Synthesis of Bio-Based Polyamide 6,5 via Enzymatic Polycondensation
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Enzymatic Synthesis of Polyamide 6,5
2.3. Analytical Techniques
3. Results and Discussion
3.1. Qualitative Identification of Bio-Based Polyamide 6,5
3.2. Synthesis Effectiveness of Bio-Based Polyamide 6,5
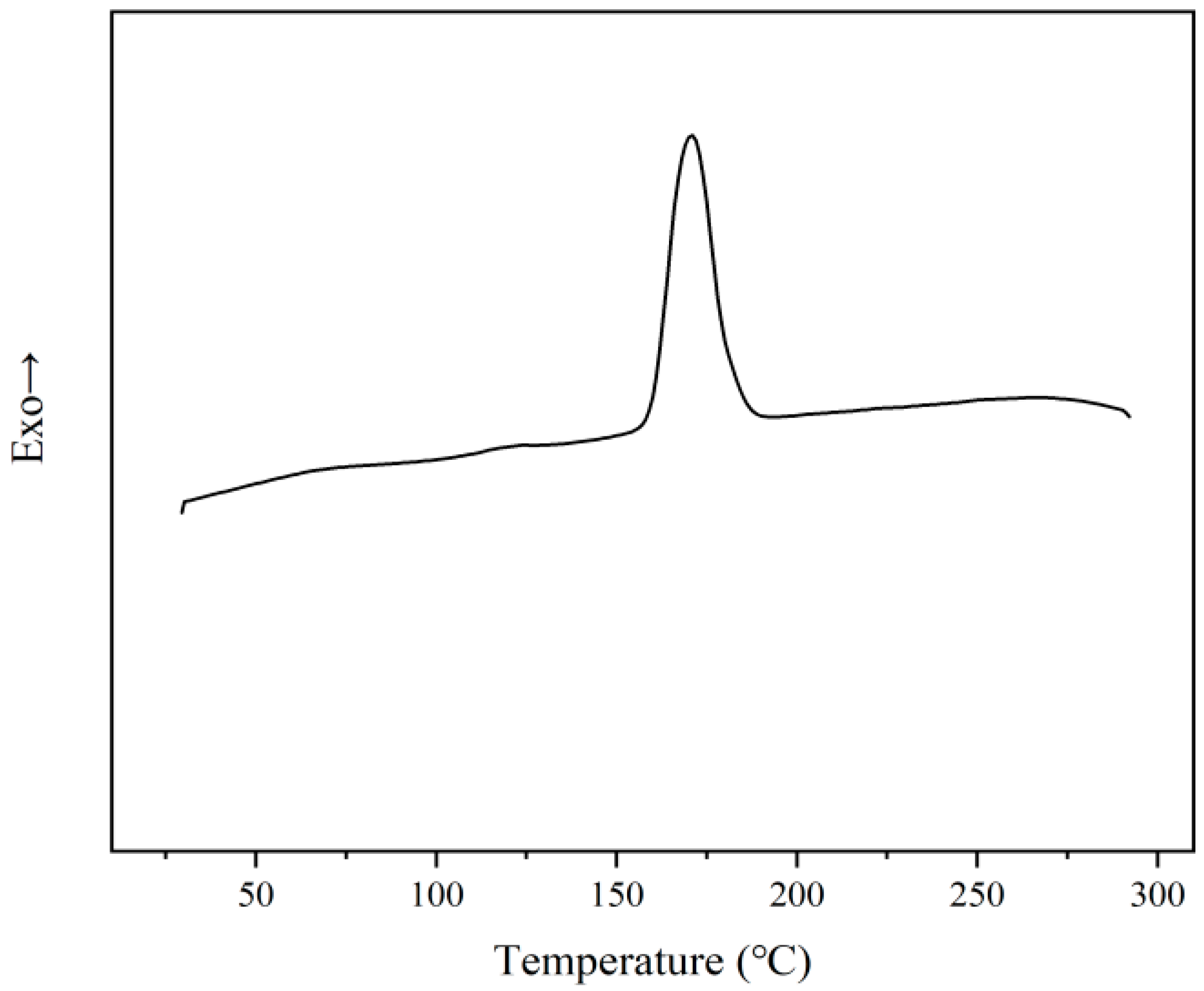
3.3. Characterization of Bio-Based Polyamide 6,5
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chanda, M.; Roy, S.K. Plastics Technology Handbook; CRC Press: Boca Raton, FL, USA, 2006. [Google Scholar]
- Carothers, W.H. Synthetie Fiber. US 2130948, 20 September 1938. [Google Scholar]
- Yokozawa, T.; Yokoyama, A. Chain-Growth Polycondensation: The Living Polymerization Process in Polycondensation. Prog. Polym. Sci. 2007, 32, 147–172. [Google Scholar] [CrossRef]
- Yokoyama, T.Y.A. Chain-Growth Polycondensation: Living Polymerization Nature in Poly- Condensation and Approach to Condensation Polymer Architecture. Polym. J. 2004, 36, 65–83. [Google Scholar]
- Qian, Z.; Chen, X.; Xu, J.; Guo, B. Chain Extension of Pa1010 by Reactive Extrusion by Diepoxide 711 and Diepoxide Tde85 as Chain Extenders. J. Appl. Polym. Sci. 2004, 94, 2347–2355. [Google Scholar] [CrossRef]
- Vouyiouka, S.N.; Filgueiras, V.; Papaspyrides, C.D.; Lima, E.L.; Pinto, J.C. Morphological Changes of Poly(Ethylene Terephthalate-Co-Isophthalate) During Solid State Polymerization. J. Appl. Polym. Sci. 2012, 124, 4457–4465. [Google Scholar] [CrossRef]
- Wang, Y.; Cui, L.; Zhang, J.; Shen, J.; Xu, H.; Zhou, Z.; Li, Y.; Zhu, M. Fire Retardant Treatments for Polyamide 6 and 66: Advances and Trends Over the Last Five Years. Acs Appl. Polym. Mater. 2025, 7, 4677–4693. [Google Scholar] [CrossRef]
- Nakajima, H.; Dijkstra, P.; Loos, K. The Recent Developments in Biobased Polymers Toward General and Engineering Applications: Polymers that are Upgraded from Biodegradable Polymers, Analogous to Petroleum-Derived Polymers, and Newly Developed. Polymers 2017, 9, 523. [Google Scholar] [CrossRef]
- Ragauskas, A.J.; Williams, C.K.; Davison, B.H.; Britovsek, G.; Cairney, J.; Eckert, C.A.; Frederick, W.J.; Hallett, J.P.; Leak, D.J.; Liotta, C.L.; et al. The Path Forward for Biofuels and Biomaterials. Science 2006, 311, 484–489. [Google Scholar] [CrossRef]
