Biological Hydrogen Energy Production by Novel Strains Bacillus paramycoides and Cereibacter azotoformans through Dark and Photo Fermentation
Abstract
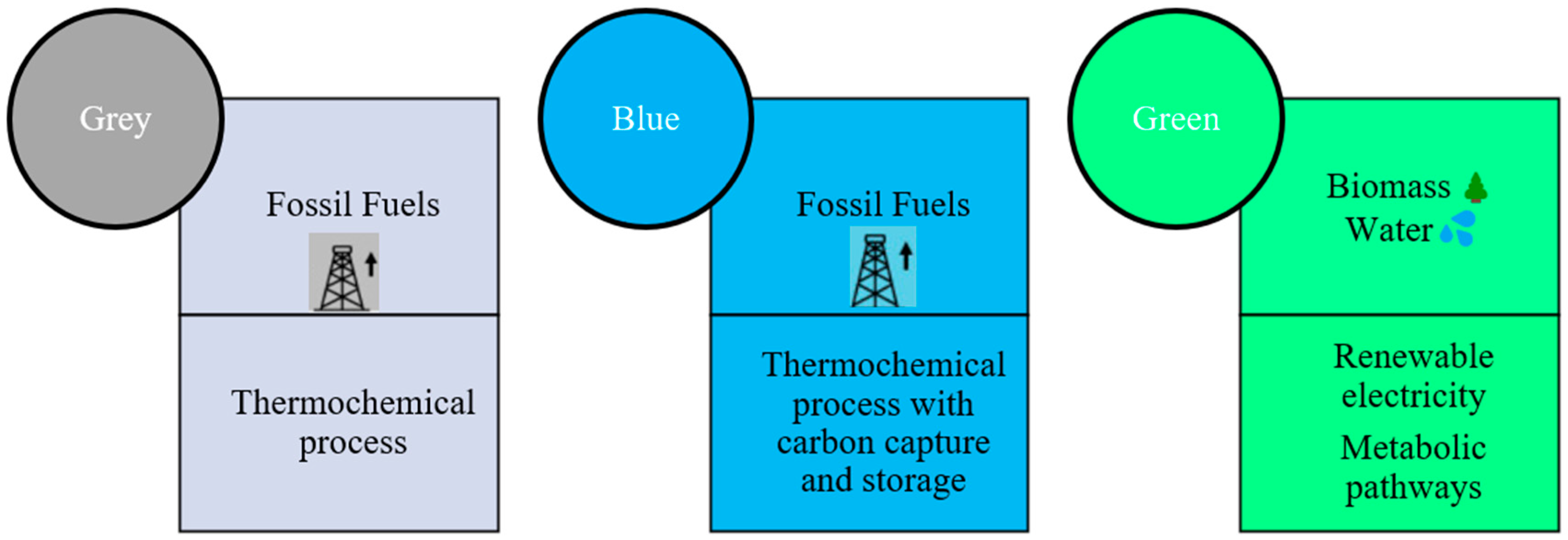
1. Introduction
2. Materials and Methods
2.1. Microorganisms and Culture Medium
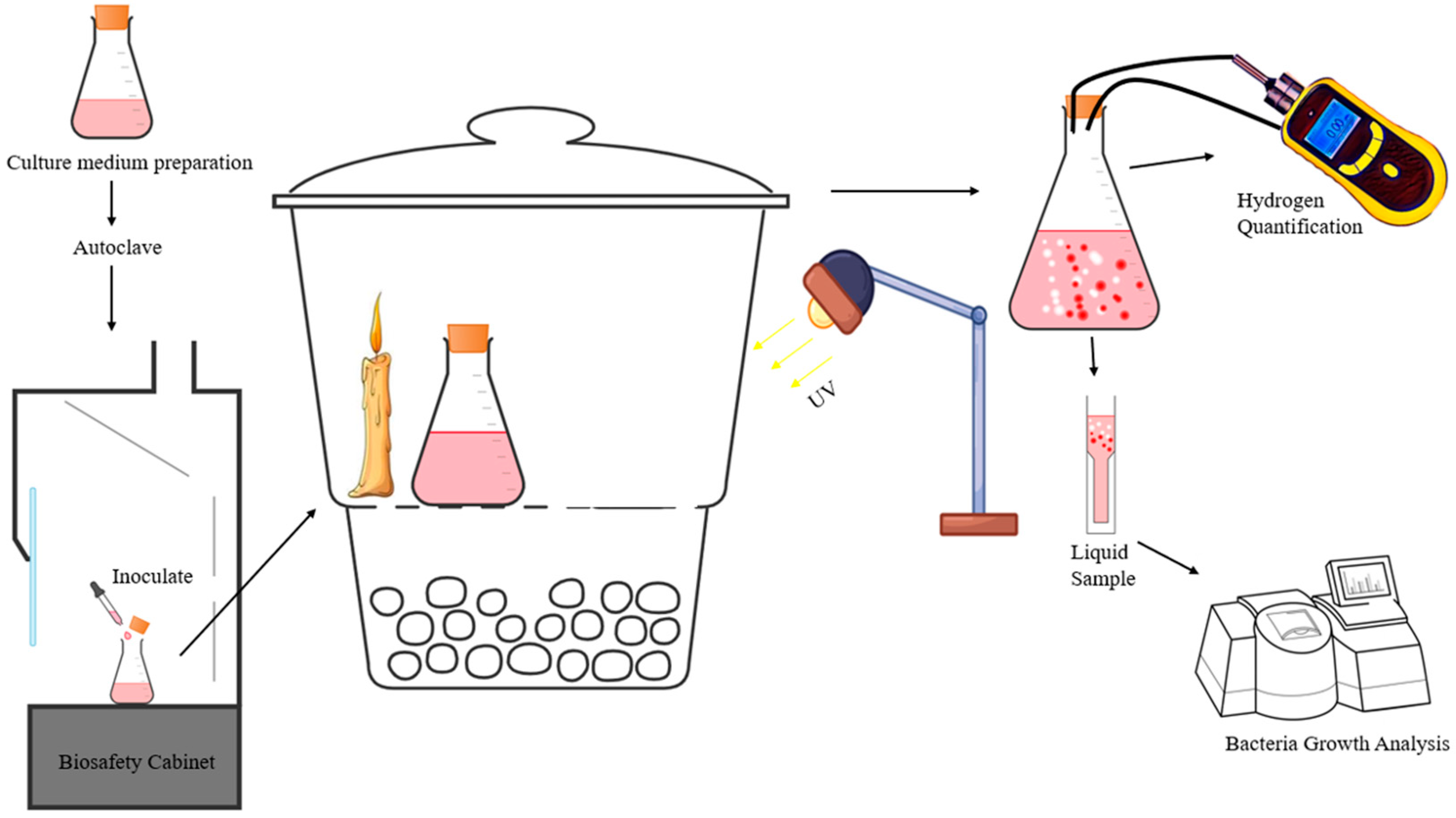
2.2. Experimental Setup and Analytical Method
2.3. Gram Staining Procedure
3. Results and Discussion
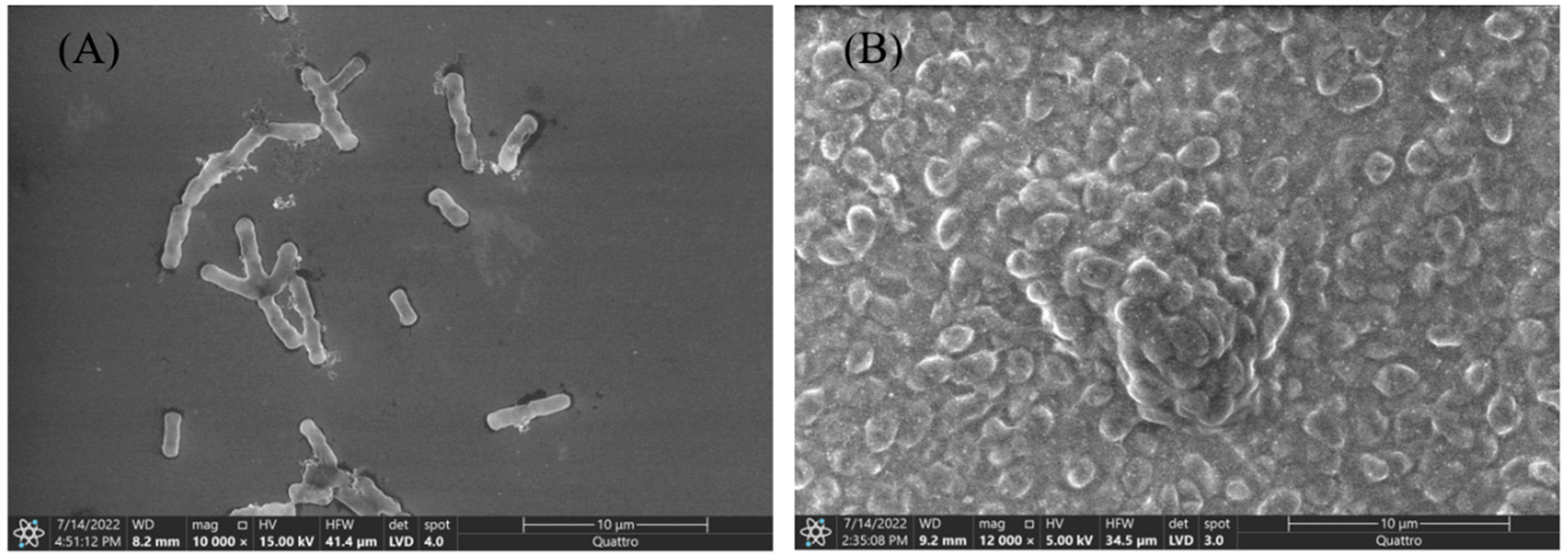
3.1. Morphology and Gram Staining
3.2. DNA Sequencing
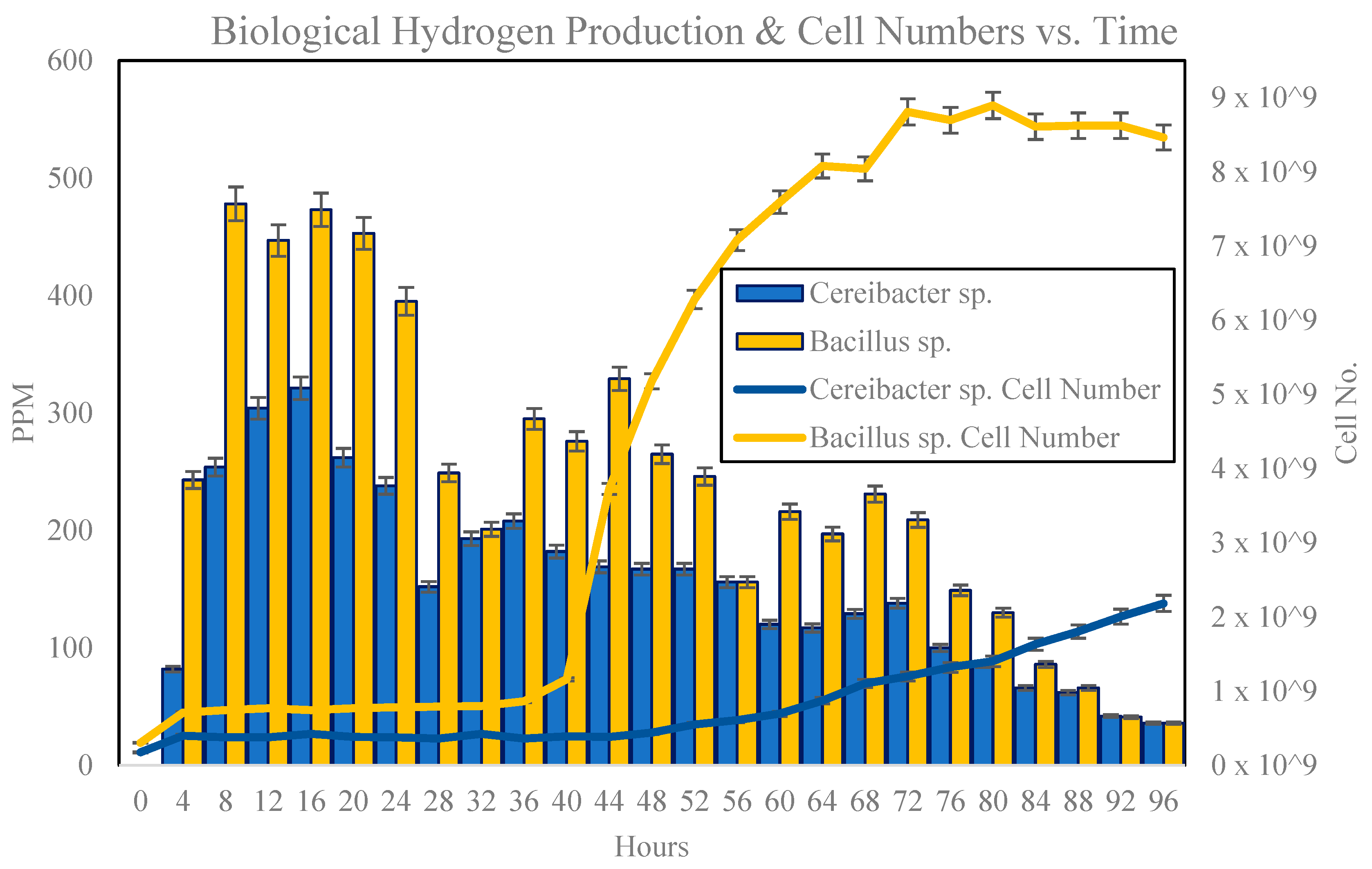
3.3. Growth Behaviour Study and Biological Hydrogen Production with 1% Inoculum
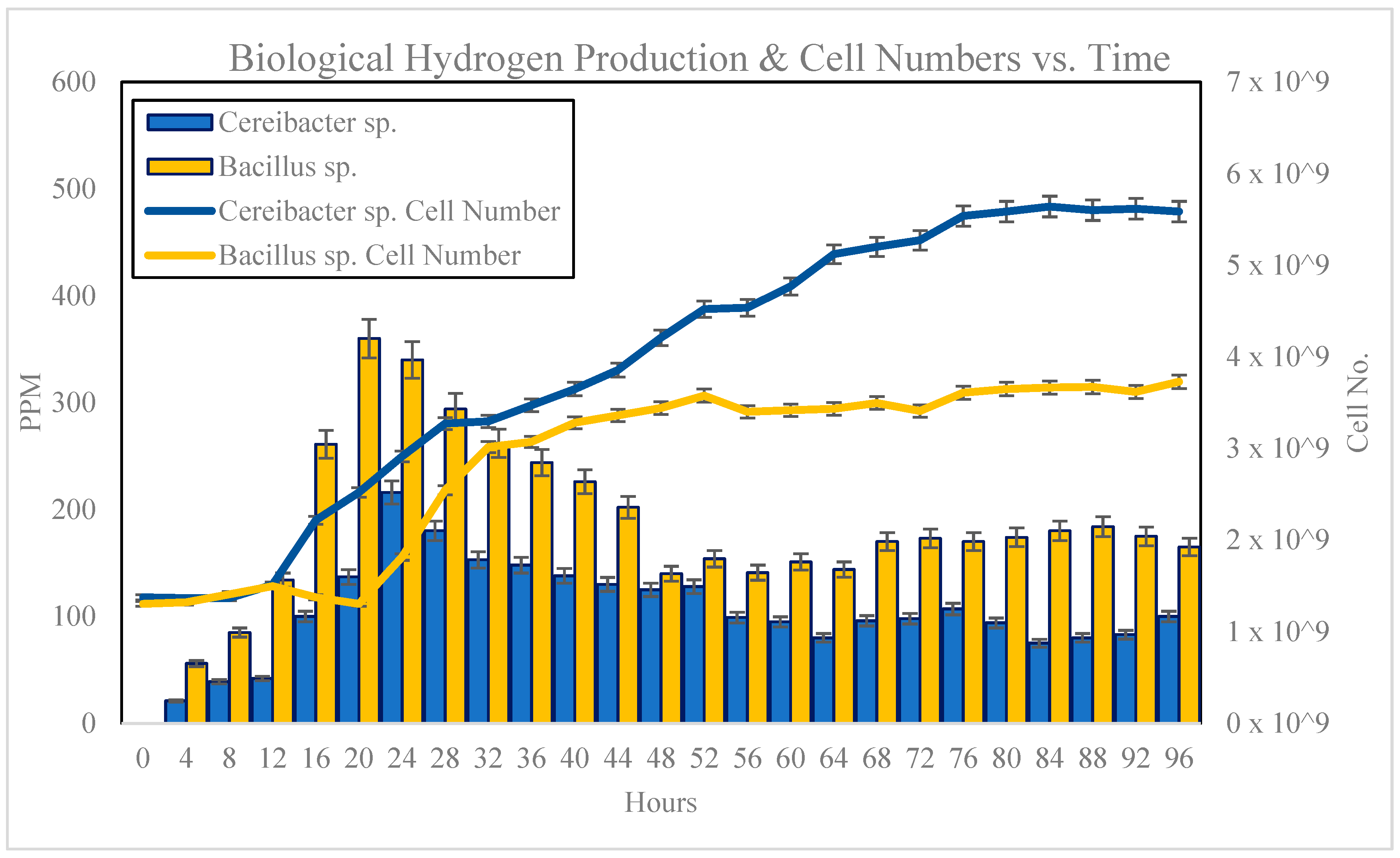
3.4. Growth Behaviour Study and Biological Hydrogen Production with 10% Inoculum
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Adeosun, O.A.; Tabash, M.I.; Anagreh, S. Oil price and economic performance: Additional evidence from advanced economies. Resour. Policy 2022, 77, 102666. [Google Scholar] [CrossRef]
- Karmaker, A.K.; Rahman, M.M.; Hossain, M.A.; Ahmed, M.R. Exploration and corrective measures of greenhouse gas emission from fossil fuel power stations for Bangladesh. J. Clean. Prod. 2020, 244, 118645. [Google Scholar] [CrossRef]
- Wang, W.-K.; Hu, Y.-H.; Liao, G.-Z.; Zeng, W.-L.; Wu, S.-Y. Hydrogen fermentation by photosynthetic bacteria mixed culture with silicone immobilization and metagenomic analysis. Int. J. Hydrogen Energy 2022, 47, 40590–40602. [Google Scholar] [CrossRef]
