Abstract
Microbial electrochemical technologies have revealed the opportunity of electrochemical enrichment for specific bacterial groups that are able to catalyze reactions of interest. However, there are unsolved challenges towards their application under aggressive environmental conditions, such as in the sea. This study demonstrates the impact of surface electrochemical potential on community composition and its corrosivity. Electrochemical bacterial enrichment was successfully carried out in natural seawater without nutrient amendments. Experiments were carried out for ten days of exposure in a closed-flow system over 316L stainless steel electrodes under three different poised potentials (−150 mV, +100 mV, and +310 mV vs. Ag/AgCl). Weight loss and atomic force microscopy showed a significant difference in corrosion when +310 mV (vs. Ag/AgCl) was applied in comparison to that produced under the other tested potentials (and an unpoised control). Bacterial community analysis conducted using 16S rRNA gene profiles showed that poised potentials are more positive as +310 mV (vs. Ag/AgCl) resulted in strong enrichment for Rhodobacteraceae and Sulfitobacter. Hence, even though significant enrichment of the known electrochemically active bacteria from the Rhodobacteraceae family was accomplished, the resultant bacterial community could accelerate pitting corrosion in 316 L stainless steel, thereby compromising the durability of the electrodes and the microbial electrochemical technologies.
1. Introduction
Microbial electrochemical technologies (METs) take advantage of microbial communities to catalyze redox reactions over electrodes [1,2,3,4]. Even though the first generation of METs were designed to produce electricity, their efficiency as wastewater treatment systems has also been proven [3,5,6,7]. The next generation of METs were then conceived to accomplish selective and energy-efficient treatment, remediation, and recovery of nutrients, metals, and by-products with industrial value [8]. However, many of these novel and disruptive METs have only been tested at the laboratory scale and under controlled conditions. Therefore, some of the main ongoing challenges for METs are to overcome the practical limitations associated with (i) the performance requirements of cost-efficient materials used as electrodes, and (ii) the enrichment of specific microbial communities which can catalyze reactions of interest in a MET under natural, uncontrolled environmental conditions [9,10], which is related to the use of METs in the implementation of bioremediation strategies.
The durability of electrode materials under exposure to aggressive natural environments where complex microbial communities can develop (e.g., coastal areas) is a major concern for the scaling-up of METs. Due to their biocompatibility and corrosion resistance, carbon-based materials (e.g., graphite and carbon felt) are commonly used as electrode materials in laboratory scale METs. However, in pilot systems, they exhibit limitations in terms of resistivity, mechanical strength, and cost [10,11,12]. Stainless steel (SS) is a common industrial material characterized by its good mechanical properties, corrosion resistance, conductivity, cost efficiency, and easy scalability [13,14,15]. Pocaznoi et al. (2012) [16] compared the abilities of carbon cloth, graphite plate, and SS as bioanodes in bioelectrochemical reactors (BERs), concluding that SS electrodes presented the highest current density among the tested materials. Furthermore, SS electrodes have been successfully proven to be effective as both anodes and cathodes in marine METs [14], producing power levels similar to those produced with graphite electrodes (0.2 W m−2) [17]. Thus, although the material evaluation of METs under real service conditions is in its infancy, SS emerges as a good candidate as an electrode material for the scaling-up of METs in marine environments.
Poised potential experiments have been used to study the effect of electrode potential on microbial colonization of the electrode in vitro [9,18,19,20,21]. This strategy has been proven by several authors to be able to modulate and enrich the presence of particular electrochemically active bacteria (EAB) which can transform contaminants in a BER. Indeed, while negative potentials have been shown to enrich heterotrophic exoelectrogenic bacteria [18,21], the use of a positive potential tends to enrich autotrophic microorganisms that are able to directly or indirectly receive electrons from an electrode [20]. In general, these types of experiments have been developed in culture media under controlled conditions using an inoculum obtained from wastewater treatment plants or anaerobic soil, where the microorganisms of interest are found in high abundance [22]. In consequence, it is necessary to evaluate the effectiveness of the application of an overpotential to an electrode as a tool to enrich the presence of EAB in METs. To determine if this tool could help with the scalability and applicability of MET in real environments, this evaluation should be accomplished under exposure to complex natural environments such as coastal marine environments.
In this work, we investigated whether the durability of SS electrodes exposed to a marine environment changes according to the applied overpotential and the enrichment of specific bacteria, as well as the application of an overpotential as a microbial enrichment tool to improve the applicability and scalability of MET. To address the proposed objective, a complete and multi-technique approach was used to evaluate the effect of different overpotentials on both the corrosion and the biological colonization of SS as an electrode material for METs operating in a coastal marine environment. Our results demonstrated differential electrochemical bacterial enrichment under the different overpotentials tested. In particular, the highest positive potential led to clear enrichment in members of the Roseobacter/Rhodobacter clade, and exerted a strong effect on deterioration of the SS electrodes.
2. Materials and Methods
2.1. Reactor Configuration
In this work, two sets of experiments were conducted. The first experiment was carried out to determine the effect of overpotential application on microbial enrichment from natural seawater samples, and the second experiment was conducted to isolate the effects of both the applied potential (abiotic) and the enriched microorganisms (biotic) on the metal corrosion.
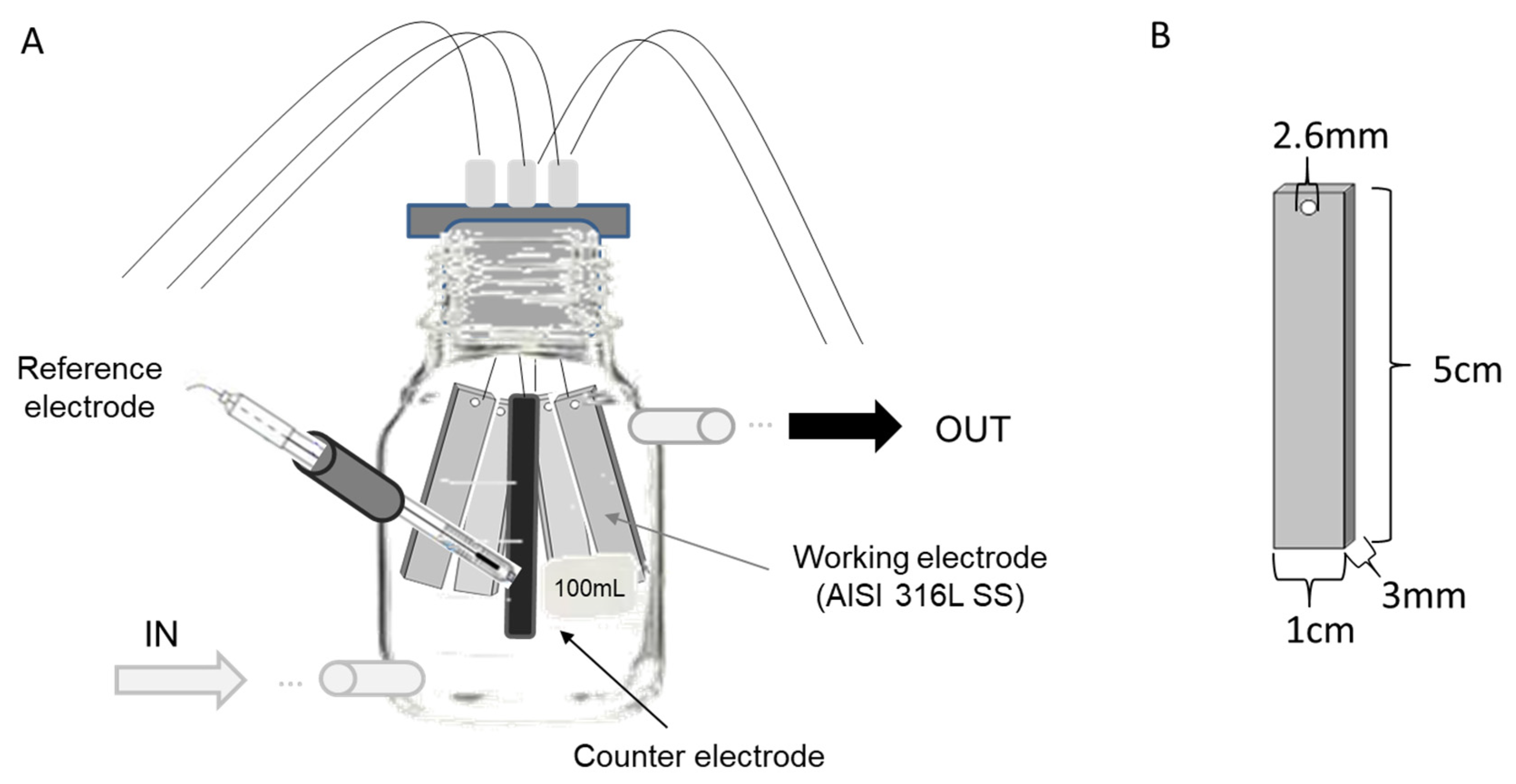
The first experiment was run using four 100 mL glass bottles as three-electrode electrochemical cells, which were modified to allow a continuous closed flow of natural seawater during the experiment (Figure 1). Each bottle contained four AISI 316L (X2CrNiMo17-12-2, material number 1.4404) SS electrodes as the working electrode (1.5 cm3 each), one graphite rod as the counter electrode (2 cm3), and one Ag/AgCl electrode as the reference electrode (3 M KCl) (Figure 1). This experimental design was based on the reactor designed by [21]. Before seawater exposure, each SS plate was polished using emery paper (grit 240), rinsed with distilled water, degreased with acetone, and finally dried by blowing hot air. An Interface 1000TM potentiostat (GAMRY, Warminster, PA, USA) connected to an ECM8™ electrochemical multiplexer (GAMRY, Warminster, PA, USA) was used to poise each cell at one of three specific potentials over the ten-day course of the experiments. The potentials were: (i) −150 mV (vs. Ag/AgCl), representing the reported optimal applied potential for enriching marine EAB [9]; (ii) +100 mV (vs. Ag/AgCl), corresponding to the open circuit potential (OCP) reported for 316L SS under marine conditions after 10 days [23] and allowing for the enrichment of autotrophic exoelectrotrophic microorganisms on the electrodes [20]; and (iii) +310 mV (vs. Ag/AgCl), representing a near potential of one of most-positive potentials reported for the oxygen reduction reaction [24]. As a control, a fourth electrochemical cell was run under OCP conditions.
