A Deep Learning Model for Data-Driven Discovery of Functional Connectivity
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.1.1. Fbirn
2.1.2. Preprocessing
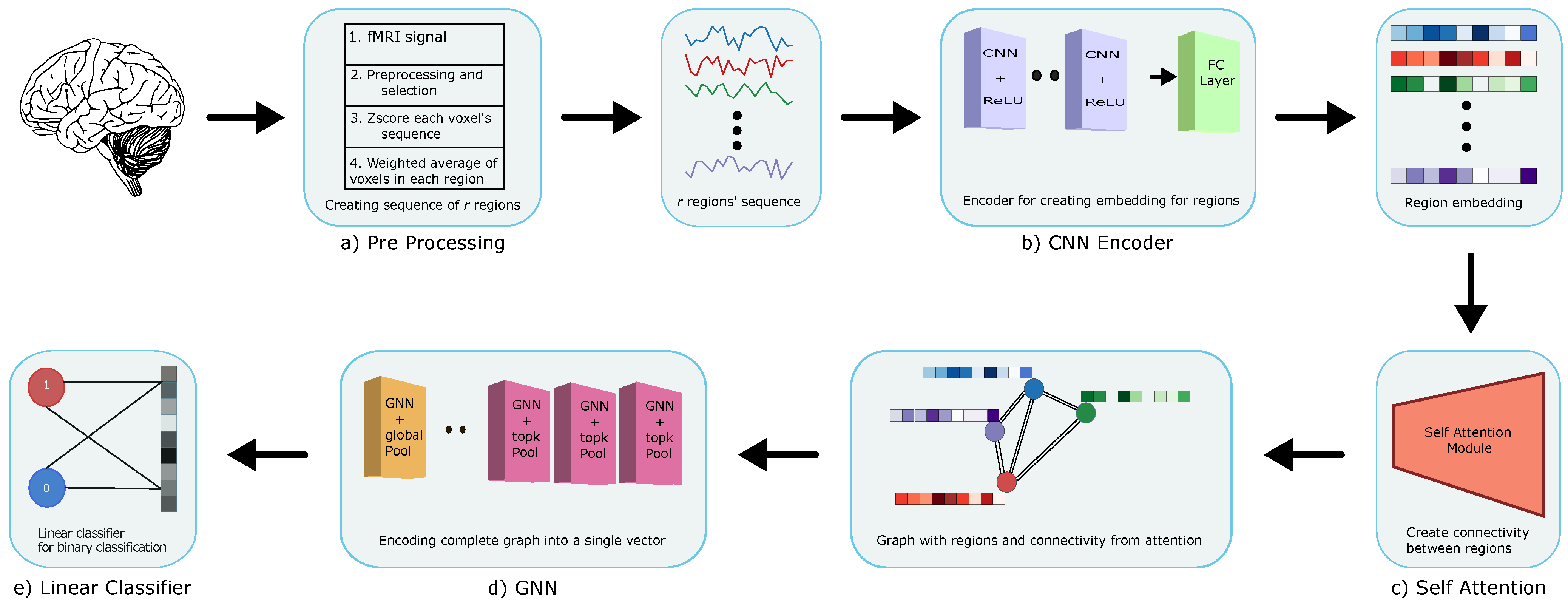
2.2. Method
2.2.1. Cnn Encoder
2.2.2. Self Attention
2.2.3. GNN
2.2.4. Training and Testing
3. Results
3.1. Classification
3.2. Functional Connectivity
3.3. Region Selection
4. Discussion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Liu, Y.; Liang, M.; Zhou, Y.; He, Y.; Hao, Y.; Song, M.; Yu, C.; Liu, H.; Liu, Z.; Jiang, T. Disrupted small-world networks in schizophrenia. Brain 2008, 131, 945–961. [Google Scholar] [CrossRef]
- Lynall, M.E.; Bassett, D.S.; Kerwin, R.; McKenna, P.J.; Kitzbichler, M.; Muller, U.; Bullmore, E. Functional Connectivity and Brain Networks in Schizophrenia. J. Neurosci. 2010, 30, 9477–9487. [Google Scholar] [CrossRef]
- Yu, R.; Zhang, H.; An, L.; Chen, X.; Wei, Z.; Shen, D. Connectivity strength-weighted sparse group representation-based brain network construction for MCI classification. Hum. Brain Mapp. 2017, 38, 2370–2383. [Google Scholar] [CrossRef]
- Gadgil, S.; Zhao, Q.; Pfefferbaum, A.; Sullivan, E.V.; Adeli, E.; Pohl, K.M. Spatio-Temporal Graph Convolution for Resting-State fMRI Analysis. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2020; Martel, A.L., Abolmaesumi, P., Stoyanov, D., Mateus, D., Zuluaga, M.A., Zhou, S.K., Racoceanu, D., Joskowicz, L., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 528–538. [Google Scholar] [CrossRef]
- Yan, W.; Plis, S.; Calhoun, V.D.; Liu, S.; Jiang, R.; Jiang, T.Z.; Sui, J. Discriminating schizophrenia from normal controls using resting state functional network connectivity: A deep neural network and layer-wise relevance propagation method. In Proceedings of the 2017 IEEE 27th International Workshop on Machine Learning for Signal Processing (MLSP), Tokyo, Japan, 25–28 September 2017; pp. 1–6. [Google Scholar] [CrossRef]
- Parisot, S.; Ktena, S.I.; Ferrante, E.; Lee, M.; Guerrero, R.; Glocker, B.; Rueckert, D. Disease prediction using graph convolutional networks: Application to Autism Spectrum Disorder and Alzheimer’s disease. Med. Image Anal. 2018, 48, 117–130. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, J.; Brown, C.; Miller, S.; Booth, B.; Chau, V.; Grunau, R.; Zwicker, J.; Hamarneh, G. BrainNetCNN: Convolutional Neural Networks for Brain Networks; Towards Predicting Neurodevelopment. NeuroImage 2016, 146. [Google Scholar] [CrossRef]
- Ktena, S.I.; Parisot, S.; Ferrante, E.; Rajchl, M.; Lee, M.; Glocker, B.; Rueckert, D. Distance Metric Learning Using Graph Convolutional Networks: Application to Functional Brain Networks. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2017; Descoteaux, M., Maier-Hein, L., Franz, A., Jannin, P., Collins, D.L., Duchesne, S., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 469–477. [Google Scholar] [CrossRef]
