1. Introduction
Available studies have confirmed that cancer cells exhibit multiple genetic mutations at the somatic level, the nature of which is related to the genetic susceptibility of the individual organisms. Since then, many susceptibility genes and their polymorphisms have been identified for esophageal cancer, and polymorphisms of susceptibility genes have been identified, most of which are concentrated in gene regions related to carbohydrate and lipid metabolism and vitamin synthesis [
1,
2]. In patients with esophageal cancer, the rapid proliferation of tumor cells compared to normal cells and the increased energy requirements result in significant differences between the metabolism of tumor and normal tissues [
3]. Previous studies have revealed that genes involved in lipid metabolism are highly expressed in malignant tumors and are considered as oncogenes [
4,
5]. Epidemiological studies have shown that obese individuals are at increased risk of malignancies, including esophageal squamous cell carcinoma, suggesting that fatty acid metabolism plays a key role in tumorigenesis and progression [
5,
6,
7]. Fatty acids are one of the main energy sources in mammals and are crucial for the normal growth and metabolism of cells. However, the relationship between fatty acid metabolism and the pathological development of esophageal cancer remains unclear, and the relationship between lipid metabolism and the potential clinical prognosis of esophageal cancer has not been explored.
Opa-interacting protein 5 (OIP5), an Opa-interacting protein associated with eisseria gonorrhoeae opacity [
8], is highly expressed in the human testis [
9]. The upregulation of OIP5 is associated with adverse clinical features in multiple cancer types, including gastric cancer [
10,
11]. Downregulation of OIP5 has been reported to inhibit the proliferation of cultured colorectal and gastric cells [
10]. Given the limited evidence for the association of OIP5 with oncogenic events, the mechanisms of OIP5-promoted tumorigenesis in esophageal cancer needs to be further investigated.
Our study demonstrated for the first time that OIP5 promotes the occurrence and development of esophageal cancer cells by mediating intracellular lipid metabolism. The study of the target gene OIP5 provides reference for the subsequent research of its function, mechanism of action, and clinical significance. However, the effect of OIP5 on fatty acid metabolism in esophageal squamous cell carcinoma has not been investigated. This study provides a new perspective on the role of intracellular fatty acid metabolism in the development of esophageal squamous cell carcinoma and may provide new insights into the diagnosis and potential treatment of esophageal cancer. It also presents a theoretical basis for the improvement of ESCC prognostic models, the elucidation of drug resistance mechanisms, and the development of corresponding targeted therapies.
2. Materials and Methods
2.1. Download ESCC-Related Expression Profile Dataset
Data from public databases were used in this study to analyze gene expression in patients with esophageal squamous cell carcinoma and healthy/control patients. GSE20347 and GSE75241 datasets were downloaded from the GEO database (
https://www.ncbi.nlm.nih.gov/geo/ (accessed on 1 July 2020)). The GSE20347 dataset is based on the GPL571 platform ( [HG-U133A_2] Affymetrix Human Genome U133A 2.0 Array), while the GSE75241 dataset is based on the GPL5175 platform ([HuEx-1_0-st] Affymetrix Human Exon 1.0 ST Array [transcript (gene) version]). The array data for GSE20347 include 17 esophageal squamous cells and their matched normal tissues as control. The GSE75241 dataset include 15 esophageal squamous cell carcinoma samples and 15 non-malignant mucosal samples. In addition to these, in order to obtain a sufficient number of ESCC samples, we also obtained 80 ESCC samples and 79 normal control tissues to be compared against them from The Cancer Genome Atlas (TCGA) using the USCS Refseq Gene Array (
https://www.gtexportal.org/ (accessed on 2 July 2020)) and 1 adjacent non-tumor tissue sample (
https://tcga-data.nci.nih.gov/tcga/ (accessed on 2 July 2020)).
