Assessment of Soft Computing Techniques for the Prediction of Compressive Strength of Bacterial Concrete
Abstract
:1. Introduction
2. Research Significance
3. Review of Regression and Soft Computing Techniques
3.1. Multiple Linear Regression
3.2. Random Forest
3.3. M5P Model
- K—set of instances that attain the node.
- Ki—the subset of illustrations that have the product of the possible set.
- and —the standard deviation
3.4. Random Tree Model
3.5. Reduced Error Pruning Tree (REP Tree)
3.6. Support Vector Regression (SVR)
3.7. Performance Evaluation Indices
- values of the actual observations in a sample
- mean of the values of the actual observations
- values of the predicted observations in a sample
- mean of the values of the predicted observations
4. Materials and Methodology
4.1. Design Proportions
4.2. Data Set
5. Results and Discussion
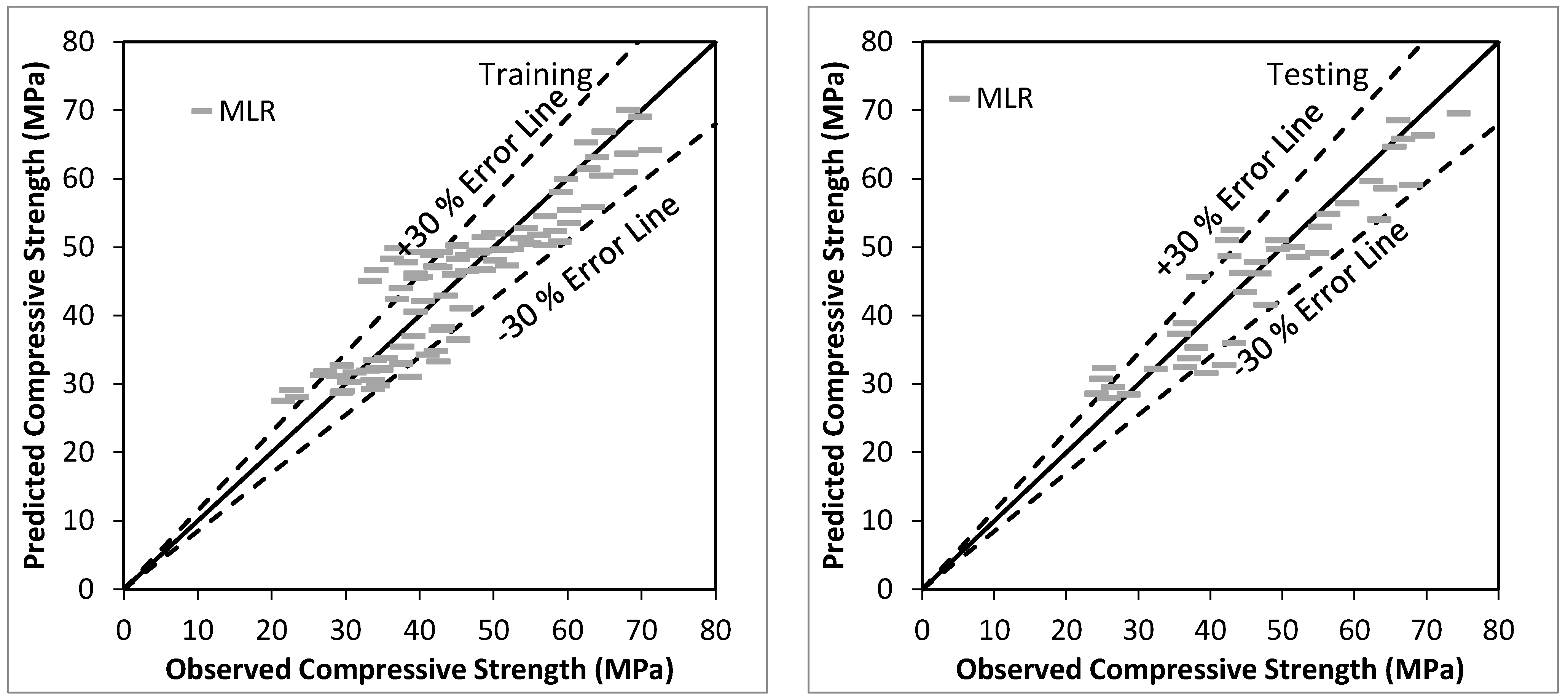
5.1. Results of Multiple Linear Regression
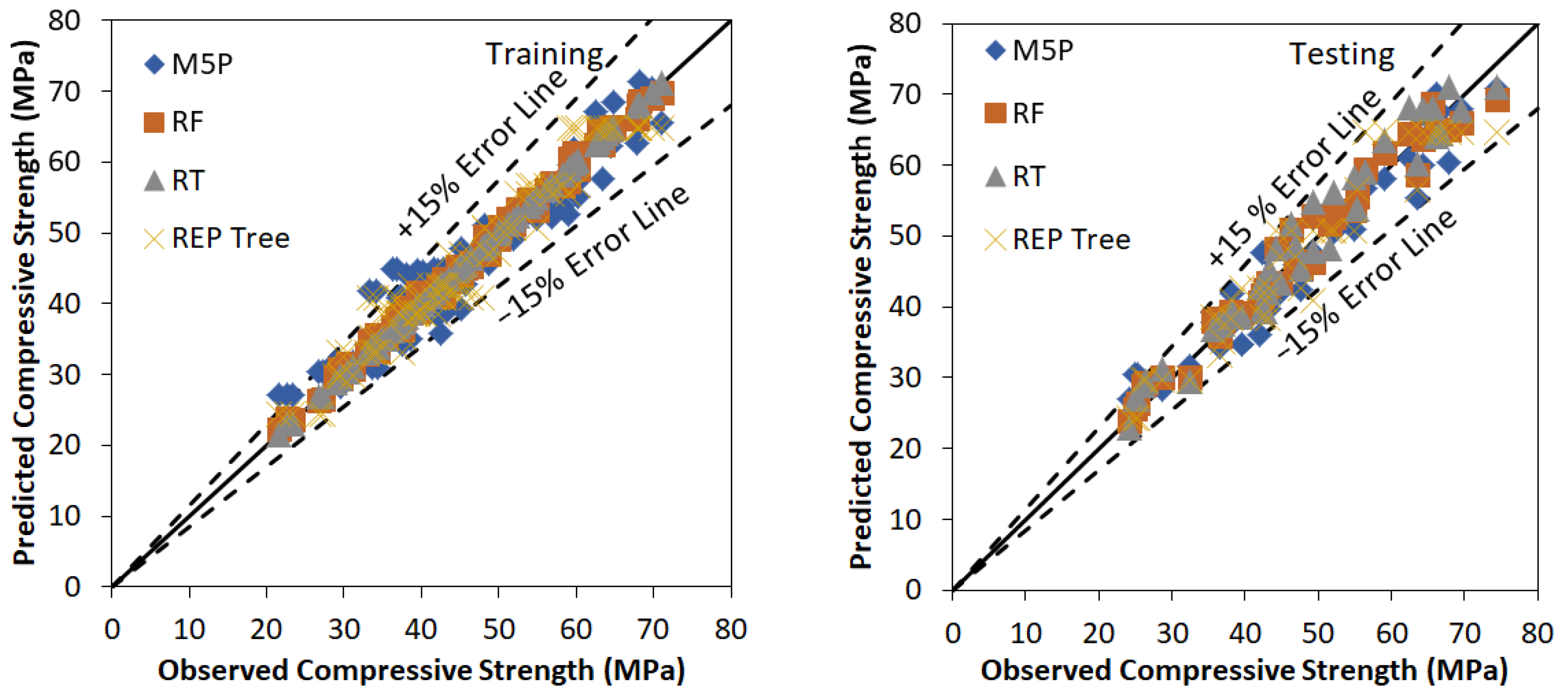
5.2. Results of the Tree and Forest-Based Models
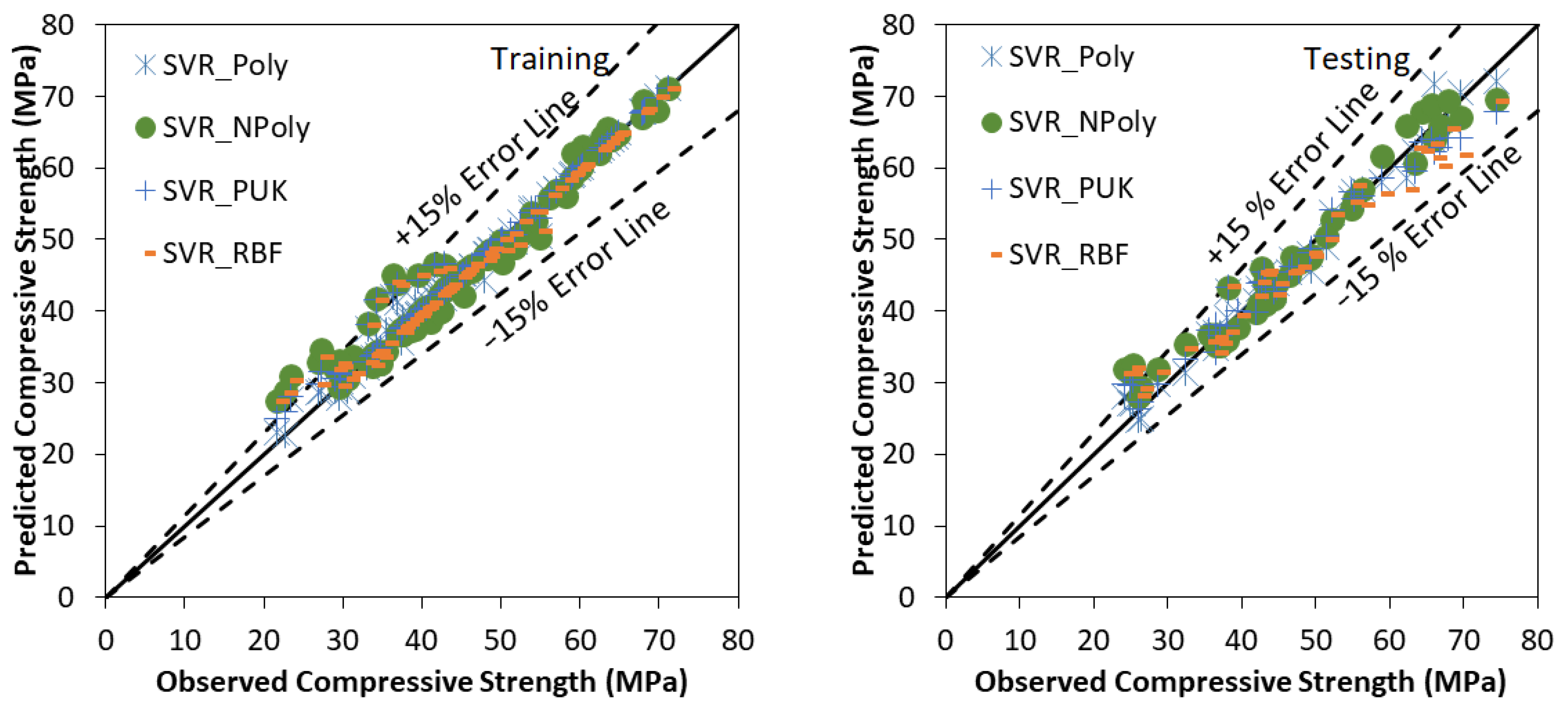
5.3. Results of SVR Based Models
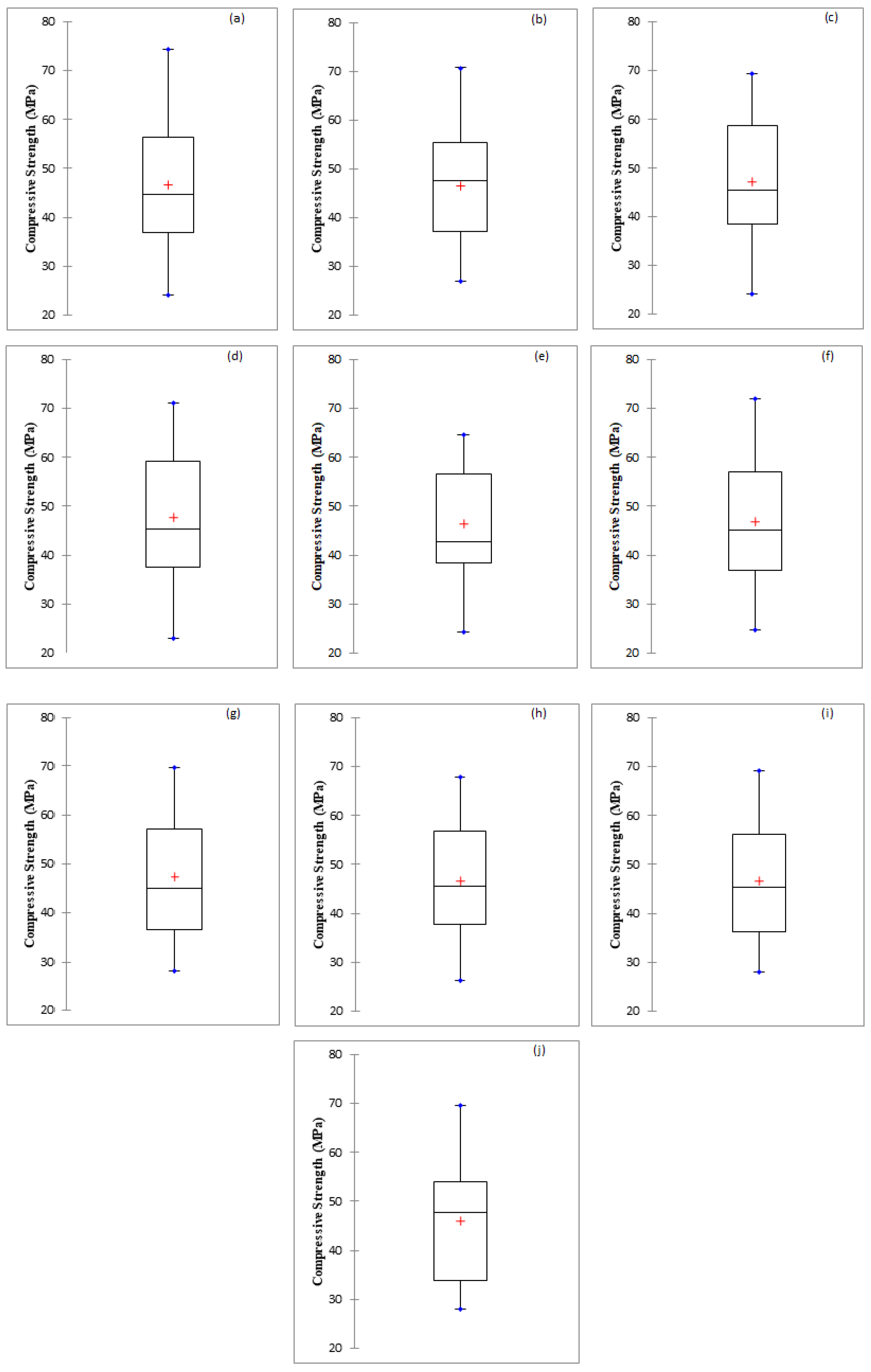
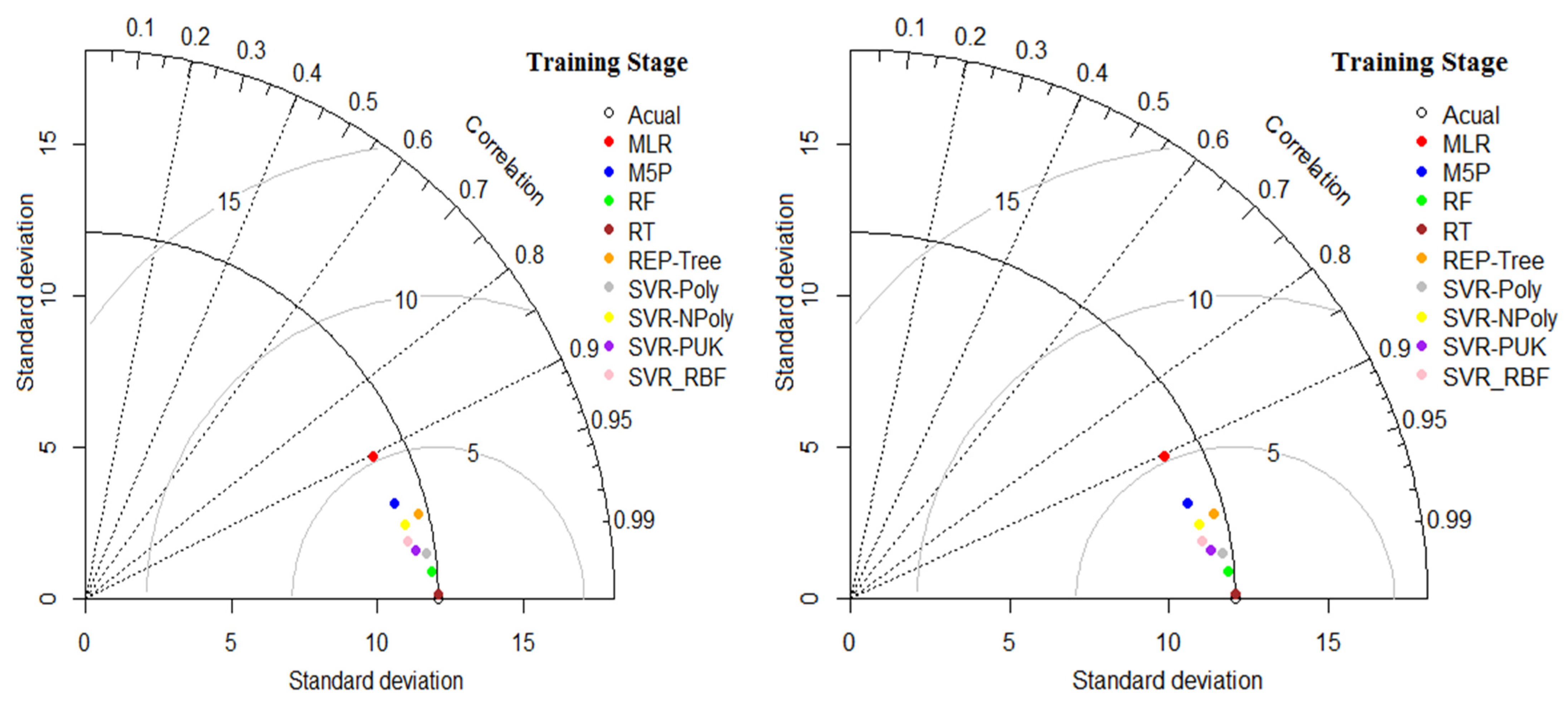
5.4. Comparison among Regression and Soft Computing-Based Models
5.5. Sensitivity Analysis
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Franzoni, E. Materials selection for green buildings: Which tools for engineers and architects? Procedia Eng. 2011, 21, 883–890. [Google Scholar] [CrossRef] [Green Version]
- Melchert, L. The Dutch sustainable building policy: A model for developing countries? Build. Environ. 2007, 42, 893–901. [Google Scholar] [CrossRef]
- Mindeguia, J.-C.; Hager, I.; Pimienta, P.; Borderie, C.L.; Carré, H. Parametrical study of transient thermal strain of high performance concrete. Cem. Concr. Res. 2013, 48, 40–52. [Google Scholar] [CrossRef]
