The Influence of Micro-Oxygen Addition on Desulfurization Performance and Microbial Communities during Waste-Activated Sludge Digestion in a Rusty Scrap Iron-Loaded Anaerobic Digester
Abstract
:1. Introduction
2. Materials and Methods
2.1. Substrates, Inoculant and Rusty Scrap Iron
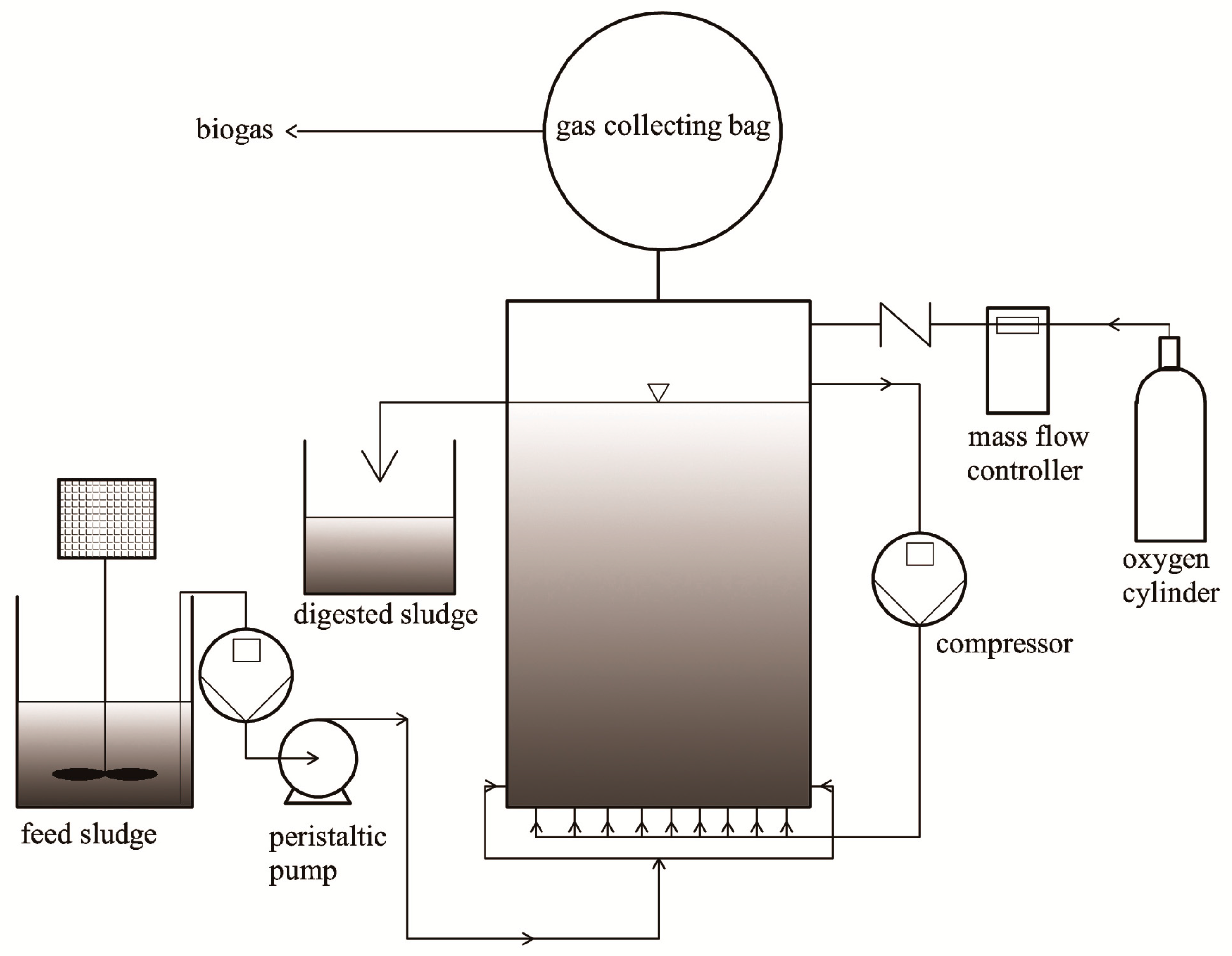
2.2. Semi-Continuous Microaerobic Digestion
2.3. Analysis
2.4. DNA Extraction, Amplification by Polymerase Chain Reaction, Denaturing Gradient Gel Electrophoresis and Sequencing
3. Results and Discussion
3.1. Sludge Digestion System with Rusty Scrap Iron Amendments during Semi-Continuous Anaerobic and Microaerobic Digestion
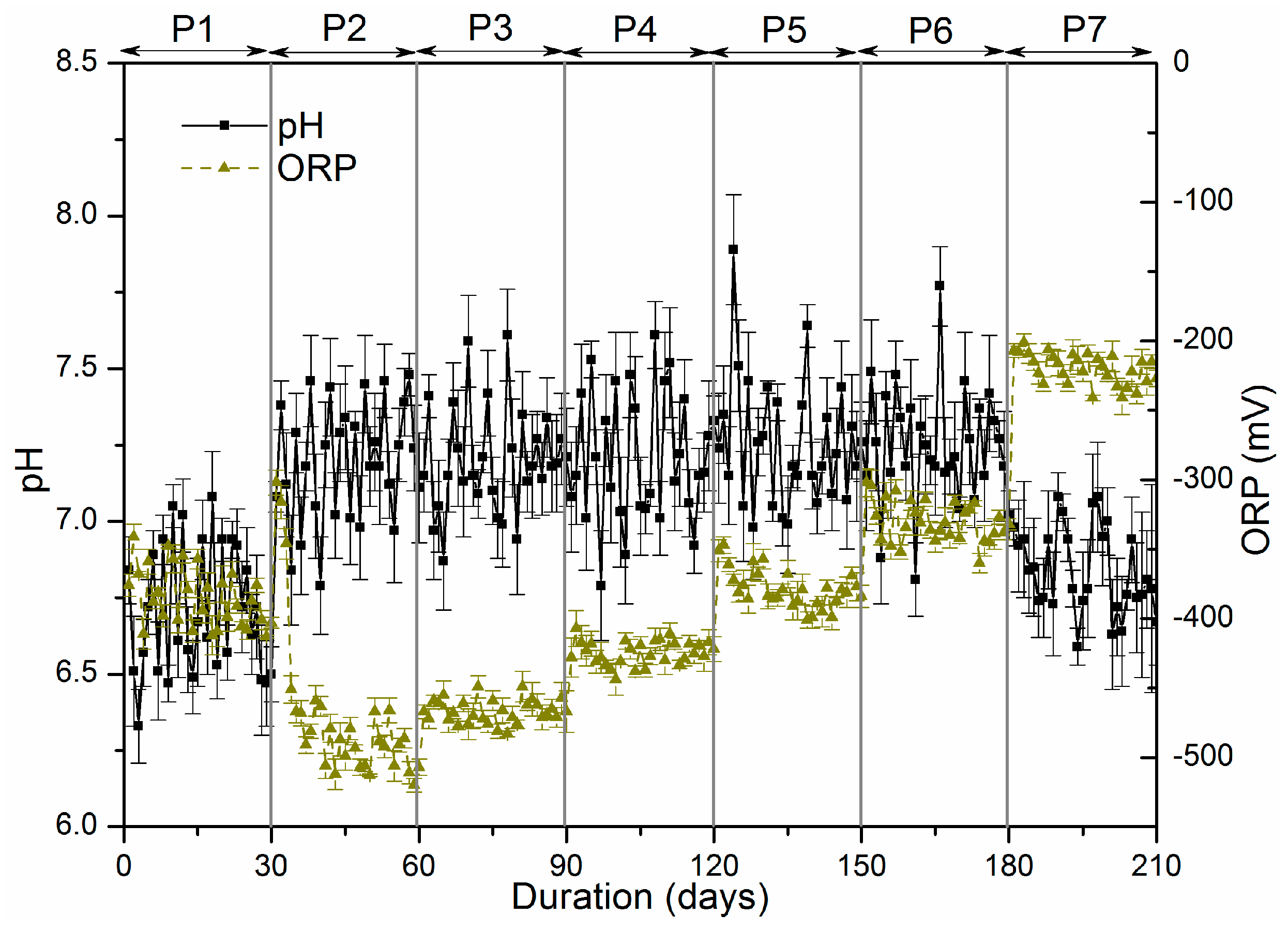
3.1.1. The Variation of pH and Oxidation-Reduction Potential
3.1.2. The Anaerobic and Microaerobic Digestion Performance
3.1.3. H2S and O2 Concentration in Biogas
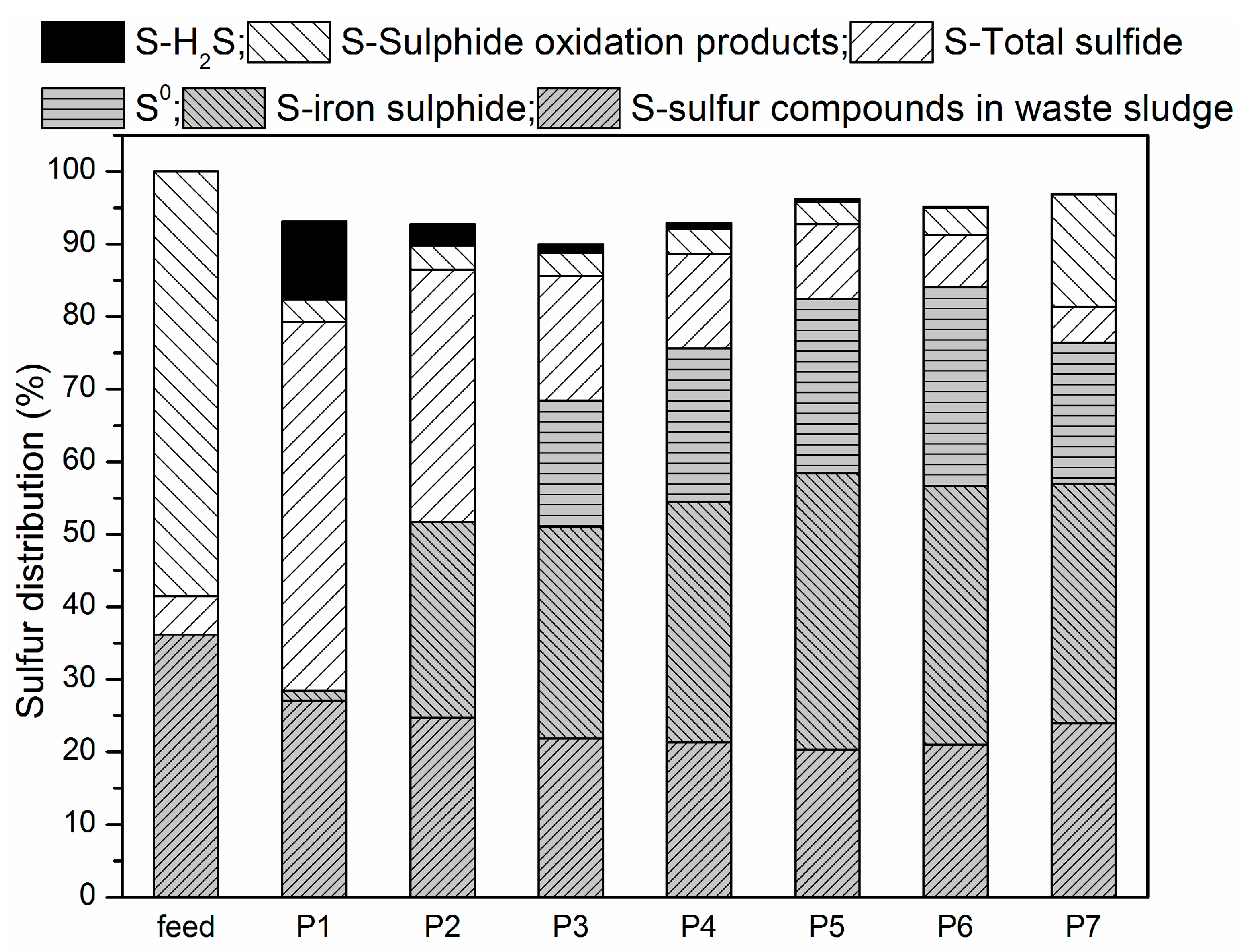
3.2. Sulfur Balance
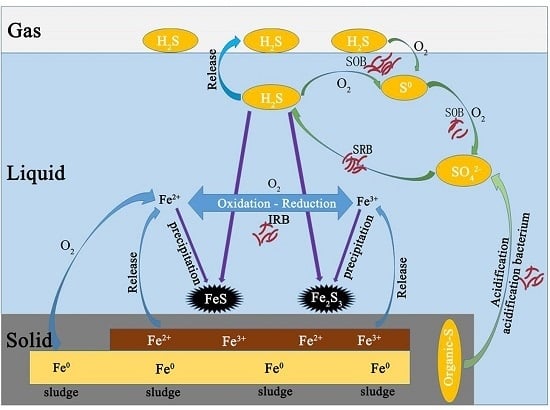
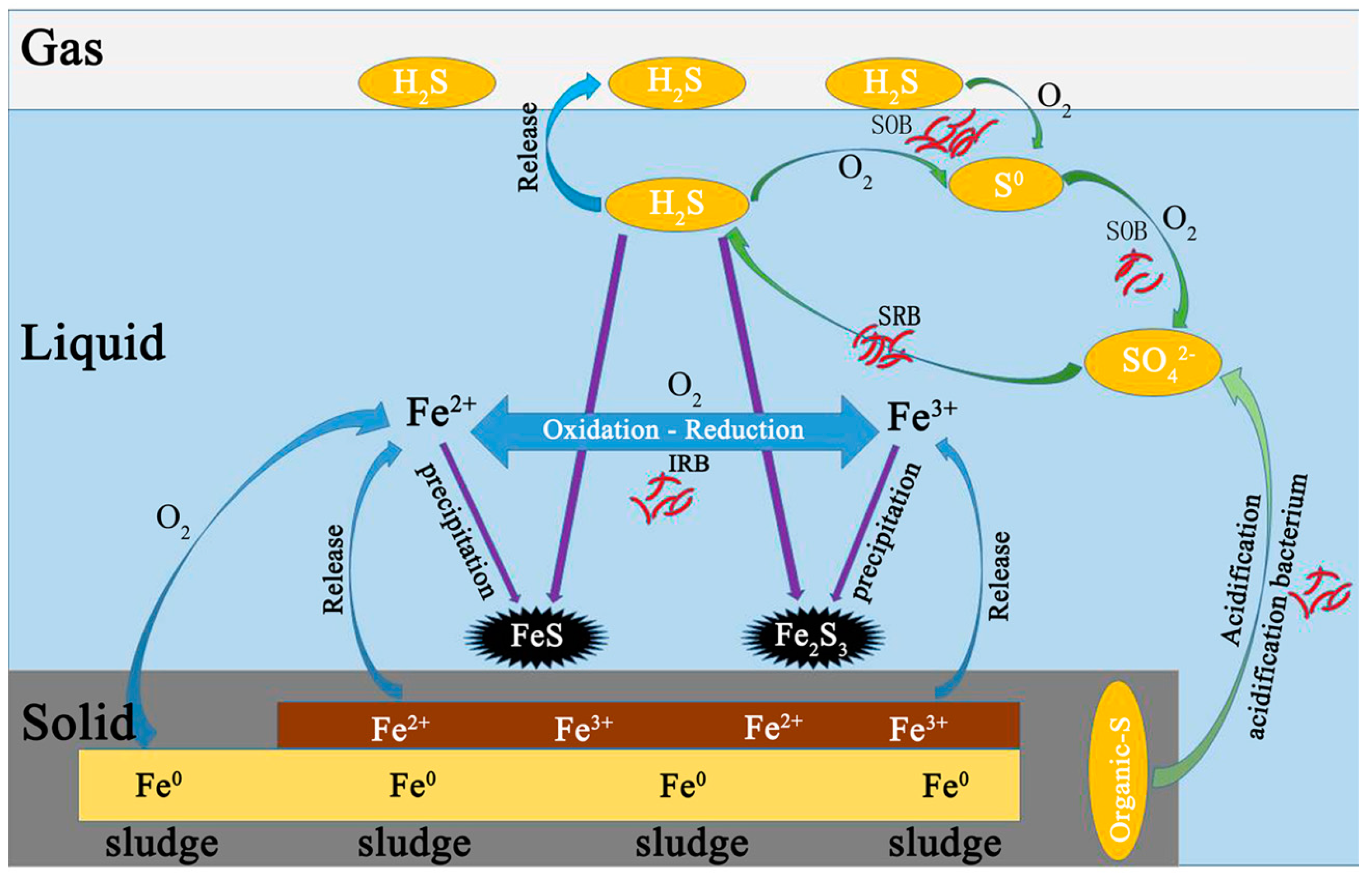
3.3. The Possible Control Mechanism of H2S in Biogas
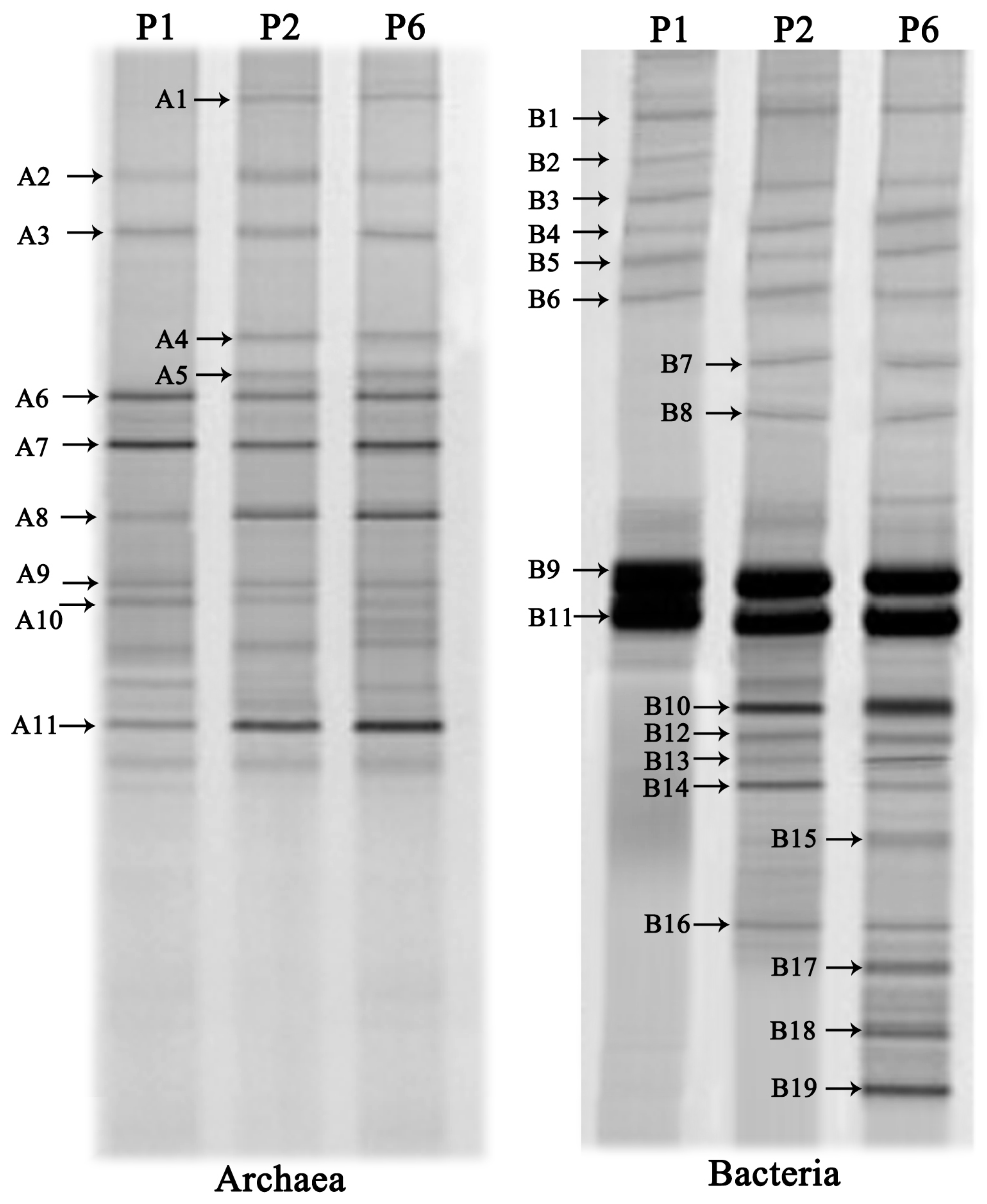
3.4. DGGE Analysis of the Microbial Communities
3.5. Economic Analysis
4. Conclusions and Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Zhang, C.; Su, H.; Baeyens, J.; Tan, T. Reviewing the anaerobic digestion of food waste for biogas production. Renew. Sustain. Energy Rev. 2014, 38, 383–392. [Google Scholar] [CrossRef]
- Agostini, A.; Battini, F.; Giuntoli, J.; Tabaglio, V.; Padella, M.; Baxter, D.; Marelli, L.; Amaducci, S. Environmentally sustainable biogas? The key role of manure co-digestion with energy crops. Energies 2015, 8, 5234–5265. [Google Scholar] [CrossRef]
- Dai, X.H.; Li, X.S.; Zhang, D.; Chen, Y.G.; Dai, L.L. Simultaneous enhancement of methane production and methane content in biogas from waste activated sludge and perennial ryegrass anaerobic co-digestion: The effects of pH and C/N ratio. Bioresour. Technol. 2016, 216, 323–330. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.L.; Rößler, B.; Zielonka, S.; Wonneberger, A.M.; Lemmer, A. Effects of organic loading rate on the performance of a pressurized anaerobic filter in two-phase anaerobic digestion. Energies 2014, 7, 736–750. [Google Scholar] [CrossRef]
- Dunfield, P.F.; Khmelenina, V.N.; Suzina, N.E.; Trotsenko, Y.A.; Dedysh, S.N. Methylocella silvestris sp. nov., a novel methanotroph isolated from an acidic forest cambisol. Int. J. Syst. Evol. Microbiol. 2003, 53, 1231–1239. [Google Scholar] [CrossRef] [PubMed]
- Appels, L.; Baeyens, J.; Degrève, J.; Dewil, R. Principles and potential of the anaerobic digestion of waste-activated sludge. Prog. Energy. Combust. Sci. 2008, 34, 755–781. [Google Scholar] [CrossRef]