2.2. Differential Expression Genes Recognition
We used the downloaded data for background correction, using R to convert gene probe IDs from microarray data into genomic symbols in platform files. The dataset was normalized accordingly using the
t-test ((Version 3.3,
http://www.bioconductor.org (accessed on 14 October 2022))) of the linear model of the LIMMA package. We set the intercept value to |Log 2 FC| ≥ 1 (adjusted
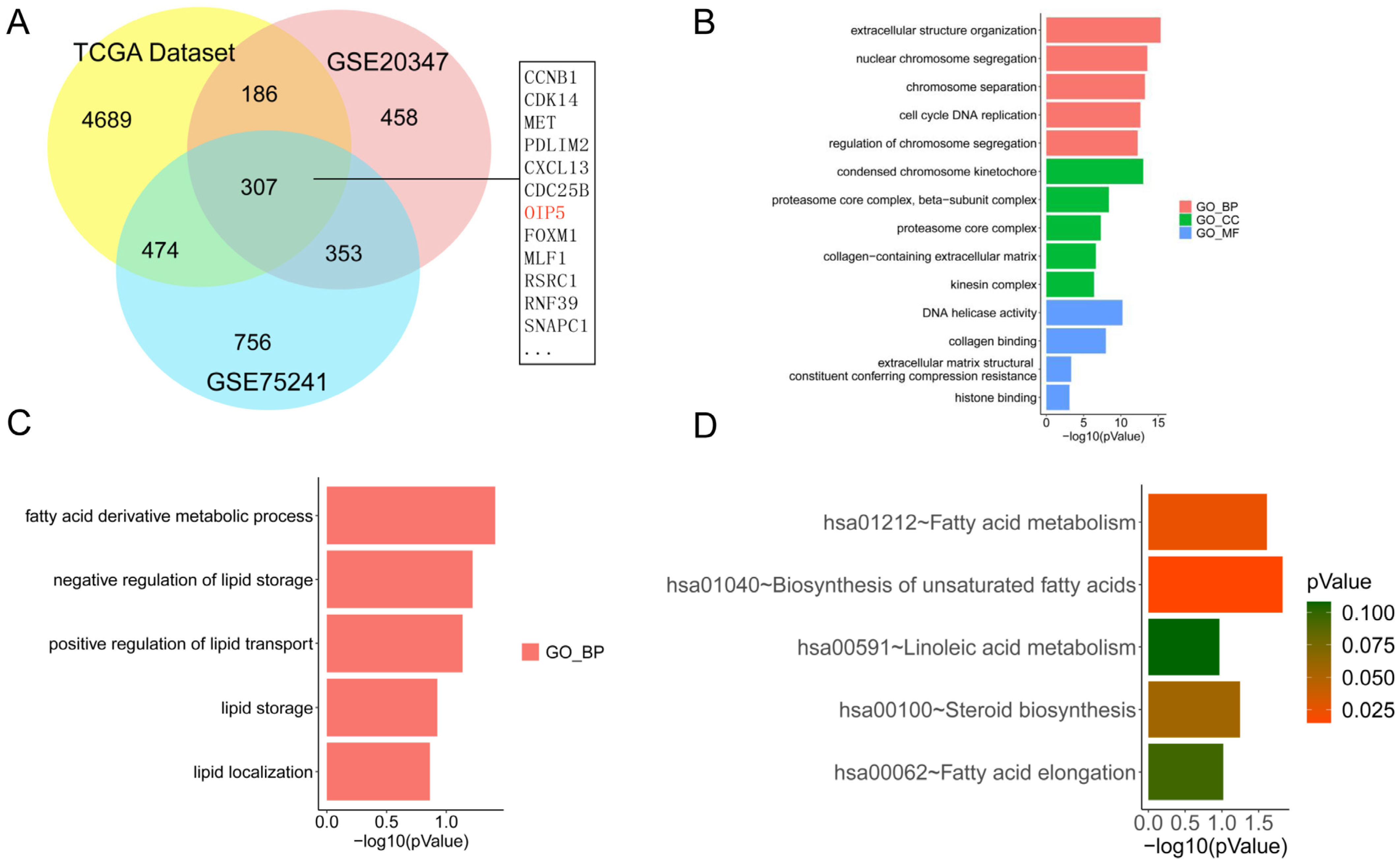
p-value < 0.05) to select differentially expressed genes between ESCC samples and normal control tissues. Next, we plotted the overlapping parts of the three datasets and found 307 differentially expressed genes (DEGs) by using a Venn plot (
http://bioinformatics.psb.ugent.be/webtools/Venn/ (accessed on 26 August 2022)). At the same time, we performed gene ontology (GO) enrichment analysis on these differentially expressed genes to elucidate the GO functions of DEGs, such as biological processes (BP), molecular function (MF), and cellular components (CC). We also selected the KEGG enrichment pathway related to lipid metabolism to determine whether these DEGs were related to lipid metabolism in the development of esophageal cancer. A
p < 0.05 was used as the cut-off standard for significant functions and pathways. We also performed a GSEA analysis to determine whether these differential genes are enriched in lipid metabolism in esophageal squamous cell carcinoma. Finally, we present a protein–protein interaction (PPI) network encoded by lipid metabolism-related genes OIP5 using the STRING online database (Version 10.0,
http://string-db.org (accessed on 26 August 2022)) and the Cytoscape software (Version 3.8.0,
http://www.cytoscape.org/ (accessed on 26 August 2022)).
