Biomechanical Features of Graphene-Augmented Inorganic Nanofibrous Scaffolds and Their Physical Interaction with Viruses
Abstract
1. Introduction
2. Materials and Methods
2.1. Graphene-Augmentation on Alumina Nanofibers
2.2. Biomechanical Properties
2.3. Viruses and Their Preparation
2.4. Virus Binding to GAIN Scaffolds
2.5. Plaque Assay for IAV
2.6. Statistical Analysis
3. Results and Discussion
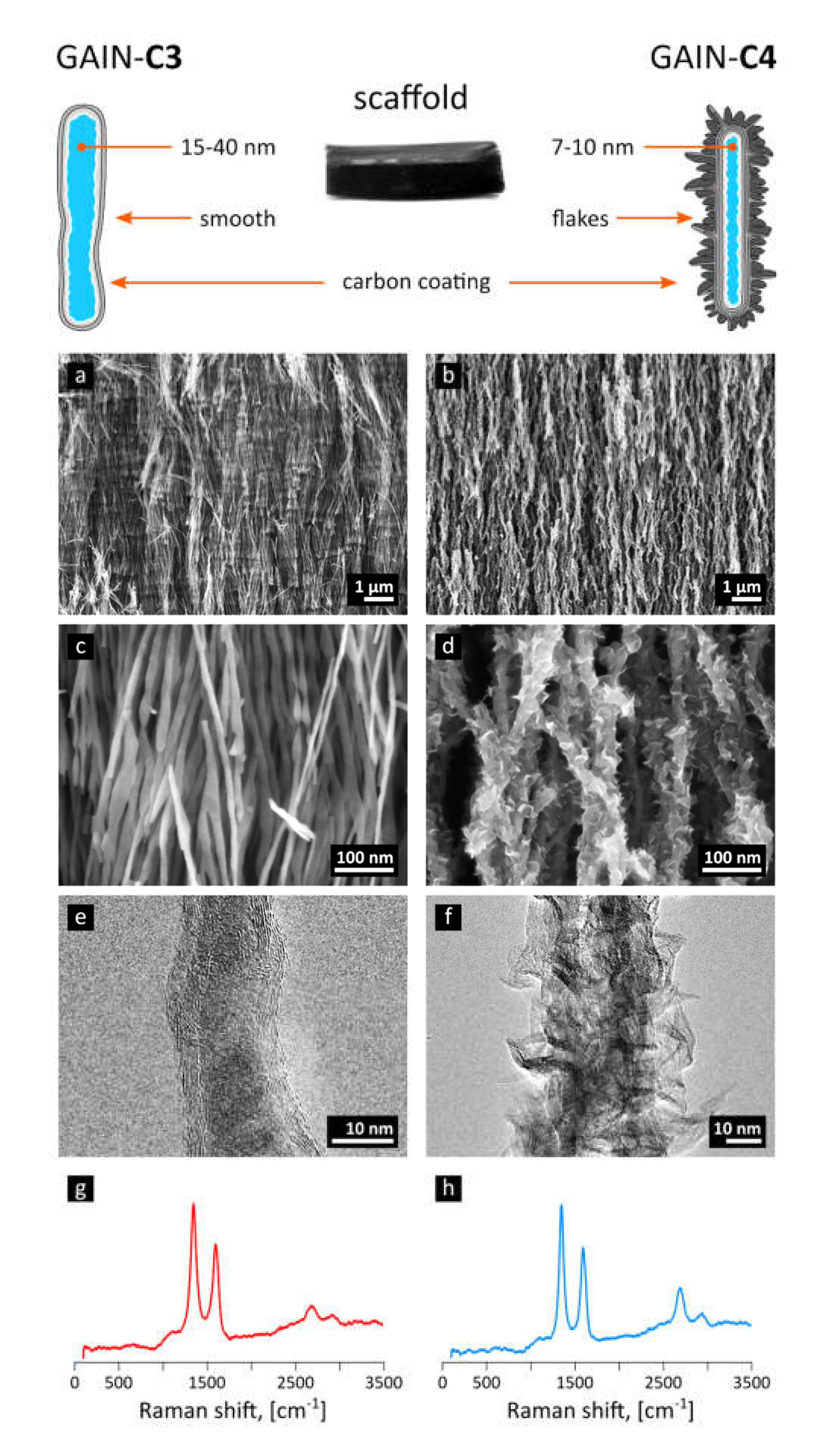
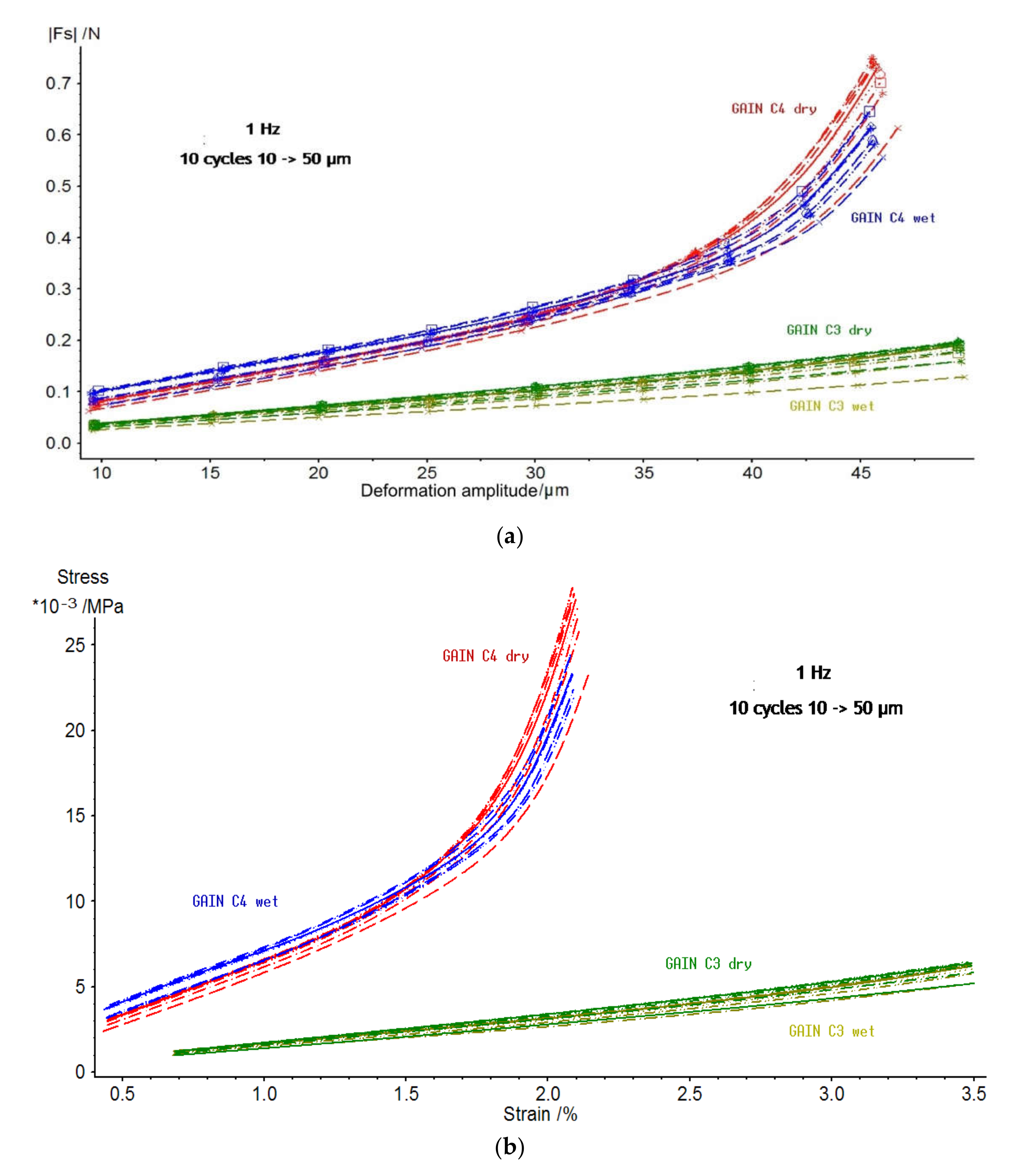
3.1. Properties of GAIN Scaffolds
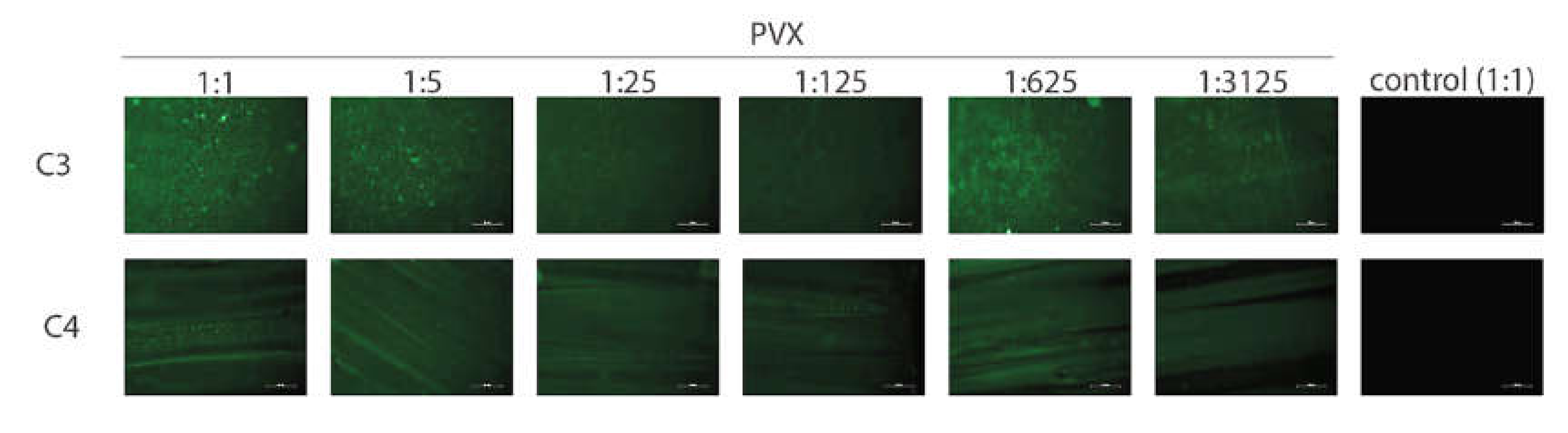
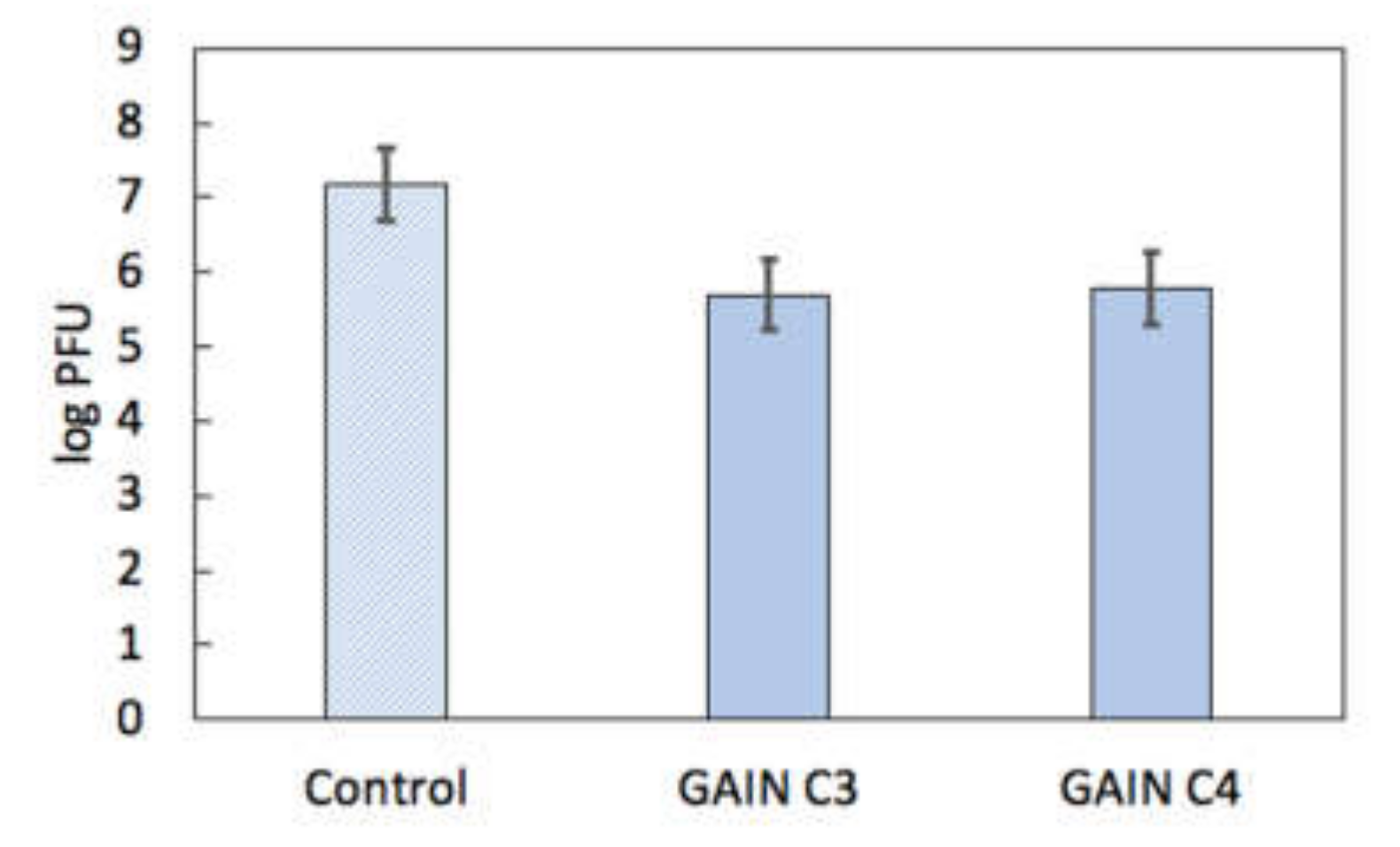
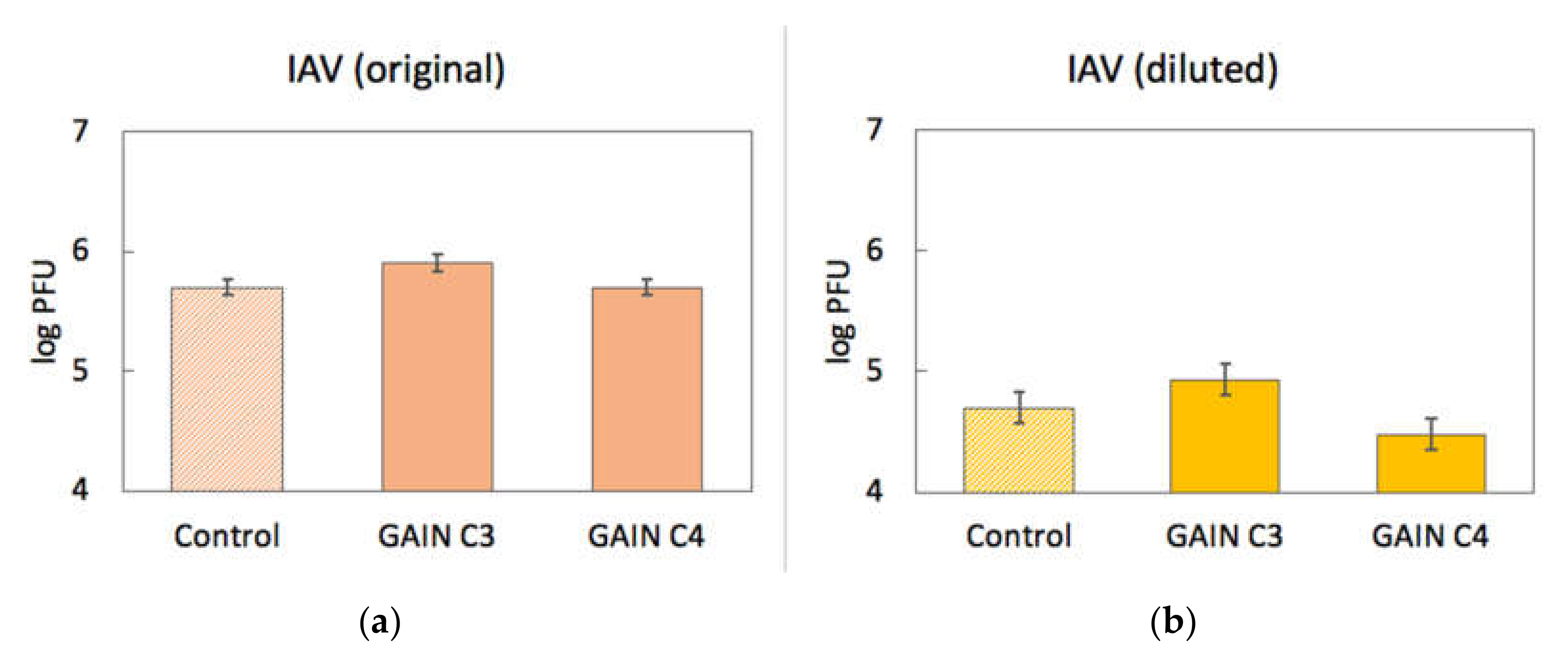
3.2. Virus Interactions with GAIN Scaffolds
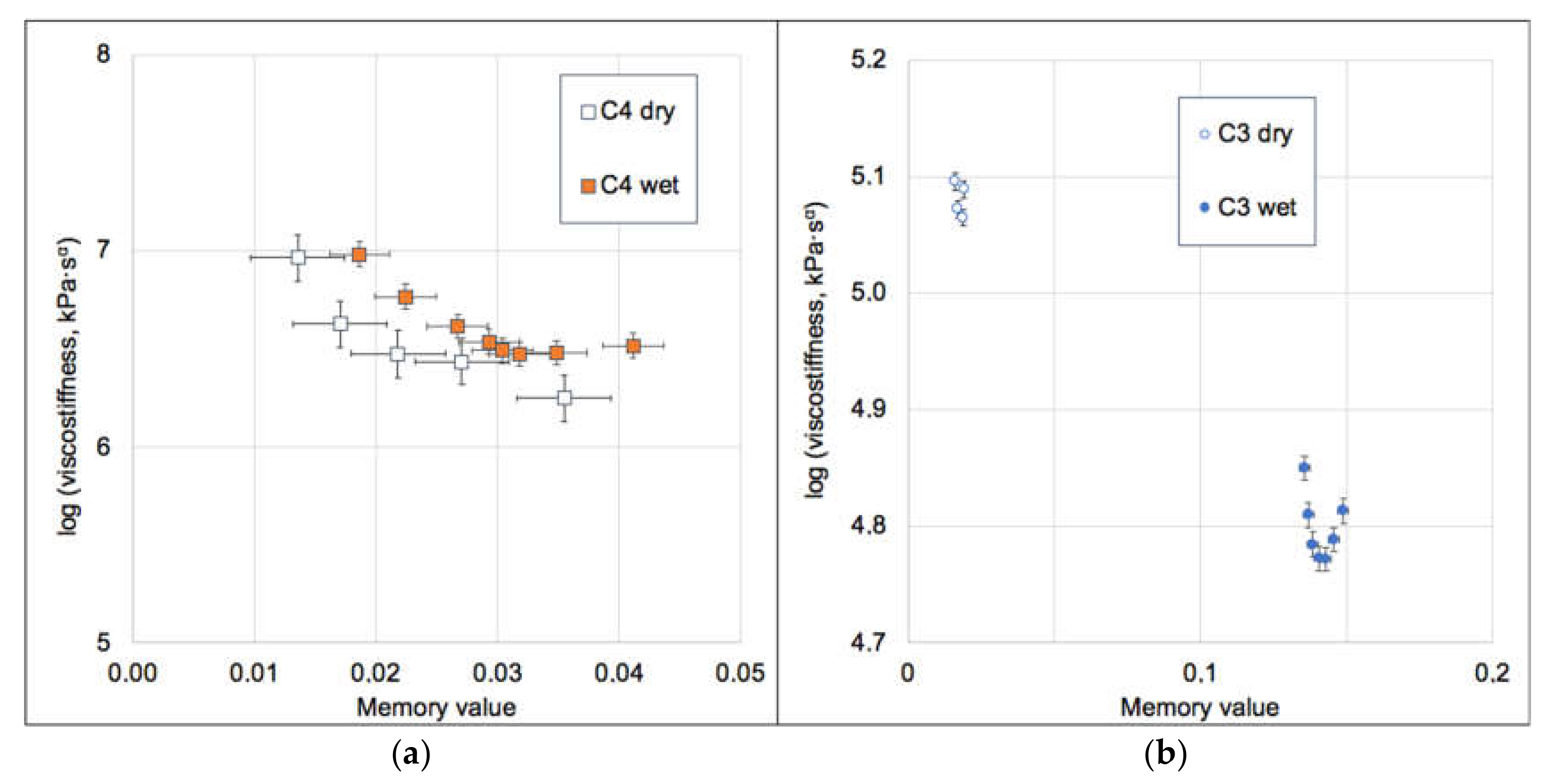
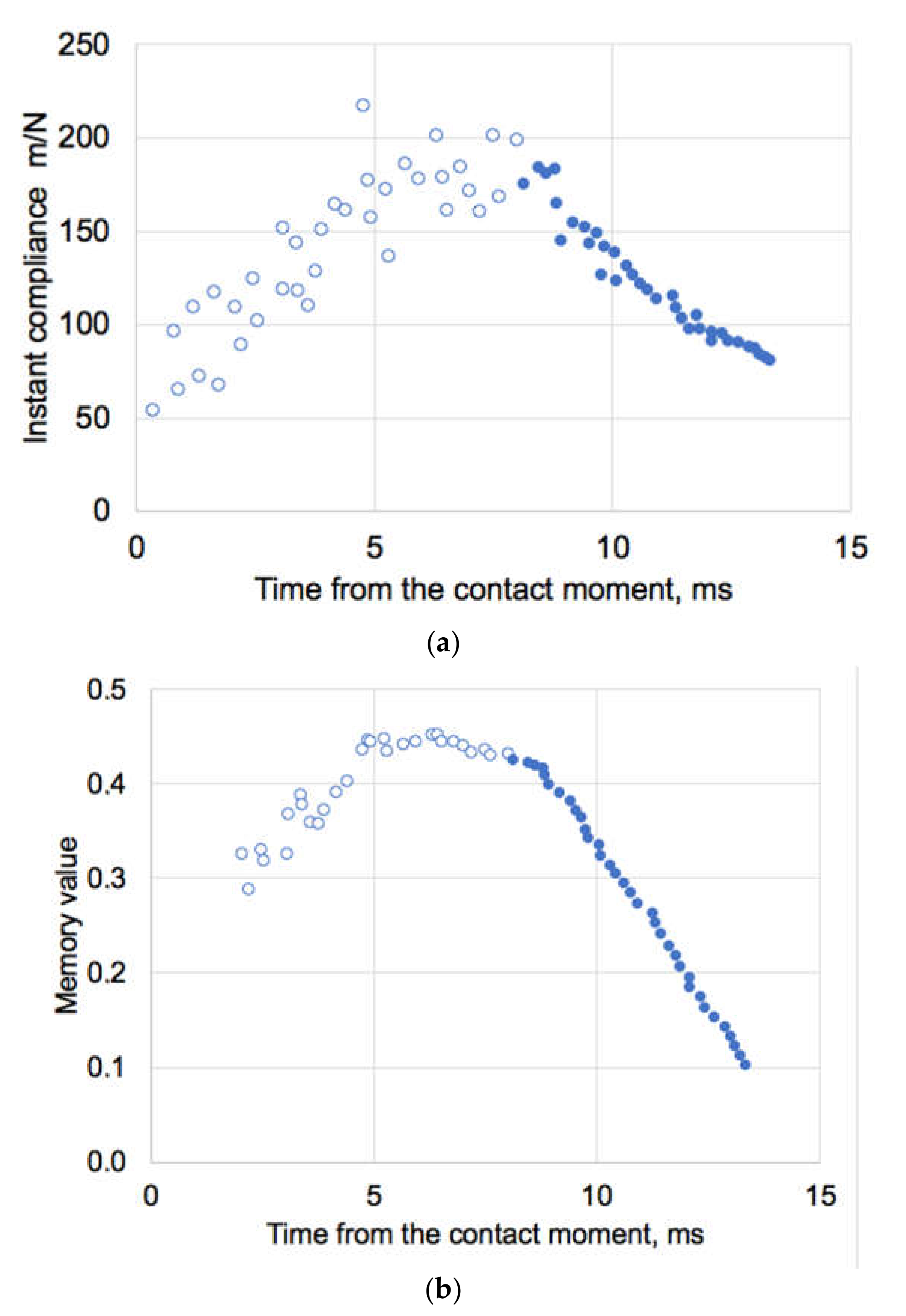
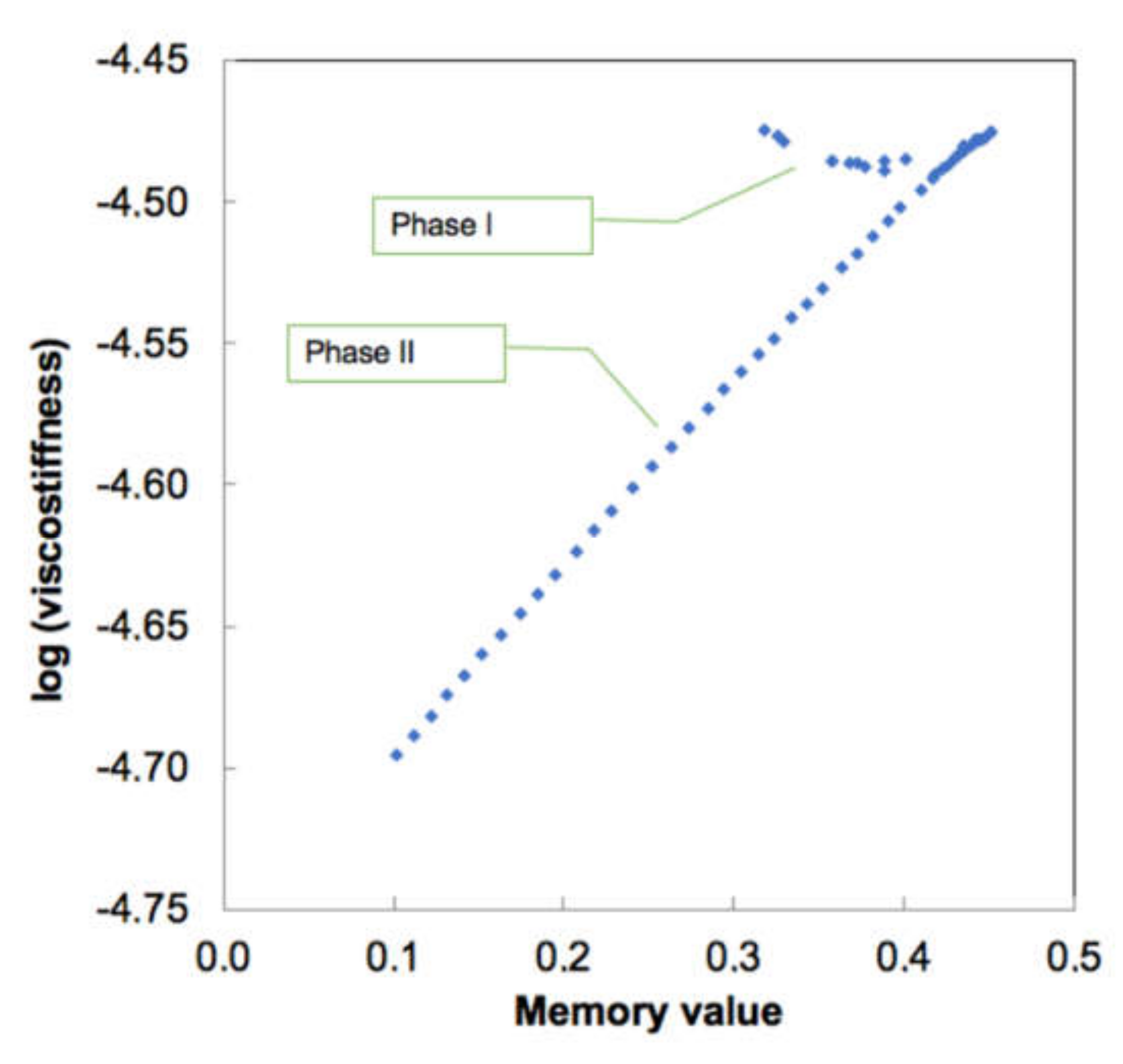
3.3. Biomechanics of IAV and Its Adherence
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- An, J.; Chua, C.K.; Yu, T.; Li, H.; Tan, L.P. Advanced nanobiomaterial strategies for the development of organized tissue en-gineering constructs. Nanomedicine 2013, 8, 591–602. [Google Scholar] [CrossRef] [PubMed]
- Di Marzio, N.; Eglin, D.; Serra, T.; Moroni, L. Bio-Fabrication: Convergence of 3D bioprinting and nano-biomaterials in tissue engineering and regenerative medicine. Front. Bioeng. Biotechnol 2020, 8, 326. [Google Scholar] [CrossRef] [PubMed]
