossRef]
- Shwartz-Ziv, R.; Armon, A. Tabular data: Deep learning is not all you need. Inf. Fusion 2022, 81, 84–90. [Google Scholar] [CrossRef]
- Hollmann, N.; Müller, S.; Eggensperger, K.; Hutter, F. TABPFN: A transformer that solves small tabular classification problems in a second. arXiv 2022, arXiv:2207.01848. [Google Scholar] [CrossRef]
- Hollmann, N.; Müller, S.; Purucker, L.; Krishnakumar, A.; Körfer, M.; Hoo, S.B.; Schirrmeister, R.T.; Hutter, F. Accurate predictions on small data with a tabular foundation model. Nature 2025, 633, 792–799. [Google Scholar] [CrossRef]
- Ruiz, L.M.; Fernández, M.; Genaro, A.; Martín-Pascual, J.; Zamorano, M. Multi-parametric analysis based on physico-chemical characterization and biochemical methane potential estimation for the selection of industrial wastes as co-substrates in anaerobic digestion. Energies 2023, 16, 5444. [Google Scholar] [CrossRef]
- Zhang, W. The Impact of Nitrogen Control Strategies and of Biopackaging Degradation on the Implementation of the Anaerobic Digestion of Selected MSW Fractions. Ph.D. Thesis, University of Southampton, Southampton, UK, 2019. [Google Scholar]
- Rossi, A.; Morlino, M.S.; Gaspari, M.; Campanaro, S.; Basile, A.; Kougias, P.; Treu, L. Analysis of the anaerobic digestion metagenome under environmental stresses stimulating prophage induction. Microbiome 2022, 10, 125. [Google Scholar] [CrossRef]
- Ganeshan, P.; Bose, A.; Lee, J.; Barathi, S.; Rajendran, K. Machine learning for high solid anaerobic digestion: Performance prediction and optimization. Bioresour. Technol. 2024, 400, 130665. [Google Scholar] [CrossRef] [PubMed]
- Yildirim, O.; Ozkaya, B. Prediction of biogas production of industrial-scale anaerobic digestion plant by machine learning algorithms. Chemosphere 2023, 335, 138976. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Heaven, S.; Banks, C.J. Validation of two theoretically derived equations for predicting pH in CO2 biomethanisation. Processes 2023, 11, 113. [Google Scholar] [CrossRef]
- Madsen, M.; Holm-Nielsen, J.B.; Esbensen, K.H. Monitoring of anaerobic digestion processes: A review perspective. Renew. Sustain. Energy Rev. 2011, 15, 3141–3155. [Google Scholar] [CrossRef]
- Boe, K.; Batstone, D.J.; Steyer, J.-P.; Angelidaki, I. State indicators for monitoring the anaerobic digestion process. Water Res. 2010, 44, 5973–5980. [Google Scholar] [CrossRef]
- Yang, J.; Zhang, J.; Du, X.; Gao, T.; Cheng, Z.; Fu, W.; Wang, S. Ammonia inhibition in anaerobic digestion of organic waste: A review. Int. J. Environ. Sci. Technol. 2025, 22, 3927–3942. [Google Scholar] [CrossRef]
- Ahring, B.K.; Sandberg, M.; Angelidaki, I. Volatile fatty acids as indicators of process imbalance in anaerobic digestors. Appl. Microbiol. Biotechnol. 1995, 43, 559–565. [Google Scholar] [CrossRef]
- Stekhoven, D.J.; Bühlmann, P. MissForest—Non-parametric missing value imputation for mixed-type data. Bioinformatics 2012, 28, 112–118. [Google Scholar] [CrossRef]
- Liu, L.; Chen, X.; Petinrin, O.O.; Zhang, W.; Rahaman, S.; Tang, Z.-R.; Wong, K.-C. Machine learning protocols in early cancer detection based on liquid biopsy: A survey. Life 2021, 11, 638. [Google Scholar] [CrossRef]
- Kraskov, A.; Stögbauer, H.; Grassberger, P. Estimating mutual information. Phys. Rev. E 2004, 69, 066138. [Google Scholar] [CrossRef]
- Givisis, I.; Kalatzis, D.; Christakis, C.; Kiouvrekis, Y. Comparing explainable AI models: SHAP, LIME, and their role in electric field strength prediction over urban areas. Electronics 2025, 14, 4766. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, H. Random forest-based machine failure prediction: A performance comparison. Appl. Sci. 2025, 15, 8841. [Google Scholar] [CrossRef]