2.3. Cell Culture, Cell Transfection and Construction of Gene Downregulation, and Overexpression Lentiviral Vectors
ESC-410 and KYSE-150 ESCC cell lines were purchased from the Shanghai Chinese Academy of Sciences Type Culture Collection Cell Bank. The two cell lines were cultured in culture flasks in growth medium RPMI1640 and incubated in an incubator (5% CO2, 37 °C) with 10% fetal bovine serum (Invitrogen, Waltham, MA, USA) + 1% penicillin streptomycin (Gibco, Gaithersburg, MD, USA). All experiments were repeated at least 3 times with both cell lines.
Plasmid-transfected viruses carrying the empty vector (pLVX-IRES-puro) and OIP5 and acyl-CoA synthetase long-chain family member 1 (ACSL1) gene vectors (three plasmid systems including pSAX2, pMD2G, and pLVX-IRES-puro-OIP5/ACSL1) were transfected using the helper dye Lipo3000 (Invitrogen, USA). The transfection was divided into 5 groups including the negative control group (Negative control, NC), the OIP5 and ACSL1 gene-overexpression groups (shRNA + RNA), and the OIP5 and ACSL1 gene-silencing groups (shRNA). Different lentiviruses (pLVX-IRES-puro-OIP5/ACSL1) and the same volume of control viral vector were added to them. Stable OIP5 and ACSL1 gene-silencing cell lines were obtained by screening ESC-410 and KYSE-150 ESCC cells with puromycin (Gibco, Gaithersburg, MD, USA) (2 ug/mL).
2.4. Tissue Microarray and Immunohistochemistry
To investigate the expression pattern of OIP5 in gastric cancer tissues, we used immunohistochemistry to detect tissue chips (chips were purchased from Shanghai Xinchao Biotechnology Co., Ltd. (Shanghai, China), including 2 normal esophageal mucosal tissues, 29 ESCC tissues, and the expression of OIP5 protein in paired 29 paracancerous tissues). It was found that OIP5 protein expression was significantly higher in ESCC tissues than in paraneoplastic tissues as well as in normal esophageal mucosal tissues. The results showed that there is significant difference in OIP5 expression between ESCC tissues and adjacent tissues.
2.5. RNA Extraction, cDNA (Complementary DNA) Synthesis and Real-Time PCR Analysis
Nude-mouse tumor tissue (50–100 mg) was ground into powder, added to 1 mL TRIzol lysis solution, and allowed to stand for 5 min at room temperature. After dissolving, it was transferred to a sterile centrifuge tube, and the supernatant was collected by centrifugation. The dead cells were washed in pre-chilled PBS and placed on ice after the cell culture media aspirated from the plate. We added 300–400 µL of TRIzol reagent to each well, mixed thoroughly until it became a homogeneous and translucent liquid state, and incubated it on ice for 5 min. The cells containing lysates in each well were pipetted into the EP tube, and the experimental group and the experimental group number were marked. We added 1/5 Trizol volume of chloroform, shook vigorously, and let it stand at room temperature for 3 min. The centrifuge was frozen at 4 °C, and centrifuged at 12,000 rpm for 30 min. The supernatant was transferred to a new EP tube, 50% triazole volume of isopropanol was added, gently mixed, and left at room temperature for 10 min. The centrifuge was set to 4 °C, at 12,000 rpm × 30 min, and the supernatant was discarded and then continued at 12,000 rpm × 5 min. A white RNA precipitate was observed in the EP tube. The same amount of 75% DEPC H2O ethanol was then added to the EP tube for washing. The centrifuge was set to 4 °C, at 7500 rpm × 5 min, and the supernatant was discarded, and the process was repeated twice. Then, the particles were dried, and a proper amount of DEPC H2O was added and incubated at 55–60 °C for 10 min to dissolve the precipitate and to obtain an RNA solution. RNA concentration was measured by NanoDrop spectrophotometer (Eppendorf, Hamburg, Germany).
Then, we used reverse transcription kits, which reverse transcribe RNA samples to cDNA.(Tiangen Biochemical Technology Co., Ltd., Beijing, China).