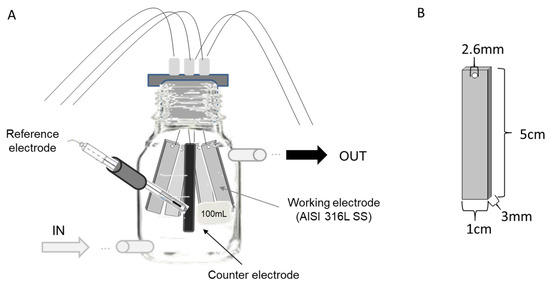
Figure 1.
Experimental design. (A) Design of the bottles, indicating seawater inlet and outlet and the arrangement of the AISI 316L Stainless Steel (SS) electrodes, reference electrode, and counter electrode. (B) Dimensions of SS electrodes.
The second experiment involved the same experimental setup as that used in the first, but two cells were set under abiotic conditions, with 0.2 µm-filtered natural seawater. Inside of one of these two cells the working electrode was polarized at +310 mV (vs. Ag/AgCl) and the other was run as the control under OCP conditions. A third electrochemical cell was tested under biotic conditions (using unfiltered seawater) and the working electrode was poised at +310 mV (vs. Ag/AgCl).
For both experiments, fresh, natural coastal seawater was collected from the Estación Costera de Investigaciones Marinas (ECIM) of the Pontificia Universidad Católica de Chile, located in the Eastern Pacific Ocean at 33°30′16′′ S, 71°38′23′′ W. Water was stored in a temperature-controlled room at 20 °C. Air pumps and a light schedule (12 h/12 h) were used to maintain saturated conditions of phototrophic activity and dissolved oxygen (8 mg L−1) during the experiment. Electrochemical cells were connected to a tank in a closed-loop system. A peristaltic pump was used to maintain a constant flow of seawater (0.5 mL s−1) throughout the 10 days of the experiment.
2.2. Surface Analysis for Corrosion Evaluation
2.2.1. Weight Loss
To assess weight loss, each coupon was weighed before the assembly of the experiments with an analytical balance with a precision of 0.1 mg (Shimadzu ATY224, Kyoto, Japan). The samples analyzed were the three SS electrodes after they were first sonicated for community analysis (described in Section 2.3). After sonication, the SS electrodes were washed, degreased with acetone, and dried with hot air to be reweighed. A one-way analysis of variance (ANOVA), followed by an a posteriori Tukey test, were performed using GraphPad Prism version 6 to check for significant differences between treatments [25].
2.2.2. Atomic Force Microscopy (AFM)
For the second experiment, after the cleaning of coupons to remove the biofilm, the topographies of the surfaces of the SS electrodes were observed using atomic force microscopy (AFM). The measurements were performed with an Innova® Atomic Force Microscope (Bruker, Billerica, MA, USA). Each coupon was measured in triplicate at different points to obtain a representative analysis. The topography was measured in air, using the tapping mode with a silicon nitride probe. Roughness analysis was carried out on the images obtained, over an area of 35 mm × 35 mm.
2.3. Microbial Community Analysis
2.3.1. DNA Extraction
After the exposure time, three of the four SS electrodes in each bottle were removed and sonicated in 50 mL of sterile seawater for 5 min (Elmasonic S 30H, Elma Schmidbauer GmbH, Frechen, Germany). The sonicated products were filtered through a 0.2-µm membrane filter to collect the biomass and perform the subsequent DNA extraction with phenol:chloroform based on a previously reported protocol [26]. Additionally, at the end of the experiment, 2 L of water from the tank was filtered through a 0.2-µm membrane filter to assess changes in the microbial community of the water column present in the tank. DNA quantification was performed with a Qubit® 2.0 fluorometer (InvitrogenTM, Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s protocol.
2.3.2. Fragment Analysis (FA) of 16S rRNA Genes
To identify the microbial community composition developed on the tested electrodes, amplification of 16S rRNA genes was performed in all DNA samples. The 16S rRNA genes were amplified by polymerase chain reaction (PCR), using the 5′-NED-labeled primer 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) and the primer 1492R (5′-GGYTACCTTGTTACGACTT-3′), without fluorescent labels [27]. The final concentrations in the PCR mix were 1.2 mM MgCl2, 1X KAPA PCR buffer B (KB1002), 0.3 mM dNTPs, 0.3 μM for each primer, and 1 U KAPA Taq polymerase (KAPABiosystems, Cape Town, South Africa). PCR conditions were as follows: 95 °C for 5 min, 25 cycles of 95 °C for 45 s, 56 °C for 45 s, and 72 °C for 1 min, and a final elongation step at 72 °C for 7 min. 16S rRNA gene PCR amplicons (1492 bp in length) were checked by electrophoresis on a 1% agarose gel. PCR products (17.5 μL) were digested independently with 2.5 U of HaeIII restriction endonuclease and incubated overnight at 37 °C. Terminal restriction fragments (T-RFs) were detected using capillary electrophoresis (Macrogen, Seúl, Korea), and data were obtained using Peak Scanner software (v1.0, AB Applied Biosystems, Foster City, CA, USA). The sizes of the T-RFs were determined using the internal standard LIZ1200 (Applied Biosystems, Rotkreuz, Switzerland). Data analysis included the T-RFs between 50 and 900 nucleotides and excluded those representing less than 0.5% of the total area/abundance [28], thus recalculating the area of each selected T-RF as a proportion of the total area.
2.3.3. 16S rRNA Gene Clone Library Construction
To identify the different operational taxonomic units (OTUs) observed in the Terminal Restriction Fragment Length Polymorphism (T-RFLP) profiles, a clone library was created for the 16S rRNA gene. For this purpose, amplification and subsequent cloning of the 16S rRNA gene were performed from the DNA of each sample using primers 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) and 1492R (5′-GGYTACCTTGTTACGACTT-3′) [27]. The PCR products obtained were cloned into the vector TOPO-TA using a mixture containing 4 µL of PCR product, 1 µL of salt solution, and 1 µL of cloning vector (Cloning Kit R for Sequencing, Invitrogen). This mixture was incubated at room temperature for 20 min, and then 45 µL of competent cells of Escherichia coli (DH5α) were added and incubated in ice for 20 min. The mixture was then placed in the thermoregulated bath at 42 °C for 50 s and finally placed in ice for 2 min. After transformation, the mixture was incubated in 950 µL of Super Optimal Broth (SOB) medium (20 mM glucose, 20 g L−1 tryptone, 5 g L−1 yeast extract, 0.5 g L−1 NaCl, 2.5 mM KCl, 5 mM MgCl2, and 5 mM MgSO4, pH 7.0) for 1.5 h at 37 °C with constant agitation. Afterward, 100 µL of the incubated solution was collected and seeded on LB agar plates supplemented with ampicillin (50 µg mL−1), 10 µL of X-gal (80 mg mL−1), and 40 µL of IPTG (500 mM). The seeded plates were incubated at 37 °C overnight.
To corroborate the presence of the fragment of interest in the cloning vector, cloned sequences were amplified by PCR using the M13 forward (M13F) (5′-GTAAAACGACGGCCAG-3′) and M13 reverse (M13R) (5′-CAGGAAACAGCTATGAC-3′) primers. The final concentrations in the PCR mix (final volume 25 µL) were: 1.2 mM MgCl2; 1X buffer without Mg2+; 0.25 mM dNTPs; 0.5 μM each of M13F and M13R; and 1.25 U KAPA Taq polymerase. PCR conditions were: 94 °C for 10 min; 25 cycles at 94 °C for 45 s; 50 °C for 1 min; 72 °C for 1 min; and a final elongation step at 72 °C for 7 min. PCR amplicons were checked by electrophoresis on a 1.5% agarose gel. Positive clones were confirmed based on their length of about 780 bp. Positive clones were sequenced at Macrogen Inc., Seoul, Korea. The obtained clone sequences are available in the Sequence Read Archive database (Table S1).
