Exploring the Dynamic Organization of Random and Evolved Boolean Networks
Abstract
1. Introduction
2. Materials and Methods
2.1. The Relevance Index
2.2. Random Boolean Networks
2.3. Genetic Algorithm
3. Results
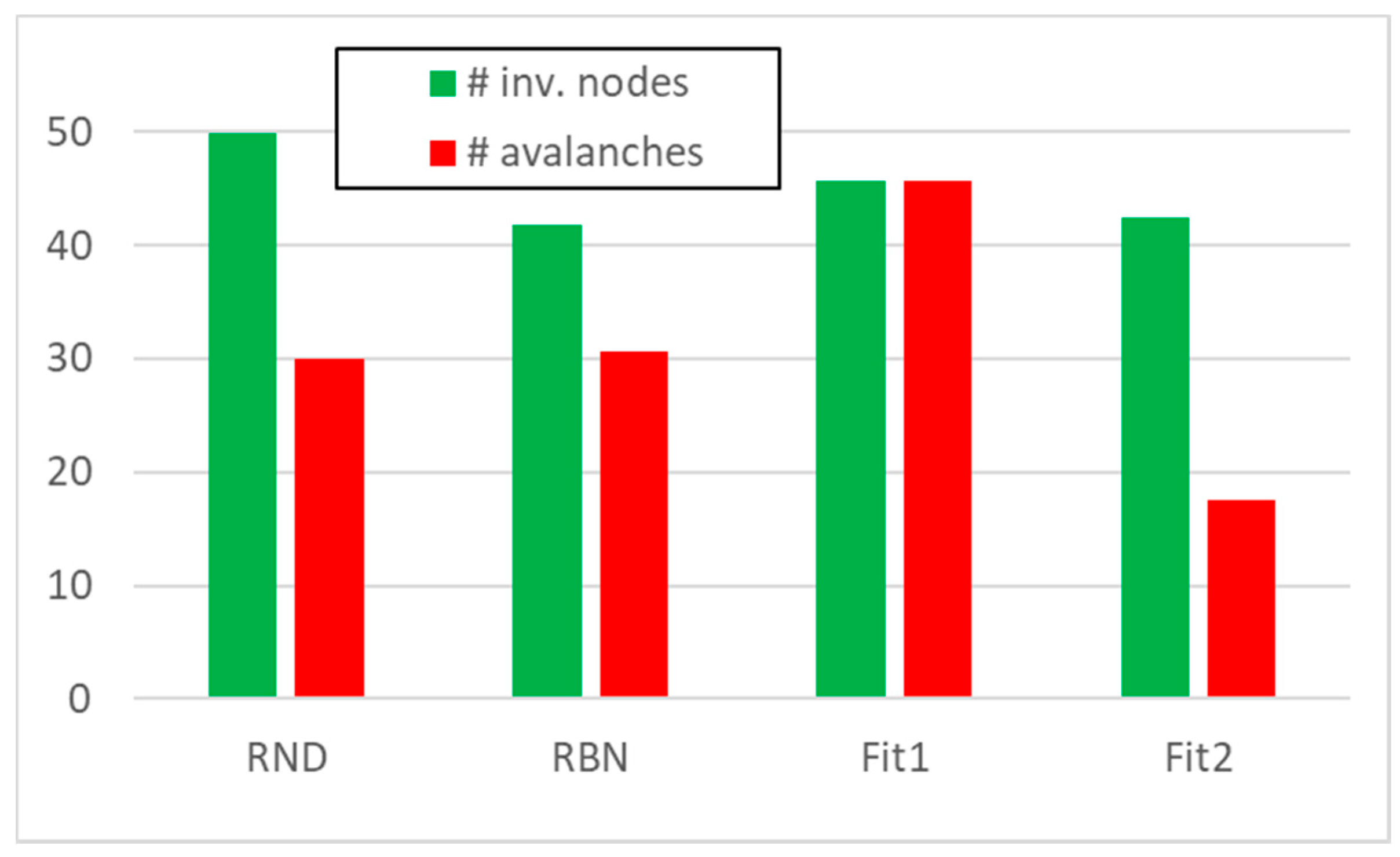
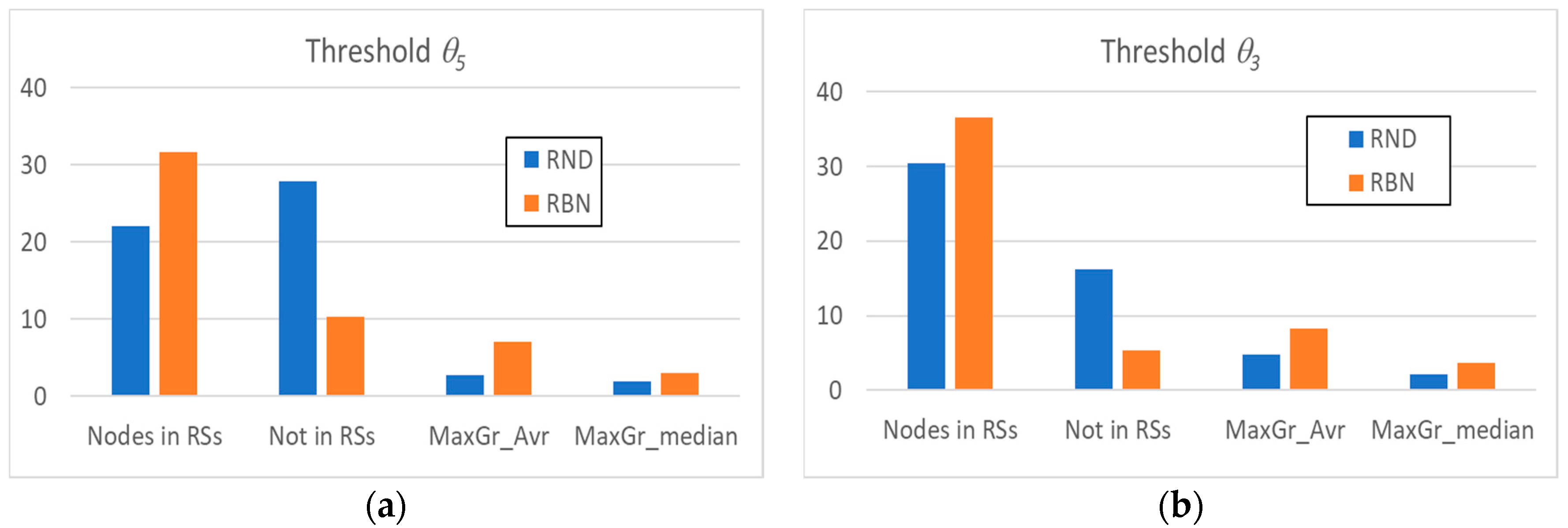
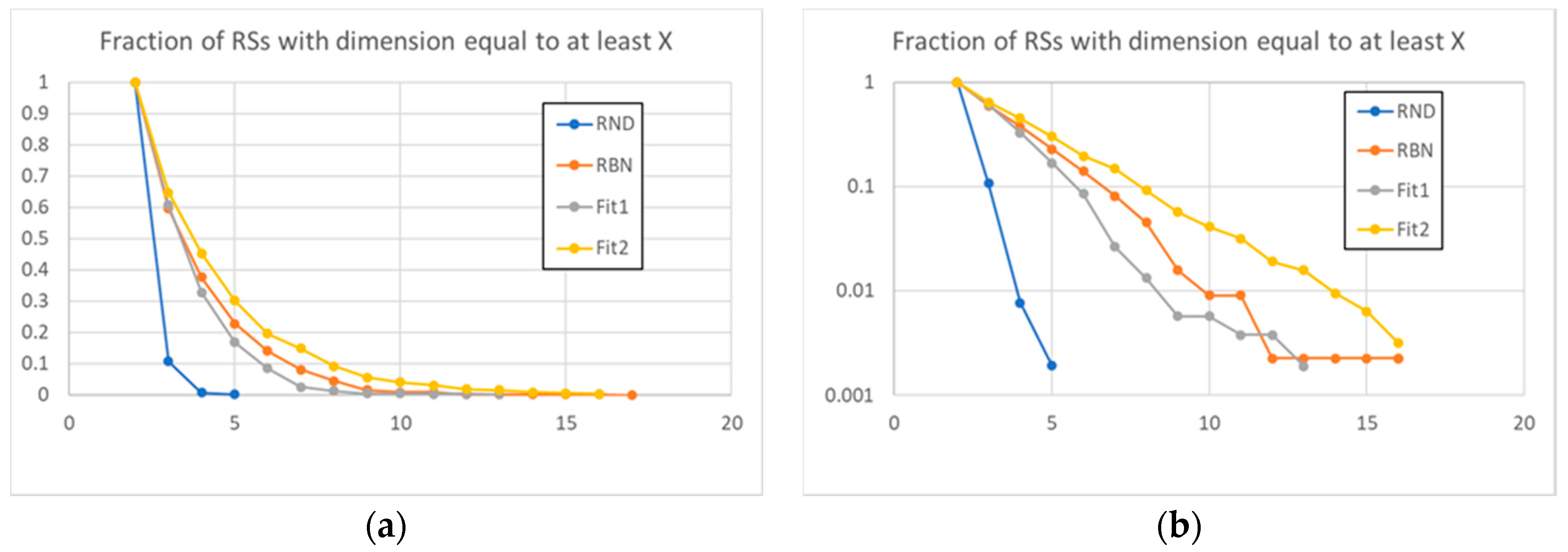
3.1. Random Avalanches and Avalanches in Random Boolean Networks
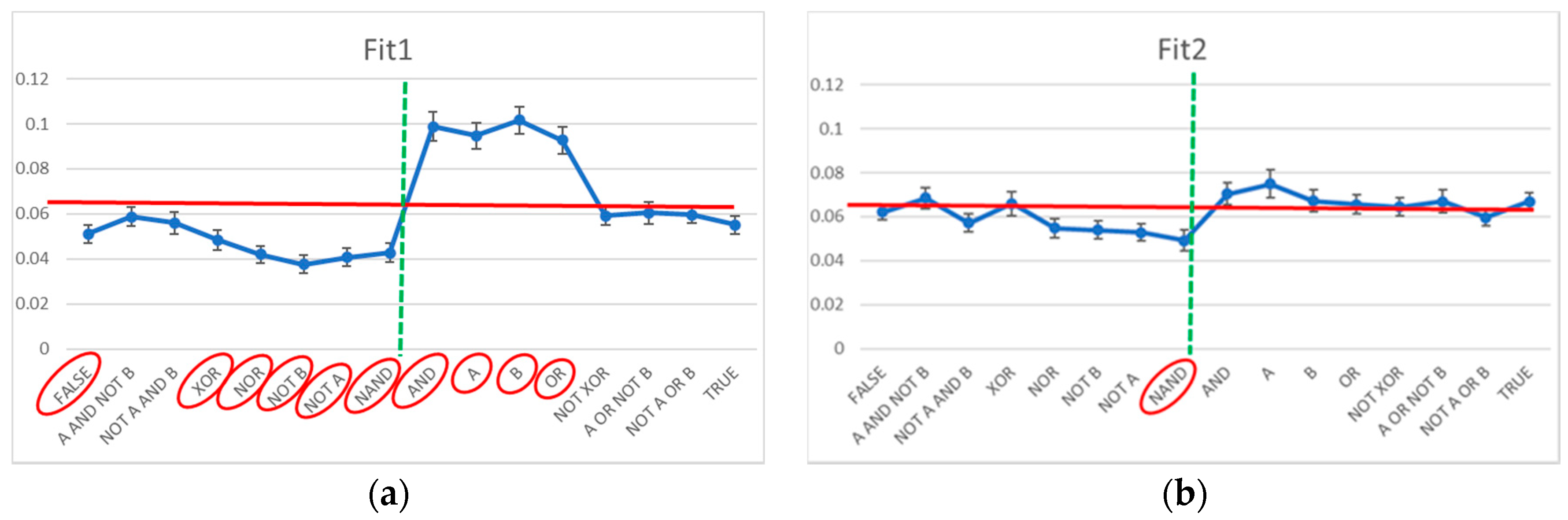
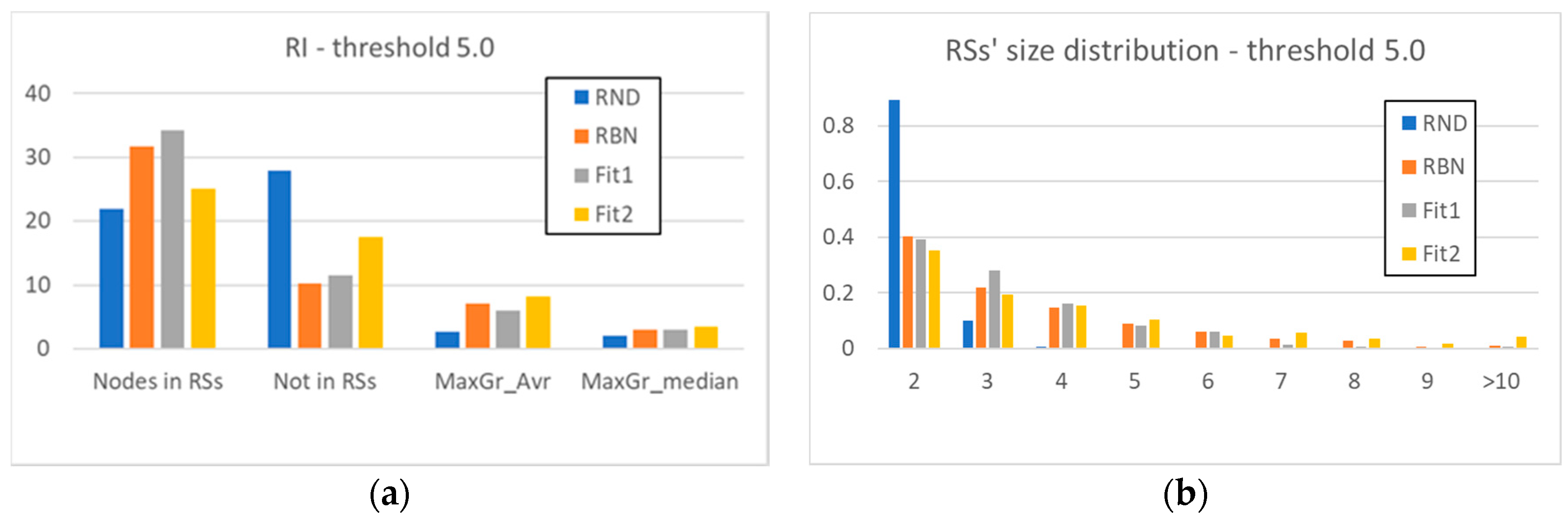
3.2. Static and Dynamic Characteristics in Evolved Systems
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Albert, R.; Barabási, A.L. Statistical mechanics of complex networks. Rev. Mod. Phys. 2002, 74, 47. [Google Scholar] [CrossRef]
- Albert, R.; Barabási, A.L. Emergence of Scaling in Random Networks. Science 1999, 286, 509–512. [Google Scholar] [CrossRef]
- Watts, D.; Strogatz, S. Collective dynamics of ‘small-world’ networks. Nature 1998, 393, 440–442. [Google Scholar] [CrossRef] [PubMed]
- Barrát, A.; Barthélemy, M.; Vespignani, A. Dynamical Processes on Complex Networks; Cambridge University Press: Cambridge, UK, 2008; ISBN 978-0-521-87950-7. [Google Scholar]
- Buldyrev, S.; Parshani, R.; Paul, G.; Stanley, H.E.; Havlin, S. Catastrophic cascade of failures in interdependent networks. Nature 2010, 464, 1025–1028. [Google Scholar] [CrossRef]
- Kitsak, M.; Gallos, L.K.; Havlin, S.; Liljeros, F.; Muchnik, L.; Stanley, H.E.; Makse, H.A. Identification of influential spreaders in complex networks. Nat. Phys. 2010, 6, 888–893. [Google Scholar] [CrossRef]
- Newman, M. Networks, 2nd ed.; Oxford University Press: Oxford, UK, 2018; ISBN 9780198805090. [Google Scholar]
- Pastor-Satorras, R.; Vespignani, A. Epidemic Spreading in Scale-Free Networks. Phys. Rev. Lett. 2001, 86, 3200. [Google Scholar] [CrossRef]
- Vespignani, A. The fragility of interdependency. Nature 2010, 464, 984–985. [Google Scholar] [CrossRef] [PubMed]
