Scale Reduction Techniques for Computing Maximum Induced Bicliques
Abstract
1. Introduction
2. Integer Programming Formulations
3. Exact Algorithm Based on the Proposed Scale Reduction
- Find a lower bound. Use a heuristic algorithm to obtain a lower bound on the optimal solution.
- Apply scale reduction. Given a heuristic solution, recursively identify and remove vertices that cannot be included in a globally optimal solution, until no further reduction is possible.
- Solve. Apply a standard exact algorithm to find a globally optimal solution in the residual graph.
3.1. Finding a Lower Bound
| Algorithm 1 Greedy induced star construction heuristic. |
|
3.2. Scale Reduction Techniques
3.2.1. Scale Reduction for the MB Problem
- 1.
- 2.
- for all k.
| Algorithm 2 Scale reduction algorithm. |
|
3.2.2. Scale Reduction for the MEB Problem
4. Computational Experiments
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| IP | integer programming |
| MIP | mixed integer programming |
| MB | maximum biclique |
| MEB | maximum edge biclique |
| RDS | Russian Doll Search |
References
- Gagneur, J.; Krause, R.; Bouwmeester, T.; Casari, G. Modular decomposition of protein-protein interaction networks. Genome Biol. 2004, 5, R57. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.; Mason, S.; Barabási, A.; Oltvai, Z.N. Lethality and certainty in protein networks. Nature 2001, 411, 41–42. [Google Scholar] [CrossRef] [PubMed]
- Spirin, V.; Mirny, L.A. Protein complexes and functional modules in molecular networks. Proc. Natl. Acad. Sci. USA 2003, 100, 12123–12128. [Google Scholar] [CrossRef] [PubMed]
- Madeira, S.C.; Oliveira, A.L. Biclustering Algorithms for Biological Data Analysis: A Survey. IEEE/ACM Trans. Comput. Biol. Bioinform. 2004, 1, 24–45. [Google Scholar] [CrossRef] [PubMed]
- Nussbaum, D.; Pu, S.; Sack, J.; Uno, T.; Zarrabi-Zadeh, H. Finding maximum edge bicliques in convex bipartite graphs. Algorithmica 2012, 64, 311–325. [Google Scholar] [CrossRef]
- Faloutsos, M.; Faloutsos, P.; Faloutsos, C. On power-law relationships of the Internet topology. In Proceedings of the ACM-SIGCOMM Conference on Applications, Technologies, Architectures, and Protocols for Computer Communication, Cambridge, MA, USA, 30 August–3 September 1999; pp. 251–262. [Google Scholar]
- Pastor-Satorras, R.; Vazquez, A.; Vespignani, A. Dynamical and correlation properties of the Internet. Phys. Rev. Lett. 2001, 87, 258701. [Google Scholar] [CrossRef] [PubMed]
- Acuña, V.; Ferreira, C.; Freire, A.; Moreno, E. Solving the maximum edge biclique packing problem on unbalanced bipartite graphs. Discrete Appl. Math. 2014, 164, 2–12. [Google Scholar] [CrossRef]
- Amilhastre, J.; Vilarem, M.; Janssen, P. Complexity of minimum biclique cover and minimum biclique decomposition for bipartite domino-free graphs. Discrete Appl. Math. 1998, 86, 125–144. [Google Scholar] [CrossRef]
- Cheng, Y.; Church, G. Biclustering of expression data. In Proceedings of the 8th International Conference on Intelligent Systems for Molecular Biology, La Jolla, CA, USA, 19–23 August 2000; pp. 93–100. [Google Scholar]
- Dawande, M.; Keskinocak, P.; Swaminathan, J.; Tayur, S. On Bipartite and Multipartite Clique Problems. J. Algorithms 2001, 41, 388–403. [Google Scholar] [CrossRef]
