A Novel Approach for Cognitive Clustering of Parkinsonisms through Affinity Propagation
Abstract
1. Introduction
2. Materials and Methods
2.1. Participants
2.2. MMSE Assessment
2.3. Statistical Analysis
2.3.1. Residuals Calculation
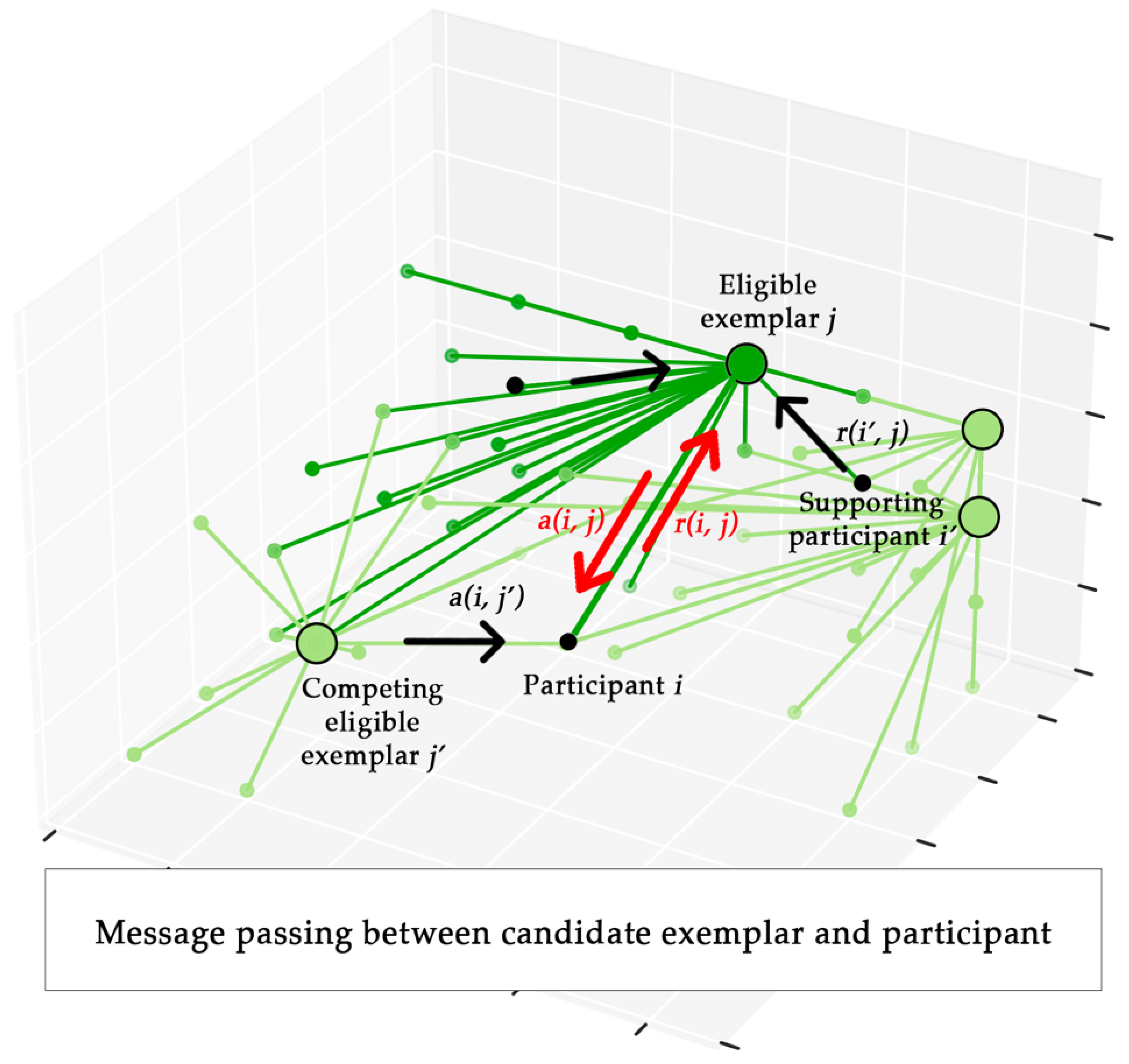
2.4. Affinity Propagation
| Algorithm 1 | Cognitive clustering of regression residuals trough Affinity Propagation |
| Input: | MMSE subscale scores matrix Y(n × m), covariates matrix X(k), damping factor d |
| Output: | Number of exemplars ex, cluster assignment vector ξ |
| 1: | initialization: E(n × m), matrix of regression residuals; S(n × n), matrix of similarities; p(n × 1), preference vector of Affinity Propagation; |
| 2: | forj = 1 to m do |
| 3: | fitj = ols(yj ~ x1 + … + xk); fitted model of the j-th column in Y, with x1, …, xk as the columns in X |
| 4: | for i = 1 to n−1 do |
| 5: | eij = yij—fitj.predict(yij); regression residual = the difference between yij and its prediction by fitj |
| 6: | fori = 1 to n do |
| 7: | for j = 1 to m do |
| 8: | for z = 1 to m do |
| 9: | sij = −; negative squared Euclidean distance between eiz and ejz |
| 10: | pi = Smin OR pi = Smedian |
| 11: | ap = AffinityPropagation(S, p, d); |
| 12: | ex = size(ap.exemplars); |
| 13: | ξ = ap.cluster_membership; |
2.5. Clustering Accuracy Assessment
3. Results
3.1. Statistical Analysis
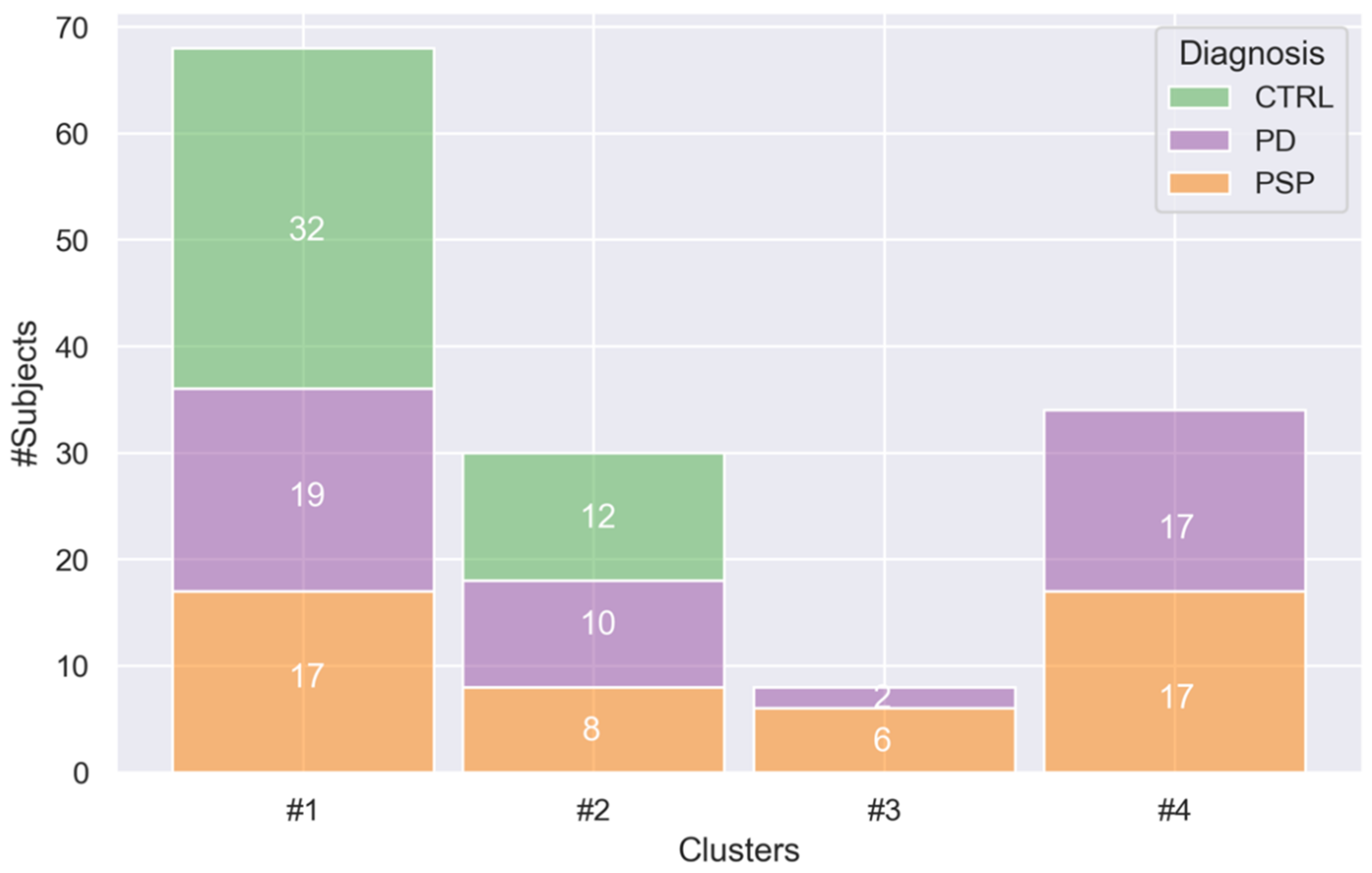
3.2. Cluster Analysis
- Cluster #1 CTRL: male, age 62, education 16, MMSE subscales = [5, 5, 3, 5, 3, 2, 1, 3, 1, 1, 1];
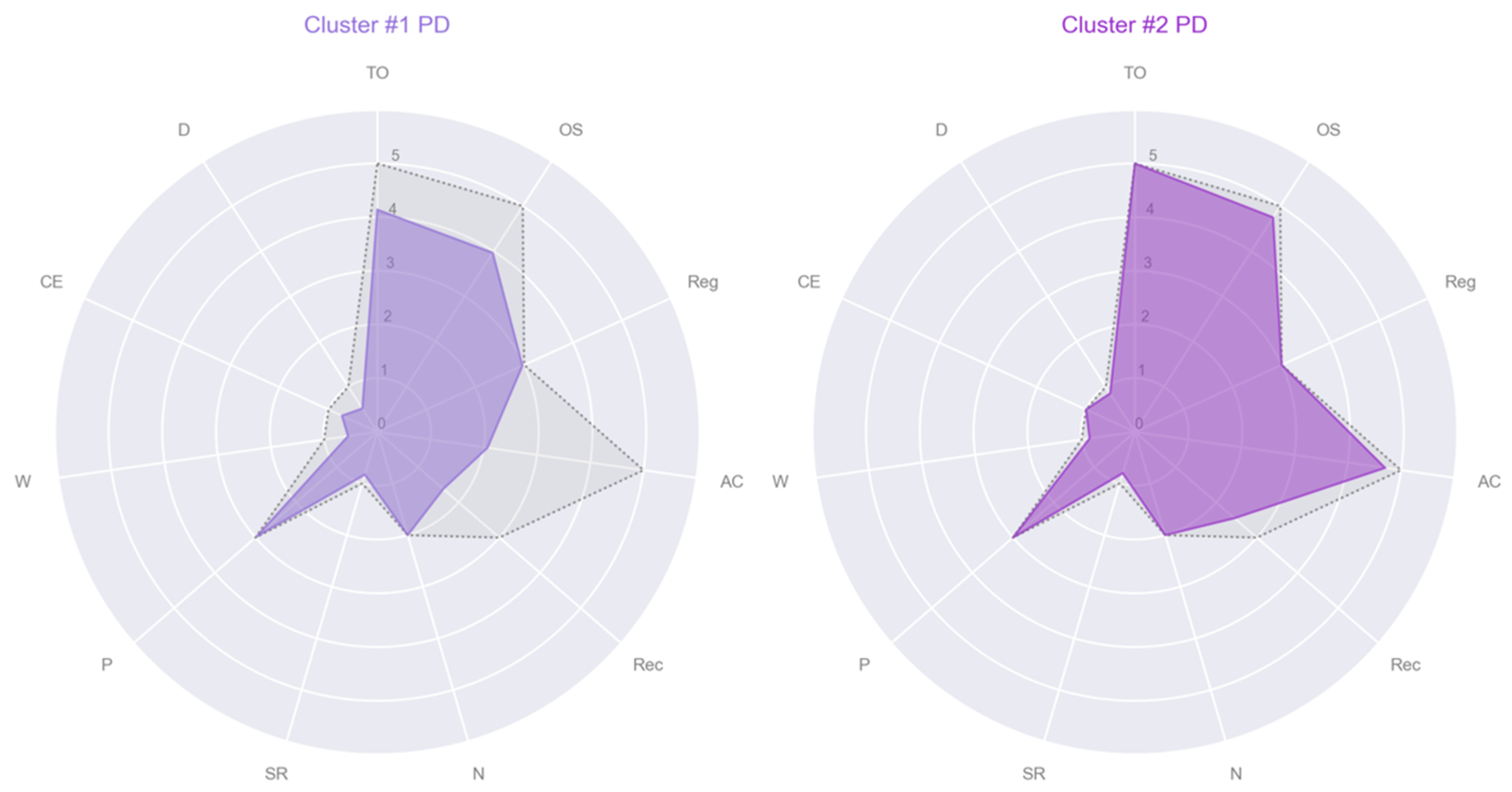
- Cluster #1 PD: male, age 43, education 13, MMSE subscales = [5, 5, 3, 4, 3, 2, 1, 3, 1, 1, 1];