- Lee, D.-H. Biohydrogen yield efficiency and the benefits of dark, photo and dark-photo fermentative production technology in circular Asian economies. Int. J. Hydrogen Energy 2020, 46, 13908–13922. [Google Scholar] [CrossRef]
- Li, X.; Sui, K.; Zhang, J.; Liu, X.; Xu, Q.; Wang, D.; Yang, Q. Revealing the mechanisms of rhamnolipid enhanced hydrogen production from dark fermentation of waste activated sludge. Sci. Total. Environ. 2021, 806, 150347. [Google Scholar] [CrossRef] [PubMed]
- Abad, A.V.; Dodds, P. Production of Hydrogen. In Encyclopedia of Sustainable Technologies; Elsevier: Berlin/Heidelberg, Germany, 2017; pp. 293–304. [Google Scholar] [CrossRef]
- Hermesmann, M.; Müller, T. Green, Turquoise, Blue, or Grey? Environmentally friendly Hydrogen Production in Transforming Energy Systems. Prog. Energy Combust. Sci. 2022, 90, 100996. [Google Scholar] [CrossRef]
- Srivastava, N.; Srivastava, M.; Malhotra, B.D.; Gupta, V.K.; Ramteke, P.; Silva, R.N.; Shukla, P.; Dubey, K.K.; Mishra, P. Nanoengineered cellulosic biohydrogen production via dark fermentation: A novel approach. Biotechnol. Adv. 2019, 37, 107384. [Google Scholar] [CrossRef]
- Vernès, L.; Li, Y.; Chemat, F.; Abert-Vian, M. Biorefinery Concept as a Key for Sustainable Future to Green Chemistry—The Case of Microalgae. In Plant Based “Green Chemistry 2.0”; Springer: Singapore, 2019; pp. 15–50. [Google Scholar] [CrossRef]