Finally, a real-time quantitative polymerase chain reaction analysis was performed using a real-time quantitative PCR kit (Tiangen Biochemical Technology Co., Ltd., Beijing, China). To perform RT-qPCR, 1 µL of cDNA was mixed with 1 µL of gene-specific forward primers, 1 µL of gene-specific reverse primers, and 10 µL of SYBR Green Real-time PCR Master Mix, and reactions were detected using the LightCycler 480 II instrument (Roche, Basel, Switzerland). The detailed RT-PCR reaction is shown below: 1 cycle of pre-denaturation at 95 °C for 30 min, followed by 40 cycles of denaturation at 95 °C for 15 s, annealing at 60 °C for 30 s, and extension at 72 °C for 30 s. Finally, the temperature was slowly raised to the range of 55–95 °C, and at each temperature, the sample was cycled for 30 s, for a total of 40 cycles. The CT values and related parameters of each group were recorded, and the relative gene expression levels were calculated by the comparative CT method (ΔΔ-CT). The primer sequences are shown in
Table 1.
2.6. Quantification of Associated Lipids in ESCC
Different experimental groups were designed, and each group of cells was inoculated in 6-well plates at 5 × 105 per well and placed in an incubator (37 °C, 5% CO2) for 24 h to reach 40–50% confluence. Cells were lysed in 5% Triton X-100 Cell Lysis Buffer. Intracellular neutral lipids were detected using BODIPY 493/503 dye (Thermo Fisher, Waltham, MA, USA) according to the manufacturer’s instructions. Intracellular triglyceride (TAG) and phospholipid (PL) content was measured using a triglyceride quantitative detection kit (BioAssay Systems, Hayward, CA, USA) and a phospholipid quantitative detection kit (BioAssay Systems), respectively.
2.7. Cell Proliferation Assay (CCK-8 Method)
To determine whether OIP5 affects the colony-forming ability of ESCC cells, we designed three groups including control groups (sh-NC), a silenced-OIP5 gene group (sh-OIP5), and an exogenously overexpressing-OIP5 gene group after silencing the OIP5 gene (sh-OIP5 + OIP5). Stably transfected ESCC cells were removed, digested, centrifuged, and resuspended, and 100 µL of cell suspension was inoculated in 96-well culture plates at a density of 2 × 103 cells/mL. The plates were pre-incubated in an incubator (37 °C, 5% CO2) for 24 h and detected at 0, 24, 48, and 72 h. In addition, at the corresponding incubation time points, we added CCK8 solution (10 µL) to each well and placed the plate in an incubator (37 °C, 5% CO2) for 1 h. Finally, OD values (450 nm) were detected by a microplate reader, and cell growth curves were plotted in combination with the assay results at each time point.
2.8. Clone Formation Assay
Logarithmic growth phase cells were routinely digested, pipetted into individual cells and resuspended in DMEM medium (Gibco, USA) containing 10% fetal bovine serum. We diluted the cell suspension to 100 cells per dish. The density was inoculated in dishes containing 10 mL of pre-warmed medium, and the dishes were placed in an incubator (37 °C, 5% CO2) for 2 weeks. When macroscopic clones appeared in the petri dish, the culture process was considered to be terminated. The supernatant was discarded, and PBS was used for washing twice. We fixed with 4% paraformaldehyde for 15 min, discarded the fixative, and stained the colonies with Giemsa solution (Sinopharm Chemical Reagent, Shanghai, China) for 15 min, washed the staining solution with flowing water, and dried at room temperature. Colonies with more than 50 cells were counted under a microscope and histograms were drawn.
2.9. Mouse Xenograft
ESC-410 ESCC cells were stably transfected with OIP5 overexpression plasmid or empty vector, and untransfected ESC-410 cells were set as the control group. After PBS digestion and centrifugation of ESC-410 cells, cells were washed twice with PBS to remove serum. Cell pellets were pipetted to the appropriate concentration using serum-free medium.
