Immune Biomarkers in Blood from Sarcoma Patients: A Pilot Study
Abstract
:1. Introduction
2. Materials and Methods
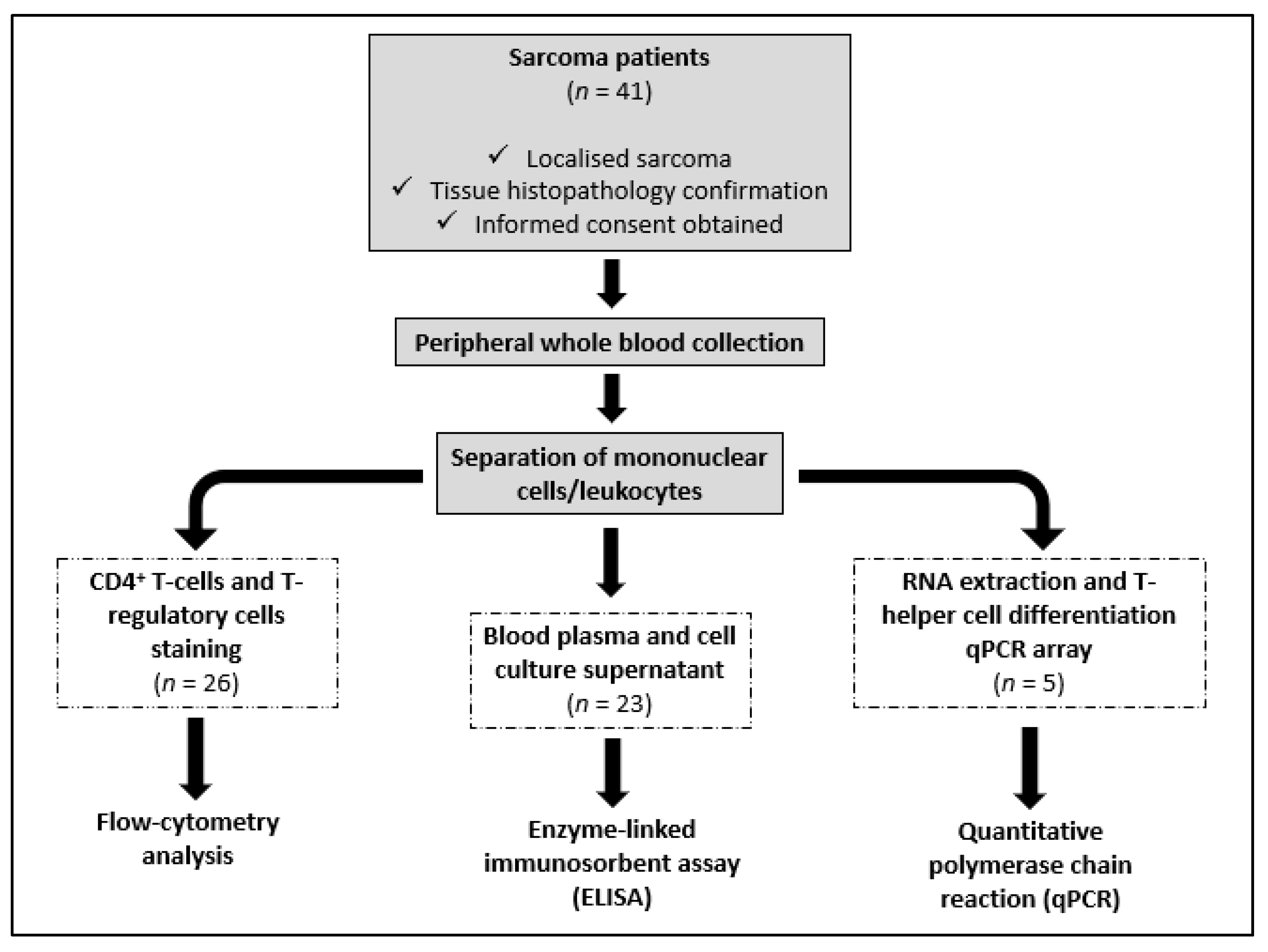
2.1. Sarcoma Patients and Healthy Volunteers
2.2. Analysis of CD4+ T Cells and T-Regulatory Cells in Peripheral Blood
2.3. Peripheral Blood Leukocyte Culture and Quantification of Cytokines
2.4. RNA Isolation and Quantitative PCR Array
2.5. Statistical Analysis
3. Results
3.1. Demographic Data
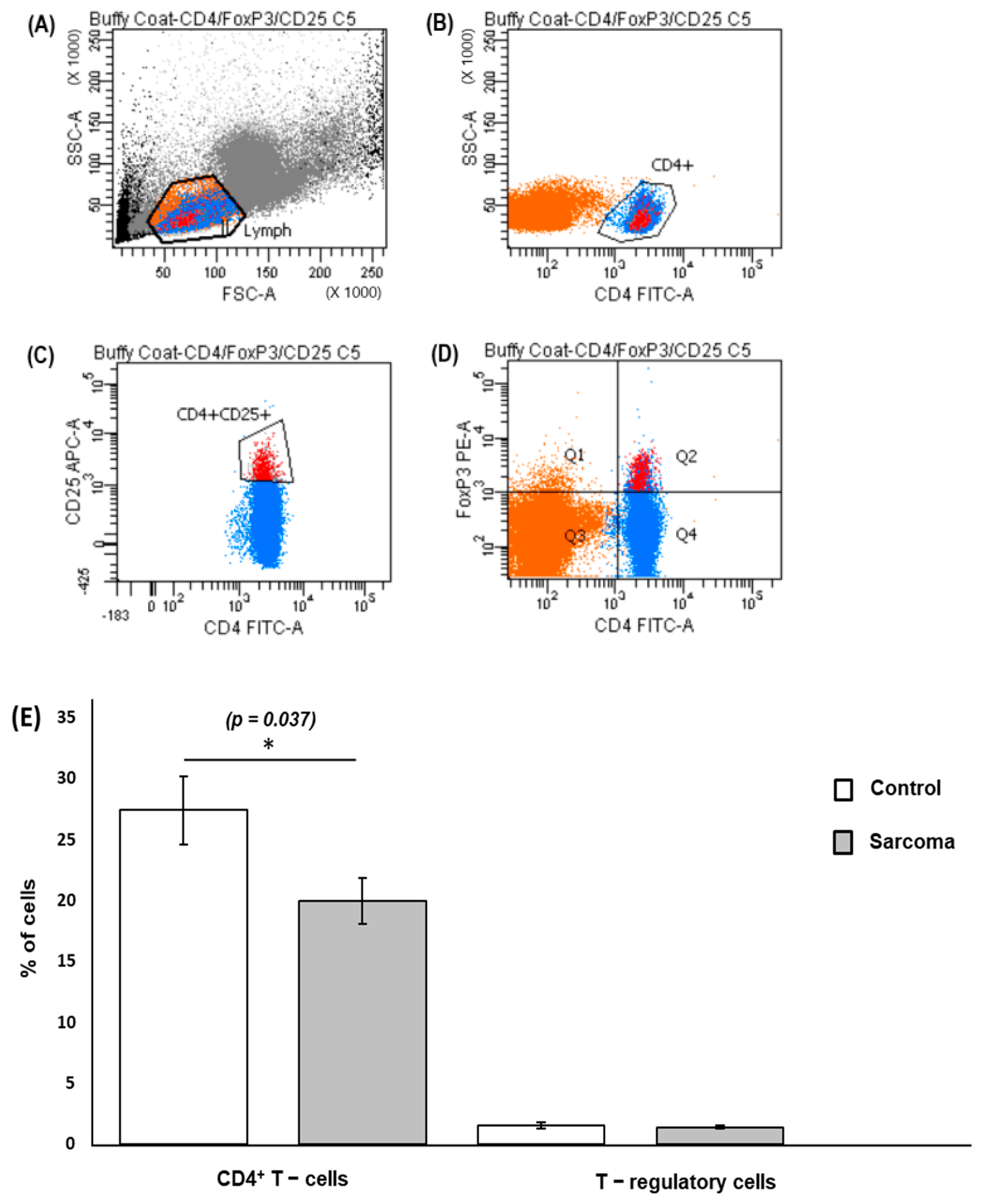
3.2. CD4+ T Cells and Treg Cells in Peripheral Blood
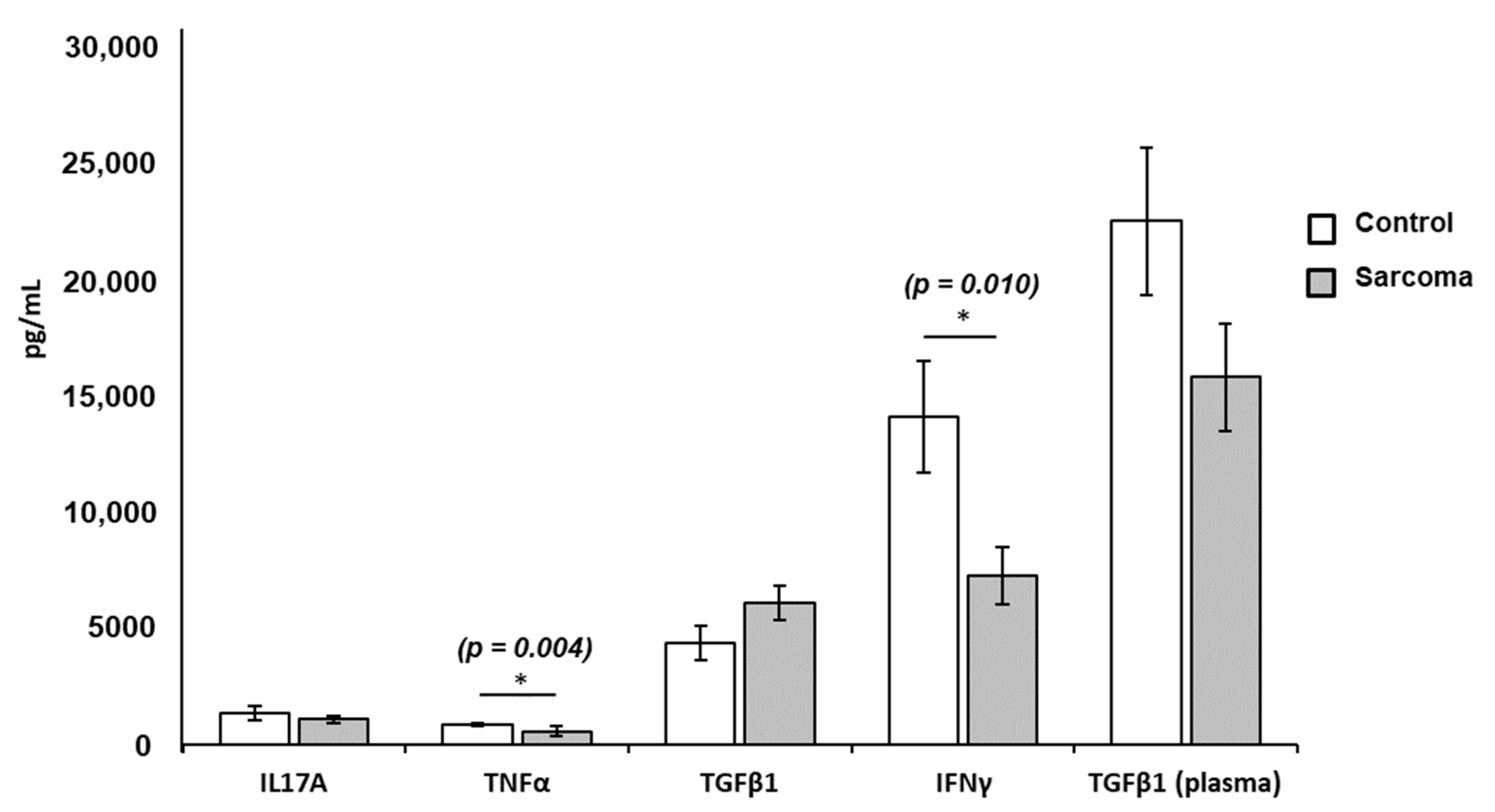
3.3. Cytokine Levels in Culture Supernatants and Plasma
3.4. Correlation among the Levels of Cytokines and T Cells
3.5. Hematological Analysis
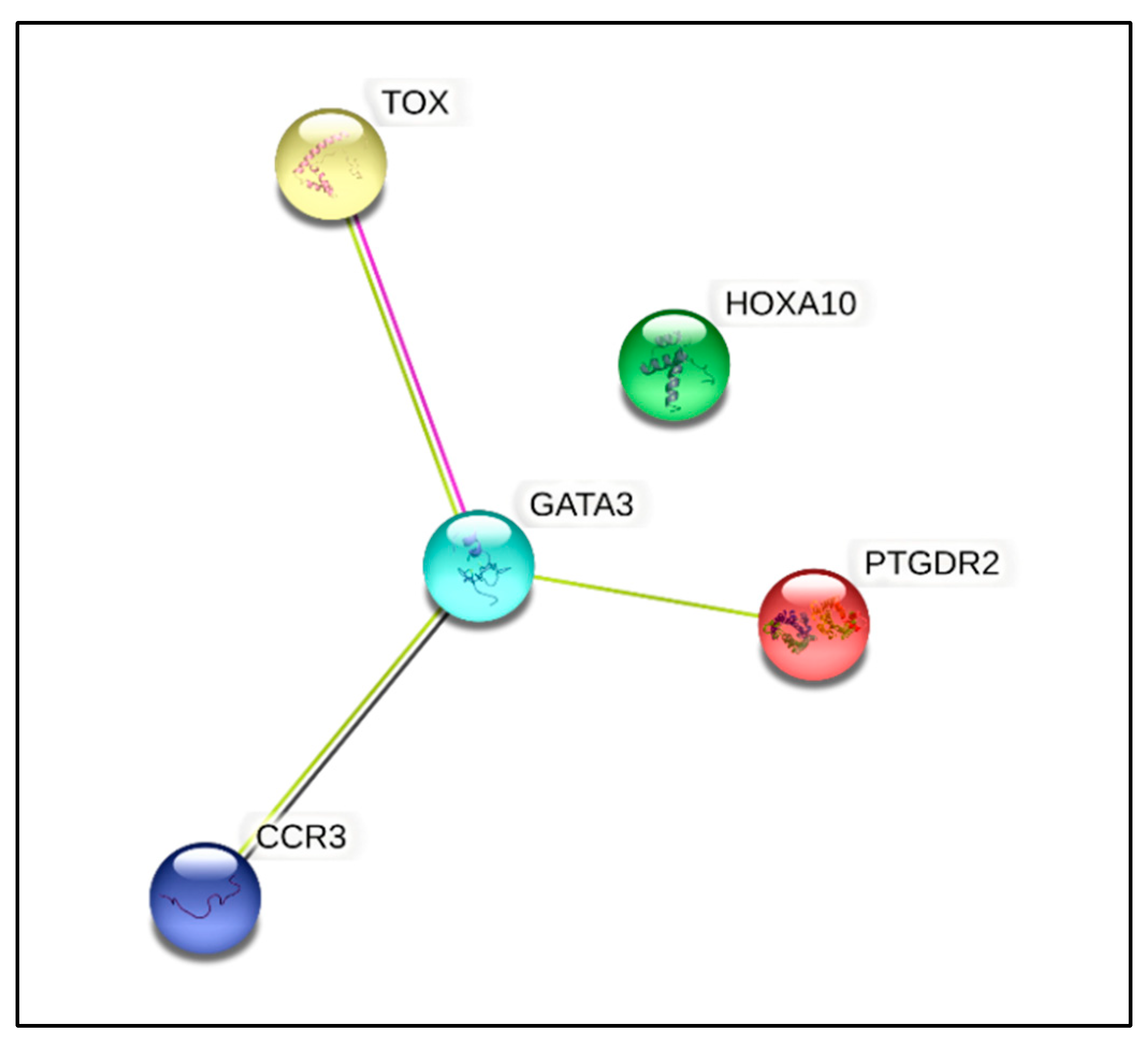
3.6. Gene Expression Studies
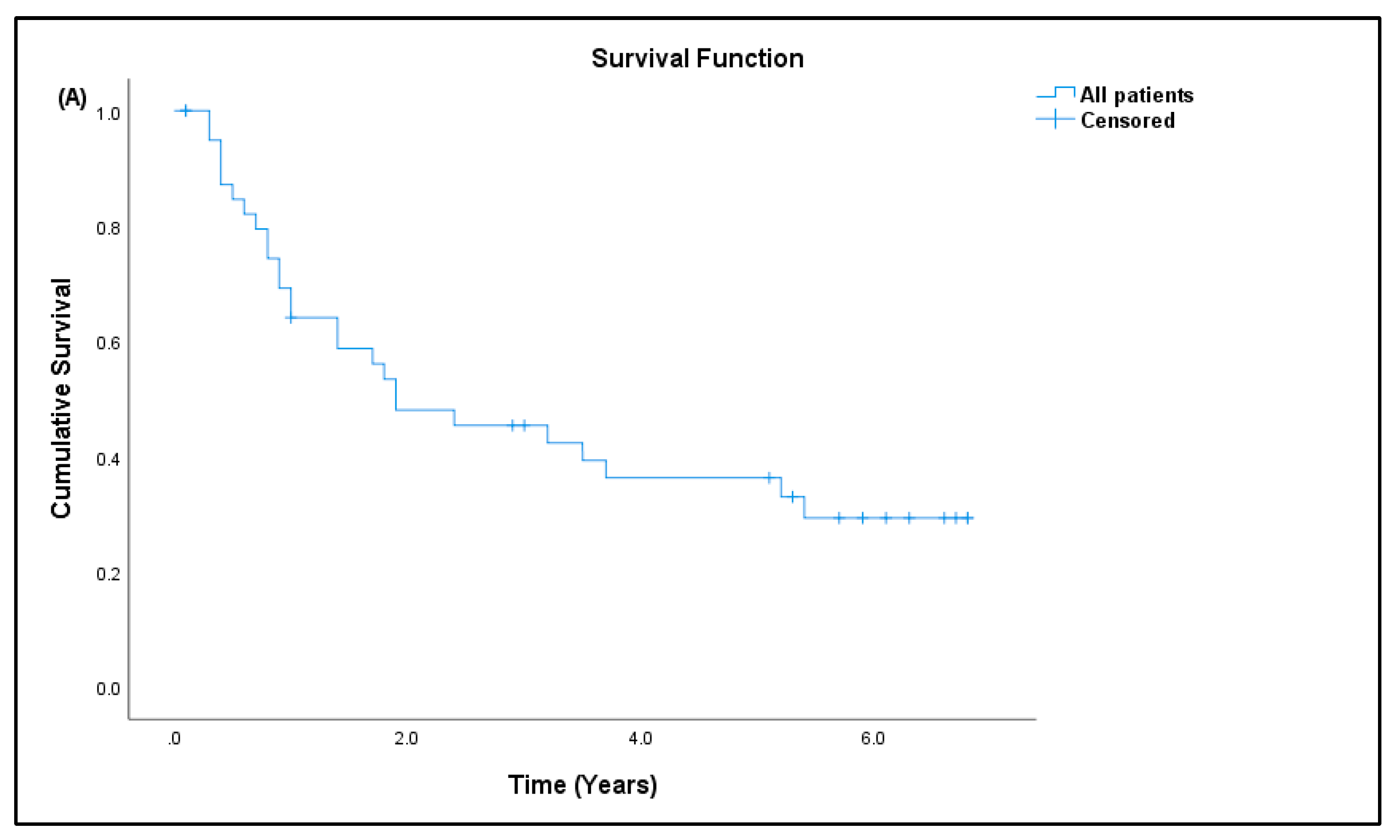
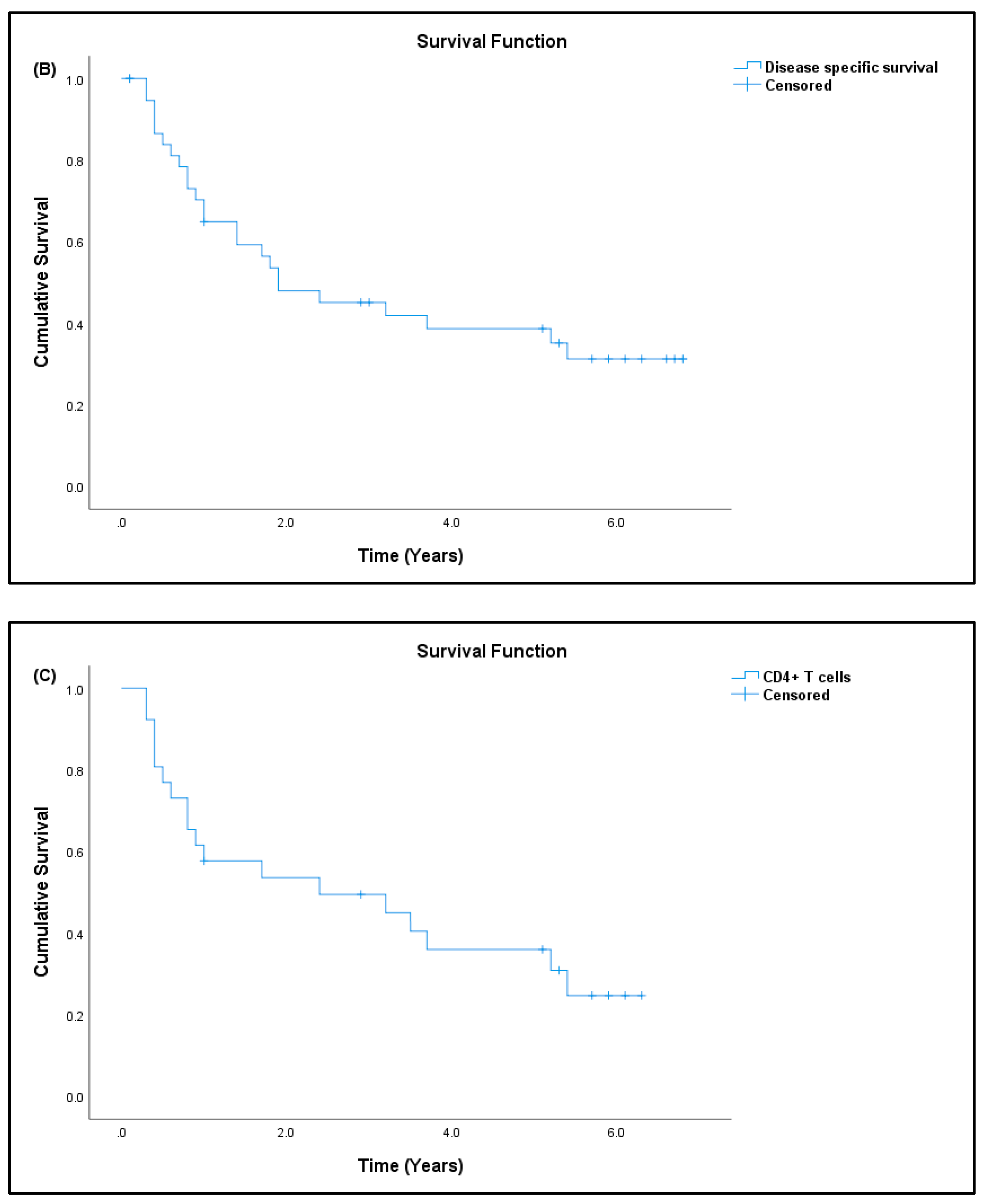
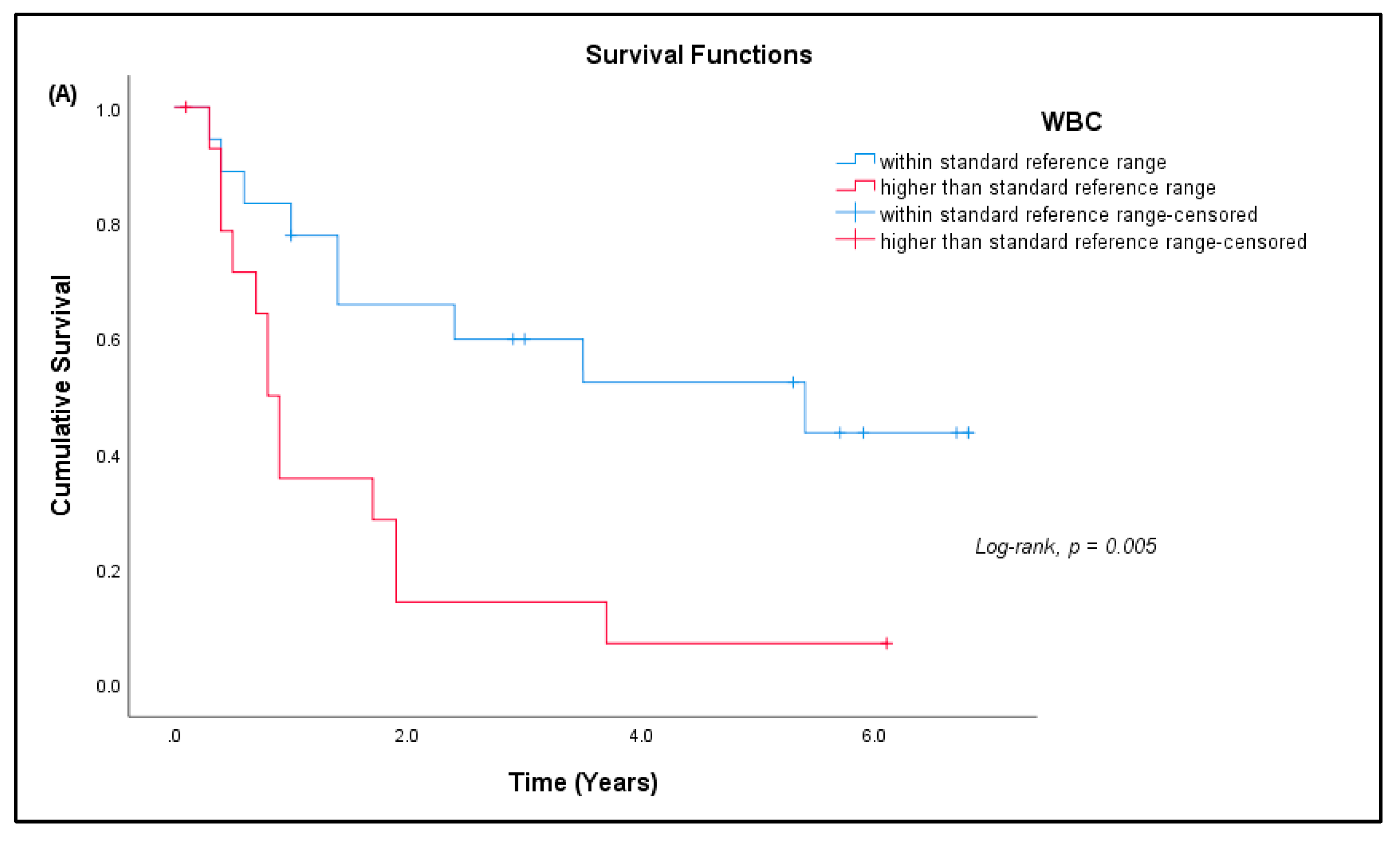
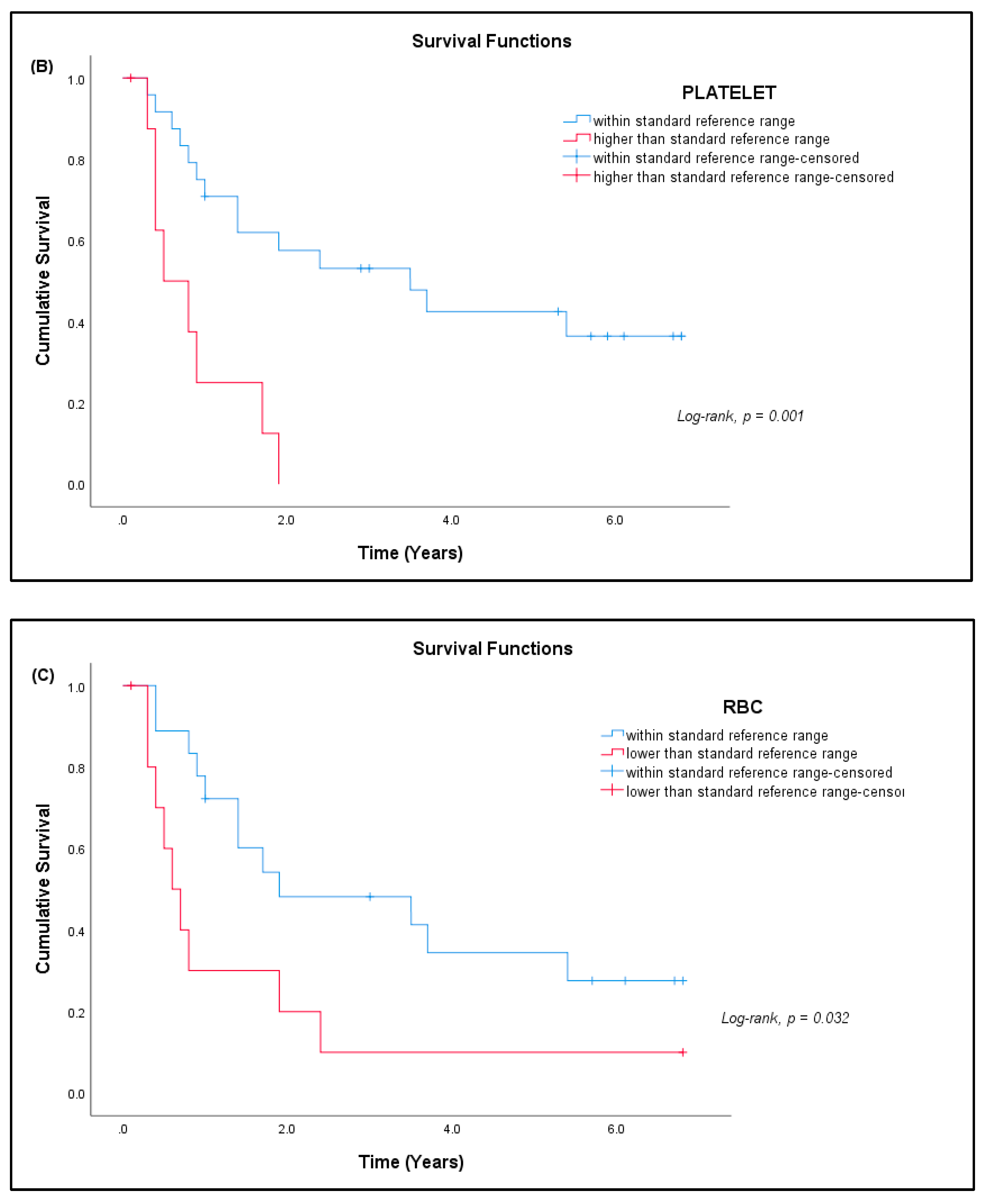
3.7. Patient Prognosis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nicholson, L.B. The Immune System. Essays Biochem. 2016, 60, 275–301. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, L.; Chong, M.M.W.; Littman, D.R. Plasticity of CD4+ T Cell Lineage Differentiation. Immunity 2009, 30, 646–655. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zaenker, P.; Gray, E.S.; Ziman, M.R. Autoantibody Production in Cancer-The Humoral Immune Response toward Autologous Antigens in Cancer Patients. Autoimmun. Rev. 2016, 15, 477–483. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vinay, D.S.; Ryan, E.P.; Pawelec, G.; Talib, W.H.; Stagg, J.; Elkord, E.; Lichtor, T.; Decker, W.K.; Whelan, R.L.; Kumara, H.M.C.S.; et al. Immune Evasion in Cancer: Mechanistic Basis and Therapeutic Strategies. Semin. Cancer Biol. 2015, 35, S185–S198. [Google Scholar] [CrossRef]
- Schreiber, R.D.; Old, L.J.; Smyth, M.J. Cancer Immunoediting: Integrating Immunity’s Roles in Cancer Suppression and Promotion. Science 2011, 331, 1565–1570. [Google Scholar] [CrossRef] [Green Version]
- den Haan, J.M.M.; Arens, R.; van Zelm, M.C. The Activation of the Adaptive Immune System: Cross-Talk between Antigen-Presenting Cells, T Cells and B Cells. Immunol. Lett. 2014, 162, 103–112. [Google Scholar] [CrossRef]
- Turvey, S.E.; Broide, D.H. Innate Immunity. J. Allergy Clin. Immunol. 2010, 125, S24–S32. [Google Scholar] [CrossRef]
- Durgeau, A.; Virk, Y.; Corgnac, S.; Mami-Chouaib, F. Recent Advances in Targeting CD8 T-Cell Immunity for More Effective Cancer Immunotherapy. Front. Immunol. 2018, 9, 14. [Google Scholar] [CrossRef]
- Niu, H.Q.; Zhao, X.C.; Li, W.; Xie, J.F.; Liu, X.Q.; Luo, J.; Zhao, W.P.; Li, X.F. Characteristics and Reference Ranges of CD4+T Cell Subpopulations among Healthy Adult Han Chinese in Shanxi Province, North China. BMC Immunol. 2020, 21, 1–10. [Google Scholar] [CrossRef]