- Ménard-Moyon, C. Applications of carbon nanotubes in the biomedical field. In Smart Nanoparticles Biomedicine; Ciofani, G., Ed.; Elsevier: Amsterdam, The Netherlands, 2018; pp. 83–101. [Google Scholar]
- Kazantseva, J.; Ivanov, R.; Gasik, M.M.; Neuman, T.; Hussainova, I. Graphene-augmented nanofiber scaffolds demonstrate new features in cells behaviour. Sci. Rep. 2016, 6, 30150. [Google Scholar] [CrossRef] [PubMed]
- Kazantseva, J.; Hussainova, I.; Ivanov, R.; Neuman, T.; Gasik, M. Hybrid graphene–ceramic nanofiber network for sponta-neous neural differentiation of stem cells. Interface Focus 2018, 8, 20170037. [Google Scholar] [CrossRef] [PubMed]
- Kazantseva, J.; Ivanov, R.; Gasik, M.; Neuman, T.; Hussainova, I. Graphene-augmented nanofiber scaffolds trigger gene ex-pression switching of four cancer cell types. ACS Biomater. Sci. Eng. 2018, 4, 1622–1629. [Google Scholar] [CrossRef]
- Hussainova, I.; Gasik, M.; Ivanov, R. Self-Aligned Fibrous Scaffolds for Auto-Mechanoinduction of Cell Cultures. U.S. Patent 10370640 B2, 6 August 2019. [Google Scholar]
- Gasik, M. Understanding biomaterial-tissue interface quality: Combined in vitro evaluation. Sci. Technol. Adv. Mater. 2017, 18, 550–562. [Google Scholar] [CrossRef]
- Melo-Fonseca, F.; Miranda, G.; Domingues, H.S.; Mendes Pinto, I.; Gasik, M.; Silva, F.S. Reengineering bone-implant interfaces for improved mechanotransduction and clinical outcomes. Stem Cell Rev. 2020. [Google Scholar] [CrossRef]