- Jung, J.-H.; Sim, Y.-B.; Ko, J.; Park, S.Y.; Kim, G.-B.; Kim, S.-H. Biohydrogen and biomethane production from food waste using a two-stage dynamic membrane bioreactor (DMBR) system. Bioresour. Technol. 2022, 352, 127094. [Google Scholar] [CrossRef]
- Mejía-Saucedo, C.; Buitrón, G.; León-Galván, M.F.; Carrillo-Reyes, J. Biomass purge strategies to control the bacterial community and reactor stability for biohydrogen production from winery wastewater. Int. J. Hydrogen Energy 2021, 47, 5891–5900. [Google Scholar] [CrossRef]
- Zhang, L.; Wang, Y.-Z.; Zhao, T.; Xu, T. Hydrogen production from simultaneous saccharification and fermentation of lignocellulosic materials in a dual-chamber microbial electrolysis cell. Int. J. Hydrogen Energy 2019, 44, 30024–30030. [Google Scholar] [CrossRef]
- Rorke, D.C.S.; Lekha, P.; Kana, G.E.B.; Sithole, B.B. Effect of pharmaceutical wastewater as nitrogen source on the optimization of simultaneous saccharification and fermentation hydrogen production from paper mill sludge. Sustain. Chem. Pharm. 2022, 25, 100619. [Google Scholar] [CrossRef]
- Wang, J.; Yin, Y. Progress in microbiology for fermentative hydrogen production from organic wastes. Crit. Rev. Environ. Sci. Technol. 2019, 49, 825–865. [Google Scholar] [CrossRef]
- Pu, Y.; Tang, J.; Wang, X.C.; Hu, Y.; Huang, J.; Zeng, Y.; Ngo, H.H.; Li, Y. Hydrogen production from acidogenic food waste fermentation using untreated inoculum: Effect of substrate concentrations. Int. J. Hydrogen Energy 2019, 44, 27272–27284. [Google Scholar] [CrossRef]