- Cluster #2 PD: female, age 61, education 8, MMSE subscales = [5, 5, 3, 5, 3, 2, 1, 3, 1, 1, 1];
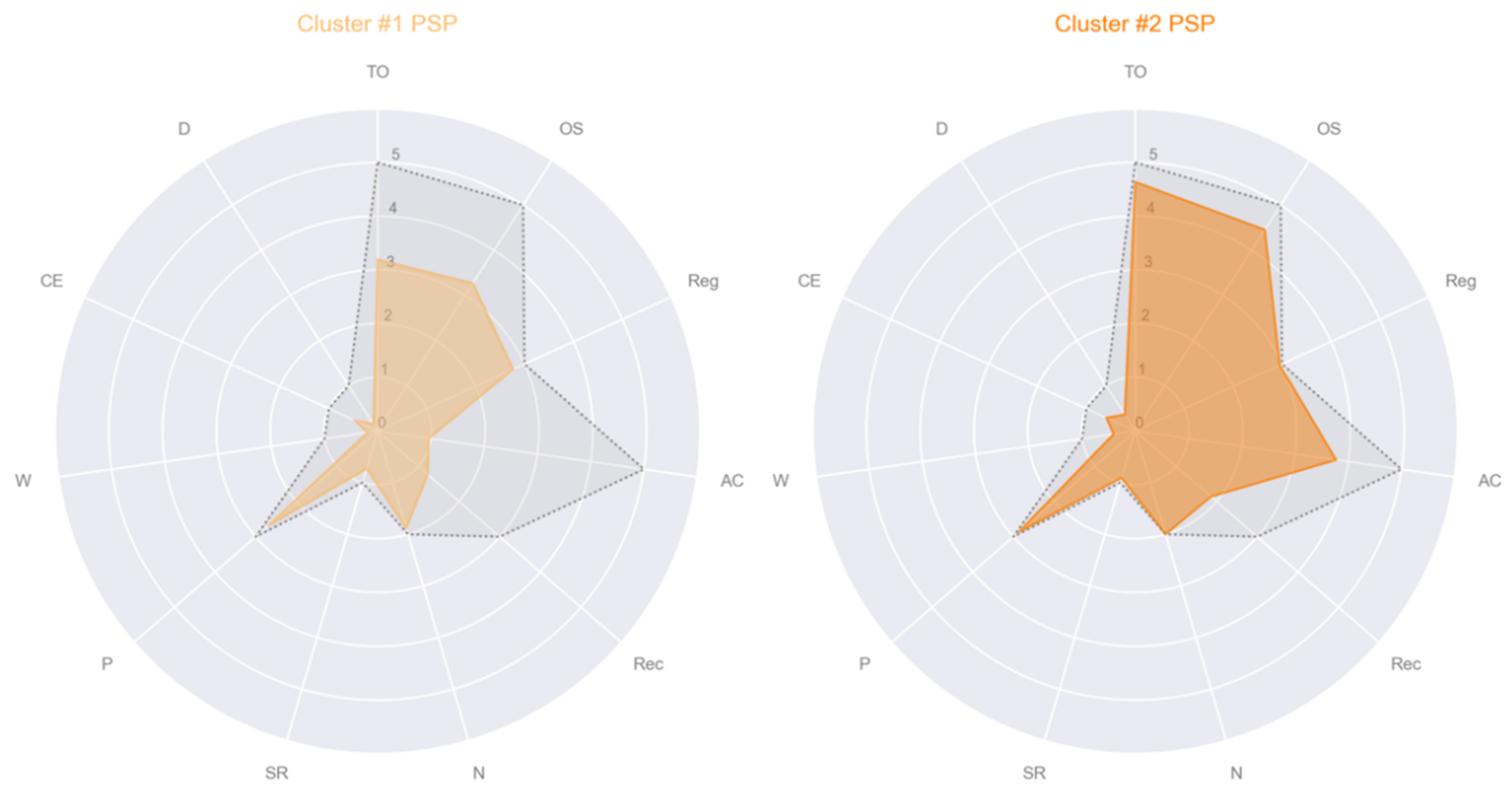
- Cluster #1 PSP: male, age 57, education 17, MMSE subscales = [5, 5, 3, 2, 1, 2, 1, 3, 0, 1, 0];
- Cluster #2 PSP: male, age 78, education 13, MMSE subscales = [5, 5, 3, 5, 2, 2, 1, 3, 0, 1, 0];
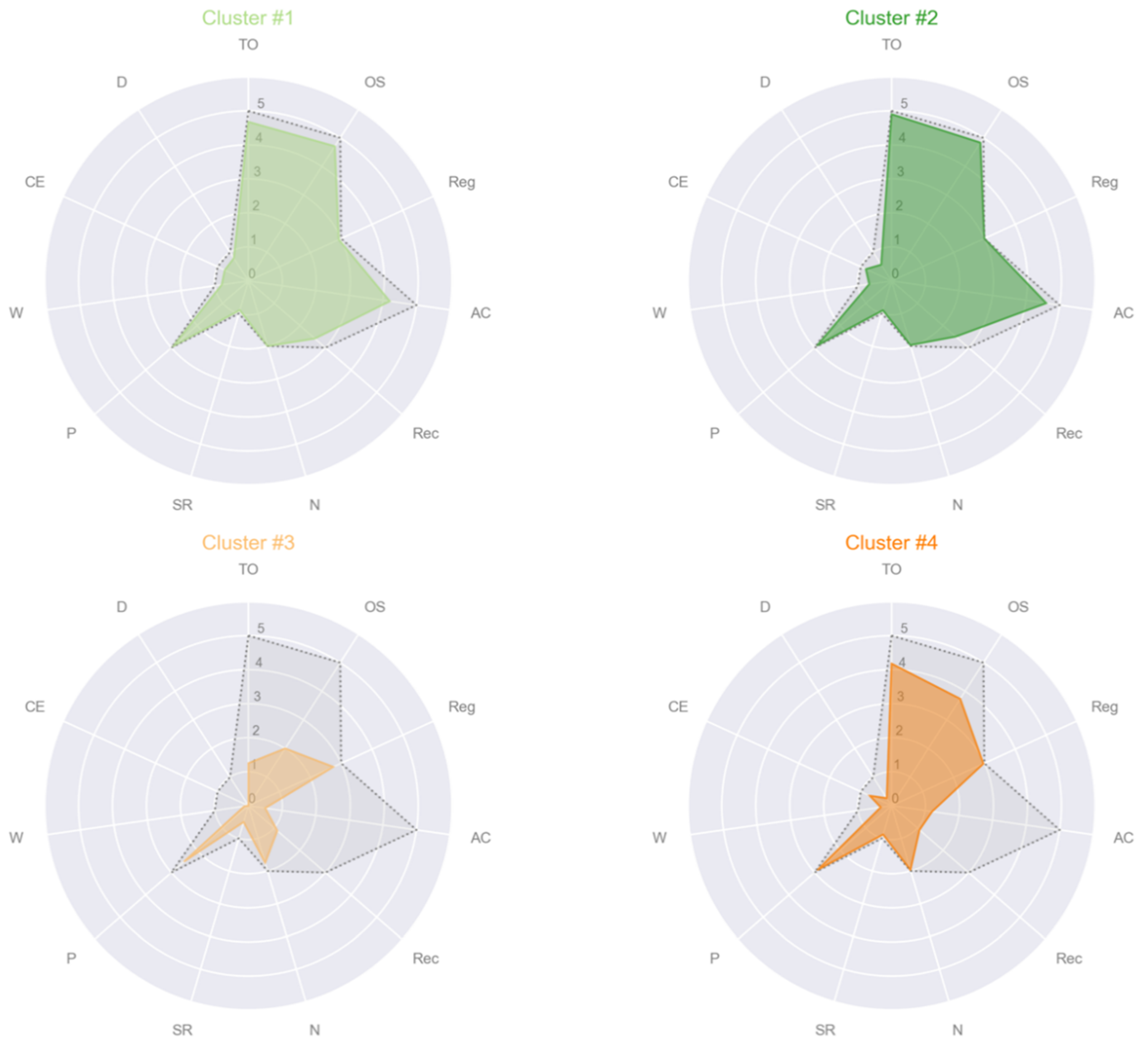
- Cluster #1: CTRL, female, age 59, education 16, MMSE subscales = [5, 5, 3, 5, 3, 2, 1, 3, 1, 1, 1];
- Cluster #2: PD, female, age 61, education 8, MMSE subscales = [5, 5, 3, 5, 3, 2, 1, 3, 1, 1, 1];
- Cluster #3: PD, female, age 70, education 5, MMSE subscales = [1, 2, 3, 1, 1, 2, 1, 3, 0, 0, 0];
- Cluster #4: PSP, male, age 78, education 8, MMSE subscales = [4, 4, 3, 1, 1, 2, 1, 3, 0, 1, 0].
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
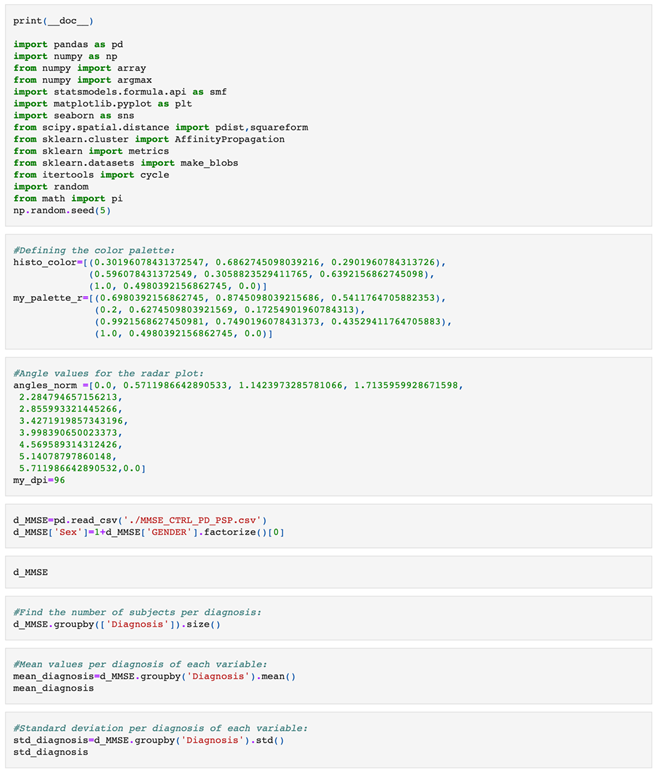 |
 |
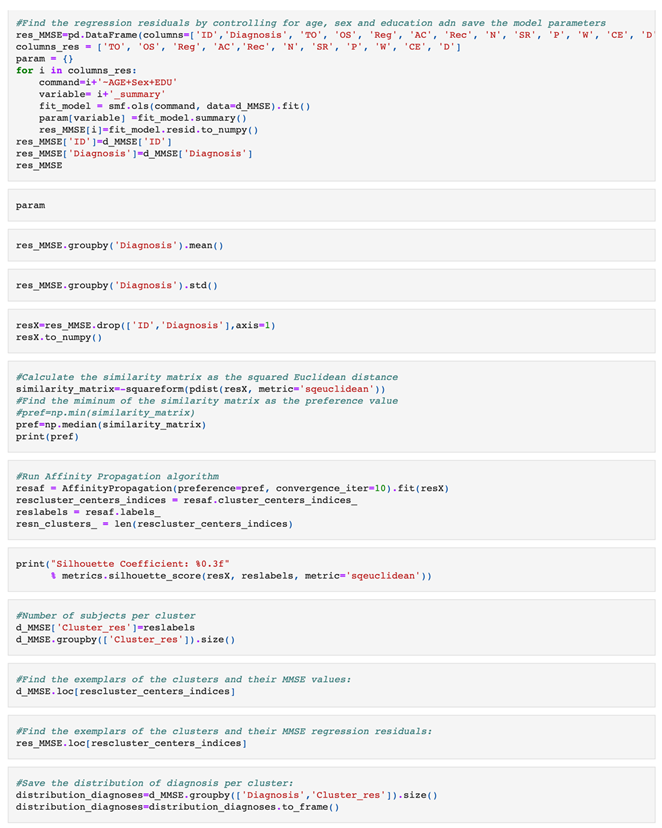 |
 |
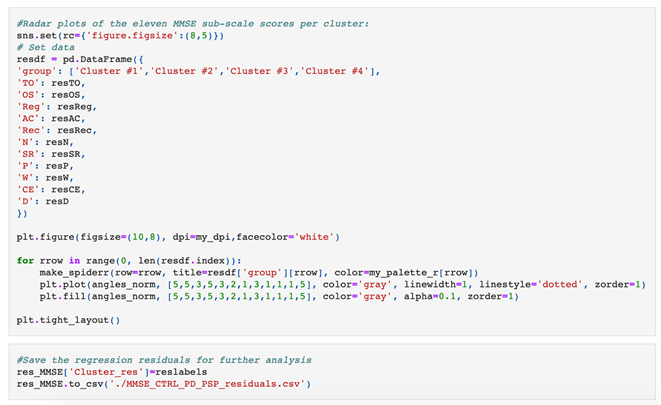 |
Appendix B
| R2 | F | Intercept (β0, p-Value) | Age (β1, p-Value) | Sex (β2, p-Value) | Education (β3, p-Value) | |
|---|---|---|---|---|---|---|
| TO | 0.193 | 10.8 | 4.9218, <0.001 | −0.0162, 0.104 | −0.1011, 0.563 | 0.0739, <0.001 |
| OS | 0.309 | 20.3 | 4.0724, <0.001 | −0.0142, 0.110 | 0.1743, 0.264 | 0.0983, <0.001 |
| Reg | 0.074 | 3.6 | 3.0925, <0.001 | −0.0029, 0.331 | −0.0489, 0.344 | 0.0118, 0.039 |
| AC | 0.312 | 20.6 | 3.6018, 0.004 | −0.0267, 0.080 | −0.0646, 0.809 | 0.1675, <0.001 |
| Rec | 0.114 | 5.8 | 2.9579, <0.001 | −0.0159, 0.092 | −0.1284, 0.437 | 0.0426, 0.020 |