Cells from different groups (approximately 2 × 10
7 cells) were inoculated subcutaneously onto the back of BALB/c nude mice (5 weeks old, male). The date of inoculation was recorded as D0, and the tumor volume was measured on D5, D10, D15, D20, D25, and D30. The tumor volume was calculated using the equation: tumor volume (mm
3) = 0.5 × (a × b
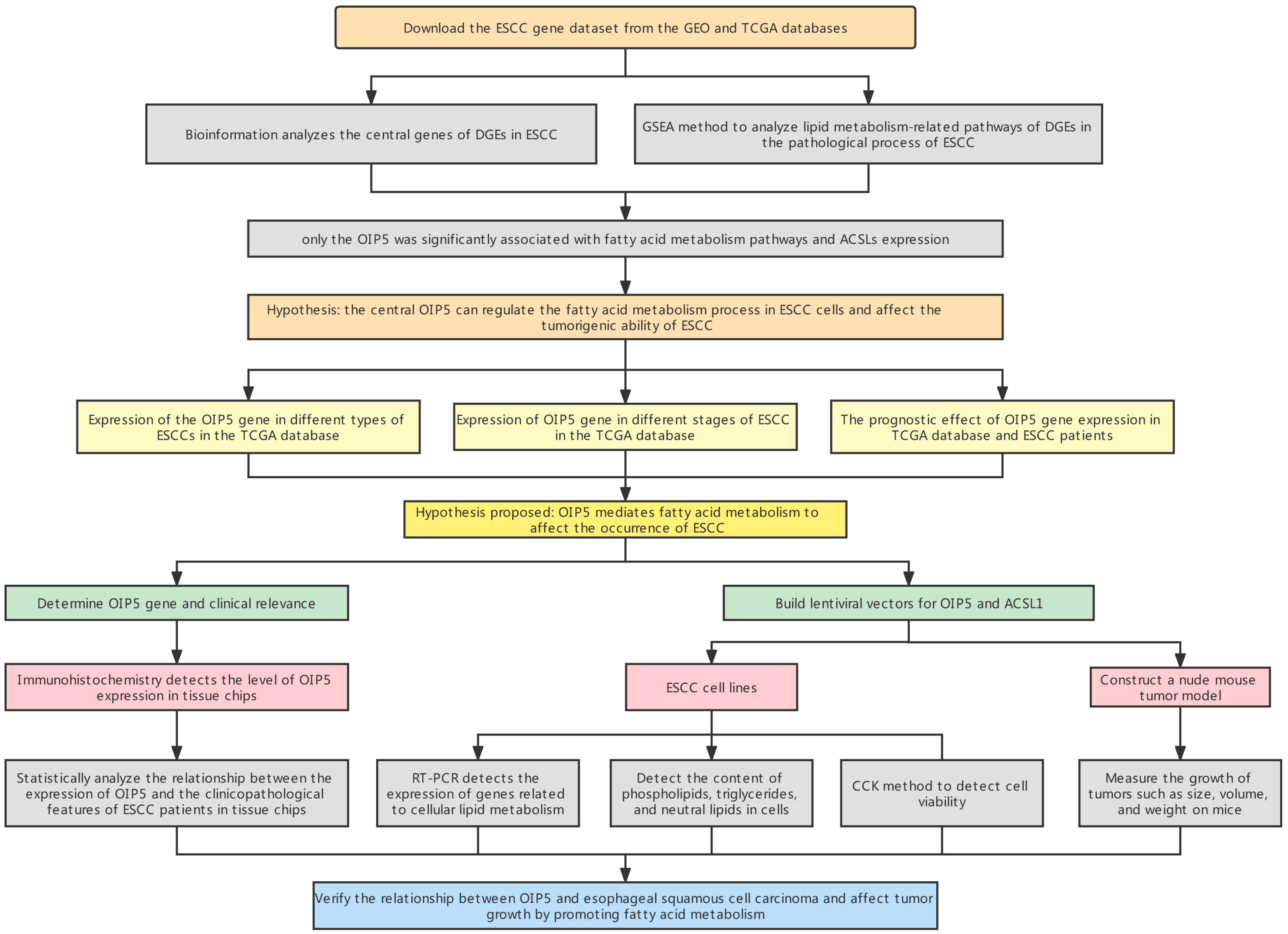
2), where a corresponds to the tumor’s long diameter and b corresponds to the tumor’s short diameter. Tumor size was measured, and growth curves were plotted. After 30 days, nude mice were executed, subcutaneous tumors on the back of nude mice were excised for photographing and recording, and curves were plotted using tumor volume data recorded on D0, D5, D10, D15, D20, D25, and D30. Tumor size, weight, and gene expression patterns were further analyzed. All procedures were approved by Ethics Committee of Changzhou First People’s Hospital (The specific flow chart is shown in
Figure 1).
2.10. Statistical Analysis
Data were statistically analyzed with Graphpad Prism 9.0 and expressed as mean ± standard deviation of at least three independent experiments. The data were analyzed using R software.