- Kumar, R.; Raghavan, P.; Rajagopalan, S.; Tomkins, A. Trawling the Web for Emerging Cyber-Communities. In Proceedings of the 8th international conference on World Wide Web, Toronto, ON, Canada, 11–14 May 1999; pp. 1481–1493. [Google Scholar]
- Liu, G.; Sim, K.; Li, J. Efficient Mining of Large Maximal Bicliques. In Proceedings of the International Conference on Data Warehousing and Knowledge Discovery, Krakow, Poland, 4–8 September 2006; Springer: Berlin/Heidelberg, Germany, 2006; Volume 4081, pp. 437–448. [Google Scholar]
- Sanderson, M.J.; Driskell, A.C.; Ree, R.H.; Eulenstein, O.; Langley, S. Obtaining maximal concatenated phylogenetic data sets from large sequence databases. Mol. Biol. Evol. 2003, 20, 1036–1042. [Google Scholar] [CrossRef] [PubMed]
- Alexe, G.; Alexe, S.; Crama, Y.; Foldes, S.; Hammer, P.L.; Simeone, B. Consensus algorithms for the generation of all maximal bicliques. Discrete Appl. Math. 2004, 145, 11–21. [Google Scholar] [CrossRef]
- Gély, A.; Nourine, L.; Sadi, B. Enumeration aspects of maximal cliques and bicliques. Discrete Appl. Math. 2009, 157, 1447–1459. [Google Scholar] [CrossRef]
- Mukherjee, A.P.; Tirthapura, S. Enumerating maximal bicliques from a large graph using MapReduce. In Proceedings of the 2014 IEEE International Congress on Big Data, Anchorage, AK, USA, 27 June–2 July 2014; pp. 707–716. [Google Scholar]
- Eppstein, D. Arboricity and bipartite subgraph listing algorithms. Inf. Process. Lett. 1994, 51, 207–211. [Google Scholar] [CrossRef]
- McCreesh, C.; Prosser, P. An exact branch and bound algorithm with symmetry breaking for the maximum balanced induced biclique problem. In Proceedings of the Integration of AI and OR Techniques in Constraint Programming: 11th International Conference, CPAIOR 2014, Cork, Ireland, 19–23 May 2014; Simonis, H., Ed.; Springer International Publishing: Cham, Switzerland, 2014; pp. 226–234. [Google Scholar]
- Binkele-Raible, D.; Fernau, H.; Gaspers, S.; Liedloff, M. Exact exponential-time algorithms for finding bicliques. Inf. Process. Lett. 2010, 111, 64–67. [Google Scholar] [CrossRef]
- Hochbaum, D.S. Approximating clique and biclique problems. J. Algorithms 1998, 29, 174–200. [Google Scholar] [CrossRef]
- Sim, K.; Li, J.; Gopalkrishnan, V.; Liu, G. Mining Maximal Quasi-Bicliques to Co-Cluster Stocks and Financial Ratios for Value Investment. In Proceedings of the Sixth International Conference on Data Mining (ICDM’06), Hong Kong, China, 18–22 December 2006; pp. 1059–1063. [Google Scholar]
- Maulik, U.; Mukhopadhyay, A.; Bhattacharyya, M.; Kaderali, L.; Brors, B.; Bandyopadhyay, S.; Eils, R. Mining Quasi-Bicliques from HIV-1-Human Protein Interaction Network: A Multiobjective Biclustering Approach. IEEE/ACM Trans. Comput. Biol. Bioinform. 2013, 10, 423–435. [Google Scholar] [CrossRef] [PubMed]
- Ganter, B.; Wille, R. Formale Begriffsanalyse-Mathematische Grundlagen; Springer: New York, NY, USA, 1996. [Google Scholar]
- Peeters, R. The maximum edge biclique problem is NP-complete. Discrete Appl. Math. 2003, 131, 651–654. [Google Scholar] [CrossRef]
- Ambühl, C.; Mastrolilli, M.; Svensson, O. Inapproximability results for maximum edge biclique, minimum linear arrangement, and sparsest cut. SIAM J. Comput. 2011, 40, 567–596. [Google Scholar] [CrossRef]
- Tan, J. Inapproximability of Maximum Weighted Edge Biclique and Its Applications. In Proceedings of the 5th International Conference on Theory and Application of Models of Computation, Xi’an, China, 25–29 April 2008; Springer: Berlin/Heidelberg, Germany, 2008; pp. 282–293. [Google Scholar]