| N | 0.083 | 4.1 | 1.7130, <0.001 | 0.0010, 0.598 | 0.0586, 0.085 | 0.0106, 0.005 |
| SR | 0.022 | 1 | 0.7640, 0.004 | 0.0007, 0.821 | −0.0240, 0.674 | 0.0098, 0.119 |
| P | 0.071 | 3.4 | 3.1548, <0.001 | −0.0069, 0.089 | 0.0584, 0.410 | 0.0111, 0.153 |
| W | 0.252 | 15.3 | 1.2908, <0.001 | −0.0125, 0.003 | −0.0789, 0.278 | 0.0283, <0.001 |
| CE | 0.150 | 8 | 0.9553, 0.004 | −0.0054, 0.183 | −0.0788, 0.270 | 0.0253, 0.010 |
| D | 0.379 | 27.7 | 1.3470, <0.001 | −0.0180, <0.001 | 0.0741, 0.271 | 0.0321, <0.001 |
References
- Tryon, R.C. Cluster analysis. Edwards Brothers. Ann. Arbor Mich. 1939, 122. [Google Scholar]
- Le, N.Q.K.; Do, D.T.; Chiu, F.Y.; Yapp, E.K.Y.; Yeh, H.Y.; Chen, C.Y. XGBoost Improves Classification of MGMT Promoter Methylation Status in IDH1 Wildtype Glioblastoma. J. Pers. Med. 2020, 10, 128. [Google Scholar] [CrossRef]
- Ho Thanh Lam, L.; Le, N.H.; Van Tuan, L.; Tran Ban, H.; Nguyen Khanh Hung, T.; Nguyen, N.T.K.; Huu Dang, L.; Le, N.Q.K. Machine Learning Model for Identifying Antioxidant Proteins Using Features Calculated from Primary Sequences. Biology 2020, 9, 325. [Google Scholar] [CrossRef]
- Johnson, S.C. Hierarchical clustering schemes. Psychometrika 1967, 32, 241–254. [Google Scholar] [CrossRef]
- Ward, J.H., Jr. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 1963, 58, 236–244. [Google Scholar] [CrossRef]
- Rao, M. Cluster analysis and mathematical programming. J. Am. Stat. Assoc. 1971, 66, 622–626. [Google Scholar] [CrossRef]
- Steinhaus, H. Sur la division des corp materiels en parties. Bull. Acad. Pol. Sci. 1956, 1, 801. [Google Scholar]