- Palafox-Félix, M.; Huerta-Ocampo, J.; Hernández-Ortíz, M.; Encarnación-Guevara, S.; Vázquez-Moreno, L.; Guzmán-Partida, A.M.; Cabrera, R. Proteomic analysis reveals the metabolic versatility of Amycolatopsis sp. BX17: A strain native from milpa agroecosystem soil. J. Proteom. 2021, 253, 104461. [Google Scholar] [CrossRef]
- Tang, T.; Chen, Y.; Liu, M.; Du, Y.; Tan, Y. Effect of pH on the performance of hydrogen production by dark fermentation coupled denitrification. Environ. Res. 2022, 208, 112663. [Google Scholar] [CrossRef] [PubMed]
- Gadow, S.I.; Jiang, H.; Watanabe, R.; Li, Y.-Y. Effect of temperature and temperature shock on the stability of continuous cellulosic-hydrogen fermentation. Bioresour. Technol. 2013, 142, 304–311. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Xin, X.; Liu, H.; Wu, Y.; Zhong, N.; Chang, H. Monitoring Biohydrogen Production and Metabolic Heat in Biofilms by Fiber Bragg Grating Sensors. Anal. Chem. 2019, 91, 7842–7849. [Google Scholar] [CrossRef]
- Yamamura, S.; Ike, M.; Fujita, M. Dissimilatory arsenate reduction by a facultative anaerobe, Bacillus sp. strain SF-1. J. Biosci. Bioeng. 2003, 96, 454–460. [Google Scholar] [CrossRef]
- Arai, H.; Roh, J.H.; Kaplan, S. Transcriptome Dynamics during the Transition from Anaerobic Photosynthesis to Aerobic Respiration in Rhodobacter sphaeroides 2.4.1. J. Bacteriol. 2008, 190, 286–299. [Google Scholar] [CrossRef]
- Zagrodnik, R.; Laniecki, M. The role of pH control on biohydrogen production by single stage hybrid dark- and photo-fermentation. Bioresour. Technol. 2015, 194, 187–195. [Google Scholar] [CrossRef]