2.3.4. Analysis of Clone Sequences
Sequence Scanner Software v1.0 (AB Applied Biosystems, Foster City, CA, USA) was used to check the quality of the sequences. Sequences were then analyzed with VECTOR NTI software Version 11.5.4 (Thermo Fisher Scientific, Waltham, MA, USA) to eliminate the remaining vector sequence and leave the sequences in the 5′–3′ direction, considering the primer 27F as the beginning. Through this software, an in-silico analysis was also performed with the restriction enzyme HaeIII for the subsequent taxonomic assignment of the observed T-RFs. The T-RFs of each condition were assigned with the in-silico analysis of the sequenced clones for each condition, except in the +100 mV (vs. Ag/AgCl) condition. In this condition no clones were obtained, and only taxonomic assignment was made using the T-RFs. All edited sequences were compared to the published database (nr/nt) using BLASTn [29] from the National Center for Biotechnology and Information (NCBI, December 2018).
2.3.5. Amplicon Analysis of 16 rRNA Gene Sequences
To obtain a deeper taxonomic analysis of the microbial community enriched for under each set of conditions, amplicon analysis (AA) using next-generation sequencing (NGS) of the V4 hypervariable region of the 16S rRNA gene was performed with the Illumina MiSeq platform (Integrated Microbiome Resource (IMR), Toronto, ON, Canada). The sequences obtained in the present study are publicly available in the Sequence Read Archive database under the accession number PRJNA603906. The 16S rRNA gene amplicon sequences were processed using Mothur [30]. Sequences were de-multiplexed, assembled, and assigned to samples by matching them to barcode sequences using the make.contigs script and primers were removed using cutadapt [31]. Sequences with undesired lengths (200–300 bp), ambiguous nucleotides, and homopolymers longer than 8 bp were removed before further analysis. Afterward, sequences were aligned using the recreated Silva SEED v119 [32] as the reference. Chloroplast, eukaryotic, archaeal, and mitochondrial sequences were discarded. Sequences were also checked for PCR chimeras using UCHIME version 4.2.40 (mybiosoftware) [33]. High-quality sequences were clustered into operational taxonomic units (OTUs) with the furthest-neighbor algorithm, with a minimum sequence identity cut-off of 97%. Taxonomic assignments were performed against Silva v119 [32]. OTUs formed by 20 or fewer reads were not considered in subsequent analyses. For the OTUs that could not be identified at the gender level, a manual BLASTn was performed, wherein each sequence was compared to the published database (December 2018) (nr/nt) using BLASTn [29] from NCBI.
2.3.6. Statistical Analysis of Fragment and Amplicon Analysis Profiles
To determine the similarity between the samples and the robustness of the results obtained by FA and AA, fragment and amplicon profiles were statistically analyzed via multivariate analysis using Primer-E software version 6 (Primer-E, Plymouth, UK). For each dataset (FA and AA), the average of the relative abundance profile of each replicate of the OTU was transformed to its square root, and then a similarity matrix was obtained using the Bray–Curtis coefficient [34]. With the similarity matrix thus generated, a cluster was performed for visual interpretation of the grouping, where the proximity between samples corresponded to their similarity. To evaluate the statistical significance of the differences between and within communities, a one-way ANOSIM and a SIMPROF analysis were performed [35].
2.3.7. Similarity Analysis between Cloning and Amplicon Sequences
The sequences obtained via cloning and amplicon sequences were compared to determine the similarity between sequences and therefore the robustness of the community composition results. For this, the sequences of the most abundant OTUs, associated with the same bacterial genera in cloning and amplicon sequences, were used to perform a BLAST between sequences. With these results, a literature search was carried out to identify the metabolism and natural environment reported for each detected genus.
To determine the difference and/or similarity in bacterial identification and metabolic inference between samples, main bacterial taxa were identified using 16S rRNA gene sequences of clone library and AA results. Each sequence was taxonomically assigned by manual BLASTn of the sequence against the NCBI nt/nr database. For this analysis, only the families with a relative abundance higher than 1% of the total abundance in each sample were used.
2.3.8. Microscopy Analysis
The SS coupons used for microscopy were cut into two parts. One part was used for epifluorescence microscopy and the other for scanning electron microscopy (SEM). For epifluorescence microscopy, each coupon was immersed in 0.1% w/v acridine orange (Sigma Chemical Co., St. Louis, MO, USA) for 5 min at room temperature. Then, each coupon was washed with Phosphate Buffered Saline (PBS). Each coupon was examined using an Olympus CX31 microscope (Olympus, Tokyo, Japan). The other halves of the coupons were used for SEM analysis. For SEM analysis, samples were fixed with 2% glutaraldehyde, treated by critical-point drying, and coated with a thin gold film. Each coupon was examined using a LEO 1420VP microscope (LEO Electron Microscopy Inc., New York, NY, USA) [23]. For each coupon, three pictures were taken at randomly selected areas.
3. Results and Discussion
3.1. Electrode Deterioration
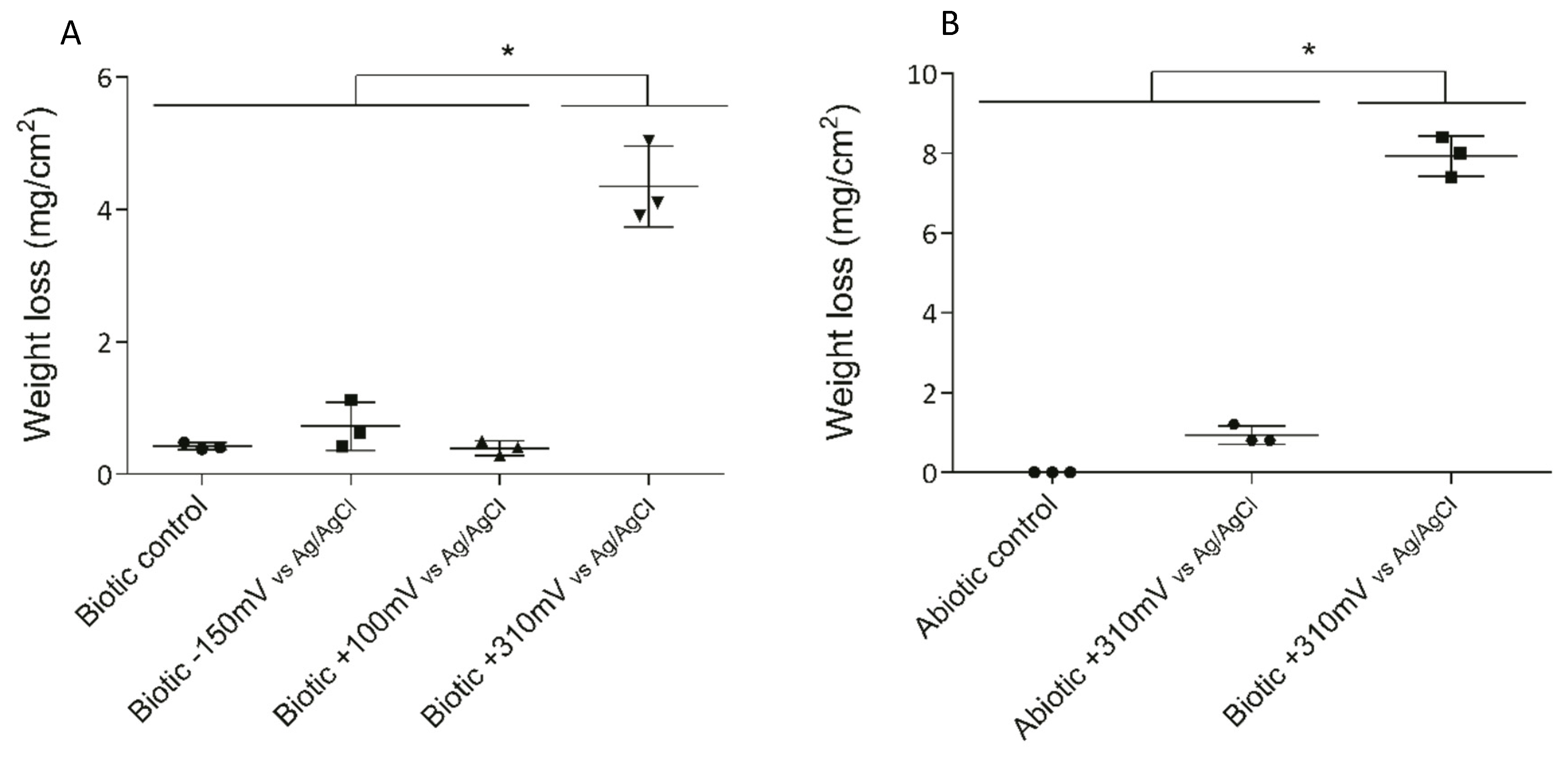
Three different potentials were applied to the 316L SS electrodes to determine the effect of overpotential application on electrode deterioration and bacterial enrichment. After 10 days of exposure, the maximum weight loss (as an indicator of deterioration) was observed for the +310 mV (vs. Ag/AgCl) treatment, with significant differences (P < 0.05) from that which occurred under the other conditions (Figure 2A).
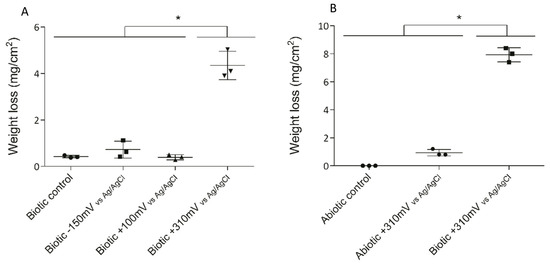
Figure 2.