- Yannakakis, M. Node and edge-deletion NP-complete problems. In Proceedings of the Tenth Annual ACM Symposium on Theory of Computing, San Diego, CA, USA, 1–3 May 1987; ACM: New York, NY, USA, 1978; pp. 253–264. [Google Scholar]
- Trukhanov, S.; Balasubramaniam, C.; Balasundaram, B.; Butenko, S. Algorithms for detecting optimal hereditary structures in graphs, with application to clique relaxations. Comput. Optim. Appl. 2013, 56, 113–130. [Google Scholar] [CrossRef]
- Verfaillie, G.; Lemaitre, M.; Schiex, T. Russian doll search for solving constraint optimization problems. In Proceedings of the National Conference on Artificial Intelligence, Portland, OR, USA, 4–8 August 1996; pp. 181–187. [Google Scholar]
- Abello, J.; Pardalos, P.M.; Resende, M.G.C. On maximum clique problems in very large graphs. In External Memory Algorithms; American Mathematical Society: Boston, MA, USA, 1999; pp. 119–130. [Google Scholar]
- Verma, A.; Buchanan, A.; Butenko, S. Solving the maximum clique and vertex coloring problems on very large sparse networks. INFORMS J. Comput. 2015, 27, 164–177. [Google Scholar] [CrossRef]
- Strash, D. On the power of simple reductions for the maximum independent set problem. In Proceedings of the International Computing and Combinatorics Conference (COCOON 2016), Ho Chi Minh City, Vietnam, 2–4 August 2016; Dinh, T.N., Thai, M.T., Eds.; Springer: Cham, Switzerland, 2016; pp. 345–356. [Google Scholar]
- Akiba, T.; Iwata, Y. Branch-and-reduce exponential/FPT algorithms in practice: A case study of vertex cover. Theor. Comput. Sci. 2016, 609, 211–225. [Google Scholar] [CrossRef]
- Cygan, M.; Fomin, F.; Kowalik, L.; Lokshtanov, D.; Marx, D.; Pilipczuk, M.; Pilipczuk, M.; Saurabh, S. Parameterized Algorithms; Springer: Cham, Switzerland, 2015. [Google Scholar]
- Downey, R.G.; Fellows, M.R. Parameterized Complexity; Springer: Cham, Switzerland, 1999. [Google Scholar]
- Lin, B. The parameterized complexity of k-biclique. In Proceedings of the Twenty-Sixth Annual ACM-SIAM Symposium on Discrete Algorithms, San Diego, CA, USA, 4–6 January 2015; SIAM: Philadelphia, PA, USA, 2015; pp. 605–615. [Google Scholar]
- Feng, Q.; Zhou, Z.; Wang, J. Parameterized Algorithms for Maximum Edge Biclique and Related Problems. In Proceedings of the Frontiers in Algorithmics: 10th International Workshop, FAW 2016, Qingdao, China, 30 June 30–2 July 2016; Zhu, D., Bereg, S., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 75–83. [Google Scholar]
- Östergård, P.R.J. A fast algorithm for the maximum clique problem. Discrete Appl. Math. 2002, 120, 197–207. [Google Scholar] [CrossRef]
- Tomita, E.; Sutani, Y.; Higashi, T.; Takahashi, S.; Wakatsuki, M. A Simple and Faster Branch-and-Bound Algorithm for Finding a Maximum Clique. In Proceedings of the WALCOM: Algorithms and Computation: 4th International Workshop, WALCOM 2010, Dhaka, Bangladesh, 10–12 February 2010; Rahman, M.S., Fujita, S., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 191–203. [Google Scholar]
- SNAP. Stanford Large Network Dataset Collection. 2012. Available online: http://snap.stanford.edu/data/ (accessed on 20 May 2017).
- DIMACS. Graph Partitioning and Graph Clustering: Tenth DIMACS Implementation Challenge. 2012. Available online: http://cc.gatech.edu/dimacs10/ (accessed on 20 May 2017).