- Orsi, E.; Folch, P.L.; Monje-López, V.T.; Fernhout, B.M.; Turcato, A.; Kengen, S.W.M.; Eggink, G.; Weusthuis, R.A. Characterization of heterotrophic growth and sesquiterpene production by Rhodobacter sphaeroides on a defined medium. J. Ind. Microbiol. Biotechnol. 2019, 46, 1179–1190. [Google Scholar] [CrossRef]
- Jang, W.J.; Lee, K.-B.; Jeon, M.-H.; Lee, S.-J.; Hur, S.W.; Lee, S.; Lee, B.-J.; Lee, J.M.; Kim, K.-W.; Lee, E.-W. Characteristics and biological control functions of Bacillus sp. PM8313 as a host-associated probiotic in red sea bream (Pagrus major) aquaculture. Anim. Nutr. 2023, 12, 20–31. [Google Scholar] [CrossRef]
- Eyring, M.; Martin, P. Spectroscopy in Forensic Science. In Reference Module in Chemistry, Molecular Sciences and Chemical Engineering; Elsvier: Amsterdam, The Netherlands, 2013. [Google Scholar] [CrossRef]
- O’Toole, G.A. Classic Spotlight: How the Gram Stain Works. J. Bacteriol. 2016, 198, 3128. [Google Scholar] [CrossRef] [PubMed]
- Ruhal, R.; Kataria, R. Biofilm patterns in gram-positive and gram-negative bacteria. Microbiol. Res. 2021, 251, 126829. [Google Scholar] [CrossRef]
- Sizar, O.; Unakal, C.G. Gram Positive Bacteria. In Management of Antimicrobials in Infectious Diseases; Humana Press: Totowa, NJ, USA, 2022; pp. 29–41. [Google Scholar] [CrossRef]
- Carter, G.R. Corynebacterium. In Diagnostic Procedure in Veterinary Bacteriology and Mycology; Academic Press: Cambridge, MA, USA, 1990; pp. 263–270. [Google Scholar] [CrossRef]