- Filho, A.M.; Jammal, M.P.; CÔbo, E.; de CastroCôbo, E.; Silveira, T.P.; Adad, S.J.; Murta, E.F.C.; Nomelini, R.S. Correlation of Cytokines and Inducible Nitric Oxide Synthase Expression with Prognostic Factors in Ovarian Cancer. Immunol. Lett. 2014, 158, 195–199. [Google Scholar] [CrossRef]
- Geginat, J.; Paroni, M.; Facciotti, F.; Gruarin, P.; Kastirr, I.; Caprioli, F.; Pagani, M.; Abrignani, S. The CD4-Centered Universe of Human T Cell Subsets. Semin. Immunol. 2013, 25, 252–262. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, E.; Klein, M.; Bopp, T. Th9 Cells, New Players in Adaptive Immunity. Trends Immunol. 2014, 35, 61–68. [Google Scholar] [CrossRef] [PubMed]
- McGeachy, M.J.; Cua, D.J. Th17 Cell Differentiation: The Long and Winding Road. Immunity 2008, 28, 445–453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, D.; Rebekka, G.; David, J.; Antonio, L.; Sallusto, F. Production of Interleukin 22 but Not Interleukin 17 by a Subset of Human Skin-Homing Memory T Cells. Nat. Immunol. 2009, 10, 857. [Google Scholar] [CrossRef]
- Chtanova, T.; Tangye, S.G.; Newton, R.; Frank, N.; Hodge, M.R.; Rolph, M.S.; Mackay, C.R. T Follicular Helper Cells Express a Distinctive Transcriptional Profile, Reflecting Their Role as Non-Th1/Th2 Effector Cells That Provide Help for B Cells. J. Immunol. 2004, 173, 68–78. [Google Scholar] [CrossRef] [PubMed]
- Goto, S.; Sato, M.; Kaneko, R.; Itoh, M.; Sato, S.; Takeuchi, S. Analysis of Th1 and Th2 Cytokine Production by Peripheral Blood Mononuclear Cells as a Parameter of Immunological Dysfunction in Advanced Cancer Patients. Cancer Immunol. Immunother. 1999, 48, 435–442. [Google Scholar] [CrossRef]
- Gaur, P.; Singh, A.K.; Shukla, N.K.; Das, S.N. Inter-Relation of Th1, Th2, Th17 and Treg Cytokines in Oral Cancer Patients and Their Clinical Significance. Hum. Immunol. 2014, 75, 330–337. [Google Scholar] [CrossRef]
- Miggelbrink, A.M.; Jackson, J.D.; Lorrey, S.J.; Srinivasan, E.S.; Waibl-Polania, J.; Wilkinson, D.S.; Fecci, P.E. CD4 T-Cell Exhaustion: Does It Exist and What Are Its Roles in Cancer? Clin. Cancer Res. 2021, 27, 5742–5752. [Google Scholar] [CrossRef]
- Zamarron, B.F.; Chen, W. Dual Roles of Immune Cells and Their Factors in Cancer Development and Progression. Int. J. Biol. Sci. 2011, 7, 651. [Google Scholar] [CrossRef]
- Toomer, K.H.; Chen, Z. Autoimmunity as a Double Agent in Tumor Killing and Cancer Promotion. Front. Immunol. 2014, 5, 116. [Google Scholar] [CrossRef] [Green Version]
- Sakurai, T.; Yoshiga, D.; Ariyoshi, W.; Okinaga, T.; Kiyomiya, H.; Furuta, J.; Yoshioka, I.; Tominaga, K.; Nishihara, T. Essential Role of Mitogen-Activated Protein Kinases in IL-17A-Induced MMP-3 Expression in Human Synovial Sarcoma Cells. BMC Res. Notes 2016, 9, 68. [Google Scholar] [CrossRef] [Green Version]