- Imbierowicz, M.; Chacuk, A. Kinetic model of excess activated sludge thermohydrolysis. Water Res. 2012, 46, 5747–5755. [Google Scholar] [CrossRef] [PubMed]
- Ibeid, S.; Elektorowicz, M.; Oleszkiewicz, J. Modification of activated sludge properties caused by application of continuous and intermittent current. Water Res. 2013, 47, 903–910. [Google Scholar] [CrossRef] [PubMed]
- Nah, I.W.; Kang, Y.W.; Hwang, K.Y.; Song, W.K. Mechanical pretreatment of waste activated sludge for anaerobic digestion process. Water Res. 2000, 34, 2362–2368. [Google Scholar] [CrossRef]
- Wu, D.L.; Shen, Y.H.; Ding, A.Q.; Mahmood, Q.; Liu, S.; Tu, Q.P. Effects of nanoscale zero-valent iron particles on biological nitrogen and phosphorus removal and microorganisms in activated sludge. J. Hazard. Mater. 2013, 262, 649–655. [Google Scholar] [CrossRef] [PubMed]
- Suanon, F.; Sun, Q.; Li, M.Y.; Cai, X.; Zhang, Y.C.; Yan, Y.J.; Yu, C.P. Application of nanoscale zero valent iron and iron powder during sludge anaerobic digestion: Impact on methane yield and pharmaceutical and personal care products degradation. J. Hazard. Mater. 2016, 321, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, Y.; Quan, X.; Li, Y.; Zhao, Z.; Meng, X.; Chen, S. Optimization of anaerobic acidogenesis by adding Fe0 powder to enhance anaerobic wastewater treatment. Chem. Eng. J. 2012, 192, 179–185. [Google Scholar] [CrossRef]
- Rasi, S.; Läntelä, J.; Rintala, J. Trace compounds affecting biogas energy utilization-a review. Energy Convers. Manag. 2011, 52, 3369–3375. [Google Scholar] [CrossRef]
- Yang, S.; Tang, Y.; Gou, M.; Jiang, X. Effect of sulfate addition on methane production and sulfate reduction in a mesophilic acetatefed anaerobic reactor. Appl. Microbiol. Biotechnol. 2015, 99, 3269–3277. [Google Scholar] [CrossRef] [PubMed]
- Lar, J.S.; Li, X. Removal of H2S during anaerobic bioconversion of dairy manure. Chin. J. Chem. Eng. 2009, 17, 273–277. [Google Scholar] [CrossRef]
- Syed, M.; Soreanu, G.; Faletta, P.; Béland, M. Removal of hydrogen sulfide from gas streams using biological processes—A review. Can. Biosyst. Eng. 2006, 48, 2.1–2.14. [Google Scholar]
- Kelly, D.P.; Shergill, J.K.; Lu, W.P.; Wood, A.P. Oxidative metabolism of inorganic sulfur compounds by bacteria. Antonie Van Leeuwenhoek 1997, 71, 95–107. [Google Scholar] [CrossRef] [PubMed]
- Van der Zee, F.P.; Villaverde, S.; García-Encina, P.A.; Fdz-Polanco, F. Sulphide removal by moderate oxygenation of anaerobic sludge environments. Bioresour. Technol. 2007, 98, 518–524. [Google Scholar] [CrossRef] [PubMed]