Weight loss of SS electrodes under each treatment after 10 days. To determine the significant differences between the treatments, a posteriori Tukey test was performed. Significant differences between treatments are indicated with asterisks, where each asterisk group treatments according to similarity. (A) Weight losses in the initial experiment under biotic conditions. (B) Weight losses under the abiotic and biotic conditions with the +310 mV (vs. Ag/AgCl) potential applied.
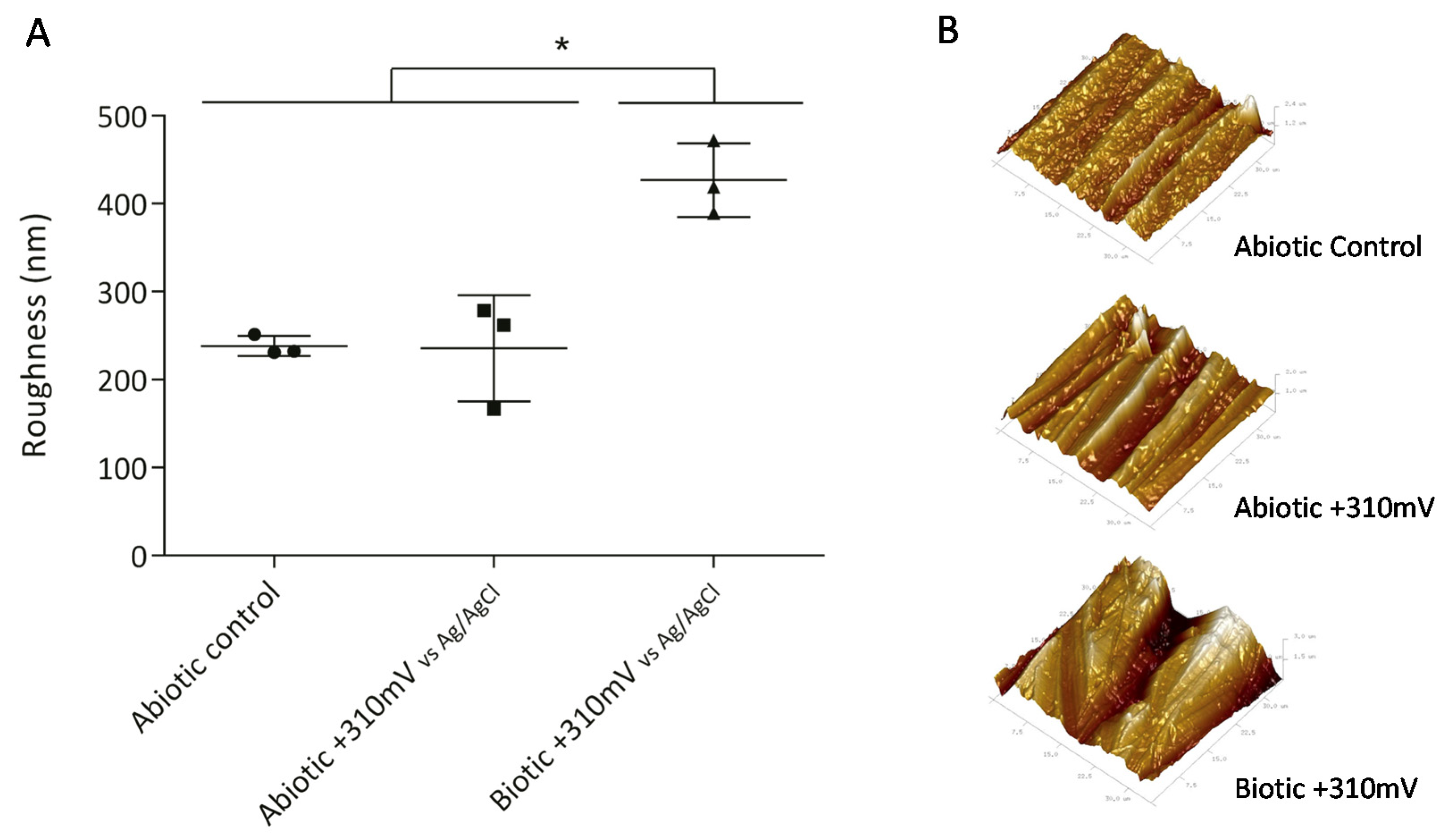
To determine whether the significant weight loss observed under the +310 mV (vs. Ag/AgCl) potential was due to the effect of the electrochemically enriched bacteria or the potential itself, an abiotic control experiment was conducted. Figure 2B shows that the maximum weight loss, which occurred in the SS electrodes poised at +310 mV (vs. Ag/AgCl) under biotic conditions, was significantly different from the corrosion produced under abiotic conditions both with and without the overpotential (P < 0.05). This result suggests that the observed deterioration was associated more with the enrichment of specific bacteria than with abiotic corrosion. Besides, AFM analysis of the electrodes showed that the applied potential of +310 mV (vs Ag/AgCl) significantly increased the surface roughness, probably due to electrode corrosion (Figure 3).
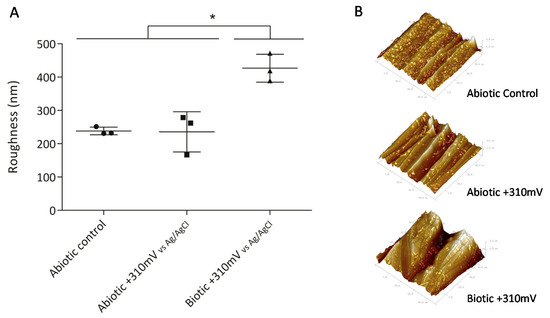
Figure 3.
Roughness of SS electrodes (second experiment), measured by atomic force microscopy (AFM). Asterisks represent significant differences between treatments (P < 0.05). (A) Average roughness of SS electrodes with standard deviation and (B) 3D Atomic Force Microscope (AFM) images of SS electrodes obtained over an area of 35 µm × 35 µm.
Indeed, biocorrosion of SS has been linked with the action of autotrophic metabolism [36,37,38]. This type of corrosion is frequently associated with the ennoblement of the OCP involving iron-oxidizing bacteria, manganese-oxidizing bacteria [36,38], and, more recently, electro-autotrophic bacteria able to use an electrode (i.e., a SS coupon) as the electron donor to reduce oxygen [37,39,40]. Furthermore, the ennoblement of SS plates exposed to natural seawater typically shifts the OCP by +200 to +300 mV (vs. Ag/AgCl), bringing it closer to the pitting corrosion potential [37]. This microbially driven phenomenon results in the initiation of SS pitting and crevice corrosion. Pitting corrosion of 316L SS coupons exposed to seawater has been reported under an applied potential ranging from +500 to +600 mV (vs. Ag/AgCl) after a few days of immersion [41]. Therefore, the significant differences in weight loss (Figure 2) and roughness (Figure 3) exhibited by the coupons exposed to the +310 mV (vs. Ag/AgCl) potential, in comparison to those produced the other tested conditions, may be linked with the enrichment for particular bacterial groups that could catalyze electron transfer, rather than with the applied potential itself.
Under the tested conditions, no significative damage was observed on SS electrodes polarized at +100 mV (vs. Ag/AgCl) that correspond to OCP reported for 316L SS under marine conditions after 10 days [23]. These results suggest that SS is a good candidate to be used as electrode material for the scaling-up of METs in marine environments. Hence, this finding opens the opportunity to explore long-term evaluation of this material in marine METs involving biocathodic reactions.
3.2. Bacterial Enrichment Using Overpotentials
To corroborate the microbial community development on the tested electrodes, epifluorescence microscopy and SEM were performed (Supplementary Materials Figures S1 and S2). Microscopic analysis revealed the presence of microorganisms attached to the surface of the electrode, with the presence of extracellular polymeric-like substances (Figure S2). These results confirm early biofilm development on all the coupons.
During the experiment, three different enrichment levels were expected because of the experimental design, which would be influenced by (i) the effect of water storage (bottle effect), (ii) the effect of the SS surface (i.e., material composition and roughness), and (iii) the effect of the overpotential application. The storage conditions and the surface were transversal in all the conditions studied, allowing the isolation of the specific effect of microbial enrichment upon application of the overpotential.