- Pattillo, J.; Youssef, N.; Butenko, S. On clique relaxation models in network analysis. Eur. J. Oper. Res. 2013, 226, 9–18. [Google Scholar] [CrossRef]
- Gschwind, T.; Irnich, S.; Podlinski, I. Maximum weight relaxed cliques and Russian Doll Search revisited. Discrete Appl. Math. 2017. (in press) [CrossRef]

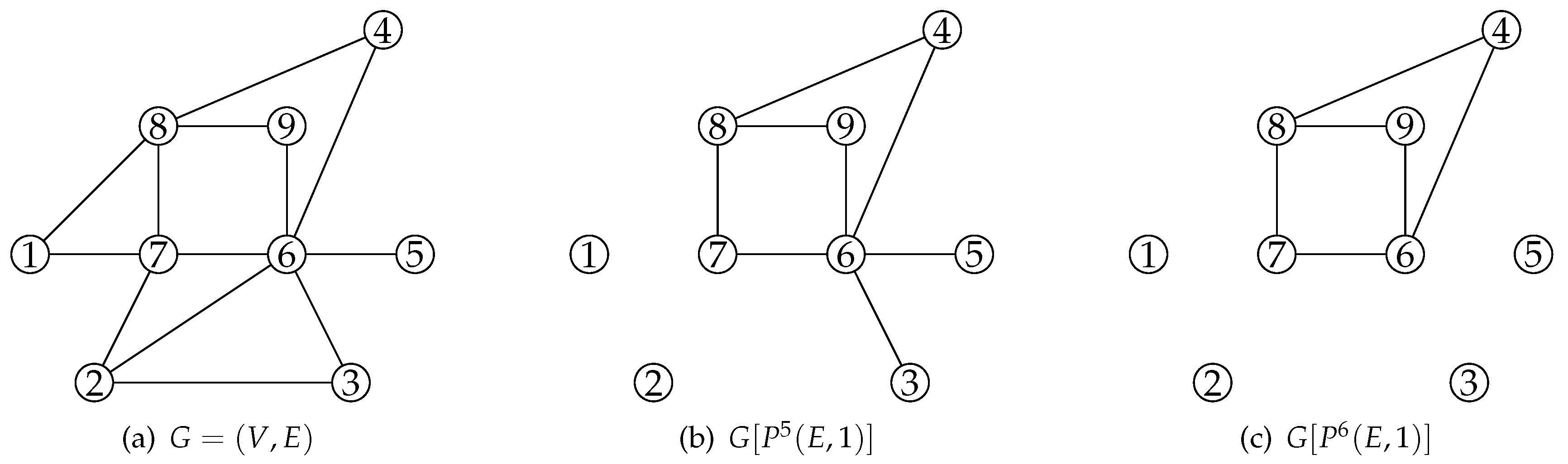
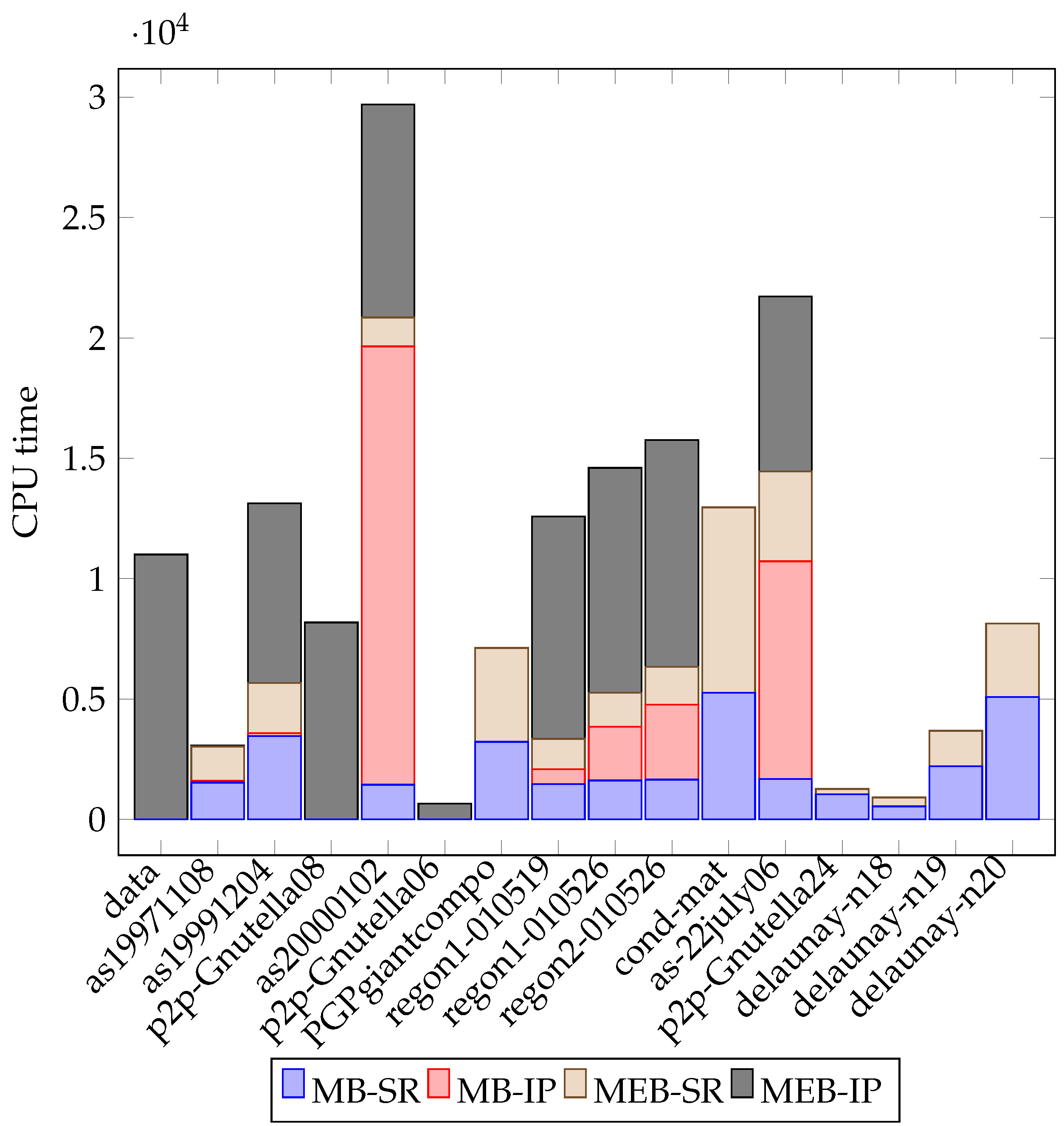
| Graph | LB | SR-CPU(s) | Opt. Obj. | CPU (s) | ||||
|---|---|---|---|---|---|---|---|---|
| jazz | 198 | 2742 | 18 | 85 | 704 | 0.44 | 20 | 0.67 |
| 1133 | 5451 | 34 | 35 | 35 | 0.61 | 34 | 0.64 | |
| netscience | 1589 | 2742 | 16 | 26 | 42 | 0.02 | 16 | 0.03 |
| add20 | 2395 | 7462 | 30 | 68 | 186 | 464.66 | 30 | 464.73 |
| data | 2851 | 15,093 | 8 | 41 | 99 | 1.09 | 8 | 1.36 |
| as19971108 | 3015 | 5347 | 540 | 583 | 740 | 1531.77 | 540 | 1602.59 |
| add32 | 4960 | 9462 | 16 | 66 | 108 | 0.17 | 16 | 0.42 |
| CA-GrQC | 5242 | 14,490 | 29 | 35 | 41 | 0.39 | 29 | 0.42 |
| as19991204 | 6296 | 12,830 | 1294 | 1407 | 2234 | 3461.69 | 1294 | 3582.89 |
| p2p-Gnutella08 | 6301 | 20,777 | 88 | 88 | 87 | 1.36 | 88 | 1.48 |
| as20000102 | 6474 | 13,233 | 1338 | 1454 | 2430 | 1440.82 | 1340 | 19,645.00 |
| p2p-Gnutella09 | 8114 | 26,013 | 98 | 98 | 97 | 1.09 | 98 | 1.47 |
| hep-th | 8361 | 15,751 | 22 | 81 | 151 | 0.40 | 23 | 0.66 |
| p2p-Gnutella06 | 8717 | 31,525 | 104 | 113 | 121 | 0.21 | 104 | 0.44 |
| p2p-Gnutella05 | 8846 | 31,839 | 87 | 88 | 88 | 0.76 | 87 | 1.11 |
| CA-HepTH | 9877 | 25,973 | 32 | 43 | 59 | 4.18 | 32 | 4.42 |