- Figueiredo, G.G.O.; Lopes, V.R.; Romano, T.; Camara, M.C. Clostridium. In Beneficial Microbes in Agro-Ecology: Bacteria and Fungi; Academic Press: Cambridge, MA, USA, 2020; pp. 477–491. [Google Scholar] [CrossRef]
- Kollipara, A.; Lee, D.; Darville, T. Sexually Transmitted Infections and the Urgent Need for Vaccines: A Review of Four Major Bacterial STI Pathogens. In Mucosal Vaccines: Innovation for Preventing Infectious Diseases; Academic Press: Cambridge, MA, USA, 2020; pp. 625–647. [Google Scholar] [CrossRef]
- Wu, M.; Li, X. Klebsiella pneumoniae and Pseudomonas aeruginosa. In Molecular Medical Microbiology; Academic Press: Cambridge, MA, USA, 2020; pp. 1547–1564. [Google Scholar] [CrossRef]
- Do, Y.S.; Schmidt, T.M.; Zahn, J.A.; Boyd, E.S.; de la Mora, A.; DiSpirito, A.A. Role of Rhodobacter sp. Strain PS9, a Purple Non-Sulfur Photosynthetic Bacterium Isolated from an Anaerobic Swine Waste Lagoon, in Odor Remediation. Appl. Environ. Microbiol. 2003, 69, 1710–1720. [Google Scholar] [CrossRef] [PubMed]
- Swinnen, I.A.M.; Bernaerts, K.; Dens, E.J.J.; Geeraerd, A.H.; Van Impe, J.F. Predictive modelling of the microbial lag phase: A review. Int. J. Food Microbiol. 2004, 94, 137–159. [Google Scholar] [CrossRef] [PubMed]
- Raju, D.V.; Nagarajan, A.; Pandit, S.; Nag, M.; Lahiri, D.; Upadhye, V. Effect of bacterial quorum sensing and mechanism of antimicrobial resistance. Biocatal. Agric. Biotechnol. 2022, 43, 102409. [Google Scholar] [CrossRef]