- Gasik, M. Biomechanical characterization of engineered tissues and implants for tissue/organ replacement applications. In Biomaterials for Organ and Tissue Regeneration; Vrana, N., Knopf-Marques, H., Barthes, J., Eds.; Woodhead Publishing: Cambridge, UK, 2020; pp. 599–627. [Google Scholar]
- Shin, J.W. Mooney DJ. Extracellular matrix stiffness causes systematic variations in proliferation and chemosensitivity in myeloid leukemias. Cell Stem Cell 2016, 18, 16–19. [Google Scholar] [CrossRef]
- Rimondini, L.; Gasik, M. Biomaterials and Immune Response, 1st ed.; CRC Press: Boca Raton, FL, USA, 2018. [Google Scholar]
- Kilian, K.A.; Bugarija, B.; Lahn, B.T.; Mrksich, M. Geometric cues for directing the differentiation of mesenchymal stem cells. Proc. Natl. Acad. Sci. USA 2010, 107, 4872–4877. [Google Scholar] [CrossRef]
- Carpenter, P.L. Microbiology, 3rd ed.; W. B. Saunders Co. Publ.: Pennsylvania, PA, USA, 1972; 494p. [Google Scholar]
- Tuthill, T.J.; Groppelli, E.; Hogle, J.M.; Rowlands, D.J. Picornaviruses. Curr. Top. Microbiol. Immunol. 2010, 343, 43–89. [Google Scholar] [CrossRef]
- Schwarz, B.; Uchida, M.; Douglas, T. Biomedical and catalytic opportunities of virus-like particles in nanotechnology. Adv. Virus Res. 2017, 97, 1–60. [Google Scholar] [PubMed]
- Wena, A.M.; Steinmetz, N.F. Design of virus-based nanomaterials for medicine, biotechnology, and energy. Chem. Soc. Rev. 2016, 45, 4074–4126. [Google Scholar] [CrossRef] [PubMed]