- Brusco, M.J.; Steinley, D.; Stevens, J.; Cradit, J.D. Affinity propagation: An exemplar-based tool for clustering in psychological research. Br. J. Math. Stat. Psychol. 2019, 72, 155–182. [Google Scholar] [CrossRef] [PubMed]
- Allen, D.N.; Goldstein, G. Cluster Analysis in Neuropsychological Research; Springer: Berlin/Heidelberg, Germany, 2013. [Google Scholar]
- Morris, R.; Blashfield, R.; Satz, P. Neuropsychology and cluster analysis: Potentials and problems. J. Clin. Exp. Neuropsychol. 1981, 3, 79–99. [Google Scholar] [CrossRef]
- Cotrena, C.; Damiani Branco, L.; Ponsoni, A.; Milman Shansis, F.; Paz Fonseca, R. Neuropsychological Clustering in Bipolar and Major Depressive Disorder. J. Int. Neuropsychol. Soc. 2017, 23, 584–593. [Google Scholar] [CrossRef] [PubMed]
- Hermens, D.F.; Hodge, M.A.R.; Naismith, S.L.; Kaur, M.; Scott, E.; Hickie, I.B. Neuropsychological clustering highlights cognitive differences in young people presenting with depressive symptoms. J. Int. Neuropsychol. Soc. 2011, 17, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Burdick, K.; Russo, M.; Frangou, S.; Mahon, K.; Braga, R.; Shanahan, M.; Malhotra, A. Empirical evidence for discrete neurocognitive subgroups in bipolar disorder: Clinical implications. Psychol. Med. 2014, 44, 3083. [Google Scholar] [CrossRef]
- Lewandowski, K.; Sperry, S.; Cohen, B.; Öngür, D. Cognitive variability in psychotic disorders: A cross-diagnostic cluster analysis. Psychol. Med. 2014, 44, 3239. [Google Scholar] [CrossRef]
- Kim, S.Y.; Lim, T.S.; Lee, H.Y.; Moon, S.Y. Clustering mild cognitive impairment by mini-mental state examination. Neurol. Sci. 2014, 35, 1353–1358. [Google Scholar] [CrossRef] [PubMed]