- Son, J.; Sohn, Y.J.; Baritugo, K.; Jo, S.Y.; Song, H.M.; Park, S.J. Recent Advances in Microbial Production of Diamines, Aminocarboxylic Acids, and Diacids as Potential Platform Chemicals and Bio-Based Polyamides Monomers. Biotechnol. Adv. 2023, 62, 108070. [Google Scholar] [CrossRef] [PubMed]
- Wendisch, V.F.; Mindt, M.; Pérez-García, F. Biotechnological Production of Mono- And Diamines Using Bacteria: Recent Progress, Applications, and Perspectives. Appl. Microbiol. Biotechnol. 2018, 102, 3583–3594. [Google Scholar] [CrossRef]
- Bah, C.S.F.; Bekhit, A.E.A.; Carne, A.; McConnell, M.A. Production of Bioactive Peptide Hydrolysates From Deer, Sheep and Pig Plasma Using Plant and Fungal Protease Preparations. Food Chem. 2015, 176, 54–63. [Google Scholar] [CrossRef]
- Radzik, P.; Leszczyńska, A.; Pielichowski, K. Modern Biopolyamide-Based Materials: Synthesis and Modification. Polym. Bull. 2020, 77, 501–528. [Google Scholar] [CrossRef]
- Zhang, P.; Fan, H.; Tian, S.; Chen, Y.; Yan, J. Synergistic Effect of Phosphorus–Nitrogen and Silicon-Containing Chain Extenders On the Mechanical Properties, Flame Retardancy and Thermal Degradation Behavior of Waterborne Polyurethane. Rsc Adv. 2016, 6, 72409–72422. [Google Scholar] [CrossRef]
- Douka, A.; Vouyiouka, S.; Papaspyridi, L.; Papaspyrides, C.D. A Review On Enzymatic Polymerization to Produce Polycondensation Polymers: The Case of Aliphatic Polyesters, Polyamides and Polyesteramides. Prog. Polym. Sci. 2018, 79, 1–25. [Google Scholar] [CrossRef]
- Heise, A.; Palmans, A.R.A. Hydrolases in Polymer Chemistry: Chemoenzymatic Approaches to Polymeric Materials; Palmans, A.R.A., Heise, A., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 79–113. [Google Scholar]
- Marchildon, K. Polyamides–Still Strong After Seventy Years. Macromol. React. Eng. 2011, 5, 22–54. [Google Scholar] [CrossRef]
- Müllen, K. Book Review On Enzymatic Polymerization Towards Green Polymer Chemistry. J. Polym. Environ. 2020, 28, 737–738. [Google Scholar] [CrossRef]
- Panova, A.; Dicosimo, R.; Brugel, E.G.; Tam, W.; EIDP Inc. Enzymatic Production of Macrocyclic Amide Oligomers. US7507560B2, 24 March 2009. [Google Scholar]