- Goncalves, J.J.; Govind, R. Enhanced biofiltration using cell attachment promotors. Environ. Sci. Technol. 2009, 43, 1049–1054. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, M.; Gomez, J.M.; Aroca, G.; Cantero, D. Removal of hydrogen sulfide by immobilized Thiobacillus thioparus in a biotrickling filter packed with polyurethane foam. Bioresour. Technol. 2009, 100, 4989–4995. [Google Scholar] [CrossRef] [PubMed]
- Janssen, A.J.H.; Ruitenberg, R.; Buisman, C.J.N. Industrial applications of new sulphur biotechnology. Water Sci. Technol. 2001, 44, 85–90. [Google Scholar] [PubMed]
- Meyer-Kohlstock, D.; Haupt, T.; Heldt, E.; Heldt, N.; Kraft, E. Biochar as additive in biogas-production from bio-waste. Energies 2016, 9, 247–256. [Google Scholar] [CrossRef]
- Ciotola, R.J.; Martin, J.F.; Tamkin, A.; Castańo, J.M.; Rosenblum, J.; Bisesi, M.S.; Lee, J. The influence of loading rate and variable temperatures on microbial communities in anaerobic digesters. Energies 2014, 7, 785–803. [Google Scholar] [CrossRef]
- Chu, L.; Yan, S.; Xing, X.H.; Sun, X.; Jurcik, B. Progress and perspectives of sludge ozonation as a powerful pretreatment method for minimization of excess sludge production. Water Res. 2009, 43, 1811–1822. [Google Scholar] [CrossRef] [PubMed]
- Ramos, I.; Díaz, I.; Fdz-Polanco, M. The role of the headspace in hydrogen sulfide removal during microaerobic digestion of sludge. Water Sci. Technol. 2012, 66, 2258–2264. [Google Scholar] [CrossRef] [PubMed]
- American Public Health Association (APHA). Standard Methods for the Examination of Water and Wastewater, 20th ed.; Clescerl, L.S., Greenberg, A.E., Eaton, A.D., Eds.; APHA: Washington, DC, USA, 1998. [Google Scholar]
- Buchholz-Cleven, B.E.; Rattunde, B.; Straub, K.L. Screening for genetic diversity of isolates of anaerobic Fe (II)-oxidizing bacteria using DGGE and whole-cell hybridization. Syst. Appl. Microbiol. 1997, 20, 301–309. [Google Scholar] [CrossRef]
- O’Reilly, J.; Lee, C.; Chinalia, F.; Collins, G.; Mahony, T.; O’Flaherty, V. Microbial community dynamics associated with biomass granulation in low temperature (15 °C) anaerobic wastewater treatment bioreactors. Bioresour. Technol. 2010, 101, 6336–6344. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.B.; Feng, Y.H.; Yu, Q.L.; Xu, Z.B.; Quan, X. Enhanced high-solids anaerobic digestion of waste activated sludge by the addition of scrap iron. Bioresour. Technol. 2014, 159, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Murto, M.; Björnsson, L.; Mattiasson, B. Impact of food industrial waste on anaerobic co-digestion of sewage sludge and pig manure. J. Environ. Manag. 2004, 70, 101–107. [Google Scholar] [CrossRef]
- Ogejo, J.A.; Wen, Z.; Ignosh, J.; Bendfeldt, E.; Collins, E.R. Biomethane Technology; Virginia Cooperative Extension: Blacksburg, VA, USA, 2009; pp. 442–881. [Google Scholar]
