Synaptotagmin 13 Is Highly Expressed in Estrogen Receptor-Positive Breast Cancer
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Quantitative Real-Time Reverse Transcription-Polymerase Chain Reaction (qRT-PCR)
2.3. Western Blotting Analysis
2.4. PCR Array Analysis
2.5. Public Datasets of BC Cell Lines and Patients
2.6. Statistical Analysis
3. Results
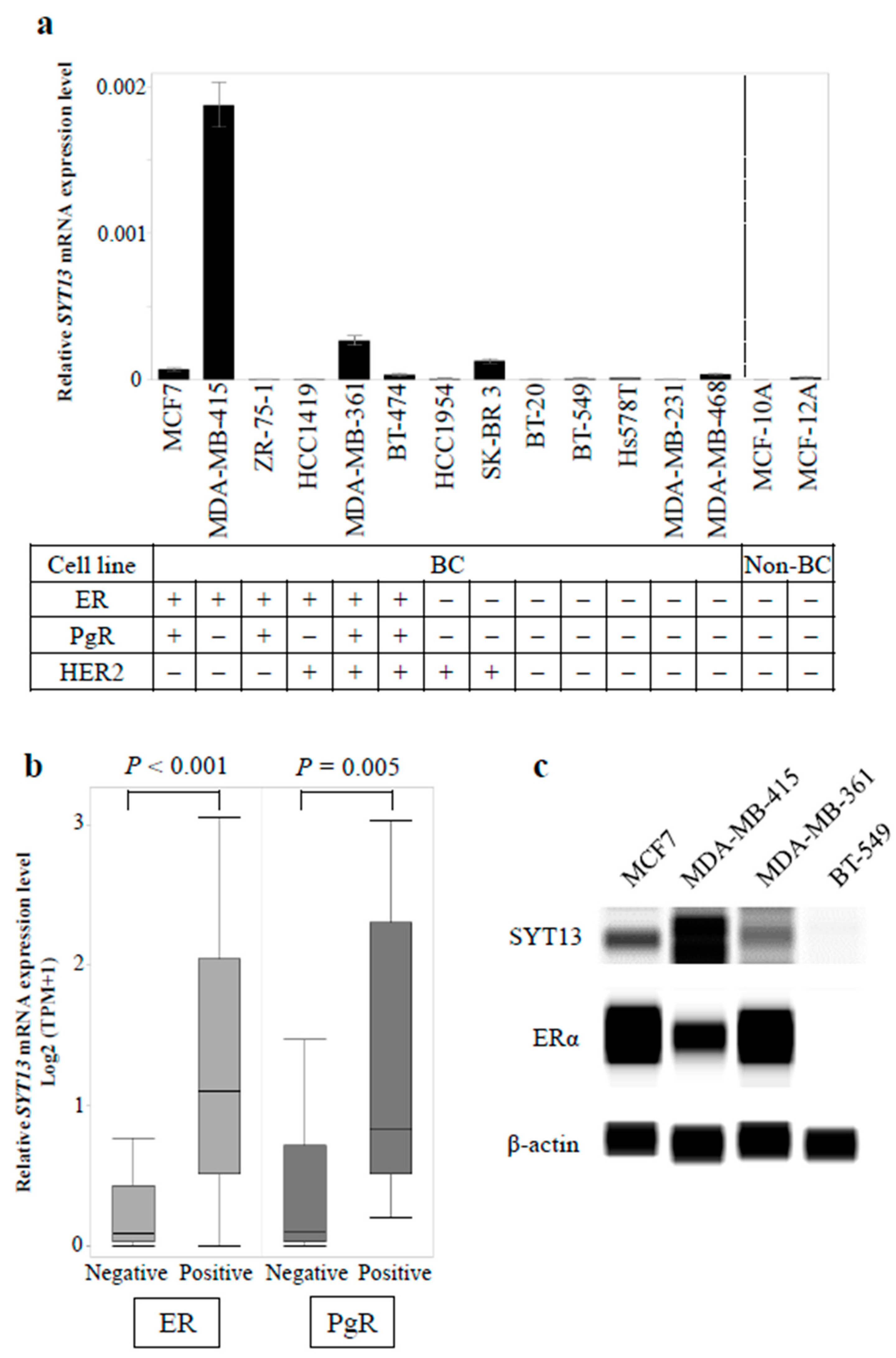
3.1. SYT13 mRNA Expression Levels in Cell Lines
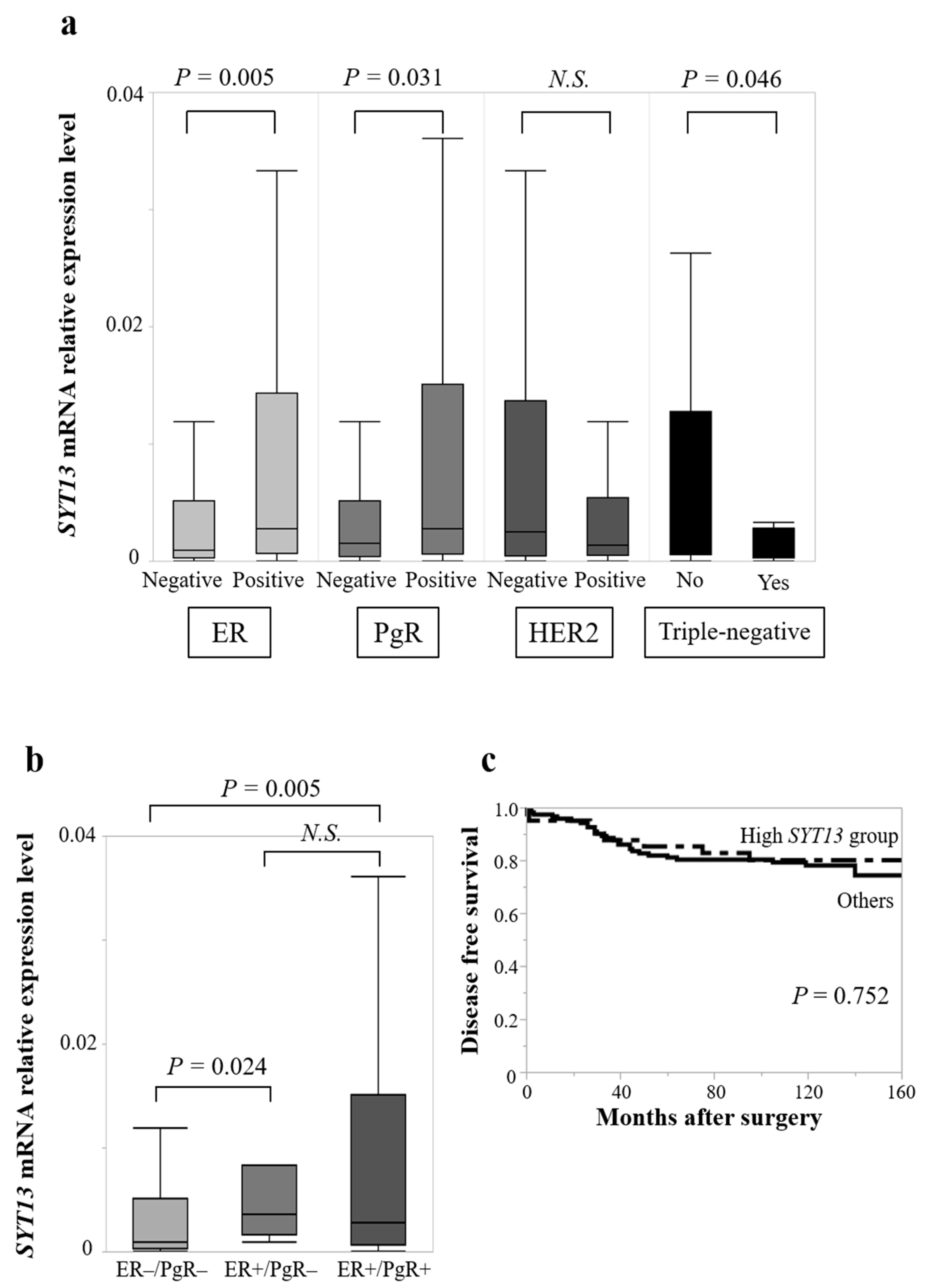
3.2. Association of SYT13 mRNA Expression Levels with Clinicopathological Factors in BC Patients
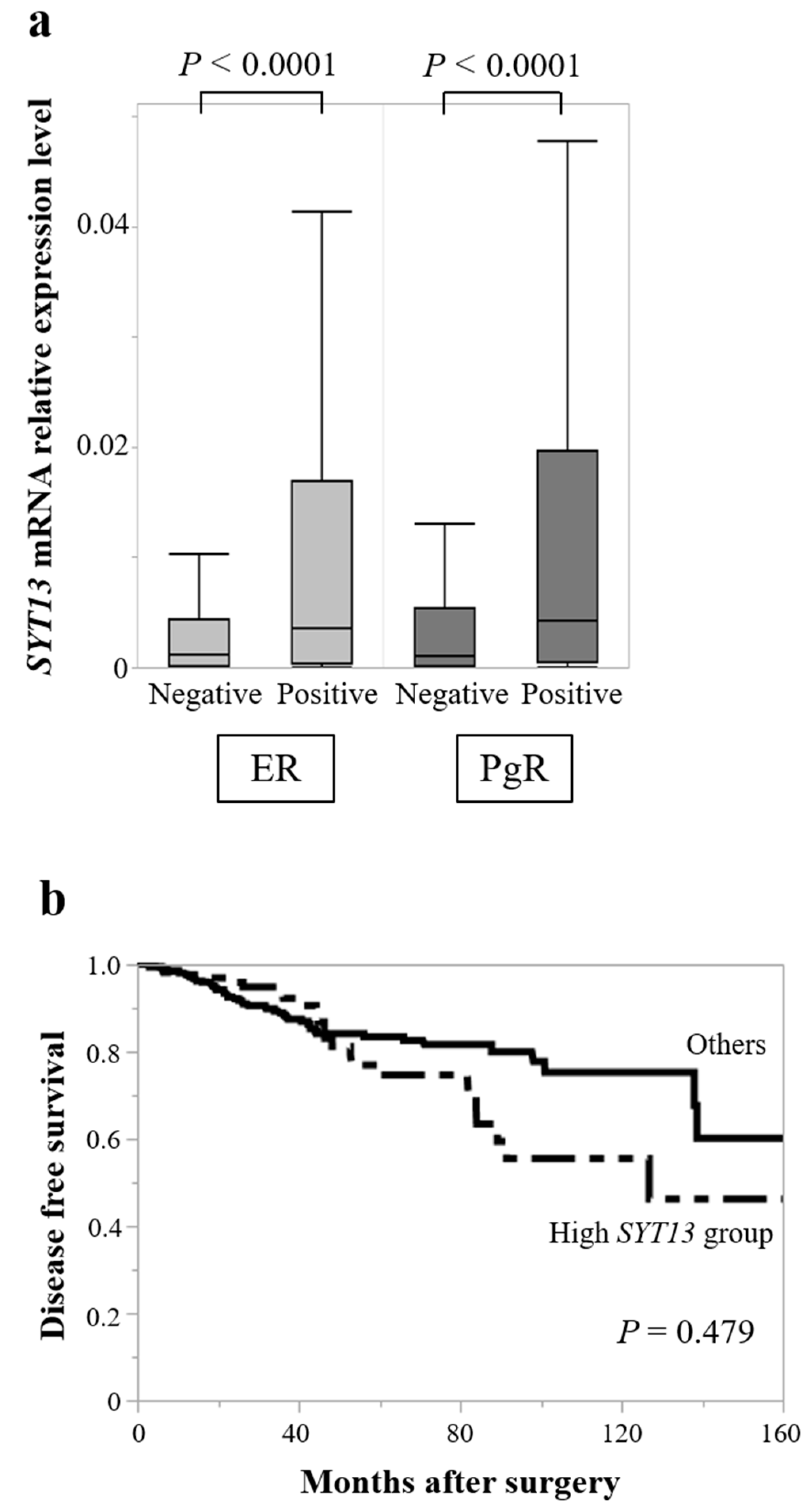
3.3. TCGA Database Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics. CA Cancer J. Clin. 2019, 69, 7–34. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iwata, H. Future treatment strategies for metastatic breast cancer: Curable or incurable? Breast Cancer 2011, 19, 200–205. [Google Scholar] [CrossRef]