- Raja, I.S.; Kim, C.; Song, S.J.; Shin, Y.C.; Kang, M.S.; Hyon, S.H.; Oh, J.W.; Han, D.W. Virus-incorporated biomimetic nanocompo-sites for tissue regeneration. Nanomaterials 2019, 9, 14. [Google Scholar] [CrossRef] [PubMed]
- Kutuzov, M. Method and System for Alumina Nanofibers Synthesis from Molten Aluminum. U.S. Patent Application No. 13/756,366, 1 August 2013. [Google Scholar]
- Aghayan, M.; Hussainova, I.; Gasik, M.; Kutuzov, M.; Friman, M. Coupled thermal analysis of novel alumina nanofibers with ultrahigh aspect ratio. Thermoch. Acta. 2013, 574, 140–144. [Google Scholar] [CrossRef]
- Stamatin, S.N.; Hussainova, I.; Ivanov, R.; Colavita, P.E. Quantifying Graphitic Edge Exposure in Graphene-Based Materials and Its Role in Oxygen Reduction Reactions. ACS Catal. 2016, 6, 5215–5221. [Google Scholar] [CrossRef]
- Taleb, M.; Ivanov, R.; Bereznev, S.; Kazemi, S.H.; Hussainova, I. Ultra-sensitive voltammetric simultaneous determination of dopamine, uric acid and ascorbic acid based on a graphene-coated alumina electrode. Microchim. Acta. 2017, 184, 4603–4610. [Google Scholar] [CrossRef]
- Ivanov, R.; Hussainova, I.; Aghayan, M.; Drozdova, M.; Perez-Coll, D.; Rodriguez, M.A.; Rubio-Marcos, F. Graphene-encapsulated aluminium oxide nanofibers as a novel type of nanofillers for electroconductive ceramics. J. Eur. Ceram. Soc. 2015, 35, 4017–4021. [Google Scholar] [CrossRef]