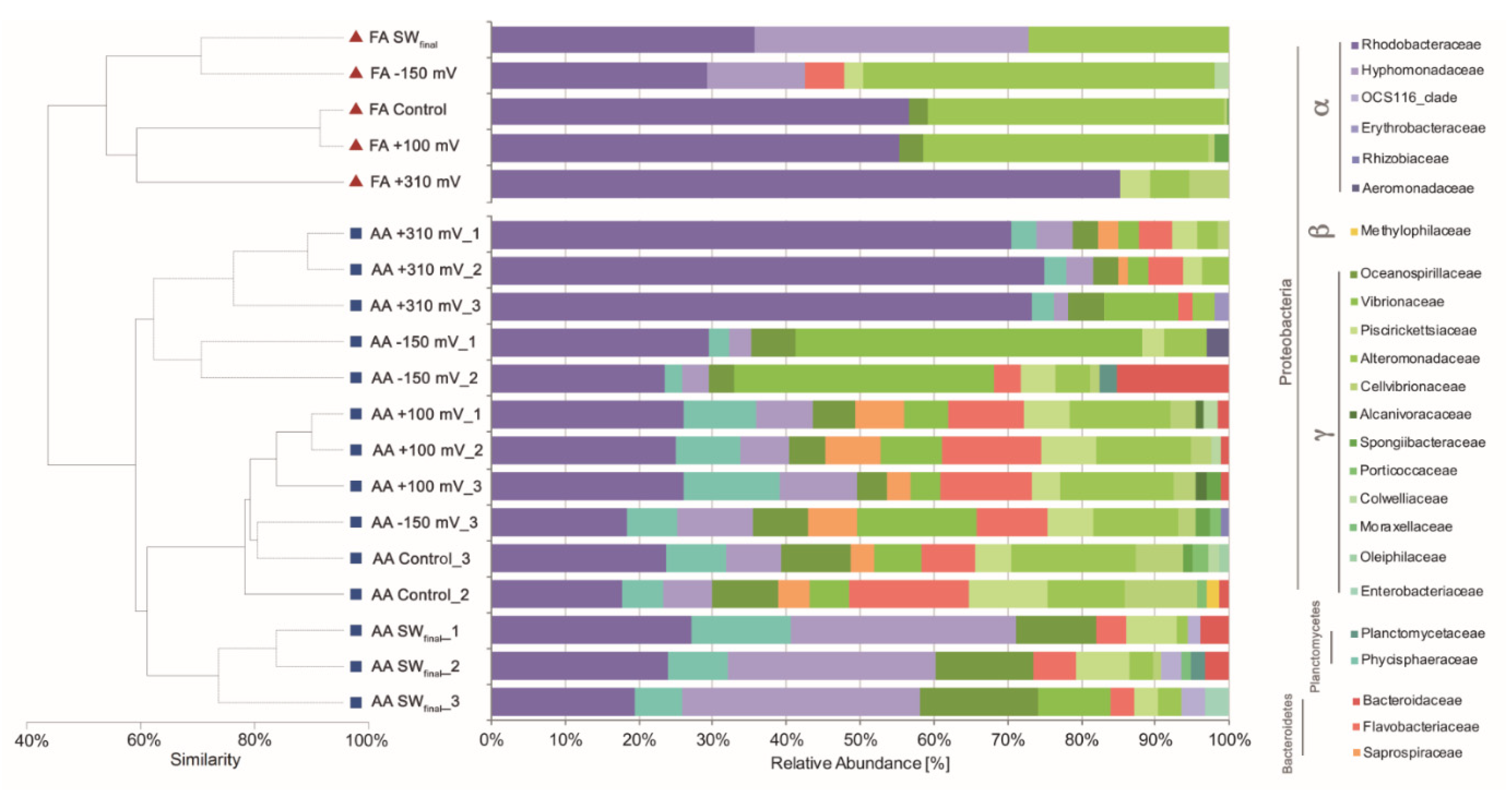
To evaluate the effect of the applied potentials on specific microbial enrichment, the bacterial community developed under each treatment was characterized at the end of the experiment using its 16S rRNA gene profiles. Cluster analysis based on the Bray–Curtis similarity coefficients obtained from both FA and AA datasets indicated a similar and consistent pattern, with the samples exposed at +310 mV (vs. Ag/AgCl) being the most different in each dataset (Figure 4), with 65% and 80% similarity, respectively. Differences were supported by SIMPROF (Pi 4,114, P < 0.05) and ANOSIM (R: 0.718, P < 0.05) analyses.
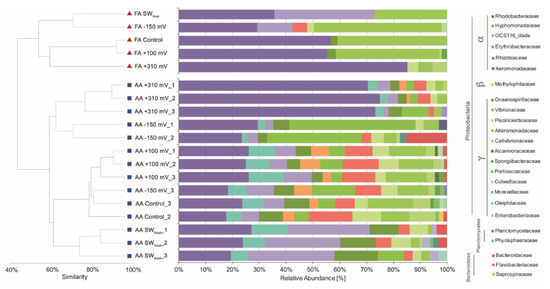
Figure 4.
Cluster and relative abundance family-level taxonomic analysis of the bacterial communities associated with each treatment.
Cluster analyses were complemented with taxonomic assignation to the family level of the main OTUs detected by cloning (i.e., FA) and AA. In the case of the samples exposed at +310 mV (vs. Ag/AgCl), the Rhodobacteraceae bacterial family was highly enriched from a natural inoculum of seawater without any nutrients added. Both techniques, FA and AA, gave similar results in terms of relative abundance of Rhodobacteraceae in the +310 mV (vs. Ag/AgCl) samples, with an average relative abundance of 81.80% by FA and 69.31% (±2.81) by AA.
On the other hand, the +100 mV, −150 mV (vs. Ag/AgCl) polarization, and control treatments resulted in similar bacterial compositions (Figure 4), suggesting that the effect of applying +100 mV, −150 mV (vs. Ag/AgCl) to an electrode, on the enrichment of specific marine bacteria, was not greater than the effect generated by the material itself. Sulfitobacter and Glaciecola, two of the most abundant microbial genera identified in the +100 mV (vs. Ag/AgCl) and −150 mV (vs. Ag/AgCl) samples (Table 1), were previously identified as the most abundant genera in microbial communities developed over SS surfaces [42]. In the case of the control, wherein no potential was applied, typical microbial genera previously associated with electroactive microbial communities were identified. When exposed to seawater, the 316L SS surface potential naturally reached +100 mV (vs. Ag/AgCl) after 10 days [23], and as is shown in Figure 4, the control and +100 mV (vs. Ag/AgCl) treatments resulted in similar microbial community compositions. These results suggest that microbial settling in natural complex environments, as well as in reactors run under controlled laboratory conditions [18,19,20,21], depends on the value of the SS electrode potential.

Table 1.
Relative abundance (%) of most-abundant genre identified by amplicon analysis (AA) in each sample (+100 mV, −150 mV, +310 mV vs. Ag/AgCl and Control).
3.3. Metabolic Inference of Electrochemically Enriched Bacterial Communities
To gain more information about the microbial community identified in each sample, the most abundant genera were characterized, identifying their metabolisms and the environments in which they have been found, according to previous reports in the literature. Table 2 lists, in decreasing order of abundance, the most abundant genera of each family that were identified for each treatment. Furthermore, the similarity between the sequences obtained by FA and AA, associated with each gender, was analyzed. All pairs of sequences had similarities of >80% and were related to the same bacterial genera, confirming the correspondence between a fingerprinting technique (such as FA) and AA analysis [43,44].

Table 2.
Similarity analysis between sequences obtained by fragment analysis (FA) and amplicon analysis (AA), and their metabolisms and the environments wherein have been reported. The similarity between sequences obtained by AA and FA is expressed as % similarity.
For the discussion of the relative abundance of the most abundant microbial genera identified in each condition, only the AA analyzes were used. The reason for this is due to the low number of clones obtained by condition, which generates an oversize of the relative abundance values of the genera identified by the clones in each condition (Table S2). The genera with higher relative abundances were Roseobacter, Pheobacter, and Sulfitobacter, with relative abundances of 16.8%, 10.5%, and 9.9%, respectively (Table 1). These genera showed the highest relative abundances in the +310 mV (vs. Ag/AgCl) samples. All of these genera have been related to autotrophic microorganisms and reported in electrochemically active microbial communities [42,45,46,47]. Other genera that have not been previously implicated in biocorrosion of SS, including Aestuariicella, Lewinella, and Spongiibacter, were also detected, with relative abundances of 2.43%, 3.18%, and 0.5%, respectively (Table 1). The majority of the genera shown in Table 2 have been previously associated with electroactive microbial communities developed on SS plates or electrodes [42,45,48,49,50,51,52,53,54,55,56,57,58].
As described in Materials and Methods, the potential of −150 mV (vs. Ag/AgCl) was selected as a representative midpoint potential capable to enrich for reported marine EAB from marine sediments [9]. The groups of microorganisms reported by Rowe et al. (2015) at −150 mV (vs. Ag/AgCl) were not found in our experiments. Nevertheless, Sulfitobacter and Glaciecola have been reported as the predominant bacteria in electroactive microbial communities on SS electrodes [42] (Table 2), and these were identified as two of the most abundant microbial genera enriched on the electrodes polarized at −150 mV (vs. Ag/AgCl) in our experiments (Table 1).
Poised potentials around +100 mV (vs. Ag/AgCl) have been identified as selective pressures for specific microbial communities mostly composed by Marinobacter, Chromatiaceae, and Labrenzia (MCL). This particular community, identified as a MCL-biocathode, has been reported to be capable of using the electrons supplied by the electrochemical system to drive CO2 fixation and O2 reduction [20,59,60]. Although Marinobacter and Labrenzia were not the most abundant genera identified in our electrodes tested over +100 mV (vs. Ag/AgCl), the abundances of both genera were significantly higher in comparison with their abundances in the other samples (Figure S3). It is important to highlight that the aforementioned MCL-biocathode cluster was obtained using marine sediment as the inoculum [61]. In this study, electrochemical enrichment was accomplished directly from natural seawater. In consequence, differences in bacterial composition and abundance between these experiments were expected.