- Urvoy, M.; Lami, R.; Dreanno, C.; Delmas, D.; L’helguen, S.; Labry, C. Quorum Sensing Regulates the Hydrolytic Enzyme Production and Community Composition of Heterotrophic Bacteria in Coastal Waters. Front. Microbiol. 2021, 12, 3831. [Google Scholar] [CrossRef] [PubMed]
- Ulhiza, T.A.; Puad, N.I.M.; Azmi, A.S. Preliminary optimization of process conditions for biohydrogen production from sago wastewater. In Proceedings of the 2017 IEEE 3rd International Conference on Engineering Technologies and Social Sciences, ICETSS, Bangkok, Thailand, 7–8 August 2017; Volume 2018, pp. 1–4. [Google Scholar] [CrossRef]
- Liao, Q.; Zhong, N.; Zhu, X.; Huang, Y.; Chen, R. Enhancement of hydrogen production by optimization of biofilm growth in a photobioreactor. Int. J. Hydrogen Energy 2015, 40, 4741–4751. [Google Scholar] [CrossRef]
- Barer, M.R. Bacterial growth, physiology and death. In Medical Microbiology, 18th ed.; Churchill Livingstone: London, UK, 2012; pp. 39–53. [Google Scholar] [CrossRef]
- Preiss, L.; Hicks, D.B.; Suzuki, S.; Meier, T.; Krulwich, T.A. Alkaliphilic bacteria with impact on industrial applications, concepts of early life forms, and bioenergetics of ATP synthesis. Front. Bioeng. Biotechnol. 2015, 3, 75. [Google Scholar] [CrossRef]
- Jaishankar, J.; Srivastava, P. Molecular Basis of Stationary Phase Survival and Applications. Front. Microbiol. 2017, 8, 2000. [Google Scholar] [CrossRef] [PubMed]





| Sample | 16S rRNA Sequenced Gene Size (Base Pair) | GenBank Accession Number | % Identity | % Query Cover | Scientific Name |