- Ostrowski, K.; Sadowski, Ł.; Stefaniuk, D.; Wałach, D.; Gawenda, T.; Oleksik, K.; Usydus, I. The effect of the shape of coarse aggregate on the 2 properties of self-compacting high-performance 3 fibre-reinforced concrete. Materials 2018, 11, 1372. [Google Scholar] [CrossRef] [Green Version]
- Ostrowski, K. The influence of coarse aggregate shape on the properties of high-performance, self-compacting concrete. Tech. Trans. Civ. Eng. 2017, 5, 25–33. [Google Scholar] [CrossRef] [Green Version]
- Häkkinen, T.; Belloni, K. Barriers and drivers for sustainable building. Build. Res. Inf. 2011, 39, 239–255. [Google Scholar] [CrossRef]
- Chen, Z.S.; Martinez, L.; Chang, J.P.; Wang, X.J.; Xionge, S.H.; Chin, K.S. Sustainable building material selection: A QFD-and ELECTRE III-embedded hybrid MCGDM approach with consensus building. Eng. Appl. Artif. Intell. 2019, 85, 783–807. [Google Scholar] [CrossRef]
- Invidiata, A.; Lavagna, M.; Ghisi, E. Selecting design strategies using multi-criteria decision making to improve the sustainability of buildings. Build. Environ. 2018, 139, 58–68. [Google Scholar] [CrossRef]
- Roodman, D.M.; Lenssen, N.K.; Peterson, J.A. A Building Revolution: How Ecology and Health Concerns Are Transforming Construction; Worldwatch Institute: Washington, DC, USA, 1995; p. 11. [Google Scholar] [CrossRef]
- Wang, J.; Jonkers, H.M.; Boon, N.; De Belie, N. Bacillus sphaericus LMG 22257 is physiologically suitable for self-healing concrete. Appl. Microbiol. Biotechnol. 2017, 101, 5101–5114. [Google Scholar] [CrossRef]
- Stanaszek-Tomal, E. Bacterial Concrete as a Sustainable Building Material? Sustainability 2020, 12, 696. [Google Scholar] [CrossRef] [Green Version]
- Achal, V.; Mukherjee, A. A review of microbial precipitation for sustainable construction. Constr. Build. Mater. 2015, 93, 1224–1235. [Google Scholar] [CrossRef]