- Dionisi, D.; Adams, T.; Dempster, C. Open Mixed Cultures for Bioethanol Production From Lignocellulosic Materials. New Biotech. 2014, 31, S31. [Google Scholar] [CrossRef]
- Yang, B.; Wang, G.; Xia, B.; Zhong, M.; Fan, P.; Chen, F.; Fei, Z. Enzymatic Synthesis of Chiral Polyamide Via Condensation of Natural Source Amino Acid Diesters and Diamine. Macromol. Chem. Phys. 2021, 222, 2100162. [Google Scholar] [CrossRef]
- Dordick, J.S. Enzymatic and Chemoenzymatic Approaches to Polymer Synthesis. Trends. Biotechnol. 1992, 10, 287–290. [Google Scholar] [CrossRef] [PubMed]
- Frampton, M.B.; Zelisko, P.M. Synthesis of Lipase-Catalysed Silicone-Polyesters and Silicone-Polyamides at Elevated Temperatures. Chem. Commun. 2013, 49, 9269. [Google Scholar] [CrossRef]
- Jiang, Y.; Loos, K. Enzymatic Synthesis of Biobased Polyesters and Polyamides. Polymers 2016, 8, 243. [Google Scholar] [CrossRef]
- Ragupathy, L.; Ziener, U.; Dyllick-Brenzinger, R.; von Vacano, B.; Landfester, K. Enzyme-Catalyzed Polymerizations at Higher Temperatures: Synthetic Methods to Produce Polyamides and New Poly(Amide-Co-Ester)S. J. Mol. Catal. B Enzym. 2012, 76, 94–105. [Google Scholar] [CrossRef]
- Claudia Ortiz, M.L.F. Novozym 435: The “Perfect” Lipase Immobilized Biocatalyst? Catal. Sci. Technol. 2019, 9, 2380–2420. [Google Scholar] [CrossRef]
- Kumar, A.; Gross, R.A. Candida antartica Lipase B Catalyzed Polycaprolactone Synthesis: Effects of Organic Media and Temperature. Biomacromolecules 2000, 1, 133–138. [Google Scholar] [CrossRef]
- Schwab, L.W.; Kroon, R.; Schouten, A.J.; Loos, K. Enzyme-Catalyzed Ring-Opening Polymerization of Unsubstituted β-Lactam. Macromol. Rapid Commun. 2008, 29, 794–797. [Google Scholar] [CrossRef]
- Stavila, E.; Arsyi, R.Z.; Petrovic, D.M.; Loos, K. Fusarium Solani Pisi Cutinase-Catalyzed Synthesis of Polyamides. Eur. Polym. J. 2013, 49, 834–842. [Google Scholar] [CrossRef]
- Uvenilir, S.E.G.U. Lipase Catalyzed Synthesis of Bio-Based Polyamide 5.6: An Alternative Route. J. Macromol. Sci. Part A Pure Appl. Chem. 2024, 1, 69–78. [Google Scholar]
- Sharma, S.; Belluati, A.; Kumar, M.; Dhiman, S. Enzymatic Reaction Network-Driven Polymerization-Induced Transient Coacervation. Angew. Chem. Int. Ed. 2025, 64, e202421620. [Google Scholar] [CrossRef] [PubMed]