- Phillips, M.L.; Stage, E.C., Jr.; Lane, K.A.; Gao, S.; Risacher, S.L.; Goukasian, N.; Saykin, A.J.; Carrillo, M.C.; Dickerson, B.C.; Rabinovici, G.D.; et al. Neurodegenerative Patterns of Cognitive Clusters of Early-Onset Alzheimer’s Disease Subjects: Evidence for Disease Heterogeneity. Dement. Geriatr. Cogn. Disord. 2019, 48, 131–142. [Google Scholar] [CrossRef] [PubMed]
- Alashwal, H.; El Halaby, M.; Crouse, J.J.; Abdalla, A.; Moustafa, A.A. The Application of Unsupervised Clustering Methods to Alzheimer’s disease. Front. Comput. Neurosci. 2019, 13, 31. [Google Scholar] [CrossRef] [PubMed]
- Van Rooden, S.M.; Heiser, W.J.; Kok, J.N.; Verbaan, D.; Van Hilten, J.J.; Marinus, J. The identification of Parkinson’s disease subtypes using cluster analysis: A systematic review. Mov. Disord. 2010, 25, 969–978. [Google Scholar] [CrossRef]
- Mu, J.; Chaudhuri, K.R.; Bielza, C.; de Pedro-Cuesta, J.; Larrañaga, P.; Martinez-Martin, P. Parkinson’s Disease Subtypes Identified from Cluster Analysis of Motor and Non-motor Symptoms. Front. Aging Neurosci. 2017, 9. [Google Scholar] [CrossRef]
- Liu, P.; Feng, T.; Wang, Y.-j.; Zhang, X.; Chen, B. Clinical heterogeneity in patients with early-stage Parkinson’s disease: A cluster analysis. J. Zhejiang Univ. Sci. B 2011, 12, 694. [Google Scholar] [CrossRef]
- van Rooden, S.M.; Colas, F.; Martinez-Martin, P.; Visser, M.; Verbaan, D.; Marinus, J.; Chaudhuri, R.K.; Kok, J.N.; van Hilten, J.J. Clinical subtypes of Parkinson’s disease. Mov. Disord. 2011, 26, 51–58. [Google Scholar] [CrossRef]