| PGPgiantcompo | 10,680 | 24,316 | 105 | 114 | 122 | 3223.57 | 105 | 3223.85 |
| p2p-Gnutella04 | 10,876 | 39,994 | 97 | 100 | 102 | 0.12 | 97 | 0.42 |
| oregon1-010519 | 11,051 | 22,724 | 2203 | 2384 | 4289 | 1466.45 | 2207 | 2085.01 |
| oregon1-010526 | 11,173 | 23,409 | 2199 | 2385 | 4308 | 1619.41 | 2203 | 3852.24 |
| oregon2-010526 | 11,460 | 16,365 | 2230 | 2428 | 4676 | 1652.65 | 2234 | 4762.25 |
| cond-mat | 16,726 | 47,594 | 41 | 55 | 72 | 5257.99 | 41 | 5258.05 |
| p2p-Gnutella25 | 22,687 | 54,705 | 64 | 66 | 67 | 0.22 | 64 | 0.33 |
| as-22july06 | 22,963 | 48,436 | 2243 | 2387 | 4125 | 1678.23 | 2245 | 10,722.34 |
| p2p-Gnutella24 | 26,518 | 65,369 | 304 | 356 | 426 | 1040.93 | 304 | 1043.72 |
| p2p-Gnutella30 | 36,682 | 88,328 | 54 | 54 | 53 | 0.59 | 54 | 0.70 |
| p2p-Gnutella31 | 62,586 | 147,892 | 90 | 95 | 100 | 0.41 | 90 | 0.70 |
| delaunay-n14 | 16,384 | 49,122 | 9 | 26 | 39 | 4.15 | 9 | 4.23 |
| delaunay-n16 | 65,536 | 196,575 | 9 | 18 | 34 | 61.05 | 9 | 61.13 |
| delaunay-n17 | 131,072 | 393,176 | 9 | 67 | 123 | 243.29 | 10 | 244.00 |
| delaunay-n18 | 262,144 | 786,396 | 11 | 65 | 123 | 541.73 | 12 | 542.29 |
| delaunay-n19 | 524,288 | 1,572,823 | 11 | 54 | 92 | 2205.50 | 11 | 2206.09 |
| delaunay-n20 | 1,048,576 | 3,145,686 | 12 | 24 | 45 | 5086.75 | 13 | 5087.06 |
| Graph | LB | SR-CPU(s) | Opt. Obj. | CPU (s) | ||||
|---|---|---|---|---|---|---|---|---|
| jazz | 198 | 2742 | 17 | 85 | 704 | 0.67 | 19 | 2.67 |
| 1133 | 5451 | 33 | 35 | 35 | 1.42 | 33 | 1.45 | |