- Hu, T.; Yuan, Y.; Wang, J.; Hu, H. Enzyme-Crosslinked Soy Protein Aerogels: Structural and Functional Modulations by Pre-Freezing Process. Food Hydrocoll. 2026, 170, 111698. [Google Scholar] [CrossRef]
- Tordi, P.; Tamayo, A.; Jeong, Y.; Han, B.; Al Kayal, T.; Cavallo, A.; Bonini, M.; Samorì, P. Fully Bio-Based Gelatin Organohydrogels Via Enzymatic Crosslinking for Sustainable Soft Strain and Temperature Sensing. Adv. Funct. Mater. 2025, 36, e20762. [Google Scholar] [CrossRef]
- Chu, R.; Wang, Z.; Liu, J.; Hu, G.; Liu, L.; Li, X.; Gao, C. Metabolic Engineering of Plasmid-Free Escherichia Coli for Enhanced Glutarate Biosynthesis From Glucose. Chem. Eng. J. 2025, 513, 163077. [Google Scholar] [CrossRef]
- Haagen, V.; Iser, J.; Schörner, M.; Weber, D.; Franken, T.; Wasserscheid, P.; Schühle, P. Synthesis of Methanol by Hydrogenolysis of Biobased Methyl Formate Using Highly Stable and Active Cu-Spinel Catalysts in Slurry and Gas Phase Reactions. Green Chem. 2023, 25, 2338–2348. [Google Scholar] [CrossRef]
- Navarro, E.; Franco, L.; Subirana, J.A.; Puiggali, J. Nylon 65 Has a Unique Structure with Two Directions of Hydrogen Bonds. Macromolecules 1995, 28, 8742–8750. [Google Scholar] [CrossRef]
- Dixon, E.G.; Ciano, L.; Pordea, A.; Alvey, K.; Taresco, V. Harnessing Enzymes for Greener Polymerisations: Advances in Chain and Step Growth Processes. Polym. Chem. 2025, 16, 2997–3029. [Google Scholar] [CrossRef]
- Kobayashi, S.; Uyama, H.; Kimura, S. Enzymatic Polymerization. Chem. Rev. 2001, 101, 3793–3818. [Google Scholar] [CrossRef]
- Kohutiar, M.; Kakošová, L.; Krbata, M.; Janík, R.; Fekiač, J.J.; Breznická, A.; Eckert, M.; Mikuš, P.; Timárová, Ľ. Comprehensive Review: Technological Approaches, Properties, and Applications of Pure and Reinforced Polyamide 6 (Pa6) and Polyamide 12 (Pa12) Composite Materials. Polymers 2025, 17, 442. [Google Scholar] [CrossRef]
- Stavila, E.; Loos, K. Synthesis of Polyamides and their Copolymers Via Enzymatic Polymerization. J. Renew. Mater. 2015, 3, 268–280. [Google Scholar] [CrossRef]
- Kobayashi, S. Recent Developments in Lipase-Catalyzed Synthesis of Polyesters. Macromol. Rapid Commun. 2009, 30, 237–266. [Google Scholar] [CrossRef] [PubMed]