Quantitative data were subjected to a one-way ANOVA or two-sided t-test under normal distribution and chi-square, otherwise, the Wilcoxon rank-sum test was performed. A value of p < 0.05 was considered as statistically significant.
4. Discussion
Despite improvements in surgical techniques and adjuvant chemoradiotherapy, esophageal squamous cell carcinoma has a poor prognosis among malignancies. The pathogenesis of esophageal cancer has not been thoroughly investigated, but abnormal gene expression is still considered as an important factor in the occurrence of esophageal cancer. Gene expression profiling (also known as transcriptomics) enables the detection of entire genomes, which may help elucidate tumorigenesis. Metabolomics provides a wealth of data information reflecting genetic and epigenetic metabolic alterations and is dedicated to the identification and development of metabolically active targets in cancer therapeutics and pharmacology [
12]. Thus, the integration of metabolomics with transcriptomics may provide a deeper understanding of tumor pathogenesis [
13].
OIP5 is a protein-coding gene located on chromosome 15 and belongs to the cancer/testis antigen (CTA) gene family [
7]. OIP5 encodes a 25 kDa protein with a helical-structured domain, originally identified as an Opa (Neisseria gonorrhoeae opacity-associated) interacting protein by a yeast two-hybrid analysis [
14]. Aberrant OIP5 expression is common in various types of cancer, including glioblastoma [
15], bladder, esophagus [
16], breast [
17], gastric, colorectal [
10], and liver cancer [
18]. The expression status and biological function of OIP5 in esophageal squamous cell carcinoma and the exact mechanism of OIP5 have not been well studied.
In the early stage, we obtained differentially expressed genes (DEGs) in esophageal squamous cell carcinoma by analyzing the GEO and TCGA databases, specifically, we analyzed the TCGA dataset by gene ensemble enrichment analysis (GSEA). We found that OIP5 in differentially expressed genes was significantly associated with fatty acid-metabolism pathways. OIP5 was identified as the central gene, and the PPI network was constructed. Epidemiological studies have shown that the risk of esophageal cancer in obese people is increased, and previous literature has reported that fatty acid metabolism is closely related to the occurrence and development of malignant tumors. This suggests that the central gene OIP5 may affect the fatty acid metabolism of esophageal squamous cell carcinoma and the occurrence and development of esophageal squamous cell carcinoma. We found that the expression of ASCL1, a key enzyme for the activation of fatty acids in ESCC, correlated significantly with the expression pattern of OIP5 through preliminary experiments. OIP5 silencing or overexpression alters the content of fatty acids and their metabolic intermediates such as PLs and TGs in ESCC cells by affecting the expression of ACSL1. Through in vitro cell experiments and tumorigenic experiments in nude mice, it was elucidated that OIP5 affects fatty acid metabolism in mice by promoting the expression of ACSL1. OIP5 is a tumor-promoting factor of esophageal squamous cell carcinoma cells, which can promote the occurrence and development of esophageal squamous cell carcinoma. Our research is of great significance for elucidating the occurrence of ESCC by identifying the central genes and key pathways of esophageal cancer and may provide new ideas for the diagnosis and treatment of ESCC. It is expected to improve the ESCC prognostic model, elucidate the drug-resistance mechanism, and develop corresponding targets to provide a rationale for treatment.
This study also has certain limitations and shortcomings, such as those of the results we obtained from the knockdown and exogenous overexpression in two ESCC cell lines of several common fatty acid-metabolizing enzymes ACSL1, ACSL2, HADH, ADH4, and ALDH1B. OIP5 did not significantly downregulate and restore the mRNA expression of ADH4 and ALDH1B. We speculate that this may be due to experimental error or because ADH4 and ALDH1B are not the key fatty acid-metabolizing enzymes for OIP5-mediated lipid metabolism in ESCC. When triglycerides, phospholipids, and neutral lipids were used as the detection targets, satisfactory positive results were obtained regardless of OIP5 knockdown or overexpression. Triglyceride, phospholipid, and neutral lipid content were significantly recovered. This indicates that OIP5 is inseparable from ESCC lipid metabolism. Although our design is well-established, there are still some inconsistencies in the experimental results. In addition, the number of ESCC tissue chips we purchased was small, and the difference between immunohistochemical ESCC and normal tissues made by the tissue chips was not significant. The conclusion that OIP5 is related to the clinicopathological factors of ESCC needs to be further confirmed by larger samples. This study can provide a basis for further verification and research for new ESCC central genes for later generations.