- Iwakura, Y.; Nakae, S.; Saijo, S.; Ishigame, H. The Roles of IL-17A in Inflammatory Immune Responses and Host Defense against Pathogens. Immunol. Rev. 2008, 226, 57–79. [Google Scholar] [CrossRef]
- Reppert, S.; Boross, I.; Koslowski, M.; Türeci, Ö.; Koch, S.; Lehr, H.A.; Finotto, S. A Role for T-Bet-Mediated Tumour Immune Surveillance in Anti-IL-17A Treatment of Lung Cancer. Nat. Commun. 2011, 2, 1609. [Google Scholar] [CrossRef]
- Flowers, L.O. The Use of PCR Arrays to Investigate Human Immune Responses. Biomed. J. Sci. Tech. Res. 2019, 21, 15847–15851. [Google Scholar] [CrossRef]
- Nalairndran, G.; Chung, I.; Abdul Razack, A.H.; Chung, F.F.L.; Hii, L.W.; Lim, W.M.; Looi, C.K.; Mai, C.W.; Leong, C.O. Inhibition of Janus Kinase 1 Synergizes Docetaxel Sensitivity in Prostate Cancer Cells. J. Cell. Mol. Med. 2021, 25, 8187–8200. [Google Scholar] [CrossRef]
- Lai, Y.; Jeng, C.; Chen, S. The Roles of CD4+ T Cells in Tumor Immunity. ISRN Immunol. 2011, 2011, 1–6. [Google Scholar] [CrossRef] [Green Version]
- Zanetti, M. Tapping CD4 T Cells for Cancer Immunotherapy: The Choice of Personalized Genomics. J. Immunol. 2015, 194, 2049–2056. [Google Scholar] [CrossRef] [Green Version]
- Perez-diez, A.; Joncker, N.T.; Choi, K.; Chan, W.F.N.; Anderson, C.C.; Lantz, O.; Matzinger, P. CD4 Cells Can Be More Efficient at Tumor Rejection than CD8 Cells. Blood J. Am. Soc. Hematol. 2007, 109, 5346–5355. [Google Scholar] [CrossRef]
- Alshaker, H.A.; Matalka, K.Z. IFN- γ, IL-17 and TGF- β Involvement in Shaping the Tumor Microenvironment: The Significance of Modulating Such Cytokines in Treating Malignant Solid Tumors. Cancer Cell Int. 2011, 11, 33. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.; Paul, W.E. CD4 T Cells: Fates, Functions, and Faults. Immunobiology 2008, 112, 1557–1569. [Google Scholar] [CrossRef] [Green Version]
- Lau, K.-M.; Cheng, S.H.; Lo, K.W.; Lee, S.A.K.W.; Woo, J.K.S.; van Hasselt, C.A.; Lee, S.P.; Rickinson, A.B.; Ng, M.H.L. Increase in Circulating Foxp3+CD4+CD25(High) Regulatory T Cells in Nasopharyngeal Carcinoma Patients. Br. J. Cancer 2007, 96, 617–622. [Google Scholar] [CrossRef] [Green Version]
- Borst, J.; Ahrends, T.; Bąbała, N.; Melief, C.J.M.; Kastenmüller, W. CD4+ T Cell Help in Cancer Immunology and Immunotherapy. Nat. Rev. Immunol. 2018, 18, 635–647. [Google Scholar] [CrossRef]
- Tay, R.E.; Richardson, E.K.; Toh, H.C. Revisiting the Role of CD4+ T Cells in Cancer Immunotherapy—New Insights into Old Paradigms. Cancer Gene Ther. 2021, 28, 5–17. [Google Scholar] [CrossRef]