| netscience | 1589 | 2742 | 15 | 26 | 42 | 0.01 | 15 | 0.03 |
| add20 | 2395 | 7462 | 29 | 68 | 186 | 306.95 | 29 | 307.45 |
| data | 2851 | 15,093 | 7 | 1944 | 11,290 | 0.78 | - | >11,000 |
| as19971108 | 3015 | 5347 | 539 | 583 | 740 | 1430.57 | 539 | 1462.28 |
| add32 | 4960 | 9462 | 15 | 66 | 108 | 0.17 | 15 | 0.89 |
| CA-GrQC | 5242 | 14,490 | 28 | 35 | 41 | 0.47 | 28 | 0.51 |
| as19991204 | 6296 | 12,830 | 1293 | 1407 | 2234 | 2081.83 | 1293 | 9542.64 |
| p2p-Gnutella08 | 6301 | 20,777 | 87 | 448 | 3194 | 9.19 | 87 | 8177.89 |
| as20000102 | 6474 | 13,233 | 1337 | 1454 | 2430 | 1201.03 | 1339 | 10,044.40 |
| p2p-Gnutella09 | 8114 | 26,013 | 97 | 98 | 97 | 12.37 | 97 | 12.51 |
| hep-th | 8361 | 15,751 | 21 | 81 | 151 | 0.50 | 22 | 1.26 |
| p2p-Gnutella06 | 8717 | 31,525 | 103 | 305 | 1283 | 2.53 | 138 | 654.47 |
| p2p-Gnutella05 | 8846 | 31,839 | 86 | 136 | 203 | 5.74 | 92 | 8.38 |
| CA-HepTH | 9877 | 25,973 | 31 | 43 | 59 | 6.19 | 31 | 6.51 |
| PGPgiantcompo | 10,680 | 24,316 | 104 | 114 | 122 | 3899.91 | 104 | 3900.27 |
| p2p-Gnutella04 | 10,876 | 39,994 | 96 | 100 | 102 | 0.56 | 96 | 0.78 |
| oregon1-010519 | 11,051 | 22,724 | 2202 | 2384 | 4289 | 1262.38 | 2206 | 10,494.20 |
| oregon1-010526 | 11,174 | 23,409 | 2198 | 2385 | 4308 | 1415.75 | 2202 | 10,750.50 |
| oregon2-010526 | 11,460 | 16,365 | 2229 | 2428 | 4676 | 1579.16 | - | >11,000 |
| cond-mat | 16,726 | 47,594 | 40 | 55 | 72 | 7703.95 | 40 | 7704.02 |
| p2p-Gnutella25 | 22,687 | 54,705 | 63 | 66 | 67 | 0.31 | 63 | 0.44 |
| as-22july06 | 22,963 | 48,436 | 2242 | 2387 | 4125 | 3735.30 | - | >11,000 |