- Kundys, A.; Białecka-Florjańczyk, E.; Fabiszewska, A.; Małajowicz, J. Candida Antarctica Lipase B as Catalyst for Cyclic Esters Synthesis, their Polymerization and Degradation of Aliphatic Polyesters. J. Polym. Environ. 2018, 26, 396–407. [Google Scholar] [CrossRef]







| Element | Found (%) | Theoretical (%) | Deviation (%) |
|---|---|---|---|
| C | 57.04 | 57.37 | 0.58 |
| 56.98 | 0.68 | ||
| 57.13 | 0.42 | ||
| H | 9.53 | 9.63 | 1.06 |
| 9.73 | 1.02 | ||
| 9.62 | 0.10 | ||
| O | 21.05 | 20.84 | 1.02 |
| 21.23 | 1.88 | ||
| 21.05 | 1.01 | ||
| N | 12.38 | 12.16 | 1.81 |
| 12.06 | 0.82 | ||
| 12.20 | 0.33 |
 |  | Cycles, RM = 0 amu |
 |  | Acid/amine, RM = 18 amu |
 |  | Ester/amine, RM = 32 amu |
 |  | Diamine, RM = 116.2 amu |
 |  | Diester, RM = 160.2 amu |
| Temperature (°C) | Time (h) | Enzyme Concentrations (wt%) | Mn | Mw | PDI | Yield (%) |
|---|---|---|---|---|---|---|
| 80 | 72 | 20 | 6900 | 11,600 | 1.68 | 63 |
| 90 | 72 | 20 | 8600 | 14,100 | 1.64 | 72 |
| 100 | 72 | 20 | 8800 | 15,500 | 1.76 | 80 |
| 110 | 72 | 20 | 8800 | 15,600 | 1.77 | 83 |
| 90 | 24 | 20 | 4300 | 7400 | 1.72 | 39 |
| 90 | 48 | 20 | 6800 | 11,500 | 1.69 | 58 |
| 90 | 72 | 20 | 8600 | 14,100 | 1.64 | 72 |
| 90 | 96 | 20 | 8400 | 14,800 | 1.76 | 75 |
| 90 | 72 | 10 | 5800 | 9200 | 1.59 | 55 |
| 90 | 72 | 15 | 6800 | 11,400 | 1.68 | 62 |
| 90 | 72 | 20 | 8600 | 14,100 | 1.64 | 72 |
| 90 | 72 | 25 | 8900 | 15,500 | 1.74 | 75 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Xia, Y.; Sun, S.; Zhang, Q.; Li, J. Synthesis of Bio-Based Polyamide 6,5 via Enzymatic Polycondensation. Materials 2026, 19, 1399. https://doi.org/10.3390/ma19071399
Xia Y, Sun S, Zhang Q, Li J. Synthesis of Bio-Based Polyamide 6,5 via Enzymatic Polycondensation. Materials. 2026; 19(7):1399. https://doi.org/10.3390/ma19071399
Chicago/Turabian StyleXia, Yiran, Shidong Sun, Qianqian Zhang, and Jinlong Li. 2026. "Synthesis of Bio-Based Polyamide 6,5 via Enzymatic Polycondensation" Materials 19, no. 7: 1399. https://doi.org/10.3390/ma19071399
APA StyleXia, Y., Sun, S., Zhang, Q., & Li, J. (2026). Synthesis of Bio-Based Polyamide 6,5 via Enzymatic Polycondensation. Materials, 19(7), 1399. https://doi.org/10.3390/ma19071399