- Ghassemi, P.; Rajabi, H.; Toufigh, V. Fatigue performance of polymer ond ordinary cement concrete under corrosive conditions: A comparative study. Eng. Fail. Anal. 2020, 111, 104493. [Google Scholar] [CrossRef]
- Ghassemi, P.; Toufigh, V. Durability of epoxy polymer and ordinary cement concrete in aggressive environments. Constr. Build. Mater. 2020, 234, 117887. [Google Scholar] [CrossRef]
- Qureshi, T.; Kanellopoulos, A.; Al-Tabbaa, A. Autogenous self-healing of cement with expansive minerals-I: Impact in early age crack healing. Constr. Build. Mater. 2018, 192, 768–784. [Google Scholar] [CrossRef] [Green Version]
- Huang, H.; Ye, G.; Damidot, D. Characterization and quantification of self-healing behaviors of microcracks due to further hydration in cement paste. Cem. Concr. Res. 2013, 52, 71–81. [Google Scholar] [CrossRef]
- Sisomphon, K.; Copuroglu, O.; Koenders, E.A.B. Self-healing of surface cracks in mortars with expansive additive and crystalline additive. Cem. Concr. Compos. 2012, 34, 566–574. [Google Scholar] [CrossRef]
- Akadiri, P.O.; Chinyio, E.A.; Olomolaiye, P.O. Design of a sustainable building: A conceptual framework for implementing sustainability in the building sector. Buildings 2012, 2, 126–152. [Google Scholar] [CrossRef] [Green Version]
- Kalra, M.; Mehmood, G. A Review paper on the Effect of different types of coarse aggregate on Concrete. In IOP Conference Series: Materials Science and Engineering, October 2018; IOP Publishing: Bristol, UK, 2018; Volume 431, p. 082001. [Google Scholar] [CrossRef]
- Irrigaray, M.A.P.; Pinto, R.D.A.; Padaratz, I.J. A new approach to estimate compressive strength of concrete by the UPV method. Rev. IBRACON Estrut. Mater. 2016, 9, 395–402. [Google Scholar] [CrossRef]
- Anyaoha, U.; Peng, X.; Liu, Z. Concrete performance prediction using boosting smooth transition regression trees (BooST). In Nondestructive Characterization and Monitoring of Advanced Materials, Aerospace, Civil Infrastructure, and Transportation; XIII; International Society for Optics and Photonics: Bellingham, WA, USA, 2019; Volume 10971, p. 109710I. [Google Scholar] [CrossRef]