- Watson, T.M.; Coleman, A.J.; Williams, G.; McMurray, H.N. The effect of oxygen partial pressure on the filiform corrosion of organic coated iron. Corros. Sci. 2014, 89, 46–58. [Google Scholar] [CrossRef]
- Mamais, D.; Pitt, P.A.; Cheng, Y.W.; Loiacono, J.; Jenkins, D. Determination of ferric chloride dose to control struvite precipitation in anaerobic sludge digesters. Water Environ. Res. 1994, 66, 912–918. [Google Scholar] [CrossRef]
- Batstone, D.J.; Puyol, D.; Flores-Alsina, X.; Rodriguez, J. Mathematical modelling of anaerobic digestion processes: Applications and future needs. Rev. Environ. Sci. Biotechnol. 2015, 14, 595–613. [Google Scholar] [CrossRef]
- Kwietniewska, E.; Tys, J. Process characteristics, inhibition factors and methane yields of anaerobic digestion process, with particular focus on microalgal biomass fermentation. Renew. Sustain. Energy Rev. 2014, 34, 491–500. [Google Scholar] [CrossRef]
- Zhang, Y.B.; Jing, Y.W.; Zhang, J.X.; Sun, L.F.; Quan, X. Performance of a ZVI-UASB reactor for azo dye wastewater treatment. J. Chem. Technol. Biotechnol. 2011, 86, 199–204. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, Y.; Quan, X.; Chen, S.; Zhao, H. Applying an electric field in a built-in zero valent iron—Anaerobic reactor for enhancement of sludge granulation. Water Res. 2011, 45, 1258–1266. [Google Scholar] [CrossRef] [PubMed]
- Jenicek, P.; Keclik, F.; Maca, J.; Bindzar, J. Use of microaerobic conditions for the improvement of anaerobic digestion of solid wastes. Water Sci. Technol. 2008, 58, 1491–1496. [Google Scholar] [CrossRef] [PubMed]
- Jenicek, P.; Koubova, J.; Bindzar, J.; Zabranska, J. Advantages of anaerobic digestion of sludge in microaerobic conditions. Water Sci. Technol. 2010, 62, 427–434. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.T.; Wang, W.; Zhu, S.L.; Wang, F.H. Comparison between the oxidation of iron in oxygen and in steam at 650–750 °C. Corros. Sci. 2013, 75, 309–317. [Google Scholar] [CrossRef]
- Karri, S.; Sierra-Alvarez, R.; Field, J.A. Zero valent iron as an electron-donor for methanogenesis and sulfate reduction in anaerobic sludge. Biotechnol. Bioeng. 2005, 92, 810–819. [Google Scholar] [CrossRef] [PubMed]
- Ramos, I.; Pérez, R.; Reinoso, M.; Torio, R.; Fdz-Polanco, M. Microaerobic digestion of sewage sludge on an industrial-pilot scale: The efficiency of biogas desulphurisation under different configurations and the impact of O2 on the microbial communities. Bioresour. Technol. 2014, 164, 338–346. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.H.; Zhang, Y.B.; Quan, X.; Chen, S. Enhanced anaerobic digestion of waste activated sludge digestion by the addition of zero valent iron. Water Res. 2014, 52, 242–250. [Google Scholar] [CrossRef] [PubMed]