|---|---|---|---|---|---|
| 1 | 1509 | NR_157734.1 | 99.93 | 100 | Bacillus paramycoides |
| 2 | 1418 | NR_113300.1 | 100 | 99 | Cereibacter azotoformans |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chung Han Chua, E.; Wee, S.K.; Kansedo, J.; Lau, S.Y.; Lim, K.H.; Dol, S.S.; Lipton, A.N. Biological Hydrogen Energy Production by Novel Strains Bacillus paramycoides and Cereibacter azotoformans through Dark and Photo Fermentation. Energies 2023, 16, 3807. https://doi.org/10.3390/en16093807
Chung Han Chua E, Wee SK, Kansedo J, Lau SY, Lim KH, Dol SS, Lipton AN. Biological Hydrogen Energy Production by Novel Strains Bacillus paramycoides and Cereibacter azotoformans through Dark and Photo Fermentation. Energies. 2023; 16(9):3807. https://doi.org/10.3390/en16093807
Chicago/Turabian StyleChung Han Chua, Eldon, Siaw Khur Wee, Jibrail Kansedo, Sie Yon Lau, King Hann Lim, Sharul Sham Dol, and Anuj Nishanth Lipton. 2023. "Biological Hydrogen Energy Production by Novel Strains Bacillus paramycoides and Cereibacter azotoformans through Dark and Photo Fermentation" Energies 16, no. 9: 3807. https://doi.org/10.3390/en16093807
APA StyleChung Han Chua, E., Wee, S. K., Kansedo, J., Lau, S. Y., Lim, K. H., Dol, S. S., & Lipton, A. N. (2023). Biological Hydrogen Energy Production by Novel Strains Bacillus paramycoides and Cereibacter azotoformans through Dark and Photo Fermentation. Energies, 16(9), 3807. https://doi.org/10.3390/en16093807