- Gasik, M. In Vitro Test Method for Implant Materials. U.S. Patent 9683267 B2, 20 July 2017. [Google Scholar]
- Gasik, M.; Bilotsky, Y. In Vitro Method for Measurement and Model-Free Evaluation of Time-Invariant Biomaterials Func-Tions. U.S. Patent 10379106 B2, 13 August 2019. [Google Scholar]
- Gasik, M.; Zühlke, A.; Haaparanta, A.M.; Muhonen, V.; Laine, K.; Bilotsky, Y.; Kellomäki, M.; Kiviranta, I. The importance of controlled mismatch of biomechanical compliances of implantable scaffolds and native tissue for articular cartilage regen-eration. Frontiers Bioeng. Biotechnol. 2018, 6, 187. [Google Scholar] [CrossRef]
- Steinmetz, N.F.; Mertens, M.F.; Taurog, R.E.; Johnson, J.E.; Commandeur, U.; Fischer, R.; Manchester, M. Potato virus X as a nov-el platform for potential biomedical applications. Nano Lett. 2010, 10, 305–312. [Google Scholar] [CrossRef]
- Sokullu, E.; Abyaneh, H.S.; Gauthier, M.A. Plant/Bacterial Virus-Based Drug Discovery, Drug Delivery, and Therapeutics. Pharmaceutics 2019, 11, 211. [Google Scholar] [CrossRef]
- Lin, D.; Li, F.; Wu, Q.; Xie, X.; Wu, W.; Wu, J.; Chen, Q.; Liu, S.; He, J. A “building block” approach to the new influenza A virus entry inhibitors with reduced cellular toxicities. Sci. Rep. 2016, 6, 22790. [Google Scholar] [CrossRef] [PubMed]