- Bhanja, S.; Sengupta, B. Investigations on the compressive strength of silica fume concrete using statistical methods. Cem. Concr. Res. 2002, 32, 1391–1394. [Google Scholar] [CrossRef]
- Lee, S.C. Prediction of concrete strength using artificial neural networks. Eng. Struct. 2003, 25, 849–857. [Google Scholar] [CrossRef]
- Khalaj, G.; Pouraliakbar, H. Computer-aided modeling for predicting layer thickness of a duplex treated ceramic coating on tool steels. Ceram. Int. 2014, 40, 5515–5522. [Google Scholar] [CrossRef]
- Khalaj, G.; Nazari, A.; Khoie, S.M.M.; Khalaj, M.J. Pouraliakbar, H. Chromium carbonitride coating produced on DIN 1.2210 steel by thermo-reactive deposition technique: Thermodynamics, kinetics and modeling. Surf. Coat. Technol. 2013, 225, 1–10. [Google Scholar] [CrossRef]
- Zain, M.F.M.; Abd, S.M. Multiple regression model for compressive strength prediction of high performance concrete. J. Appl. Sci. 2009, 9, 155–160. [Google Scholar] [CrossRef]
- Hayes, T.; Usami, S.; Jacobucci, R.; McArdle, J.J. Using Classification and Regression Trees (CART) and random forests to analyze attrition: Results from two simulations. Psychol. Aging 2015, 30, 911. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yeh, I.C. Modeling of strength of high-performance concrete using artificial neural networks. Cem. Concr. Res. 1998, 28, 1797–1808. [Google Scholar] [CrossRef]
- Chopra, P.; Sharma, R.K.; Kumar, M. Prediction of compressive strength of concrete using artificial neural network and genetic programming. Adv. Mater. Sci. Eng. 2016, 2016, 7648467. [Google Scholar] [CrossRef] [Green Version]