- Villani, M.; Filisetti, A.; Benedettini, S.; Roli, A.; Lane, D.; Serra, R. The detection of intermediate level emergent structures and patterns. In Proceeding of the ECAL 2013, the 12th European Conference on Artificial Life, Sicily, Italy, 2–6 September 2013; Liò, P., Miglino, O., Nicosia, G., Nolfi, S., Pavone, M., Eds.; MIT Press: Cambridge, MA, USA, 2013; pp. 372–378. ISBN 9780262317092. [Google Scholar]
- Villani, M.; Roli, A.; Filisetti, A.; Fiorucci, M.; Serra, R.; Poli, I. The Search for Candidate Relevant Subsets of Variables in Complex Systems. Artif. Life 2015, 21, 412–431. [Google Scholar] [CrossRef]
- Villani, M.; Sani, L.; Pecori, R.; Amoretti, M.; Roli, A.; Mordonini, M.; Serra, R.; Cagnoni, S. An Iterative Information-Theoretic Approach to the Detection of Structures in Complex Systems. Complexity 2018. [Google Scholar] [CrossRef]
- Sani, L.; Pecori, R.; Mordonini, M.; Cagnoni, S. From Complex System Analysis to Pattern Recognition: Experimental Assessment of an Unsupervised Feature Extraction Method Based on the Relevance Index Metrics. Computation 2019, 7, 39. [Google Scholar] [CrossRef]
- Villani, M.; Sani, L.; Amoretti, M.; Vicari, E.; Pecori, R.; Mordonini, M.; Cagnoni, S.; Serra, R. A Relevance Index Method to Infer Global Properties of Biological Networks. In Artificial Life and Evolutionary Computation. WIVACE 2017; Pelillo, M., Poli, I., Roli, A., Serra, R., Slanzi, D., Villani, M., Eds.; Springer: Berlin/Heidelberg, Germany, 2018; Volume 830, pp. 129–141. ISBN 978-3-030-21733-4. [Google Scholar]
- Sani, L.; Lombardo, G.; Pecori, R.; Fornacciari, P.; Mordonini, M.; Cagnoni, S. Social Relevance Index for Studying Communities in a Facebook Group of Patients. In Applications of Evolutionary Computation. EvoApplications 2018; Sim, K., Kaufmann, P., Eds.; Springer: Cham, Switzerland, 2018; Volume 10784, pp. 125–140. [Google Scholar] [CrossRef]
- Sani, L.; D’Addese, G.; Graudenzi, A.; Villani, M. The Detection of Dynamical Organization in Cancer Evolution Models. In Artificial Life and Evolutionary Computation. WIVACE 2019; Cicirelli, F., Guerrieri, A., Pizzuti, C., Socievole, A., Spezzano, G., Vinci, A., Eds.; Springer: Cham, Switzerland, 2020; Volume 1200, pp. 49–61. [Google Scholar] [CrossRef]