- Erro, R.; Vitale, C.; Amboni, M.; Picillo, M.; Moccia, M.; Longo, K.; Santangelo, G.; De Rosa, A.; Allocca, R.; Giordano, F.; et al. The heterogeneity of early Parkinson’s disease: A cluster analysis on newly diagnosed untreated patients. PLoS ONE 2013, 8, e70244. [Google Scholar] [CrossRef]
- Ma, L.Y.; Chan, P.; Gu, Z.Q.; Li, F.F.; Feng, T. Heterogeneity among patients with Parkinson’s disease: Cluster analysis and genetic association. J. Neurol. Sci. 2015, 351, 41–45. [Google Scholar] [CrossRef]
- Pont-Sunyer, C.; Hotter, A.; Gaig, C.; Seppi, K.; Compta, Y.; Katzenschlager, R.; Mas, N.; Hofeneder, D.; Brücke, T.; Bayés, A. The Onset of Nonmotor Symptoms in Parkinson’s disease (The ONSET PD S tudy). Mov. Disord. 2015, 30, 229–237. [Google Scholar] [CrossRef]
- Jin, X.; Han, J. K-Means Clustering; Springer: Berlin/Heidelberg, Germany, 2010. [Google Scholar]
- Steinley, D.; Brusco, M.J. Evaluating mixture modeling for clustering: Recommendations and cautions. Psychol. Methods 2011, 16, 63. [Google Scholar] [CrossRef] [PubMed]
- Milligan, G.W.; Cooper, M.C. An examination of procedures for determining the number of clusters in a data set. Psychometrika 1985, 50, 159–179. [Google Scholar] [CrossRef]
- Frey, B.J.; Dueck, D. Clustering by passing messages between data points. Science 2007, 315, 972–976. [Google Scholar] [CrossRef]
- Mézard, M. Where are the exemplars? Science 2007, 315, 949–951. [Google Scholar] [CrossRef] [PubMed]
- Thavikulwat, P. Affinity Propagation: A Clustering Algorithm for Computer-Assisted Business Simulations and Experiential Exercises. Available online: http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.490.7628&rep=rep1&type=pdf (accessed on 3 February 2021).