- Sobhani, J.; Najimi, M.; Pourkhorshidi, A.R.; Parhizkar, T. Prediction of the compressive strength of no-slump concrete: A comparative study of regression, neural network and ANFIS models. Constr. Build. Mater. 2010, 24, 709–718. [Google Scholar] [CrossRef]
- Gupta, S.M. Support vector machines based modelling of concrete strength. World Acad. Sci. Eng. Technol. 2007, 36, 305–311. [Google Scholar]
- Yan, K.; Shi, C. Prediction of elastic modulus of normal and high strength concrete by support vector machine. Constr. Build. Mater. 2010, 24, 1479–1485. [Google Scholar] [CrossRef]
- Thakur, M.S.; Pandhiani, S.M.; Kashyap, V.; Upadhya, A.; Sihag, P. Predicting Bond Strength of FRP Bars in Concrete Using Soft Computing Techniques. Arab. J. Sci. Eng. 2021, 46, 4951–4969. [Google Scholar] [CrossRef]
- Dayev, Z.; Kairakbaev, A.; Yetilmezsoy, K.; Bahramian, M.; Sihag, P.; Kıyan, E. Approximation of the discharge coefficient of differential pressure flowmeters using different soft computing strategies. Flow Meas. Instrum. 2021, 79, 101913. [Google Scholar] [CrossRef]
- Singh, A.; Singh, R.M.; Kumar, A.S.; Kumar, A.; Hanwat, S.; Tripathi, V.K. Evaluation of soft computing and regression-based techniques for the estimation of evaporation. J. Water Clim. Chang. 2021, 12, 32–43. [Google Scholar] [CrossRef]
- Breiman, L. Bagging predictors. Mach. Learn. 1996, 24, 123–140. [Google Scholar] [CrossRef] [Green Version]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef] [Green Version]
- Quinlan, J.R. Learning with continuous classes. In Proceedings of the 5th Australian Joint Conference on Artificial Intelligence, Hobart, Australia, 16–18 November 1992; Volume 92, pp. 343–348. [Google Scholar]
- Yetilmezsoy, K.; Sihag, P.; Kıyan, E.; Doran, B. A benchmark comparison and optimization of Gaussian process regression, support vector machines, and M5P tree model in approximation of the lateral confinement coefficient for CFRP-wrapped rectangular/square RC columns. Eng. Struct. 2021, 246, 113106. [Google Scholar] [CrossRef]