- Bastolla, U.; Parisi, G. The modular structure of Kauffman networks. Phys. D 1998, 115, 219–233. [Google Scholar] [CrossRef]
- Bastolla, U.; Parisi, G. Relevant elements, magnetization and dynamical properties in Kauffman networks: A numerical study. Phys. D 1998, 115, 203–218. [Google Scholar] [CrossRef]
- Drossel, B. Random Boolean networks. In Reviews of Nonlinear Dynamics and Complexity; Schuster, H.G., Ed.; Wiley-VCH: Weinheim, Germany, 2008; Chapter 1; pp. 69–110. ISBN 9783527407293. [Google Scholar]
- Aldana, M.; Coppersmith, S.; Kadanoff, L.P. Boolean dynamics with random couplings. In Perspectives and Problems in Nonlinear Science; Kaplan, E., Marsden, J., Sreenivasan, K.R., Eds.; Springer: New York, NY, USA, 2003; pp. 23–89. ISBN 9781468495669. [Google Scholar]
- Balleza, E.; Alvarez-Buylla, E.R.; Chaos, A.; Kauffman, S.A.; Shmulevich, I.; Aldana, M. Critical dynamics in genetic regulatory networks: Examples from four kingdoms. PLoS ONE 2008, 3, e2456. [Google Scholar] [CrossRef] [PubMed]
- Hidalgo, J.; Grilli, J.; Suweis, S.; Muñoz, M.A.; Banavarc, J.R.; Amos Maritan, A. Information-based fitness and the emergence of criticality in living systems. Proc. Natl. Acad. Sci. USA 2014, 111. [Google Scholar] [CrossRef]
- Kauffman, S.A. The Origins of Order; Oxford University Press: Oxford, UK, 1993; ISBN 9780195058116. [Google Scholar]
- Kauffman, S.A. At Home in the Universe; Oxford University Press: Oxford, UK, 1995; ISBN 0195111303. [Google Scholar]
- Langton, C.G. Computation at the edge of chaos: Phase transitions and emergent computation. Phys. D 1990, 42, 12–37. [Google Scholar] [CrossRef]
- Langton, C.G. Life at the edge of chaos. In Artificial Life II; Langton, C.G., Taylor, C., Farmer, J.D., Rasmussen, S., Eds.; Addison-Wesley: Redwood City, CA, USA, 1992; pp. 41–91. [Google Scholar]
- Muñoz, M.A. Criticality and dynamical scaling in living systems. Rev. Mod. Phys. 2018, 90, 031001. [Google Scholar] [CrossRef]
- Nykter, M.; Price, N.D.; Aldana, M.; Ramsey, S.A.; Kauffman, S.A.; Hood, L.E.; Yli-Harja, O.; Shmulevich, I. Gene expression dynamics in the macrophage exhibit criticality. Proc. Natl. Acad. Sci. USA 2008, 105, 1897–1900. [Google Scholar] [CrossRef]