Roseobacter and Sulfitobacter were two of the most abundant genera identified in the +310 mV (vs. Ag/AgCl) samples (Table 1). Both groups of microorganisms have been identified as EAB with high efficiency in the catalysis of the oxygen reduction reaction [46]. As indicated in Materials and Methods, this overpotential has been related to the oxygen reduction reaction [24,62]. From a bioenergetic perspective, with higher potentials closer to the theoretical reduction potential of oxygen (i.e., +600 mV vs. Ag/AgCl at pH = 7 and pO2 = 0.2 bar), aerobic microorganisms gain less energy [24]. Using an inoculum collected from a wastewater treatment plant, Ter Heijne et al. (2010) studied the effect of positive overpotentials (+50 mV, +150 mV, and +250 mV vs. Ag/AgCl) on the development of oxygen-reducing biocathodes. While the biocathodes polarized at +50 mV and +150 mV (vs. Ag/AgCl) produced current during the first day after inoculation, that polarized at +250 mV (vs. Ag/AgCl) produced no current until day 15, probably due to slow bacterial growth and a longer start-up time [24]. On the other hand, in a bioelectrochemical study of Candidatus Tenderia electrophaga (belonging to the Chromatiaceae family), Eddie et al. (2017) showed that by applying a potential as positive as +470 mV (vs. SHE) or +250 mV (vs. Ag/AgCl), there was an increase in the expression of genes related with nitrate and oxygen reduction [40]. Luo and Moran (2014) [63] reported that about half of the marine Roseobacter clade present genes related to the nitrate reduction process, and as indicated above, Sulfitobacter and Roseobacter can reduce oxygen using electrons from a polarized electrode [46]. Our results showed that by applying a positive potential as high as +310 mV (vs. Ag/AgCl) to SS electrodes exposed to natural seawater, it was possible to establish a high enrichment pressure for enriching autotrophic oxygen-reducing EAB such as Sulfitobacter and Roseobacter.
The presence of autotrophic oxygen-reducing EAB has been related to the acceleration of biocorrosion processes [37,39,40]. The highest abundances of Sulfitobacter and Roseobacter were found in the +310 mV (vs. Ag/AgCl) treatment, and at the same time, this condition resulted in significant differences in weight loss (Figure 2) and roughness (Figure 3) in comparison with those produced under the other tested conditions. Therefore, the results of this research suggest that high abundances of Sulfitobacter and Roseobacter are directly related to electrode deterioration, specifically the biocorrosion of 316L SS.
4. Conclusions
Electrochemical bacterial enrichment from natural seawater without nutrients amended was successfully achieved for the first time to the best of our knowledge. Under the tested conditions, no significant damage was observed on SS electrodes poised at up to +100 mV (vs. Ag/AgCl). This finding opens the opportunity for long-term evaluation of SS as electrode material in marine METs. On the other hand, a significant selection for putative EAB within the Roseobacter and Sulfitobacter genera present in a marine inoculum was observed upon applying an overpotential of +310 mV (vs. Ag/AgCl), and the resultant microbial community accelerated corrosion, compromising the lifetimes of the SS electrodes. Finally, further attention should be given to the aforementioned bacterial groups as potential biological drivers of pitting corrosion in 316L SS in marine environments.
Supplementary Materials
The following are available online at https://www.mdpi.com/1996-1944/13/10/2327/s1, Figure S1: Epifluorescence microscopy of each treatment, using acridine orange 0.1% w/v. When an acridine orange binds with DNA, it will exhibit a green color, and when it binds with RNA, it will exhibit a red color. (A) +310 mV vs. Ag/AgCl, (B) −150 mV vs. Ag/AgCl, (C) +100 mV vs. Ag/AgCl, and (D) Control. All images are in 10× magnification, Figure S2: Scanning Electron Microscopy of each treatment. (A) +310 mV vs. Ag/AgCl, (B) −150 mV vs. Ag/AgCl, (C) +100 mV vs. Ag/AgCl, and (D) Control. Bar scale represents a size of 2 µm. The magnifications of these images are 13,900×, Figure S3: Relative abundance (%) of Marinobacter sp. and Labrenzia sp. in each sample, Table S1: Accession number of the clone sequences reported in this investigation and the most closely related organisms, with its accession number, Table S2: Relative abundance (%) of most abundant genre identified by amplicon analysis (AA) and fragment analysis (FA) in each sample (+100mV, −150mV, +310mV vs. Ag/AgCl and Control).
Author Contributions
Conceptualization; M.J.D.L.F., R.D.l.I., M.W., G.E.P., F.A., and I.T.V.; Methodology, M.J.D.L.F., R.D.l.I., and I.T.V.; Validation; M.J.D.L.F., L.K.D., R.D.l.I., M.W., F.A., G.E.P., and I.T.V.; Formal Analysis, M.J.D.L.F., L.K.D., R.D.l.I., and F.A.; Investigation, M.J.D.L.F., L.K.D., R.D.l.I., and I.T.V.; Resources, G.E.P., R.D.l.I., and I.T.V.; Data Curation, M.J.D.L.F., L.K.D. and R.D.l.I.; Writing—original draft preparation, M.J.D.L.F., L.K.D., R.D.l.I., and I.T.V.; Writing—review and editing, M.J.D.L.F., L.K.D., R.D.l.I., M.W., F.A., and I.T.V.; Funding acquisition, G.E.P., R.D.l.I., and I.T.V. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Vicerrectoría de Investigación de la Pontificia Universidad Católica de Chile (Project VRI-PUC Nº01/2013), the Office of Naval Research (Grant NICOP-N62909-17-1-2012), FONDECYT 1160917, and FONDECYT 1181326. This study was conducted at the Marine Energy Research and Innovation Center (MERIC) (Project CORFO 14CEI2-28228).
Acknowledgments
We give special thanks to Benjamin Lagos, Mauricio Medel, and Evelyn Cruz for their assistance in the laboratory and help with the experimental setup.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Rao, J.R.; Richter, G.J.; Von Sturm, F.; Weidlich, E. The Performance of Glucose Electrodes and the Characteristics of Different. Biofuel Cell Constructions. Bioelectrochemistry Bioenerg. 1976, 3, 139–150. [Google Scholar] [CrossRef]
- Roller, S.D.; Bennetto, H.P.; Delaney, G.M.; Mason, J.R.; Stirling, J.L.; Thurston, C.F. Electron-transfer coupling in microbial fuel cells: 1. Comparison of redox-mediator reduction rates and respiratory rates of bacteria. J. Chem. Technol. Biotechnol. 1984, 34, 3–12. [Google Scholar] [CrossRef]
- He, Z.; Kan, J.; Wang, Y.; Huang, Y.; Mansfeld, F.; Nealson, K.H. Electricity production coupled to ammonium in a microbial fuel cell. Environ. Sci. Technol. 2009, 43, 3391–3397. [Google Scholar] [CrossRef] [PubMed]
- Berks, B.C.; Ferguson, S.J.; Moir, J.W.B.; Richardson, D.J. Enzymes and associated electron transport systems that catalyse the respiratory reduction of nitrogen oxides and oxyanions. Biochim. Biophys. Acta 1995, 2728, 97–173. [Google Scholar] [CrossRef]
- Li, W.-W.; Yu, H.-Q. Stimulating sediment bioremediation with benthic microbial fuel cells. Biotechnol. Adv. 2015, 33, 1–12. [Google Scholar] [CrossRef]
- Jiang, C.; Yang, Q.; Wang, D.; Zhong, Y.; Chen, F.; Li, X.; Zeng, G.; Li, X. Simultaneous perchlorate and nitrate removal coupled with electricity generation in autotrophic denitrifying biocathode microbial fuel cell. Chem. Eng. J. 2017, 308, 783–790. [Google Scholar] [CrossRef]
- Virdis, B.; Rabaey, K.; Rozendal, R.A.; Yuan, Z.; Keller, J. Simultaneous nitrification, denitrification and carbon removal in microbial fuel cells. Water Res. 2010, 44, 2970–2980. [Google Scholar] [CrossRef]
- Wang, H.; Ren, Z.J. A comprehensive review of microbial electrochemical systems as a platform technology. Biotechnol. Adv. 2013, 31, 1796–1807. [Google Scholar] [CrossRef]
- Rowe, A.R.; Chellamuthu, P.; Lam, B.; Okamoto, A.; Nealson, K.H. Marine sediments microbes capable of electrode oxidation as a surrogate for lithotrophic insoluble substrate metabolism. Front. Microbiol. 2015, 6, 1–15. [Google Scholar] [CrossRef]