- Botheju, D.; Samarakoon, G.; Chen, C.; Bakke, R. An experimental study on the effects of oxygen in bio-gasification: Part 1. In Proceedings of the International Conference on Renewable Energies and Power Quality, Granada, Spain, 23–25 March 2010.
- Lim, J.W.; Wang, J.Y. Enhanced hydrolysis and methane yield by applying microaeration pretreatment to the anaerobic co-digestion of brown water and food waste. Waste Manag. 2013, 33, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Annachhatre, A.P.; Suktrakoolvait, S. Biological sulfide oxidation in a fluidized bed reactor. Environ. Technol. 2001, 22, 661–672. [Google Scholar] [CrossRef] [PubMed]
- Díaz, I.; Lopes, A.C.; Pérez, S.I.; Fdz-Polanco, M. Performance evaluation of oxygen, air and nitrate for the microaerobic removal of hydrogen sulphide in biogas from sludge digestion. Bioresour. Technol. 2010, 101, 7724–7730. [Google Scholar] [CrossRef] [PubMed]
- Duangmanee, T. Micro-Aeration for Hydrogen Sulfide Removal from Biogas. Ph.D. Thesis, Iowa State University, Ames, IA, USA, 2009. [Google Scholar]
- Hungate, R.E. Chapter IV a roll tube method for cultivation of strict anaerobes. In Methods in Microbiology; Norris, J.R., Ribbons, D.W., Eds.; Academic Press: Millbrae, CA, USA, 1969; Volume 3, Part B; pp. 117–132. [Google Scholar]
- Peu, P.; Sassi, J.F.; Girault, R.; Picard, S.; Saint-Cast, P.; Béline, F.; Dabert, P. Sulphur fate and anaerobic biodegradation potential during co-digestion of seaweed biomass (Ulva sp.) with pig slurry. Bioresour. Technol. 2011, 102, 10794–10802. [Google Scholar] [CrossRef] [PubMed]
- Cadillo-Quiroz, H.; Yavitt, J.B.; Zinder, S.H. Methanosphaerula palustris gen. nov., sp. nov., a hydrogenotrophic methanogen isolated from a minerotrophic fen peatland. Int. J. Syst. Evol. Microbiol. 2009, 59, 928–935. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.H.; Zhang, Y.B.; Chen, S.; Quan, X. Enhanced production of methane from waste activated sludge by the combination of high-solid anaerobic digestion and microbial electrolysis cell with iron–graphite electrode. Chem. Eng. J. 2015, 259, 787–794. [Google Scholar] [CrossRef]
- Tarlera, S.; Stams, A.J.M. Degradation of proteins and amino acids by Caloramator proteoclasticus in pure culture and in coculture with Methanobacterium thermoformicicum Z245. Appl. Microbiol. Biotechnol. 1999, 53, 133–138. [Google Scholar] [CrossRef]
- Zeikus, J.; Weimer, P.; Nelson, D.; Daniels, L. Bacterial methanogenesis: Acetate as a methane precursor in pure culture. Arch. Microbiol. 1975, 104, 129–134. [Google Scholar] [CrossRef]