- Packard, N.H. Adaptation toward the edge of chaos. In Dynamic Patterns in Complex Systems; World Scientific: Singapore, 1988; pp. 293–301. [Google Scholar] [CrossRef]
- Shmulevich, I.; Kauffman, S.A.; Aldana, M. Eukaryotic cells are dynamically ordered or critical but not chaotic. Proc. Natl. Acad. Sci. USA 2005, 102, 13439–13444. [Google Scholar] [CrossRef]
- Torres-Sosa, C.; Huang, S.; Aldana, M. Criticality is an emergent property of genetic networks that exhibit evolvability. PLoS Comput. Biol. 2012, 8, e1002669. [Google Scholar] [CrossRef]
- Beggs, J.M.; Timme, N. Being critical of criticality in the brain. Front. Physiol. 2012, 3, 163. [Google Scholar] [CrossRef]
- Roli, A.; Villani, M.; Filisetti, A.; Serra, R. Dynamical criticality: Overview and open questions. J. Syst. Sci. Complex 2018, 31, 647–663. [Google Scholar] [CrossRef]
- Villani, M.; Magrì, S.; Roli, A.; Serra, R. Evolving Always-Critical Networks. Life 2020, 10, 22. [Google Scholar] [CrossRef] [PubMed]
- Tononi, G.; Sporns, O.; Edelman, G.M. A measure for brain complexity: Relating functional segregation and integration in the nervous system. Proc. Natl. Acad. Sci. USA 1994, 91, 5033–5037. [Google Scholar] [CrossRef] [PubMed]
- Tononi, G.; McIntosh, A.; Russel, D.; Edelman, G. Functional clustering: Identifying strongly interactive brain regions in neuroimaging data. Neuroimage 1998, 7, 133–149. [Google Scholar] [CrossRef]
- Silvestri, G.; Sani, L.; Amoretti, M.; Pecori, R.; Vicari, E.; Mordonini, M.; Cagnoni, S. Searching Relevant Variable Subsets in Complex Systems Using K-Means PSO. In Artificial Life and Evolutionary Computation. WIVACE 2017; Pelillo, M., Poli, I., Roli, A., Serra, R., Slanzi, D., Villani, M., Eds.; Springer: Cham, Switzerland, 2018. [Google Scholar] [CrossRef]
- Sani, L.; Pecori, R.; Vicari, E.; Amoretti, M.; Mordonini, M.; Cagnoni, S. Can the Relevance Index be Used to Evolve Relevant Feature Sets? In Applications of Evolutionary Computation. EvoApplications 2018; Sim, K., Kaufmann, P., Eds.; Springer: Cham, Switzerland, 2018. [Google Scholar] [CrossRef]