- Grinsztajn, Y.; Oyallon, E.; Varoquaux, G. Why do tree-based models still outperform deep learning on tabular data? arXiv 2022, arXiv:2207.08815. [Google Scholar] [CrossRef]
- Cação, J.; Santos, J.; Antunes, M. Explainable AI for industrial fault diagnosis: A systematic review. J. Ind. Inf. Integr. 2025, 47, 100905. [Google Scholar] [CrossRef]
- López-Trujillo, J.; Mellado-Bosque, M.; Ascacio-Valdés, J.A.; Prado-Barragán, L.A.; Hernández-Herrera, J.A.; Aguilera-Carbó, A.F. Temperature and pH optimization for protease production fermented by Yarrowia lipolytica from agro-industrial waste. Fermentation 2023, 9, 819. [Google Scholar] [CrossRef]









| Parameter | Mean ± SD |
|---|---|
| pCO2 | 0.358 ± 0.009 |
| TAN (M) | 0.1273 ± 0.0005 |
| pH | 7.498 ± 0.031 |
| TVFA (M) | 0.0079 ± 0.0053 |
| Models | R2 | MSE (M2) | RMSE (M) | MAE (M) |
|---|---|---|---|---|
| RealTabPFN-v2.5 | 0.889008 ± 0.015600 | 0.000063 ± 0.000011 | 0.007924 ± 0.000723 | 0.005586 ± 0.000590 |
| TorchMLP (tuned) | 0.846959 ± 0.024775 | 0.000087 ± 0.000018 | 0.009292 ± 0.000976 | 0.006724 ± 0.000519 |
| Random Forest (tuned) | 0.828587 ± 0.034933 | 0.000096 ± 0.000011 | 0.009781 ± 0.000566 | 0.006716 ± 0.000586 |
| XGBoost (tuned) | 0.827921 ± 0.034601 | 0.000096 ± 0.000013 | 0.009805 ± 0.000635 | 0.006707 ± 0.000464 |
| Random Forest (default) | 0.811978 ± 0.045228 | 0.000105 ± 0.000011 | 0.010212 ± 0.000555 | 0.006806 ± 0.000467 |
| FastaiMLP (tuned) | 0.809788 ± 0.041055 | 0.000106 ± 0.000012 | 0.010296 ± 0.000604 | 0.007942 ± 0.000434 |
| XGBoost (default) | 0.796295 ± 0.042203 | 0.000114 ± 0.000010 | 0.010655 ± 0.000464 | 0.007269 ± 0.000548 |
| RealMLP (tuned) | 0.779525 ± 0.037754 | 0.000125 ± 0.000028 | 0.011148 ± 0.001221 | 0.007952 ± 0.001003 |
| Operating Regimes | MSE | RMSE |
|---|---|---|
| Stable | 0.000014 ± 0.000001 | 0.003798 ± 0.000170 |
| Overload | 0.000107 ± 0.000057 | 0.010033 ± 0.002876 |
| Operating Regimes | 0.50 | 0.75 | 0.90 | 0.95 |
|---|---|---|---|---|
| Stable | 0.000036 | 0.000128 | 0.000413 | 0.000791 |
| overload | 0.000012 | 0.000043 | 0.000114 | 0.000227 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Amangeldy, B.; Baigarayeva, Z.; Tasmurzayev, N.; Boltaboyeva, A.; Imanbek, B.; Maulenbekov, M.; Zhussupbekov, S.; Wojcik, W.; Kozhamberdieva, M.; Konysbekova, A. Benchmarking Tabular Foundation Models for Total Volatile Fatty Acid Prediction in Anaerobic Digestion. Algorithms 2026, 19, 127. https://doi.org/10.3390/a19020127
Amangeldy B, Baigarayeva Z, Tasmurzayev N, Boltaboyeva A, Imanbek B, Maulenbekov M, Zhussupbekov S, Wojcik W, Kozhamberdieva M, Konysbekova A. Benchmarking Tabular Foundation Models for Total Volatile Fatty Acid Prediction in Anaerobic Digestion. Algorithms. 2026; 19(2):127. https://doi.org/10.3390/a19020127
Chicago/Turabian StyleAmangeldy, Bibars, Zhanel Baigarayeva, Nurdaulet Tasmurzayev, Assiya Boltaboyeva, Baglan Imanbek, Marlen Maulenbekov, Sarsenbek Zhussupbekov, Waldemar Wojcik, Mergul Kozhamberdieva, and Akzhan Konysbekova. 2026. "Benchmarking Tabular Foundation Models for Total Volatile Fatty Acid Prediction in Anaerobic Digestion" Algorithms 19, no. 2: 127. https://doi.org/10.3390/a19020127
APA StyleAmangeldy, B., Baigarayeva, Z., Tasmurzayev, N., Boltaboyeva, A., Imanbek, B., Maulenbekov, M., Zhussupbekov, S., Wojcik, W., Kozhamberdieva, M., & Konysbekova, A. (2026). Benchmarking Tabular Foundation Models for Total Volatile Fatty Acid Prediction in Anaerobic Digestion. Algorithms, 19(2), 127. https://doi.org/10.3390/a19020127