- Shakeel, S.; Westerhuis, B.M.; Domanska, A.; Koning, R.I.; Matadeen, R.; Koster, A.J.; Bakker, A.Q.; Beaumont, T.; Wolthers, K.C.; Butcher, S.J. Multiple capsid-stabilizing interactions revealed in a high-resolution structure of an emerging picornavirus causing neonatal sepsis. Nature Commun. 2016, 7, 11387. [Google Scholar] [CrossRef] [PubMed]
- Chamings, A.; Druce, J.; Caly, L.; Yoga, Y.; Britton, P.N.; Macartney, K.K.; Alexandersen, S. Evolutionary analysis of human parechovirus type 3 and clinical outcomes of infection during the 2017–18 Australian epidemic. Sci. Rep. 2019, 9, 8906. [Google Scholar] [CrossRef] [PubMed]
- ImageJ Software. Available online: https://imagej.nih.gov/ij/ (accessed on 31 December 2020).
- Bagley, R.L.; Torvik, P.J. A theoretical basis for the application of fractional calculus to viscoelasticity. J. Rheol. 1983, 27, 201–210. [Google Scholar] [CrossRef]
- Rodríguez, R.F.; Salinas-Rodríguez, E.; Fujioka, J. Fractional time fluctuations in viscoelasticity: A comparative study of correlations and elastic moduli. Entropy 2018, 20, 28. [Google Scholar] [CrossRef] [PubMed]
- Coussot, C.; Kalyanam, S.; Yapp, R.; Insana, M.F. Fractional derivative models for ultrasonic characterization of polymer and breast tissue viscoelasticity. IEEE Trans. Ultrason. Ferroelectr. Freq. Control. 2009, 56, 715–726. [Google Scholar] [CrossRef]