- Kauffman, S.A. Metabolic stability and epigenesis in randomly constructed genetic nets. J. Theor. Biol. 1969, 22, 437–467. [Google Scholar] [CrossRef]
- Serra, R.; Villani, M.; Graudenzi, A.; Kauffman, S.A. Why a simple model of genetic regulatory networks describes the distribution of avalanches in gene expression data. J. Theor. Biol. 2007, 249, 449–460. [Google Scholar] [CrossRef] [PubMed]
- Wuensche, A. Genomic regulation modeled as a network with basins of attraction. Pac. Symp. Biocomput. 1998, 3, 89–102. [Google Scholar]
- Fernández, P.; Solé, R.V. The Role of Computation in Complex Regulatory Networks. In Power Laws, Scale-Free Networks and Genome Biology; Koonin, E.V., Wolf, Y.I., Karev, G.P., Eds.; Springer: Boston, MA, USA, 2006; pp. 206–225. ISBN 9780387339160. [Google Scholar] [CrossRef]
- Gershenson, C. Introduction to Random Boolean Networks. arXiv 2004, arXiv:nlin/0408006. [Google Scholar]
- Glass, L.; Kauffman, S.A. The logical analysis of continuous, non-linear biochemical control networks. J. Theor. Biol. 1973, 39, 103–129. [Google Scholar] [CrossRef]
- Shmulevich, I.; Dougherty, E.R.; Kim, S.; Zhang, W. Probabilistic Boolean networks: A rule-based uncertainty model for gene regulatory networks. Bioinformatics 2002, 18, 261–274. [Google Scholar] [CrossRef]
- Zañudo, J.G.T.; Aldana, M.; Martínez-Mekler, G. Boolean Threshold Networks: Virtues and Limitations for Biological Modeling. In Information Processing and Biological Systems; Niiranen, S., Ribeiro, A., Eds.; Springer: Berlin/Heidelberg, Germany, 2011. [Google Scholar] [CrossRef]
- Derrida, B.; Pomeau, Y. Random networks of automata: A simple annealed approximation. Europhys. Lett. 1986, 1, 45–49. [Google Scholar] [CrossRef]
- Derrida, B.; Flyvbjerg, H. The random map model: A disordered model with deterministic dynamics. J. Phys. 1987, 48, 971–978. [Google Scholar] [CrossRef]
- Holland, J.H. Adaptation in Natural and Artificial Systems; University of Michigan Press: Ann Arbor, MI, USA; MIT Press: Cambridge, MA, USA, 1975; ISBN 9780472084609. [Google Scholar]