- Zander, R.; Schauder, D.; Xin, G.; Nguyen, C.; Wu, X.; Zajac, A.; Cui, W. CD4+ T Cell Help Is Required for the Formation of a Cytolytic CD8+ T Cell Subset That Protects against Chronic Infection and Cancer. Immunity 2019, 51, 1028–1042. [Google Scholar] [CrossRef] [PubMed]
- de Mello, R.A.; Veloso, A.F.; Esrom Catarina, P.; Nadine, S.; Antoniou, G. Potential Role of Immunotherapy in Advanced Non-Small-Cell Lung Cancer. Onco. Targets. Ther. 2016, 10, 21–30. [Google Scholar] [CrossRef] [Green Version]
- Brivio, F.; Fumagalli, L.; Parolini, D.; Messina, G.; Rovelli, F.; Rescaldani, R.; Vigore, L.; Vezzo, R.; Vaghi, M.; Di Bella, S.; et al. T-Helper/T-Regulator Lymphocyte Ratio as a New Immunobiological Index to Quantify the Anticancer Immune Status in Cancer Patients. Vivo 2008, 22, 647–650. [Google Scholar] [PubMed]
- Aune, T.M.; Collins, P.L.; Collier, S.P.; Henderson, M.A.; Chang, S. Epigenetic Activation and Silencing of the Gene That Encodes IFN-γ. Front. Immunol. 2013, 4, 112. [Google Scholar] [CrossRef] [Green Version]
- Ikeda, H.; Old, L.J.; Schreiber, R.D. The Roles of IFNgamma in Protection against Tumor Development and Cancer Immunoediting. Cytokine Growth Factor Rev. 2002, 13, 95–109. [Google Scholar] [CrossRef]
- Ni, L.; Lu, J. Interferon Gamma in Cancer Immunotherapy. Cancer Med. 2018, 7, 4509–4516. [Google Scholar] [CrossRef]
- Poncette, L.; Bluhm, J.; Blankenstein, T. The Role of CD4 T Cells in Rejection of Solid Tumors. Curr. Opin. Immunol. 2022, 74, 18–24. [Google Scholar] [CrossRef]
- Vilček, J.; Feldmann, M. Cytokines as Therapeutics and Targets of Therapeutics. Rheumatologia 2006, 20, 65–74. [Google Scholar] [CrossRef]
- Sallusto, F.; Mackay, C.R.; Lanzavecchia, A. Selective Expression of the Eotaxin Receptor CCR3 by Human T Helper 2 Cells. Science 1997, 277, 2005–2007. [Google Scholar] [CrossRef]
- Tripp, C.S.; Wolf, S.F.; Unanue, E.R. Interleukin 12 and Tumor Necrosis Factor α Are Costimulators of Interferon γ Production by Natural Killer Cells in Severe Combined Immunodeficiency Mice with Listeriosis, and Interleukin 10 Is a Physiologic Antagonist. Proc. Natl. Acad. Sci. USA 1993, 90, 3725–3729. [Google Scholar] [CrossRef] [Green Version]
- Josephs, S.F.; Ichim, T.E.; Prince, S.M.; Kesari, S.; Marincola, F.M.; Escobedo, A.R.; Jafri, A. Unleashing Endogenous TNF-Alpha as a Cancer Immunotherapeutic. J. Transl. Med. 2018, 14, 242. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Chen, Y.; Liu, H.; Que, Y.; Zhang, X.; Zheng, F. Integrated Expression Profiles Analysis Reveals Correlations between the IL-33/ST2 Axis and CD8+ T Cells, Regulatory T Cells, and Myeloid-Derived Suppressor Cells in Soft Tissue Sarcoma. Front. Immunol. 2018, 9, 1179. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Chen, B.; Huang, Z.; Liu, C.; Ye, L.; Wang, C.; Tong, Y.; Yang, J.; Zhao, C. Comprehensive Profiling of Immune-Related Genes in Soft Tissue Sarcoma Patients. J. Transl. Med. 2020, 18, 337. [Google Scholar] [CrossRef] [PubMed]