- Upadhya, A.; Thakur, M.S.; Sharma, N.; Sihag, P. Assessment of Soft Computing-Based Techniques for the Prediction of Marshall Stability of Asphalt Concrete Reinforced with Glass Fiber. Int. J. Pavement Res. Technol. 2021, 1–20. [Google Scholar] [CrossRef]
- Kumar, V.; Sihag, P.; Keshavarzi, A.; Pandita, S.; Rodríguez-Seijo, A. Soft Computing Techniques for Appraisal of Potentially Toxic Elements from Jalandhar (Punjab), India. Appl. Sci. 2021, 11, 8362. [Google Scholar] [CrossRef]
- Salmasi, F.; Nouri, M.; Sihag, P.; Abraham, J. Application of SVM, ANN, GRNN, RF, GP and RT models for predicting discharge coefficients of oblique sluice gates using experimental data. Water Supply 2021, 21, 232–248. [Google Scholar] [CrossRef]
- Sangeeta; Haji Seyed Asadollah, S.B.; Sharafati, A.; Sihag, P.; Al-Ansari, N.; Chau, K.W. Machine learning model development for predicting aeration efficiency through Parshall flume. Eng. Appl. Comput. Fluid Mech. 2021, 15, 889–901. [Google Scholar] [CrossRef]
- Sihag, P.; Al-Janabi, A.M.S.; Alomari, N.K.; Ab Ghani, A.; Nain, S.S. Evaluation of tree regression analysis for estimation of river basin discharge. Modeling Earth Syst. Environ. 2021, 1–13. [Google Scholar] [CrossRef]
- Miluccio, G.; Losanno, D.; Parisi, F.; Cosenza, E. Traffic-load fragility models for prestressed concrete girder decks of existing Italian highway bridges. Eng. Struct. 2021, 249, 113367. [Google Scholar] [CrossRef]
- Indian Standard 4031-1996. Methods of Physical Tests for Hydraulic Cement; Bureau of Indian Standards: New Delhi, India, 1996.
- Indian Standard 456-2000. Indian Standard Code of Practice for General Structural Use of Plain and Reinforced Concrete; Bureau of Indian Standards: New Delhi, India, 2000.
- Indian Standard 10262-2009. Indian Standards in Concrete Mix Design; Bureau of Indian Standards: New Delhi, India, 2009.