- Hoglinger, G.U.; Respondek, G.; Stamelou, M.; Kurz, C.; Josephs, K.A.; Lang, A.E.; Mollenhauer, B.; Muller, U.; Nilsson, C.; Whitwell, J.L.; et al. Clinical diagnosis of progressive supranuclear palsy: The movement disorder society criteria. Mov. Disord. 2017, 32, 853–864. [Google Scholar] [CrossRef]
- Quattrone, A.; Morelli, M.; Nigro, S.; Quattrone, A.; Vescio, B.; Arabia, G.; Nicoletti, G.; Nistico, R.; Salsone, M.; Novellino, F.; et al. A new MR imaging index for differentiation of progressive supranuclear palsy-parkinsonism from Parkinson’s disease. Parkinsonism Relat. Disord. 2018, 54, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Barbagallo, G.; Morelli, M.; Quattrone, A.; Chiriaco, C.; Vaccaro, M.G.; Gulla, D.; Rocca, F.; Caracciolo, M.; Novellino, F.; Sarica, A.; et al. In vivo evidence for decreased scyllo-inositol levels in the supplementary motor area of patients with Progressive Supranuclear Palsy: A proton MR spectroscopy study. Parkinsonism Relat. Disord. 2019, 62, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Goldman, J.G.; Holden, S.K.; Litvan, I.; McKeith, I.; Stebbins, G.T.; Taylor, J.P. Evolution of diagnostic criteria and assessments for Parkinson’s disease mild cognitive impairment. Mov. Disord. 2018, 33, 503–510. [Google Scholar] [CrossRef]
- Postuma, R.B.; Berg, D.; Stern, M.; Poewe, W.; Olanow, C.W.; Oertel, W.; Obeso, J.; Marek, K.; Litvan, I.; Lang, A.E.; et al. MDS clinical diagnostic criteria for Parkinson’s disease. Mov. Disord. 2015, 30, 1591–1601. [Google Scholar] [CrossRef]
- Vaccaro, M.G.; Sarica, A.; Quattrone, A.; Chiriaco, C.; Salsone, M.; Morelli, M.; Quattrone, A. Neuropsychological assessment could distinguish among different clinical phenotypes of progressive supranuclear palsy: A Machine Learning approach. J. Neuropsychol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Carpinelli Mazzi, M.; Iavarone, A.; Russo, G.; Musella, C.; Milan, G.; D’Anna, F.; Garofalo, E.; Chieffi, S.; Sannino, M.; Illario, M.; et al. Mini-Mental State Examination: New normative values on subjects in Southern Italy. Aging Clin. Exp. Res. 2020, 32, 699–702. [Google Scholar] [CrossRef] [PubMed]
- Folstein, M.F.; Folstein, S.E.; McHugh, P.R. “Mini-mental state”. A practical method for grading the cognitive state of patients for the clinician. J. Psychiatr. Res. 1975, 12, 189–198. [Google Scholar] [CrossRef]
- Grigoletto, F.; Zappalà, G.; Anderson, D.W.; Lebowitz, B.D. Norms for the Mini-Mental State Examination in a healthy population. Neurology 1999, 53, 315. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Harris, C.R.; Millman, K.J.; van der Walt, S.J.; Gommers, R.; Virtanen, P.; Cournapeau, D.; Wieser, E.; Taylor, J.; Berg, S.; Smith, N.J. Array programming with NumPy. Nature 2020, 585, 357–362. [Google Scholar] [CrossRef] [PubMed]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- McKinney, W. Data structures for statistical computing in python. In Proceedings of the 9th Python in Science Conference, Austin, TX, USA, 28 June–3 July 2010; pp. 51–56. [Google Scholar]
- Seabold, S.; Perktold, J. Statsmodels: Econometric and statistical modeling with python. In Proceedings of the 9th Python in Science Conference, Austin, TX, USA, 28 June–3 July 2010; p. 61. [Google Scholar]
- Hutcheson, G.D. Ordinary least-squares regression. In L. Moutinho and GD Hutcheson, the SAGE Dictionary of Quantitative Management Research; Sage: Southend Oaks, CA, USA, 2011; pp. 224–228. [Google Scholar]
- Brusco, M.J.; Köhn, H.-F. Exemplar-based clustering via simulated annealing. Psychometrika 2009, 74, 457–475. [Google Scholar] [CrossRef]
- Rousseeuw, P.J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 1987, 20, 53–65. [Google Scholar] [CrossRef]
- Dueck, D. Affinity Propagation: Clustering Data by Passing Messages. Ph.D. Thesis, University of Toronto, Toronto, ON, Canada, September 2009. [Google Scholar]
- Brusco, M.J.; Shireman, E.; Steinley, D. A comparison of latent class, K-means, and K-median methods for clustering dichotomous data. Psychol. Methods 2017, 22, 563–580. [Google Scholar] [CrossRef] [PubMed]
- Starczewski, A. A new validity index for crisp clusters. Pattern Anal. Appl. 2017, 20, 687–700. [Google Scholar] [CrossRef]

| CTRL (44) | PD (49) | PSP (48) | p-Value | Post-Hoc | |
|---|---|---|---|---|---|
| Age | 62.6 ± 11.5 | 66.7 ± 9.38 | 70.1 ± 8.32 | 0.002 a | CTRL < PSP b |
| Female, n | 25 | 22 | 22 | N.S. c | |
| Education | 12.5 ± 4.78 | 9.27 ± 4.74 | 7.38 ± 5.05 | <0.001 a | CTRL > PD b, PSP b |
| Total MMSE | 29.2 ± 1.46 | 24.8 ± 5.08 | 20.8 ± 5.27 | <0.001 d | CTRL > PD e, PSP e; PD > PSP e |