- Nevin, K.P.; Holmes, D.E.; Woodard, T.L.; Hinlein, E.S.; Ostendorf, D.W.; Lovley, D.R. Geobacter bemidjiensis sp. nov. and Geobacter psychrophilus sp. nov., two novel Fe (III)-reducing subsurface isolates. Int. J. Syst. Evol. Microbiol. 2005, 55, 1667–1674. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.B.; Bacelar-Nicolau, P.; Okibe, N.; Thomas, A.; Hallberg, K.B. Ferrimicrobium acidiphilum gen. nov., sp. nov. and Ferrithrix thermotolerans gen. nov., sp. nov.: Heterotrophic, iron-oxidizing, extremely acidophilic actinobacteria. Int. J. Syst. Evol. Microbiol. 2009, 59, 1082–1089. [Google Scholar] [CrossRef] [PubMed]
- Díaz, I.; Pérez, S.I.; Ferrero, E.M.; Fdz-Polanco, M. Effect of oxygen dosing point and mixing on the microaerobic removal of hydrogen sulphide in sludge digesters. Bioresour. Technol. 2010, 102, 3768–3775. [Google Scholar] [CrossRef] [PubMed]
- Kodama, Y.; Watanabe, K. Sulfuricurvum kujiense gen. nov., sp. nov., a facultatively anaerobic, chemolithoautotrophic, sulfur-oxidizing bacterium isolated from an underground crude-oil storage cavity. Int. J. Syst. Evol. Microbiol. 2004, 54, 2297–2300. [Google Scholar] [CrossRef] [PubMed]
- Potivichayanon, S.; Pokethitiyook, M.; Kruatrachue, M. Hydrogen sulfide removal by a novel fixed-film bioscrubber system. Process Biochem. 2006, 41, 708–715. [Google Scholar] [CrossRef]


| Parameters | Units | WAS | APS |
|---|---|---|---|
| TCOD | mg/L | 31,253 ± 4107 | 30,854 ± 4494 |
| SCOD | mg/L | 121 ± 25 | 721 ± 28 |
| TS | g/L | 33.6 ± 4.9 | 32.9 ± 4.2 |
| VS | g/L | 20.7 ± 2.7 | 20.3 ± 2.4 |
| TSS | g/L | 32.4 ± 4.3 | 31.5 ± 3.9 |
| VSS | g/L | 19.8 ± 2.1 | 19.2 ± 1.8 |
| pH | / | 7.16 ± 0.11 | 7.06 ± 0.04 |
| Total Fe | mg/L | 31.2 ± 0.97 | 31.2 ± 0.94 |
| Soluble Fe | mg/L | 8.7 ± 0.15 | 9.4 ± 0.16 |
| Soluble Sulphide | mg/L | 34.7 ± 1.5 | 43.9 ± 1.8 |
| SO42− | mg/L | 3.4 ± 0.18 | 5.5 ± 0.22 |
| Total Elemental sulphur | mg/L | 279.6 ± 13.1 | 279.6 ± 11.8 |
| Period | P1 | P2 | P3 | P4 | P5 | P6 | P7 |
|---|---|---|---|---|---|---|---|
| Condition | An a | An | Ma b | Ma | Ma | Ma | Ma |
| Day | 1–30 | 31–60 | 61–90 | 91–120 | 121–150 | 151–180 | 181–210 |
| Duration (day) | 30 | 30 | 30 | 30 | 30 | 30 | 30 |
| RSI dosages (g/L) | 0 | 20 | 20 | 20 | 20 | 20 | 20 |
| ABPR (mL/day) | 937 | 1671 | 1645 | 1592 | 1644 | 1591 | 1197 |
| O2 supply (mL) | 0 | 0 | 1 | 5 | 10 | 15 | 20 |
| O2/biogas c | 0 | 0 | 0.61 | 3.14 | 6.08 | 9.43 | 16.71 |
| BRT (h) | 12.8 | 7.1 | 7.2 | 7.5 | 7.2 | 7.5 | 10.0 |
| Parameter | P1 | P2 | P3 | P4 | P5 | P6 | P7 |
|---|---|---|---|---|---|---|---|
| COD in (mg/L) | 31,253 | 31,253 | 31,253 | 31,253 | 31,253 | 31,253 | 31,253 |