- Schaap, I.A.T.; Eghiaian, F.; des Georges, A.; Veigel, C. Effect of envelope proteins on the mechanical properties of influenza virus. J. Biol. Chem. 2012, 287, 41078–41088. [Google Scholar] [CrossRef]
- Wang, H.; Wang, X.; Li, T.; Lee, B. Transient viscoelasticity study of tobacco mosaic virus/Ba2+ superlattice. Nanoscale Res. Lett. 2014, 9, 300. [Google Scholar] [CrossRef]
- Cieplak, M.; Robbins, M.O. Nanoindentation of 35 virus capsids in a molecular model: Relating mechanical properties to structure. PLoS ONE 2013, 8, e063640. [Google Scholar] [CrossRef]
- Park, Y.J.; Walls, A.C.; Wang, Z.; Sauer, M.M.; Li, W.; Tortorici, M.A.; Bosch, B.J.; DiMaio, F.; Veesler, D. Structures of MERS-CoV spike glycoprotein in complex with sialoside attachment receptors. Nature Struct. Mol. Biol. 2019, 26, 1151–1157. [Google Scholar] [CrossRef]

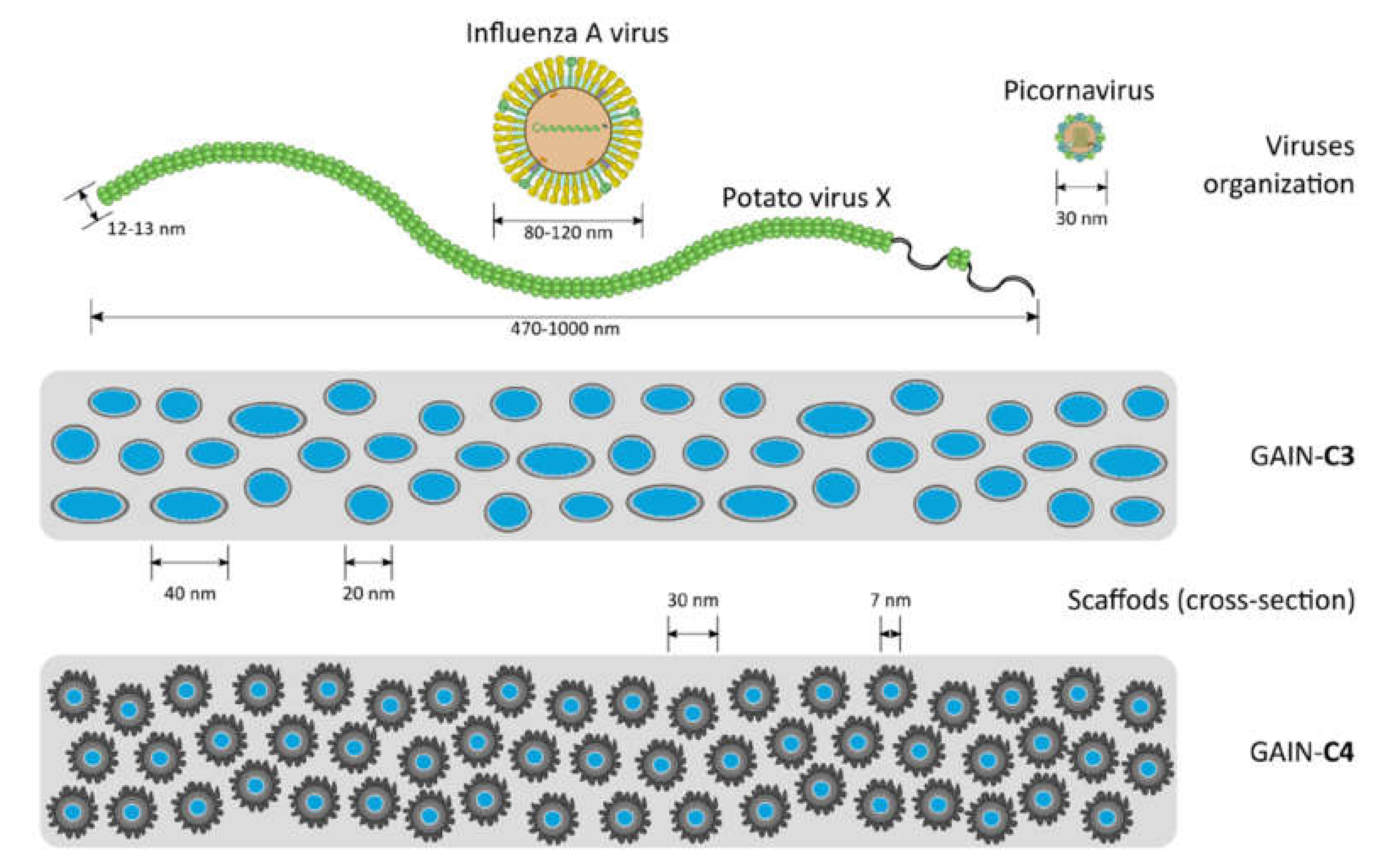

| Type | Characteristic | Envelope | Dimensions |
|---|---|---|---|
| Potato virus X (PVX) | Plant pathogen | No | 500–1000 × 10–15 nm, helical rods |
| Influenza A virus (IAV) | Orthomyxoviridae; human pathogen; ether sensitive | Yes | 80–120 nm, helical capsid |
| Human parechovirus (HPeV) | Picornaviridae; neonatal pathogen; non-sensitive to ether | No | 20–30 nm, cubic capsid |
| Condition | Dynamic Stress/Strain Ratio, kPa | |
|---|---|---|
| C3 Scaffold | C4 Scaffold | |
| Dry | 176.39 × (6.38 × 10−3 × ω)0.1455ε + 0.0152 | 1570.30 × (3.67 × 10−16 × ω) −1.2663ε + 0.0397 |