- Sivanandam, S.N.; Deepa, S.N. Introduction to Genetic Algorithms; Springer: Berlin/Heidelberg, Germany, 2008; ISBN 978-3-540-73189-4. [Google Scholar]
- Bonabeau, E.; Dorigo, M.; Theraulaz, G. Swarm Intelligence: From Natural to Artificial Systems; Oxford University Press, Santa Fe Institute Studies in the Sciences of Complexity: New York, NY, USA, 1999; ISBN 0-19-513159-2. [Google Scholar]
- Darwish, A. Bio-inspired computing: Algorithms review, deep analysis, and the scope of applications. Future Comput. Inform. J. 2018, 3, 231–246. [Google Scholar] [CrossRef]
- Dorigo, M.; Maniezzo, V.; Colorni, A. Ant system: Optimization by a colony of cooperating agents. IEEE Trans. Syst. Man Cybern. Part B 1996, 26, 29–41. [Google Scholar] [CrossRef]
- Reynolds, C.W. Flocks, Herds, and Schools: A Distributed Behavioural Model Computer Graphics. In Proceedings of the 14th Annual Conference on Computer Graphics and Interactive Techniques, Anaheim, CA, USA, 27–31 July 1987; Volume 21, pp. 25–34. [Google Scholar] [CrossRef]
- Dulebenets, M.A.; Moses, R.; Ozguven, E.E.; Vanli, A. Minimizing carbon dioxide emissions due to container handling at marine container terminals via hybrid evolutionary algorithms. IEEE Access 2017, 5, 8131–8147. [Google Scholar] [CrossRef]
- Dulebenets, M.A.; Kavoosi, M.; Abioye, O.; Pasha, J. A self-adaptive evolutionary algorithm for the berth scheduling problem: Towards efficient parameter control. Algorithms 2018, 11, 100. [Google Scholar] [CrossRef]
- Anandakumar, H.; Umamaheswari, K. A bio-inspired swarm intelligence technique for social aware cognitive radio handovers. Comput. Electr. Eng. 2018, 71, 925–937. [Google Scholar] [CrossRef]
- Slowik, A.; Kwasnicka, H. Nature inspired methods and their industry applications—Swarm intelligence algorithms. IEEE Trans. Ind. Inform. 2018, 14, 1004–1015. [Google Scholar] [CrossRef]
- Zhao, X.; Wang, C.; Su, J.; Wang, J. Research and application based on the swarm intelligence algorithm and artificial intelligence for wind farm decision system. Renew. Energy 2019, 134, 681–697. [Google Scholar] [CrossRef]