| Average COD out (mg/L) | 19,314 | 13,314 | 13,658 | 14,314 | 13,908 | 14,126 | 17,189 |
| Average COD removal (%) | 38.2 | 57.4 | 56.3 | 54.2 | 55.5 | 54.8 | 45.0 |
| VS in (g/L) | 20.3 | 20.3 | 20.3 | 20.3 | 20.3 | 20.3 | 20.3 |
| Average VS out (g/L) | 12.7 | 8.9 | 8.9 | 9.5 | 9.5 | 9.4 | 11.1 |
| Average VS removal (%) | 37.6 | 56.4 | 56.1 | 54.2 | 54.2 | 53.5 | 45.3 |
| Average Methane Yield (mL/g COD) | 218.76 | 306.00 | 300.59 | 298.54 | 298.90 | 301.13 | 261.39 |
| Average Methane Yield (mL/gVSfed) | 342.3 | 484.6 | 465.3 | 457.2 | 465.7 | 486.4 | 406.8 |
| Average H2S concentration (ppmv) | 12504 | 1933 | 776 | 484 | 234 | 75 | 68 |
| Average O2 concentration (ppmv) | / a | / | 158 | 759 | 1230 | 2083 | 8438 |
| Average CH4 concentration (ppmv) | 557,964 | 656,972 | 642,738 | 634,655 | 630,789 | 648,816 | 614,308 |
| Average CO2 concentration (ppmv) | 419,672 | 332,948 | 346,459 | 356,842 | 356,129 | 341,027 | 372,648 |
| Phase | Sulfur Species | Feed | P1 | P2 | P3 | P4 | P5 | P6 | P7 |
|---|---|---|---|---|---|---|---|---|---|
| S in solid phase (mg) | S-sulfur compounds in waste sludge | 55.9 | 41.8 | 38.3 | 33.8 | 33.1 | 31.4 | 32.5 | 37.1 |
| S-iron sulphide | / | 2.2 | 41.7 | 45.1 | 51.2 | 58.9 | 55.1 | 51.1 | |
| S0 | / | / | / | 26.9 | 32.7 | 37.2 | 42.4 | 30.1 | |
| S in liquid phase (mg) | S-Total sulfide (S2−, HS−, H2S) | 8.3 | 78.6 | 53.8 | 26.6 | 20.1 | 15.9 | 11.2 | 7.6 |
| S-Sulphide oxidation products (S2O32−, SO42−) | 90.5 | 4.7 | 5.1 | 4.9 | 5.4 | 4.8 | 5.7 | 23.9 | |
| S in gaseous phase (mg) | S-H2S | / | 16.7 | 4.6 | 1.8 | 1.1 | 0.6 | 0.3 | 0.2 |
| Total Elemental Sulphur (mg) | 154.7 | 144 | 143.5 | 139.1 | 143.6 | 148.8 | 147.2 | 150 | |
| Band No. | Closest Relatives | Similarity (%) | Classification |
|---|---|---|---|
| A1 | Methanoculleus palmolei (NR028253) | 99 | Methanomicrobia |
| A2 | Methanosphaerula palustris (NR074167) | 92 | Methanomicrobia |
| A3 | Methanoculleus marisnigri (NR074174) | 98 | Methanomicrobia |
| A4 | Methanolinea tarda (NR028163) | 97 | Methanomicrobia |
| A5 | Methanosarcina mazei (NR041956) | 100 | Methanomicrobia |
| A6 | Methanoculleus bourgensis (NR042786) | 98 | Methanomicrobia |
| A7 | Methanosarcina mazei (NR074221) | 93 | Methanomicrobia |
| A8 | Methanosaeta concilii (NR102903) | 99 | Methanomicrobia |
| A9 | Methanosarcina siciliae (NR104757) | 99 | Methanomicrobia |
| A10 | Methanosarcina barkeri (NR025303) | 97 | Methanomicrobia |
| A11 | Methanococcus aeolicus (NR029140) | 93 | Methanococci |
| B1 | Methylovirgula ligni (NR044611) | 98 | Alphaproteobacteria |
| B2 | Methylocella silvestris (NR074237) | 98 | Alphaproteobacteria |
| B3 | Uncultured bacterium (JQ726674) | 98 | Alphaproteobacteria |