| S. No. | Test Property | Result | Requirements as per IS 12269-1987 |
|---|---|---|---|
| 1 | Fineness
| ||
| 2% | Not more than 10% | ||
| 285 m2/kg | Min 225 m2/kg | ||
| 2 | Normal Consistency | 31.0% | - |
| 3 | Specific Gravity | 3.01 | - |
| 4 | Initial setting time | 95 min | Not less than 30 min |
| 5 | Final setting time | 284 min | Not more than 600 min |
| 6 | Compressive strength
| ||
| 28 N/mm2 | 27 N/mm2 (min) | ||
| 41 N/mm2 | 37 N/mm2 (min) | ||
| 56 N/mm2 | 53 N/mm2 (min) | ||
| 7 | Soundness(Le-Chatlier Exp.) | 2 mm | Not more than 10 mm |
| S. No. | Property | Test Value |
|---|---|---|
| 1 | Specific Gravity | 2.71 |
| 2 | Water absorption | 0.5% |
| 3 | Sieve Analysis Test results | Grading Curve shown in Graph 3.2 |
| 4 | Aggregate Impact Value, % | 21.50 |
| 5 | Aggregate crushing value, % | 20.40 |
| 6 | Combined Flakiness & Elongation Value, % | 21.90 |
| Mixture No | RBC00 | RBC05 | RBC10 | RBC15 | CBC00 | CBC05 | CBC10 | CBC15 |
|---|---|---|---|---|---|---|---|---|
| Cement (kg/m3) | 340 | 340 | 340 | 340 | 340 | 340 | 340 | 340 |
| River Sand (kg/m3) | 736 | 736 | 736 | 736 | - | - | - | - |
| Crushed Rock Sand (kg/m3) | - | - | - | - | 736 | 736 | 736 | 736 |
| Coarse Aggregate (kg/m3) | 1214 | 1214 | 1214 | 1214 | 1214 | 1214 | 1214 | 1214 |
| w/c ratio | 0.48 | 0.48 | 0.48 | 0.48 | 0.48 | 0.48 | 0.48 | 0.48 |
| Bacterial Cells (CFU/mL) | 105 | 105 | 105 | 105 | 105 | 105 | 105 | 105 |
| Percent of bacterial solution | 00 | 05 | 10 | 15 | 00 | 05 | 10 | 15 |
| Mixture No | RBC00 | RBC05 | RBC10 | RBC15 | CBC00 | CBC05 | CBC10 | CBC15 |
|---|---|---|---|---|---|---|---|---|
| Cement (kg/m3) | 390 | 390 | 390 | 390 | 390 | 390 | 390 | 390 |
| River Sand (kg/m3) | 642 | 642 | 642 | 642 | - | - | - | - |
| Crushed Rock Sand (kg/m3) | - | - | - | - | 642 | 642 | 642 | 642 |
| Coarse Aggregate (kg/m3) | 1261 | 1261 | 1261 | 1261 | 1261 | 1261 | 1261 | 1261 |
| w/c ratio | 0.42 | 0.42 | 0.42 | 0.42 | 0.42 | 0.42 | 0.42 | 0.42 |
| Bacterial Cells (CFU/mL) | 105 | 105 | 105 | 105 | 105 | 105 | 105 | 105 |
| Percent of bacterial solution | 00 | 05 | 10 | 15 | 00 | 05 | 10 | 15 |
| Input and Output Parameters | Mean | Standard Deviation | Minimum | Maximum | Confidence Level (95.0%) | Data Set |
|---|---|---|---|---|---|---|
| Cement | 365.00 | 25.10 | 340 | 390 | 4.39 | Overall |
| 364.71 | 25.14 | 340 | 390 | 5.36 | Training | |
| 365.61 | 25.30 | 340 | 390 | 7.99 | Testing | |
| Sand | 689.00 | 47.18 | 642 | 736 | 8.25 | Overall |
| 689.54 | 47.27 | 642 | 736 | 10.07 | Training | |
| 687.85 | 47.57 | 642 | 736 | 15.01 | Testing | |
| Aggregate | 1237.50 | 23.59 | 1214 | 1261 | 4.12 | Overall |
| 1237.23 | 23.63 | 1214 | 1261 | 5.03 | Training | |
| 1238.07 | 23.78 | 1214 | 1261 | 7.50 | Testing | |
| W/C | 0.45 | 0.03 | 0.42 | 0.48 | 0.01 | Overall |
| 0.45 | 0.03 | 0.42 | 0.48 | 0.01 | Training | |
| 0.45 | 0.03 | 0.42 | 0.48 | 0.01 | Testing | |
| Curing period | 126.75 | 124.35 | 7 | 365 | 21.75 | Overall |
| 124.97 | 124.29 | 7 | 365 | 26.49 | Training | |
| 130.54 | 125.93 | 7 | 365 | 39.75 | Testing | |
| BC | 0.08 | 0.06 | 0 | 0.15 | 0.01 | Overall |
| 0.07 | 0.06 | 0 | 0.15 | 0.01 | Training | |
| 0.08 | 0.06 | 0 | 0.15 | 0.02 | Testing | |
| Kind of Sand | 1.50 | 0.50 | 1 | 2 | 0.09 | Overall |
| 1.47 | 0.50 | 1 | 2 | 0.11 | Training | |
| 1.56 | 0.50 | 1 | 2 | 0.16 | Testing | |
| Compressive strength (MPa) | 45.70 | 12.70 | 21.56 | 74.46 | 2.22 | Overall |
| 45.17 | 12.16 | 21.56 | 71.12 | 2.59 | Training | |
| 46.82 | 13.87 | 24.16 | 74.46 | 4.38 | Testing |
| Approaches | CC | R2 | RMSE | MAE | Bias | SI | NSE |