- Yang, X.S. Flower Pollination Algorithm for Global Optimization. In Unconventional Computation and Natural Computation. UCNC 2012; Durand-Lose, J., Jonoska, N., Eds.; Springer: Berlin/Heidelberg, Germany, 2012. [Google Scholar] [CrossRef]
- Uymaz, S.A.; Tezel, G.; Yel, E. Artificial algae algorithm (AAA) for nonlinear global optimization. Appl. Soft Comput. 2015, 31, 153–171. [Google Scholar] [CrossRef]
- Yang, X.S. A New Metaheuristic Bat-Inspired Algorithm. In Nature Inspired Cooperative Strategies for Optimization (NICSO 2010). Studies in Computational Intelligence; González, J.R., Pelta, D.A., Cruz, C., Terrazas, G., Krasnogor, N., Eds.; Springer: Berlin/Heidelberg, Germany, 2010. [Google Scholar] [CrossRef]
- Pradhan, P.M.; Panda, G. Solving multiobjective problems using cat swarm optimization. Expert Syst. Appl. 2012, 39, 2956–2964. [Google Scholar] [CrossRef]
- Mirjalili, S.; Lewis, A. The whale optimization algorithm. Adv. Eng. Soft. 2016, 95, 51–67. [Google Scholar] [CrossRef]
- Deb, S.; Fong, S.; Tian, Z. Elephant search algorithm for optimization problems. In Proceedings of the Tenth International Conference on Digital Information Management (ICDIM 2015), Jeju Island, Korea, 21–23 October 2015; pp. 249–255. [Google Scholar] [CrossRef]
- Mirjalili, S.; Mirjalili, S.M.; Lewis, A. Grey wolf optimizer. Adv. Eng. Softw. 2014, 69, 46–61. [Google Scholar] [CrossRef]
- Meng, X.; Liu, Y.; Gao, X.; Zhang, H. A New Bioinspired Algorithm: Chicken Swarm Optimization; Springer: Cham, Switzerland, 2014; Part I; pp. 86–94. [Google Scholar] [CrossRef]
- Yang, X.-S.; Deb, S. Cuckoo search via Lévy flights. In Proceedings of the World Congress on Nature & Biologically Inspired Computing (NaBIC 2009), Coimbatore, India, 9–11 December 2009; pp. 210–214, ISBN 978-1-4244-5053-4. [Google Scholar]
- Elbeltagi, E.; Hegazy, T.; Dnald, G.D. Comparison among five evolutionary-based optimization algorithms. Adv. Eng. Inform. 2005, 19, 43–53. [Google Scholar] [CrossRef]
- Hughes, T.R.; Marton, M.J.; Jones, A.R.; Roberts, C.J.; Stoughton, R.; Armour, C.D.; A Bennett, H.; Coffey, E.; Dai, H.; He, Y.D.; et al. Functional discovery via a compendium of expression profiles. Cell 2000, 102, 109–126. [Google Scholar] [CrossRef]