- Shizuku, M.; Shibata, M.; Shimizu, Y.; Takeuchi, D.; Mizuno, Y. Clinical outcomes of neoadjuvant chemotherapy for patients with breast cancer: Tri-weekly nanoparticle albumin-bound paclitaxel followed by 5-fluorouracil, epirubicin, and cyclophosphamide: A retrospective observational study. Nagoya J. Med. Sci. 2020, 82, 457–467. [Google Scholar] [PubMed]
- Silvestri, M.; Cristaudo, A.; Morrone, A.; Messina, C.; Bennardo, L.; Nisticò, S.P.; Mariano, M.; Cameli, N. Emerging skin toxicities in patients with breast cancer treated with new cyclin-dependent kinase 4/6 inhibitors: A systematic review. Drug Saf. 2021, 44, 725–732. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.X.; Reinert, T.; Chmielewska, I.; Ellis, M.J. Mechanisms of aromatase inhibitor resistance. Nat. Rev. Cancer 2015, 15, 261–275. [Google Scholar] [CrossRef]
- Finn, R.S.; Martin, M.; Rugo, H.S.; Jones, S.; Im, S.-A.; Gelmon, K.; Harbeck, N.; Lipatov, O.N.; Walshe, J.M.; Moulder, S.; et al. Palbociclib and Letrozole in Advanced Breast Cancer. N. Engl. J. Med. 2016, 375, 1925–1936. [Google Scholar] [CrossRef] [PubMed]