|---|---|---|---|---|---|---|---|
| Training Stage | |||||||
| M5P | 0.96 | 0.92 | 4.90 | 2.77 | 0.20 | 0.11 | 0.92 |
| RF | 1.00 | 0.99 | 0.90 | 0.74 | 0.09 | 0.02 | 0.99 |
| RT | 1.00 | 1.00 | 0.13 | 0.06 | 0.00 | 0.00 | 1.00 |
| REP Tree | 0.97 | 0.94 | 2.89 | 2.32 | 0.00 | 0.06 | 0.94 |
| SVR_Poly | 0.99 | 0.98 | 1.57 | 0.79 | 0.24 | 0.03 | 0.98 |
| SVR_NPoly | 0.98 | 0.95 | 2.78 | 1.67 | 0.65 | 0.06 | 0.95 |
| SVR_PUK | 0.99 | 0.98 | 1.90 | 0.80 | 0.66 | 0.04 | 0.98 |
| SVR_RBF | 0.99 | 0.97 | 2.27 | 1.06 | 0.69 | 0.05 | 0.96 |
| MLR | 0.90 | 0.82 | 5.19 | 4.22 | 0.00 | 0.11 | 0.82 |
| Testing Stage | |||||||
| M5P | 0.97 | 0.94 | 4.88 | 2.88 | −0.10 | 0.10 | 0.93 |
| RF | 0.99 | 0.97 | 2.29 | 1.81 | 0.23 | 0.05 | 0.97 |
| RT | 0.98 | 0.96 | 2.82 | 2.49 | 0.86 | 0.06 | 0.96 |
| REP Tree | 0.96 | 0.92 | 3.81 | 2.97 | −0.35 | 0.08 | 0.92 |
| SVR_Poly | 0.99 | 0.98 | 1.94 | 1.52 | 0.14 | 0.04 | 0.98 |
| SVR_NPoly | 0.98 | 0.96 | 2.96 | 2.36 | 0.63 | 0.06 | 0.95 |
| SVR_PUK | 0.99 | 0.98 | 2.69 | 2.06 | −0.19 | 0.06 | 0.96 |
| SVR_RBF | 0.98 | 0.97 | 3.30 | 2.59 | −0.27 | 0.07 | 0.94 |
| MLR | 0.94 | 0.88 | 4.87 | 3.96 | −0.73 | 0.10 | 0.87 |
| Statistic | Actual | MLR | M5P | RF | RT | REP Tree | SVR_Poly | SVR_NPoly | SVR_PUK | SVR_RBF |
|---|---|---|---|---|---|---|---|---|---|---|
| Minimum | 24.16 | 27.95 | 27.08 | 23.99 | 23.02 | 24.33 | 24.78 | 28.15 | 26.27 | 28.00 |
| Maximum | 74.46 | 69.55 | 70.85 | 69.32 | 71.12 | 64.73 | 72.02 | 69.59 | 67.85 | 69.24 |
| 1st Quartile | 36.98 | 33.75 | 37.25 | 38.36 | 37.64 | 38.43 | 36.95 | 36.71 | 37.86 | 36.29 |
| Median | 44.76 | 47.82 | 47.75 | 45.36 | 45.43 | 42.66 | 45.20 | 45.07 | 45.46 | 45.37 |
| 3rd Quartile | 56.37 | 54.01 | 55.46 | 58.70 | 59.12 | 56.61 | 56.98 | 57.13 | 56.76 | 56.25 |
| Mean | 46.82 | 46.09 | 46.72 | 47.05 | 47.68 | 46.48 | 46.96 | 47.45 | 46.63 | 46.55 |
| IQR | 19.39 | 20.26 | 18.20 | 20.34 | 21.48 | 18.18 | 20.04 | 20.42 | 18.90 | 19.96 |
| Input Variable Combination | Target Variable | SVR_Poly | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Cement | Sand | Aggregate | W/C | Curing | BC | Kind of Sand | Compressive Strength (MPa) | CC | MAE | RMSE |
| 0.99 | 1.52 | 1.94 | ||||||||
| 0.98 | 1.82 | 2.47 | ||||||||
| 0.98 | 2.04 | 2.70 | ||||||||
| 0.98 | 1.82 | 2.45 | ||||||||
| 0.98 | 2.04 | 2.70 | ||||||||
| 0.80 | 7.21 | 8.59 | ||||||||
| 0.96 | 3.28 | 3.89 | ||||||||
| 0.98 | 2.48 | 2.88 | ||||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Almohammed, F.; Sihag, P.; Sammen, S.S.; Ostrowski, K.A.; Singh, K.; Prasad, C.V.S.R.; Zajdel, P. Assessment of Soft Computing Techniques for the Prediction of Compressive Strength of Bacterial Concrete. Materials 2022, 15, 489. https://doi.org/10.3390/ma15020489
Almohammed F, Sihag P, Sammen SS, Ostrowski KA, Singh K, Prasad CVSR, Zajdel P. Assessment of Soft Computing Techniques for the Prediction of Compressive Strength of Bacterial Concrete. Materials. 2022; 15(2):489. https://doi.org/10.3390/ma15020489
Chicago/Turabian StyleAlmohammed, Fadi, Parveen Sihag, Saad Sh. Sammen, Krzysztof Adam Ostrowski, Karan Singh, C. Venkata Siva Rama Prasad, and Paulina Zajdel. 2022. "Assessment of Soft Computing Techniques for the Prediction of Compressive Strength of Bacterial Concrete" Materials 15, no. 2: 489. https://doi.org/10.3390/ma15020489
APA StyleAlmohammed, F., Sihag, P., Sammen, S. S., Ostrowski, K. A., Singh, K., Prasad, C. V. S. R., & Zajdel, P. (2022). Assessment of Soft Computing Techniques for the Prediction of Compressive Strength of Bacterial Concrete. Materials, 15(2), 489. https://doi.org/10.3390/ma15020489