| Wet | 163.40 × (1.22 × 10−1× ω)−0.6330ε + 0.1546 | 1695.60 × (5.18 × 10−13 × ω) −1.4018ε + 0.0497 |
| Virus Type | Scaffold Pretreatment | Adherence to | |
|---|---|---|---|
| C3 Scaffold | C4 Scaffold | ||
| PVX | Does not affect | Very good | Very good |
| HPeV1 | - | Good | Good |
| IAV | - | No | No |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gasik, M.; Ivanov, R.; Kazantseva, J.; Bilotsky, Y.; Hussainova, I. Biomechanical Features of Graphene-Augmented Inorganic Nanofibrous Scaffolds and Their Physical Interaction with Viruses. Materials 2021, 14, 164. https://doi.org/10.3390/ma14010164
Gasik M, Ivanov R, Kazantseva J, Bilotsky Y, Hussainova I. Biomechanical Features of Graphene-Augmented Inorganic Nanofibrous Scaffolds and Their Physical Interaction with Viruses. Materials. 2021; 14(1):164. https://doi.org/10.3390/ma14010164
Chicago/Turabian StyleGasik, Michael, Roman Ivanov, Jekaterina Kazantseva, Yevgen Bilotsky, and Irina Hussainova. 2021. "Biomechanical Features of Graphene-Augmented Inorganic Nanofibrous Scaffolds and Their Physical Interaction with Viruses" Materials 14, no. 1: 164. https://doi.org/10.3390/ma14010164
APA StyleGasik, M., Ivanov, R., Kazantseva, J., Bilotsky, Y., & Hussainova, I. (2021). Biomechanical Features of Graphene-Augmented Inorganic Nanofibrous Scaffolds and Their Physical Interaction with Viruses. Materials, 14(1), 164. https://doi.org/10.3390/ma14010164