- Logan, B.E. Scaling up microbial fuel cells and other bioelectrochemical systems. Appl. Microbiol. Biotechnol. 2010, 85, 1665–1671. [Google Scholar] [CrossRef]
- Kim, J.R.; Jung, S.H.; Regan, J.M.; Logan, B.E. Electricity generation and microbial community analysis of alcohol powered microbial fuel cells. Bioresour. Technol. 2007, 98, 2568–2577. [Google Scholar] [CrossRef] [PubMed]
- Baudler, A.; Schmidt, I.; Langner, M.; Greiner, A.; Schröder, U. Does it have to be carbon? Metal anodes in microbial fuel cells and related bioelectrochemical systems. Energy Environ. Sci. 2015, 8, 2048–2055. [Google Scholar] [CrossRef]
- Selembo, P.A.; Merrill, M.D.; Logan, B.E. The use of stainless steel and nickel alloys as low-cost cathodes in microbial electrolysis cells. J. Power Sources 2009, 190, 271–278. [Google Scholar] [CrossRef]
- Dumas, C.; Mollica, A.; Féron, D.; Basséguy, R.; Etcheverry, L.; Bergel, A. Marine microbial fuel cell: Use of stainless steel electrodes as anode and cathode materials. Electrochim. Acta 2007, 53, 468–473. [Google Scholar] [CrossRef]
- Call, D.F.; Merrill, M.D.; Logan, B.E. High Surface Area Stainless Steel Brushes as Cathodes in Microbial Electrolysis Cells. Environ. Sci. Technol. 2009, 43, 2179–2183. [Google Scholar] [CrossRef] [PubMed]
- Pocaznoi, D.; Calmet, A.; Etcheverry, L.; Erable, B.; Bergel, A. Stainless steel is a promising electrode material for anodes of microbial fuel cells. Energy Environ. Sci. 2012, 5, 9645–9652. [Google Scholar] [CrossRef]
- Tender, L.M.; Reimers, C.E.; Iii, H.A.S.; Holmes, D.E.; Bond, D.R.; Lowy, D.A.; Pilobello, K.; Fertig, S.J.; Lovley, D.R. Harnessing microbially generated power on the seafloor. Nat. Biotechnol. 2002, 20, 821–825. [Google Scholar] [CrossRef]
- Miceli, J.F.; Parameswaran, P.; Kang, D.; Krajmalnik-brown, R. Enrichment and analysis of anode-respiring bacteria from diverse anaerobic inocula. Environ. Sci. Technol. 2012, 46, 10349–10355. [Google Scholar] [CrossRef]
- Zhan, G.; Zhang, L.; Tao, Y.; Wang, Y.; Zhu, X.; Li, D. Anodic ammonia oxidation to nitrogen gas catalyzed by mixed biofilms in bioelectrochemical systems. Electrochim. Acta 2014, 135, 345–350. [Google Scholar] [CrossRef]
- Wang, Z.; Leary, D.H.; Malanoski, A.H.P.; Li, R.W.; Judson Hervey, W.; Eddie, B.J.; Tender, G.S.; Yanosky, S.G.; Vora, G.J.; Tender, L.M.; et al. A previously uncharacterized, nonphotosynthetic member of the chromatiaceae is the primary CO2-fixing constituent in a self-regenerating biocathode. Appl. Environ. Microbiol. 2015, 81, 699–712. [Google Scholar] [CrossRef]
- Torres, C.I.; Krajmalnik-Brown, R.; Parameswaran, P.; Marcus, A.K.; Wanger, G.; Gorby, Y.A.; Rittmann, B.E. Selecting anode-respiring bacteria based on anode potential: Phylogenetic, electrochemical, and microscopic characterization. Environ. Sci. Technol. 2009, 43, 9519–9524. [Google Scholar] [CrossRef] [PubMed]
- Juretschko, S.; Loy, A.; Lehner, A.; Wagner, M. The microbial community composition of a nitrifying-denitrifying activated sludge from an industrial sewage treatment plant analyzed by the full-cycle rRNA approach. Syst. Appl. Microbiol. 2002, 25, 84–99. [Google Scholar] [CrossRef] [PubMed]
- Fischer, D.A.; Daille, L.; Aguirre, J.; Galarce, C.; Armijo, F.; la Iglesia, R.D.; Pizarro, G.; Vargas, I.; Walczak, M. Corrosion of stainless steel in simulated tide of fresh natural seawater of south east pacific. Int. J. Electrochem. Sci. 2016, 11, 6873–6885. [Google Scholar] [CrossRef]
- Ter Heijne, A.; Strik, D.P.B.T.B.; Hamelers, H.V.; Buisman, C.J. Cathode potential and mass transfer determine performance of oxygen reducing biocathodes in microbial fuel cells. Environ. Sci. Technol. 2010, 44, 7151–7156. [Google Scholar] [CrossRef]
- Boivin, M.E.Y.; Greve, G.D.; Kools, S.A.E.; van der Wurff, A.W.G.; Leeflang, P.; Smit, E.; Breure, A.M.; Rutgers, M.; van Straalen, N.M. Discriminating between effects of metals and natural variables in terrestrial bacterial communities. Appl. Soil Ecol. 2006, 34, 103–113. [Google Scholar] [CrossRef]
- Henríquez-castillo, C.; Rodríguez-marconi, S.; Rubio, F.; Trefault, N.; Andrade, S.; Iglesia, R. De Eukaryotic picophytoplankton community response to copper enrichment in a metal-perturbed coastal environment. Phycol. Res. 2015, 63, 189–196. [Google Scholar] [CrossRef]
- Weisburg, W.G.; Barns, S.M.; Pelletier, D.A.; Lane, D.J. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 1991, 173, 697–703. [Google Scholar] [CrossRef]
- Osborne, C.A.; Rees, G.N.; Bernstein, Y.; Janssen, P.H. New threshold and confidence estimates for terminal restriction fragment length polymorphism analysis of complex bacterial communities. Appl. Environ. Microbiol. 2006, 72, 1270–1278. [Google Scholar] [CrossRef][Green Version]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.E.W.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 2011, 17, 10. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, 590–596. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Beals, E.W. Bray-curtis ordination: An effective strategy for analysis of multivariate ecological data. Adv. Ecol. Res. 1984, 14, 1–55. [Google Scholar] [CrossRef]
- Warwick, R.M.; Clarke, K.R. Change in marine communities: An approach to statistical analysis and interpretation. Am. J. Clin. Nutr. 2007, 85, 1236–1243. [Google Scholar]
- Olesen, B.H.; Avci, R.; Lewandowski, Z. Manganese dioxide as a potential cathodic reactant in corrosion of stainless steels. Corros. Sci. 2000, 42, 211–227. [Google Scholar] [CrossRef]
- Trigodet, F.; Larché, N.; Morrison, H.G.; Jebbar, M.; Thierry, D.; Maignien, L. Electroactive bacteria associated with stainless steel ennoblement in seawater. Front. Microbiol. 2019, 10, 1–9. [Google Scholar] [CrossRef]
- Chamritski, I.G.; Burns, G.R.; Webster, B.J.; Laycock, N.J. Effect of iron-oxidizing bacteria on pitting of stainless steel. Corrosion 2004, 60, 658–669. [Google Scholar] [CrossRef]
- Eddie, B.J.; Wang, Z.; Malanoski, A.P.; Hall, R.J.; Oh, S.D.; Heiner, C.; Lin, B.; Strycharz-glaven, S.M.; Eddie, B.J. ‘Candidatus Tenderia electrophaga’, an uncultivated electroautotroph from a biocathode enrichment. Int. J. Syst. Evol. Microbiol. 2016, 66, 2178–2185. [Google Scholar] [CrossRef]
- Eddie, B.J.; Wang, Z.; Iv, W.J.H.; Leary, D.H.; Malanoski, A.P.; Tender, L.M.; Lin, B.; Strycharz-glaven, S.M. Metatranscriptomics Supports the Mechanism for Biocathode Electroautotrophy by “Candidatus Tenderia electrophaga”. Mol. Biol. Physiol. 2017, 2, 1–13. [Google Scholar] [CrossRef]
- Compère, C.; Jaffré, P.; Festy, D. Aging of type 316L stainless steel in seawater: Relationship between open-circuit potential, exposure time, and pitting potential. Corros. Sci. 1996, 52, 496–501. [Google Scholar] [CrossRef]
- Erable, B.; Byrne, N.; Etcheverry, L.; Achouak, W.; Bergel, A. Single medium microbial fuel cell: Stainless steel and graphite electrode materials select bacterial communities resulting in opposite electrocatalytic activities. Int. J. Hydrogen Energy 2017, 42, 26059–26067. [Google Scholar] [CrossRef]
- van Dorst, J.; Bissett, A.; Palmer, A.S.; Brown, M.; Snape, I.; Stark, J.S.; Raymond, B.; Mckinlay, J.; Ji, M.; Winsley, T.; et al. Community fingerprinting in a sequencing world. Fems Microbiol. Ecol. 2014, 89, 316–330. [Google Scholar] [CrossRef] [PubMed]
- Gobet, A.; Boetius, A.; Ramette, A. Ecological coherence of diversity patterns derived from classical fingerprinting and next generation sequencing techniques. Environ. Microbiol. 2014, 16, 2672–2681. [Google Scholar] [CrossRef]
- Vandecandelaere, I.; Segaert, E.; Mollica, A.; Faimali, M.; Vandamme, P. Phaeobacter caeruleus sp. nov., a blue-coloured, colony-forming bacterium isolated from a marine electroactive biofilm. Int. J. Syst. Evol. Microbiol. 2009, 59, 1209–1214. [Google Scholar] [CrossRef]
- Parot, S.; Vandecandelaere, I.; Cournet, A.; Délia, M.; Vandamme, P.; Bergé, M.; Roques, C.; Bergel, A. Catalysis of the electrochemical reduction of oxygen by bacteria isolated from electro-active biofilms formed in seawater. Bioresour. Technol. 2011, 102, 304–311. [Google Scholar] [CrossRef]