| B4 | Sphingopyxis witflariensis (NR028010) | 99 | Alphaproteobacteria |
| B5 | Novosphingobium indicum (NR044277) | 99 | Alphaproteobacteria |
| B6 | Clostridium orbiscindens DSM 6740 (NR029356) | 99 | Clostridia |
| B7 | Clostridium populeti (NR026103) | 93 | Clostridia |
| B8 | Caloramator proteoclasticus (NR026265) | 95 | Clostridia |
| B9 | Clostridium sp. (JQ670700) | 94 | Clostridia |
| B10 | Syntrophobacter fumaroxidans (NR027598) | 94 | Deltaproteobacteria |
| B11 | Acetobacter tropicalis (NR036881) | 95 | Alphaproteobacteria |
| B12 | Roseomonas lacus (NR042318) | 98 | Alphaproteobacteria |
| B13 | Sporomusa silvacetica (NR026378) | 98 | Alphaproteobacteria |
| B14 | Geobacter bemidjiensis Bem (NR042769) | 97 | Deltaproteobacteria |
| B15 | Acidimicrobium ferrooxidans (NR074390) | 92 | Acidimicrobidae |
| B16 | Ferrimicrobium acidiphilum (NR041798) | 93 | Acidimicrobidae |
| B17 | Uncultured Arcobacter sp. (HQ392839) | 99 | Epsilonproteobacteria |
| B18 | Uncultured Epsilonproteobacterium (DQ295695) | 99 | Epsilonproteobacteria |
| B19 | Uncultured Acinetobacter sp. (EU567041) | 95 | Gammaproteobacteria |
| Consumption | Price | ||||||
|---|---|---|---|---|---|---|---|
| RSI | Sodium Hydroxide | Hydrochloric Acid | Oxygen | RSI | Sodium Hydroxide | Hydrochloric Acid | Oxygen |
| g/period | g/period | mL/period | L/period | $/kg | $/kg | $/L | $/m3 |
| 8.4 | 7.2 | 17.8 | 0.45 | 1.6 | 20 | 20 | 57.7 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruan, R.; Cao, J.; Li, C.; Zheng, D.; Luo, J. The Influence of Micro-Oxygen Addition on Desulfurization Performance and Microbial Communities during Waste-Activated Sludge Digestion in a Rusty Scrap Iron-Loaded Anaerobic Digester. Energies 2017, 10, 258. https://doi.org/10.3390/en10020258
Ruan R, Cao J, Li C, Zheng D, Luo J. The Influence of Micro-Oxygen Addition on Desulfurization Performance and Microbial Communities during Waste-Activated Sludge Digestion in a Rusty Scrap Iron-Loaded Anaerobic Digester. Energies. 2017; 10(2):258. https://doi.org/10.3390/en10020258
Chicago/Turabian StyleRuan, Renjun, Jiashun Cao, Chao Li, Di Zheng, and Jingyang Luo. 2017. "The Influence of Micro-Oxygen Addition on Desulfurization Performance and Microbial Communities during Waste-Activated Sludge Digestion in a Rusty Scrap Iron-Loaded Anaerobic Digester" Energies 10, no. 2: 258. https://doi.org/10.3390/en10020258
APA StyleRuan, R., Cao, J., Li, C., Zheng, D., & Luo, J. (2017). The Influence of Micro-Oxygen Addition on Desulfurization Performance and Microbial Communities during Waste-Activated Sludge Digestion in a Rusty Scrap Iron-Loaded Anaerobic Digester. Energies, 10(2), 258. https://doi.org/10.3390/en10020258