- Rashid, B.; Arbabshirani, M.R.; Damaraju, E.; Cetin, M.S.; Miller, R.; Pearlson, G.D.; Calhoun, V.D. Classification of schizophrenia and bipolar patients using static and dynamic resting-state fMRI brain connectivity. NeuroImage 2016, 134, 645–657. [Google Scholar] [CrossRef]
- Saha, D.K.; Damaraju, E.; Rashid, B.; Abrol, A.; Plis, S.M.; Calhoun, V.D. A classification-based approach to estimate the number of resting fMRI dynamic functional connectivity states. bioRxiv 2020. [Google Scholar] [CrossRef]
- Salman, M.S.; Du, Y.; Lin, D.; Fu, Z.; Fedorov, A.; Damaraju, E.; Sui, J.; Chen, J.; Mayer, A.R.; Posse, S.; et al. Group ICA for identifying biomarkers in schizophrenia: ‘Adaptive’ networks via spatially constrained ICA show more sensitivity to group differences than spatio-temporal regression. NeuroImage 2019, 22, 101747. [Google Scholar] [CrossRef]
- Du, Y.; Fu, Z.; Calhoun, V.D. Classification and Prediction of Brain Disorders Using Functional Connectivity: Promising but Challenging. Front. Neurosci. 2018, 12, 525. [Google Scholar] [CrossRef]
- Li, Y.; Zemel, R.; Brockschmidt, M.; Tarlow, D. Gated Graph Sequence Neural Networks. In Proceedings of the ICLR’16, San Juan, Puerto Rico, 2–4 May 2016. [Google Scholar]
- Keator, D.B.; van Erp, T.G.; Turner, J.A.; Glover, G.H.; Mueller, B.A.; Liu, T.T.; Voyvodic, J.T.; Rasmussen, J.; Calhoun, V.D.; Lee, H.J.; et al. The function biomedical informatics research network data repository. Neuroimage 2016, 124, 1074–1079. [Google Scholar] [CrossRef]
- Fu, Z.; Iraji, A.; Turner, J.A.; Sui, J.; Miller, R.; Pearlson, G.D.; Calhoun, V.D. Dynamic state with covarying brain activity-connectivity: On the pathophysiology of schizophrenia. NeuroImage 2021, 224, 117385. [Google Scholar] [CrossRef]
- Fu, Z.; Caprihan, A.; Chen, J.; Du, Y.; Adair, J.C.; Sui, J.; Rosenberg, G.A.; Calhoun, V.D. Altered static and dynamic functional network connectivity in Alzheimer’s disease and subcortical ischemic vascular disease: Shared and specific brain connectivity abnormalities. Hum. Brain Mapp. 2019. [Google Scholar] [CrossRef] [PubMed]
- Tzourio-Mazoyer, N.; Landeau, B.; Papathanassiou, D.; Crivello, F.; Etard, O.; Delcroix, N.; Mazoyer, B.; Joliot, M. Automated Anatomical Labeling of Activations in SPM Using a Macroscopic Anatomical Parcellation of the MNI MRI Single-Subject Brain. NeuroImage 2002, 15, 273–289. [Google Scholar] [CrossRef]
- Lecun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, U.; Polosukhin, I. Attention is All You Need. In Proceedings of the 31st International Conference on Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017; pp. 6000–6010. [Google Scholar]
- Kiranyaz, S.; Avci, O.; Abdeljaber, O.; Ince, T.; Gabbouj, M.; Inman, D.J. 1D convolutional neural networks and applications: A survey. Mech. Syst. Signal Process. 2021, 151, 107398. [Google Scholar] [CrossRef]
- Cho, K.; van Merriënboer, B.; Bahdanau, D.; Bengio, Y. On the Properties of Neural Machine Translation: Encoder–Decoder Approaches. In Proceedings of the SSST-8, Eighth Workshop on Syntax, Semantics and Structure in Statistical Translation, Doha, Qatar, 25 October 2014; pp. 103–111. [Google Scholar] [CrossRef]