- Allgaier, M.; Uphoff, H.; Felske, A.; Wagner-do, I. aerobic anoxygenic photosynthesis in roseobacter clade bacteria from diverse marine habitats. Appl. Environ. Microbiol. 2003, 69, 5051–5059. [Google Scholar] [CrossRef]
- Li, H.; Zhou, E.; Zhang, D.; Xu, D.; Xia, J.; Yang, C.; Feng, H.; Jiang, Z.; Li, X.; Gu, T.; et al. Microbiologically influenced corrosion of 2707 hyper-duplex stainless steel by marine pseudomonas aeruginosa biofilm. Sci. Rep. 2016, 6, 1–12. [Google Scholar] [CrossRef]
- Xia, J.; Yang, C.; Xu, D.; Sun, D.; Nan, L.; Sun, Z.; Li, Q.; Gu, T.; Yang, K. Laboratory investigation of the microbiologically influenced corrosion (MIC) resistance of a novel Cu-bearing 2205 duplex stainless steel in the presence of an aerobic marine Pseudomonas aeruginosa biofilm. Biofouling 2015, 31, 481–492. [Google Scholar] [CrossRef]
- Rajasekar, A.; Ting, Y.P. Role of inorganic and organic medium in the corrosion behavior of bacillus megaterium and pseudomonas sp. in stainless steel SS 304. Ind. Eng. Chem. Res. 2011, 50, 12534–12541. [Google Scholar] [CrossRef]
- Vandecandelaere, I.; Nercessian, O.; Faimali, M.; Segaert, E.; Mollica, A.; Achouak, W.; De Vos, P.; Vandamme, P. Bacterial diversity of the cultivable fraction of a marine electroactive biofilm. Bioelectrochemistry 2010, 78, 62–66. [Google Scholar] [CrossRef] [PubMed]
- Erable, B.; Roncato, M.-A.; Achouak, W.; Bergel, A. Sampling natural biofilms: A new route to build efficient microbial anodes. Environ. Sci. Technol. 2009, 43, 3194–3199. [Google Scholar] [CrossRef] [PubMed]
- Martins, G.; Peixoto, L.; Ribeiro, D.C.; Parpot, P.; Brito, A.G.; Nogueira, R. Bioelectrochemistry towards implementation of a benthic microbial fuel cell in lake Furnas (Azores): Phylogenetic af fi liation and electrochemical activity of sediment bacteria. Bioelectrochemistry 2010, 78, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Ringelberg, D.B.; Foley, K.L.; Reynolds, C.M. Electrogenic capacity and community composition of anodic biofilms in soil-based bioelectrochemical systems. Appl. Microbiol. Biotechnol. 2011, 90, 1805–1815. [Google Scholar] [CrossRef]
- Malanoski, A.P.; Lin, B.; Brian, J.; Wang, Z.; Iv, W.J.H.; Glaven, S.M. Relative abundance of ‘Candidatus Tenderia electrophaga’ is linked to cathodic current in an aerobic biocathode community. Microb. Biotechnol. 2018, 11, 98–111. [Google Scholar] [CrossRef]
- Barco, R.A.; Emerson, D.; Sylvan, J.B.; Orcutt, B.N.; Meyers, E.J.; Ramírez, G.A.; Zhong, J.D.; Edwards, J. New insight into microbial iron oxidation as revealed by the proteomic profile of an obligate iron-oxidizing chemolithoautotroph. Appl. Environ. Microbiol. 2015, 81, 5927–5937. [Google Scholar] [CrossRef]
- Oldham, A.L.; Steinberg, M.K.; Duncan, K.E.; Makama, Z.; Duncan, K.E. Molecular methods resolve the bacterial composition of natural marine biofilms on galvanically coupled stainless steel cathodes. J. Ind. Microbiol. Biotechnol. 2017, 44, 167–180. [Google Scholar] [CrossRef]
- Holland, R.L.; Cooper, B.H.; Helgeson, N.G.; Mccracken, A.W. Automated detection of microbial growth in blood automated detection of microbial growth in blood cultures by using stainless-steel electrodes. J. Clin. Microbiol. 1980, 12, 180–184. [Google Scholar] [CrossRef]
- Strycharz-glaven, S.M.; Glaven, R.H.; Wang, Z.; Zhou, J.; Vora, G.J.; Tender, L.M. Electrochemical investigation of a microbial solar cell reveals a nonphotosynthetic biocathode catalyst. Appl. Environ. Microbiol. 2013, 79, 3933–3942. [Google Scholar] [CrossRef]
- Leary, D.H.; Hervey, J.W.; Malanoski, A.P.; Wang, Z.; Eddie, B.J.; Tender, G.S.; Vora, G.J.; Tender, L.M.; Lin, B.; Strycharz-Glaven, S.M. Metaproteomic evidence of changes in protein expression following a change in electrode potential in a robust biocathode microbiome. Proteomics 2015, 15, 3486–3496. [Google Scholar] [CrossRef]
- Malik, S.; Drott, E.; Grisdela, P.; Lee, J.; Lee, C.; Lowy, D.A.; Tender, L.M. A self-assembling self-repairing microbial photoelectrochemical solar cell. Energy Environ. Sci. 2009, 2, 292–298. [Google Scholar] [CrossRef]
- Rimboud, M.; Bergel, A.; Erable, B. Multiple electron transfer systems in oxygen reducing biocathodes revealed by different conditions of aeration/agitation. Bioelectrochemistry 2016, 110, 46–51. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Moran, M.A. Evolutionary ecology of the marine roseobacter clade. Microbiol. Mol. Biol. Rev. 2014, 78, 573–587. [Google Scholar] [CrossRef] [PubMed]
- Dang, H.; Lovell, C.R. Bacterial primary colonization and early succession on surfaces in marine waters as determined by amplified rRNA gene restriction analysis and sequence analysis of 16S rRNA genes. Appl. Environ. Microbiol. 2000, 66, 467–475. [Google Scholar] [CrossRef] [PubMed]
- Dang, H.; Li, T.; Chen, M.; Huang, G. Cross-ocean distribution of rhodobacterales bacteria as primary surface colonizers in temperate coastal marine waters. Appl. Environ. Microbiol. 2008, 74, 52–60. [Google Scholar] [CrossRef]
- Chen, G.W.; Cha, J.H.; Choi, S.J.; Lee, T.H.; Kim, C.W. Characterization of an open biocathode microbial fuel cell for electricity generation and effluent polish. Korean J. Chem. Eng. 2010, 27, 828–835. [Google Scholar] [CrossRef]
- Abraham, W.; Strompl, C.; Meyer, H.; Lindholst, S.; Moore, E.R.B.; Christ, R.; Vancanneyt, M.; Tindali, B.J.; Bennasar, A.; Smit, J.; et al. Phylogeny and polyphasic taxonomy of caulobacter species. Proposal of maricaulis gen. nov. with maricaulis maris (Poindexter) comb. nov. as the type species, and emended description of the genera brevundirnonas and caulobacter. Int. J. Syst. Bacteriol. 1999, 49, 1053–1073. [Google Scholar] [CrossRef]
- Moura, V.; Ribeiro, I.; Moriggi, P.; Capão, A.; Salles, C.; Bitati, S.; Procópio, L. The influence of surface microbial diversity and succession on microbiologically influenced corrosion of steel in a simulated marine environment. Arch. Microbiol. 2018, 200, 1447–1456. [Google Scholar] [CrossRef]
- Yuan, L.N.; Ren, L.F.; Li, Y.T.; Han, W.J.; Yu, Y.; Chu, Y.N.; Liu, G.M.; Yu, D.; Teng, M.J.; Wang, L.; et al. A complete genome assembly of glaciecola mesophila sp. nov. sequenced by using BIGIS-4 sequencer system. Sci. China Life Sci. 2011, 54, 835–840. [Google Scholar] [CrossRef][Green Version]
- Darus, L.; Ledezma, P.; Keller, J.; Freguia, S. Marine phototrophic consortia transfer electrons to electrodes in response to reductive stress. Photosynth Res. 2016, 127, 347–354. [Google Scholar] [CrossRef]
- Schwermer, C.U.; Lavik, G.; Abed, R.M.M.; Dunsmore, B.; Ferdelman, T.G.; Stoodley, P.; Gieseke, A.; De Beer, D. Impact of nitrate on the structure and function of bacterial biofilm communities in pipelines used for injection of seawater into oil fields. Appl. Environ. Microbiol. 2008, 74, 2841–2851. [Google Scholar] [CrossRef] [PubMed]
- Janvier, M.; Grimont, P.A.D. The genus Methylophaga, a new line of descent within phylogenetic branch γ of proteobacteria. Res. Microbiol. 1995, 146, 543–550. [Google Scholar] [CrossRef]
- Lo, N.; Kim, K.H.; Baek, K.; Jia, B.; Jeon, C.O. Aestuariicella hydrocarbonica gen. nov., sp. Nov., an aliphatic hydrocarbon-degrading bacterium isolated from a sea tidal flat. Int. J. Syst. Evol. Microbiol. 2015, 65, 1935–1940. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.D. Lewinella agarilytica sp. nov., a novel marine bacterium of the phylum bacteroidetes, isolated from beach sediment. Int. J. Syst. Evol. Microbiol. 2007, 57, 2814–2818. [Google Scholar] [CrossRef]
- Graeber, I.; Kaesler, I.; Borchert, M.S.; Dieckmann, R.; Pape, T.; Lurz, R.; Nielsen, P.; von Döhren, H.; Michaelis, W.; Szewzyk, U. Spongiibacter marinus gen. nov., sp. nov., a halophilic marine bacterium isolated from the boreal sponge Haliclona sp. 1. Int. J. Syst. Evol. Microbiol. 2008, 58, 585–590. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).