- Goetz, M.P.; Toi, M.; Campone, M.; Sohn, J.; Paluch-Shimon, S.; Huober, J.; Park, I.H.; Trédan, O.; Chen, S.-C.; Manso, L.; et al. MONARCH 3: Abemaciclib as initial therapy for advanced breast cancer. J. Clin. Oncol. 2017, 35, 3638–3646. [Google Scholar] [CrossRef]
- Jahn, J.E.; Best, D.H.; Coleman, W.B. Exogenous expression of synaptotagmin XIII suppresses the neoplastic phenotype of a rat liver tumor cell line through molecular pathways related to mesenchymal to epithelial transition. Exp. Mol. Pathol. 2010, 89, 209–216. [Google Scholar] [CrossRef]
- Kanda, M.; Shimizu, D.; Tanaka, H.; Tanaka, C.; Kobayashi, D.; Hayashi, M.; Takami, H.; Niwa, Y.; Yamada, S.; Fujii, T.; et al. Synaptotagmin XIII expression and peritoneal metastasis in gastric cancer. J. Br. Surg. 2018, 105, 1349–1358. [Google Scholar] [CrossRef]
- Li, Q.; Zhang, S.; Hu, M.; Xu, M.; Jiang, X. Silencing of synaptotagmin 13 inhibits tumor growth through suppressing proliferation and promoting apoptosis of colorectal cancer cells. Int. J. Mol. Med. 2019, 45, 234–244. [Google Scholar] [CrossRef]