| TO | 4.93 ± 0.45 (0.241 ± 0.57) | 4.49 ± 1.04 (0.100 ± 1.010) | 3.85 ± 1.38 (−0.321 ± 1.250) | 0.01 d | CTRL > PSP e |
| OS | 4.98 ± 0.15 (0.31 ± 0.567) | 4.29 ± 1.15 (−0.025 ± 0.992) | 3.81 ± 1.21 (−0.259 ± 0.999) | 0.003 d | CTRL > PSP e |
| Reg | 3.00 ± 0 (0.008 ± 0.084) | 2.98 ± 0.143 (0.042 ± 0.142) | 2.85 ± 0.50 (−0.050 ± 0.485) | N.S.d | N.A. |
| AC | 4.80 ± 0.60 (0.854 ± 0.846) | 3.14 ± 1.90 (−0.167 ± 1.590) | 2.25 ± 1.77 (−0.616 ± 1.690) | <0.001 d | CTRL > PD e, PSP e |
| Rec | 2.98 ± 0.15 (0.66 ± 0.39) | 1.96 ± 1.08 (−0.165 ± 0.970) | 1.52 ± 0.9 (−0.441 ± 1.010) | <0.001 d | CTRL > PD e, PSP e |
| N | 1.98 ± 0.15 (−0.02 ± 0.14) | 2.00 ± 0 (0.030 ± 0.058) | 1.94 ± 0.32 (−0.015 ± 0.303) | N.S. d | N.A. |
| SR | 1.00 ± 0 (0.10 ± 0.04) | 0.82 ± 0.39 (−0.055 ± 0.403) | 0.81 ± 0.39 (−0.038 ± 0.384) | 0.03 d | CTRL > PD e |
| P | 2.93 ± 0.45 (−0.02 ± 0.43) | 2.98 ± 0.143 (0.088 ± 0.163) | 2.77 ± 0.55 (−0.074 ± 0.540) | N.S.d | N.A. |
| W | 0.91 ± 0.30 (0.16 ± 0.30) | 0.67 ± 0.47 (0.065 ± 0.394) | 0.29 ± 0.46 (−0.210 ± 0.465) | <0.001 d | CTRL > PSP e; PD > PSP e |
| CE | 0.82 ± 0.40 (−0.003 ± 0.410) | 0.84 ± 0.37 (0.123 ± 0.344) | 0.52 ± 0.50 (−0.120 ± 0.454) | 0.01 d | PD > PSP e |
| D | 0.84 ± 0.37 (0.111 ± 0.270) | 0.65 ± 0.48 (0.083 ± 0.394) | 0.25 ± 0.44 (−0.185 ± 0.421) | <0.001 d | CTRL > PSP e; PD > PSP e |
| Median Preference | Minimum Preference | |||
|---|---|---|---|---|
| Group | #Clusters | Silhouette Index | #Clusters | Silhouette Index |
| CTRL | 8 | 0.302 | 1 | N.A. |
| PD | 7 | 0.375 | 2 | 0.675 |
| PSP | 6 | 0.387 | 2 | 0.677 |
| CTRL + PD + PSP | 16 | 0.237 | 4 | 0.601 |
| Cluster #1 (68) | Cluster #2 (30) | Cluster #3 (8) | Cluster #4 (34) | p-Value | Post-Hoc | |
|---|---|---|---|---|---|---|
| Age | 62.4 ± 10.1 | 71.3 ± 9.94 | 70.4 ± 7.93 | 69.8 ± 7.64 | <0.001 a | 1 < 2,4 b |
| Female, n | 33 | 14 | 5 | 17 | N.S. c | N.A. |
| Education | 12.1 ± 5.23 | 6.60 ± 3.23 | 5.5 ± 2.67 | 8.59 ± 4.99 | <0.001 a | 1 > 2,3,4 e |
| Total MMSE | 27.3 ± 3.93 | 27.7 ± 1.95 | 12.6 ± 2.92 | 20.1 ± 3.57 | <0.001 d | 1 > 3,4 e 2 > 1,3,4 e 4 > 3 e |
| TO | 4.68 ± 0.74 (0.026 ± 0.565) | 4.9 ± 0.55 (0.800 ± 0.550) | 1.25 ± 1.04 (−2.800 ± 0.842) | 4.18 ± 0.97 (−0.099 ± 0.888) | <0.001 d | 1 > 3 e 2 > 1,3,4 e 4 > 3 e |
| OS | 4.71 ± 0.69 (0.067 ± 0.472) | 4.83 ± 0.46 (0.856 ± 0.487) | 2 ± 0.76 (−1.85 ± 0.43) | 3.74 ± 1.24 (−0.454 ± 1.02) | <0.001 d | 1 > 3,4 e 2 > 1,3,4 e 4 > 3 e |
| Reg | 2.93 ± 0.39 (−0.056 ± 0.372) | 3 ± 0 (0.108 ± 0.040) | 2.75 ± 0.46 (−0.139 ± 0.472) | 2.97 ± 0.17 (0.049 ± 0.162) | 0.016 d | 2 > 1 e |
| AC | 4.21 ± 1.33 (0.345 ± 0.636) | 4.60 ± 0.72 (1.89 ± 0.619) | 0.50 ± 0.76 (−2.06 ± 1.11) | 1.21 ± 0.99 (−1.88 ± 0.767) | <0.001 d | 1 > 3,4 e 2 > 1,3,4 e |
| Rec | 2.59 ± 0.69 (0.3 ± 0.637) | 2.50 ± 0.73 (0.021 ± 0.18) | 1.13 ± 0.83 (−0.775 ± 0.993) | 1.09 ± 0.93 (−0.937 ± 0.889) | <0.001 d | 1 > 3,4 e 2 > 3,4 e |
| N | 2 ± 0 (0.007 ± 0.050) | 1.97 ± 0.183 (0.056 ± 0.302) | 1.75 ± 0.71 (−0.173 ± 0.675) | 1.97 ± 0.17 (0.008 ± 0.16) | N.S. d | N.A. |
| SR | 0.89 ± 0.31 (0.005 ± 0.300) | 0.9 ± 0.30 (0.072 ± 0.532) | 0.50 ± 0.53 (−0.337 ± 0.535) | 0.88 ± 0.33 (0.019 ± 0.331) | 0.025 d | 3 < 1,2,4 e |
| P | 2.94 ± 0.29 (−0.007 ± 0.268) | 2.90 ± 0.55 (0.201 ± 0.484) | 2.50 ± 0.76 (−0.312 ± 0.748) | 2.88 ± 0.41 (0.024 ± 0.416) | N.S.d | N.A. |
| W | 0.74 ± 0.41 (0.061 ± 0.316) | 0.67 ± 0.479 (0.218 ± 0.38) | 0.12 ± 0.35 (−0.333 ± 0.386) | 0.32 ± 0.47 (−0.22 ± 0.437) | <0.001 d | 1 > 3,4 e 2 > 3,4 e |
| CE | 0.76 ± 0.43 (−0.039 ± 0.376) | 0.83 ± 0.379 (0.178 ± 0.45) | 0 ± 0 (−0.605 ± 0.075) | 0.71 ± 0.46 (0.029 ± 0.418) | <0.001 d | 1 > 3 e 2 > 1,3 e 4 > 3 e |
| D | 0.79 ± 0.41 (0.070 ± 0.297) | 0.57 ± 0.50 (0.178 ± 0.450) | 0 ± 0 (−0.358 ± 0.212) | 0.26 ± 0.45 (−0.213 ± 0.402) | <0.001 d | 1 > 3,4 e 2 > 3,4 e |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sarica, A.; Vaccaro, M.G.; Quattrone, A.; Quattrone, A. A Novel Approach for Cognitive Clustering of Parkinsonisms through Affinity Propagation. Algorithms 2021, 14, 49. https://doi.org/10.3390/a14020049
Sarica A, Vaccaro MG, Quattrone A, Quattrone A. A Novel Approach for Cognitive Clustering of Parkinsonisms through Affinity Propagation. Algorithms. 2021; 14(2):49. https://doi.org/10.3390/a14020049
Chicago/Turabian StyleSarica, Alessia, Maria Grazia Vaccaro, Andrea Quattrone, and Aldo Quattrone. 2021. "A Novel Approach for Cognitive Clustering of Parkinsonisms through Affinity Propagation" Algorithms 14, no. 2: 49. https://doi.org/10.3390/a14020049
APA StyleSarica, A., Vaccaro, M. G., Quattrone, A., & Quattrone, A. (2021). A Novel Approach for Cognitive Clustering of Parkinsonisms through Affinity Propagation. Algorithms, 14(2), 49. https://doi.org/10.3390/a14020049