- Bae, J.Y.; Choi, K.U.; Kim, A.; Lee, S.J.; Kim, K.; Kim, J.Y.; Lee, I.S.; Chung, S.H.; Kim, J.I. Evaluation of Immune-biomarker Expression in High-grade Soft-tissue Sarcoma: HLA-DQA1 Expression as a Prognostic Marker. Exp. Ther. Med. 2020, 20, 107. [Google Scholar] [CrossRef] [PubMed]
- Haraguchi, T.; Miyoshi, H.; Hiraoka, K.; Yokoyama, S.; Ishibashi, Y.; Hashiguchi, T.; Matsuda, K.; Hamada, T.; Okawa, T.; Shiba, N.; et al. GATA3 Expression Is a Poor Prognostic Factor in Soft Tissue Sarcomas. PLoS ONE 2016, 11, e0156524. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yagi, R.; Zhu, J.; Paul, W.E. An Updated View on Transcription Factor GATA3-Mediated Regulation of Th1 and Th2 Cell Differentiation. Int. Immunol. 2011, 23, 415–420. [Google Scholar] [CrossRef] [Green Version]
- Cosmi, L.; Annunziato, F.; Maggi, E.; Romagnani, S.; Manetti, R. Chemoattractant Receptors Expressed on Type 2 T Cells and Their Role in Disease. Int. Arch. Allergy Immunol. 2001, 125, 273–279. [Google Scholar] [CrossRef]
- Misawa, K.; Imai, A.; Kanazawa, T.; Mima, M.; Yamada, S.; Mochizuki, D.; Yamada, T.; Shinmura, D.; Ishikawa, R.; Kita, J.; et al. G Protein-Coupled Receptor Genes, PTGDR1, PTGDR2, and PTGIR, Are Candidate Epigenetic Biomarkers and Predictors for Treated Patients with HPV-Associated Oropharyngeal Cancer. Microorganisms 2020, 8, 1504. [Google Scholar] [CrossRef] [PubMed]
- Taghon, T.; Stolz, F.; De Smedt, M.; Cnockaert, M.; Verhasselt, B.; Plum, J.; Leclercq, G. HOX-A10 Regulates Hematopoietic Lineage Commitment: Evidence for a Monocyte-Specific Transcription Factor. Blood 2002, 99, 1197–1204. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lawrence, H.J.; Sauvageau, G.; Ahmadi, N.; Lopez, A.R.; LeBeau, M.M.; Link, M.; Humphries, K.; Largman, C. Stage-and Lineage-Specific Expression of the HOXA10 Homeobox Gene in Normal and Leukemic Hematopoietic Cells. Exp. Hematol. 1995, 23, 1160–1166. [Google Scholar] [PubMed]
- Eklund, E.A.; Jalava, A.; Kakar, R. Tyrosine Phosphorylation of HoxA10 Decreases DNA Binding and Transcriptional Repression during Interferon γ-Induced Differentiation of Myeloid Leukemia Cell Lines. J. Biol. Chem. 2000, 275, 20117–20126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arora, M.; Kumari, S.; Singh, J.; Chopra, A.; Chauhan, S.S. Expression Pattern, Regulation, and Clinical Significance of TOX in Breast Cancer. Cancer Immunol. Immunother. 2021, 70, 349–363. [Google Scholar] [CrossRef]
- Guo, L.; Li, X.; Liu, R.; Chen, Y.; Ren, C.; Du, S. TOX Correlates with Prognosis, Immune Infiltration, and T Cells Exhaustion in Lung Adenocarcinoma. Cancer Med. 2020, 9, 6694–6709. [Google Scholar] [CrossRef]