| p2p-Gnutella24 | 26,518 | 65,369 | 303 | 356 | 426 | 220.50 | 303 | 225.52 |
| p2p-Gnutella30 | 36,682 | 88,328 | 53 | 54 | 53 | 2.06 | 53 | 2.12 |
| p2p-Gnutella31 | 62,586 | 147,892 | 89 | 95 | 100 | 0.98 | 89 | 1.30 |
| delaunay-n14 | 16,384 | 49,122 | 8 | 26 | 39 | 5.57 | 8 | 5.70 |
| delaunay-n16 | 65,536 | 196,575 | 8 | 18 | 34 | 80.41 | 8 | 80.54 |
| delaunay-n17 | 131,072 | 393,176 | 8 | 67 | 123 | 315.68 | 9 | 316.55 |
| delaunay-n18 | 262,144 | 786,396 | 10 | 65 | 123 | 373.68 | 11 | 374.53 |
| delaunay-n19 | 524,288 | 1,572,823 | 10 | 54 | 92 | 1478.44 | 10 | 1479.16 |
| delaunay-n20 | 1,048,576 | 3,145,686 | 11 | 24 | 45 | 3044.67 | 12 | 3045.02 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shahinpour, S.; Shirvani, S.; Ertem, Z.; Butenko, S. Scale Reduction Techniques for Computing Maximum Induced Bicliques. Algorithms 2017, 10, 113. https://doi.org/10.3390/a10040113
Shahinpour S, Shirvani S, Ertem Z, Butenko S. Scale Reduction Techniques for Computing Maximum Induced Bicliques. Algorithms. 2017; 10(4):113. https://doi.org/10.3390/a10040113
Chicago/Turabian StyleShahinpour, Shahram, Shirin Shirvani, Zeynep Ertem, and Sergiy Butenko. 2017. "Scale Reduction Techniques for Computing Maximum Induced Bicliques" Algorithms 10, no. 4: 113. https://doi.org/10.3390/a10040113
APA StyleShahinpour, S., Shirvani, S., Ertem, Z., & Butenko, S. (2017). Scale Reduction Techniques for Computing Maximum Induced Bicliques. Algorithms, 10(4), 113. https://doi.org/10.3390/a10040113