- Bresson, X.; Laurent, T. Residual Gated Graph ConvNets. arXiv 2017, arXiv:1711.07553. [Google Scholar]
- Gao, H.; Ji, S. Graph U-Nets. arXiv 2019, arXiv:1905.05178. [Google Scholar]
- Knyazev, B.; Taylor, G.W.; Amer, M.R. Understanding Attention and Generalization in Graph Neural Networks. arXiv 2019, arXiv:1905.02850. [Google Scholar]
- Vinyals, O.; Bengio, S.; Kudlur, M. Order Matters: Sequence to sequence for sets. arXiv 2016, arXiv:1511.06391. [Google Scholar]
- Mahmood, U.; Rahman, M.M.; Fedorov, A.; Lewis, N.; Fu, Z.; Calhoun, V.D.; Plis, S.M. Whole MILC: Generalizing Learned Dynamics Across Tasks, Datasets, and Populations. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2020; Martel, A.L., Abolmaesumi, P., Stoyanov, D., Mateus, D., Zuluaga, M.A., Zhou, S.K., Racoceanu, D., Joskowicz, L., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 407–417. [Google Scholar] [CrossRef]
- Jones, D.; Vemuri, P.; Murphy, M.; Gunter, J.; Senjem, M.; Machulda, M.; Przybelski, S.; Gregg, B.; Kantarci, K.; Knopman, D.; et al. Non-Stationarity in the “Resting Brain’s” Modular Architecture. PLoS ONE 2012, 7, e39731. [Google Scholar] [CrossRef]
- Fu, Z.; Sui, J.; Turner, J.A.; Du, Y.; Assaf, M.; Pearlson, G.D.; Calhoun, V.D. Dynamic functional network reconfiguration underlying the pathophysiology of schizophrenia and autism spectrum disorder. Hum. Brain Mapp. 2021, 42, 80–94. [Google Scholar] [CrossRef]
- Ebdrup, B.; Glenthøj, B.; Rasmussen, H.; Aggernaes, B.; Langkilde, A.; Paulson, O.; Lublin, H.; Skimminge, A.; Baaré, W. Hippocampal and caudate volume reductions in antipsychotic-naive first-episode schizophrenia. J. Psychiatry Neurosci. 2010, 35, 95–104. [Google Scholar] [CrossRef]
- Andreasen, N.; Pierson, R. The Role of the Cerebellum in Schizophrenia. Biol. Psychiatry 2008, 64, 81–88. [Google Scholar] [CrossRef] [PubMed]





| CV Fold | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AUC | 0.695 | 0.955 | 0.644 | 0.752 | 0.908 | 0.917 | 0.894 | 0.803 | 0.649 | 0.805 | 0.922 | 0.699 | 0.625 | 0.780 | 0.794 | 0.766 | 0.914 | 0.750 | 0.777 |
| Test | p Value |
|---|---|
| Mann-Whitney U Test | 0.0 |
| Welch’s t-test | 0.0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mahmood, U.; Fu, Z.; Calhoun, V.D.; Plis, S. A Deep Learning Model for Data-Driven Discovery of Functional Connectivity. Algorithms 2021, 14, 75. https://doi.org/10.3390/a14030075
Mahmood U, Fu Z, Calhoun VD, Plis S. A Deep Learning Model for Data-Driven Discovery of Functional Connectivity. Algorithms. 2021; 14(3):75. https://doi.org/10.3390/a14030075
Chicago/Turabian StyleMahmood, Usman, Zening Fu, Vince D. Calhoun, and Sergey Plis. 2021. "A Deep Learning Model for Data-Driven Discovery of Functional Connectivity" Algorithms 14, no. 3: 75. https://doi.org/10.3390/a14030075
APA StyleMahmood, U., Fu, Z., Calhoun, V. D., & Plis, S. (2021). A Deep Learning Model for Data-Driven Discovery of Functional Connectivity. Algorithms, 14(3), 75. https://doi.org/10.3390/a14030075