- Chung, W.; Kwabi-Addo, B.; Ittmann, M.; Jelinek, J.; Shen, L.; Yu, Y.; Issa, J.P.J. Identification of Novel Tumor Markers in Prostate, Colon and Breast Cancer by Unbiased Methylation Profiling. PLoS ONE 2008, 3, e2079. [Google Scholar] [CrossRef]
- Wang, X.; He, Q.; Shen, H.; Xia, A.; Tian, W.; Yu, W.; Sun, B. TOX Promotes the Exhaustion of Antitumor CD8+ T Cells by Preventing PD1 Degradation in Hepatocellular Carcinoma. J. Hepatol. 2019, 71, 731–741. [Google Scholar] [CrossRef]
- Yun, S.; Lee, S.H.; Yoon, S.R.; Kim, M.S.; Piao, Z.H.; Myung, P.K.; Kim, T.D.; Jung, H.; Choi, I. TOX Regulates the Differentiation of Human Natural Killer Cells from Hematopoietic Stem Cells in Vitro. Immunol. Lett. 2011, 136, 29–36. [Google Scholar] [CrossRef]
- Catton, C.N.; O’Sullivan, B.; Kotwall, C.; Cummings, B.; Hao, Y.; Fornasier, V. Outcome and Prognosis in Retroperitoneal Soft Tissue Sarcoma. Int. J. Radiat. Oncol. Biol. Phys. 1994, 29, 1005–1010. [Google Scholar] [CrossRef]
- Ruka, W.; Rutkowski, P.; Kaminska, J.; Rysinska, A.; Steffen, J. Alterations of Routine Blood Tests in Adult Patients with Soft Tissue Sarcomas: Relationships to Cytokine Serum Levels and Prognostic Significance. Ann. Oncol. 2001, 12, 1423–1432. [Google Scholar] [CrossRef] [PubMed]
- Mabuchi, S.; Matsumoto, Y.; Isohashi, F.; Yoshioka, Y.; Ohashi, H.; Morii, E.; Hamasaki, T.; Aozasa, K.; Mutch, D.G.; Kimura, T. Pretreatment Leukocytosis Is an Indicator of Poor Prognosis in Patients with Cervical Cancer. Gynecol. Oncol. 2011, 122, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Yao, M.; Chen, X.; Lu, W.; Lv, Q.; Wang, K.; Zhang, L.; Lu, F. Reduced Red Blood Cell Count Predicts Poor Survival after Surgery in Patients with Primary Liver Cancer. Medicine 2015, 94, e577. [Google Scholar] [CrossRef]
- Hwang, S.G.; Kim, K.M.; Cheong, J.H.; Kim, H.I.; An, J.Y.; Hyung, W.J.; Noh, S.H. Impact of Pretreatment Thrombocytosis on Blood-Borne Metastasis and Prognosis of Gastric Cancer. Eur. J. Surg. Oncol. 2012, 38, 562–567. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Kobayashi, E.; Suehara, Y.; Ito, A.; Kubota, D.; Tanzawa, Y.; Endo, M.; Nakatani, F.; Nakatsura, T.; Kawai, A.; et al. Immunological Status of Peripheral Blood Is Associated with Prognosis in Patients with Bone and Soft-Tissue Sarcoma. Oncol. Lett. 2021, 21, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Murata, T.; Lin, M.I.; Aritake, K.; Matsumoto, S.; Narumiya, S.; Ozaki, H.; Urade, Y.; Hori, M.; Sessa, W.C. Role of Prostaglandin D2 Receptor DP as a Suppressor of Tumor Hyperpermeability and Angiogenesis in Vivo. Proc. Natl. Acad. Sci. USA 2008, 105, 20009–20014. [Google Scholar] [CrossRef] [Green Version]