- Corder, G.W.; Foreman, D.I. Nonparametric Statistics: A Step-by-Step Approach; Wiley: New York, NY, USA, 2014; ISBN 978-1118840313. [Google Scholar]
- Beni, G.; Wang, J. Swarm Intelligence in Cellular Robotic Systems. In Robots and Biological Systems: Towards a New Bionics? NATO ASI Series (Series F: Computer and Systems Sciences); Dario, P., Sandini, G., Aebischer, P., Eds.; Springer: Berlin/Heidelberg, Germany, 1993. [Google Scholar] [CrossRef]
- Roli, A.; Manfroni, M.; Pinciroli, C.; Birattari, M. On the Design of Boolean Network Robots. In Applications of Evolutionary Computation. EvoApplications 2011; Di Chio, C., Ed.; Springer: Berlin/Heidelberg, Germany, 2011. [Google Scholar] [CrossRef]
| System | Parameter | Value |
|---|---|---|
| GA | number of generations | 100 |
| GA | number of RBNs in population | 100 |
| GA | crossover probability | 0.7 |
| GA | mutation probability per node | 0.02 |
| GA | number of best individuals directly transmitted to next generation (elitism) | 3 |
| BN | number of nodes per BN | 50 |
| BN | number of inputs per node | 2 |
| BN | average initial bias in initial population | 0.5 |
| BN | number of initial conditions per BN | 10,000 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
d’Addese, G.; Magrì, S.; Serra, R.; Villani, M. Exploring the Dynamic Organization of Random and Evolved Boolean Networks. Algorithms 2020, 13, 272. https://doi.org/10.3390/a13110272
d’Addese G, Magrì S, Serra R, Villani M. Exploring the Dynamic Organization of Random and Evolved Boolean Networks. Algorithms. 2020; 13(11):272. https://doi.org/10.3390/a13110272
Chicago/Turabian Styled’Addese, Gianluca, Salvatore Magrì, Roberto Serra, and Marco Villani. 2020. "Exploring the Dynamic Organization of Random and Evolved Boolean Networks" Algorithms 13, no. 11: 272. https://doi.org/10.3390/a13110272
APA Styled’Addese, G., Magrì, S., Serra, R., & Villani, M. (2020). Exploring the Dynamic Organization of Random and Evolved Boolean Networks. Algorithms, 13(11), 272. https://doi.org/10.3390/a13110272