- Baird, K.; Davis, S.; Antonescu, C.R.; Harper, U.L.; Walker, R.L.; Chen, Y.; Glatfelter, A.A.; Duray, P.H.; Meltzer, P.S. Gene Expression Profiling of Human Sarcomas: Insights into Sarcoma Biology. Cancer Res. 2005, 65, 9226–9235. [Google Scholar] [CrossRef] [Green Version]
| Sarcoma | Normal Controls | |
|---|---|---|
| Flow cytometer analysis | (n = 26) | (n = 10) |
| Age, median (IQR; range) | 34.50 (32.75; 13–68) | 33.00 (4.5; 23–40) |
| Female | 13 (50%) | 4 (40%) |
| Male | 13 (50%) | 6 (60%) |
| Cytokine analysis | (n = 23) | (n = 9) |
| Age, mean (SD; range) | 35.74 (17.89; 15–68) | 32.22 (4.84; 23–40) |
| Female | 8 (34.78%) | 3 (33.33%) |
| Male | 15 (65.22%) | 6 (66.67%) |
| qPCR array | (n = 5) | (n = 5) |
| Age, mean (SD; range) | 38.80 (18.60; 16–62) | 39.60 (12.30; 21–55) |
| Male | 5 (100%) | 5 (100%) |
| Female | - | - |
| Clinical Features | Bone: | 19 (100%) | Soft Tissues: | 22 (100%) |
|---|---|---|---|---|
| Type: | Osteosarcoma | 8 (42.11%) | Pleomorphic sarcoma | 7 (31.82%) |
| Chondrosarcoma | 4 (21.05%) | Liposarcoma | 5 (22.73%) | |
| Ewing’s sarcoma | 3 (15.79%) | Synovial sarcoma | 2 (9.09%) | |
| Pleomorphic sarcoma | 2 (10.53%) | Leiomyosarcoma | 2 (9.09%) | |
| Myofibroblastic sarcoma | 1 (5.26%) | Rhabdomyosarcoma | 2 (9.09%) | |
| Malignant GCT | 1 (5.26%) | Desmoid tumor | 1 (4.55%) | |
| Alveolar soft part sarcoma | 1 (4.55%) | |||
| Ewing’s sarcoma | 1 (4.55%) | |||
| Chondrosarcoma | 1 (4.55%) | |||
| Site: | Femur | 6 (31.58%) | Thigh | 12 (54.55%) |
| Pelvis | 5 (26.32%) | Chest wall | 4 (18.18%) | |
| Knee | 3 (15.79%) | Pelvis | 2 (9.09%) | |
| Fibula | 2 (10.53%) | Leg | 2 (9.09%) | |
| Scapular | 1 (5.26%) | Back | 1 (4.55%) | |
| Humerus | 1 (5.26%) | Arm | 1 (4.55%) | |
| Tibia | 1 (5.26%) |
| Cytokines | Normal Controls (n = 9) Mean ± SE (pg/mL) | Sarcoma Patients (n = 23) Mean ± SE (pg/mL) | p-Value |
|---|---|---|---|
| IL-17A | 1320.87 ± 317.07 | 1066.58 ± 149.74 | NS |
| TNF-α | 843.74 ± 85.95 | 562.87 ± 190.69 | 0.004 * |
| TGF-β1 | 4338.48 ± 720.37 | 6042.73 ± 727.75 | NS |
| IFN-γ | 14,041.66 ± 2379.13 | 7218.27 ± 1216.02 | 0.010 * |
| TGF-β1 (plasma) | 22,413.09 ± 3152.50 | 15,725.30 ± 2285.05 | NS |
| Parameters | Cytokines (pg/mL) (n = 23) | T Cells (%) (n = 26) | ||||||
|---|---|---|---|---|---|---|---|---|
| IL-17A | TNF-α | IFN-γ | TGF-β1 | TGF-β1 (Plasma) | CD4+ T-Cells | T-Reg Cells | ||
| IL-17A | r | 0.039 | 0.264 | 0.580 ** | 0.39 | 0.055 | 0.209 | |
| P | NS | NS | 0.004 | NS | NS | NS | ||
| TNF-α | r | 0.039 | −0.111 | 0.425 * | 0.225 | −0.272 | 0.207 | |
| P | NS | NS | 0.043 | NS | NS | NS | ||
| IFN-γ | r | 0.264 | −0.111 | −0.004 | −0.127 | −0.486 * | −0.079 | |
| P | NS | NS | NS | NS | 0.019 | NS | ||
| TGF-β1 | r | 0.580 ** | 0.425 * | −0.004 | −0.082 | −0.014 | 0.407 | |
| P | 0.004 | 0.043 | NS | NS | NS | NS | ||
| TGF-β1 (plasma) | r | 0.039 | 0.225 | −0.127 | −0.082 | −0.016 | 0.162 | |
| P | NS | NS | NS | NS | NS | NS | ||
| Complete Blood Count (N = 33) | ||||
|---|---|---|---|---|
| Gender: | Male: 18 | Female: 15 | ||
| Age: mean ± SD (range): | 37.79 ± 18.09 (15–68 years) | |||
| Parameters | Reference range | Mean value ± SD | Status, n (%) | |
| WBC (× 109/L) | A | 4.0–10.0 | 9.82 ± 3.46 | 15 (45.45%) * |
| Platelets (× 109/L) | A | 150–400 | 332.64 ± 120.82 | 9 (27.27%) * |
| Hemoglobin (g/L) | FA | 120.0–150.0 | 113.07 ± 18.84 | 1 (3.03%) * |
| MA | 130.0–170.0 | 135.33 ± 20.16 | 17 (51.51%) ** | |
| Hematocrit (L/L) | FA | 0.36–0.46 | 0.34 ± 0.05 | 16 (48.48%) ** |
| MA | 0.40–0.50 | 0.40 ± 0.06 | ||
| RBC (× 1012/L) | FA | 3.80–4.80 | 4.19 ± 0.51 | 4 (12.12%) * |
| MA | 4.50–5.50 | 4.76 ± 0.63 | 11 (33.33%) ** | |
| MCV (fl) | A | 77–97 | 83.00 ± 4.43 | 3 (9.09%) ** |
| MCH (pg) | A | 27.0–32.0 | 27.74 ± 2.29 | 10 (30.30%) ** |
| Genes | Descriptions | Fold Change | Fold Regulation | p-Value * |
|---|---|---|---|---|
| HOXA10 | Homeobox A10 | 2.0905 | Up | 0.019 |
| CCR3 | C-C Chemokine receptor type 3 | −3.1256 | Down | 0.045 |
| GATA3 | GATA binding protein 3 | −3.4729 | Down | 0.021 |
| PTGDR2 | Prostaglandin D2 receptor 2 | −4.7838 | Down | 0.015 |
| TOX | Thymocyte selection-associated high-mobility-group box | −2.3072 | Down | 0.040 |
| Patients (N) | Percentage (%) | |
|---|---|---|
| # Survival outcome, n = 41 (100%) | ||
| Dead | 26 | 63.41 |
| Alive | 15 | 36.59 |
| # Cause of death, n = 26 (100%) | ||
| Advanced disease | 24 | 92.31 |
| Others | 2 | 7.69 |
| Survival duration, n = 24 (100%) | ||
| <6 months | 6 | 25.00 |
| ≤1 year | 7 | 29.17 |
| <2 years | 6 | 25.00 |
| <5 years | 3 | 12.50 |
| >5 years | 2 | 8.33 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Munisamy, S.; Radhakrishnan, A.K.; Ramdas, P.; Samuel, P.J.; Singh, V.A. Immune Biomarkers in Blood from Sarcoma Patients: A Pilot Study. Curr. Oncol. 2022, 29, 5585-5603. https://doi.org/10.3390/curroncol29080441
Munisamy S, Radhakrishnan AK, Ramdas P, Samuel PJ, Singh VA. Immune Biomarkers in Blood from Sarcoma Patients: A Pilot Study. Current Oncology. 2022; 29(8):5585-5603. https://doi.org/10.3390/curroncol29080441
Chicago/Turabian StyleMunisamy, Sarmini, Ammu Kutty Radhakrishnan, Premdass Ramdas, Priscilla Josephine Samuel, and Vivek Ajit Singh. 2022. "Immune Biomarkers in Blood from Sarcoma Patients: A Pilot Study" Current Oncology 29, no. 8: 5585-5603. https://doi.org/10.3390/curroncol29080441
APA StyleMunisamy, S., Radhakrishnan, A. K., Ramdas, P., Samuel, P. J., & Singh, V. A. (2022). Immune Biomarkers in Blood from Sarcoma Patients: A Pilot Study. Current Oncology, 29(8), 5585-5603. https://doi.org/10.3390/curroncol29080441