- Zhang, L.; Fan, B.; Zheng, Y.; Lou, Y.; Cui, Y.; Wang, K.; Zhang, T.; Tan, X. Identification SYT13 as a novel biomarker in lung adenocarcinoma. J. Cell. Biochem. 2020, 121, 963–973. [Google Scholar] [CrossRef]
- Watanabe, M.; Shibata, M.; Inaishi, T.; Ichikawa, T.; Soeda, I.; Miyajima, N.; Takano, Y.; Takeuchi, D.; Tsunoda, N.; Kanda, M.; et al. MZB1 expression indicates poor prognosis in estrogen receptor-positive breast cancer. Oncol. Lett. 2020, 20, 1. [Google Scholar] [CrossRef]
- Shibata, M.; Kanda, M.; Shimizu, D.; Tanaka, H.; Umeda, S.; Hayashi, M.; Inaishi, T.; Miyajima, N.; Adachi, Y.; Takano, Y.; et al. Expression of regulatory factor X1 can predict the prognosis of breast cancer. Oncol. Lett. 2017, 13, 4334–4340. [Google Scholar] [CrossRef] [Green Version]
- Kanda, M.; Kasahara, Y.; Shimizu, D.; Miwa, T.; Umeda, S.; Sawaki, K.; Nakamura, S.; Kodera, Y.; Obika, S. Amido-Bridged Nucleic Acid-Modified Antisense Oligonucleotides Targeting SYT13 to Treat Peritoneal Metastasis of Gastric Cancer. Mol. Ther. Nucleic Acids 2020, 22, 791–802. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.S.; Dering, J.; Conklin, D.; Kalous, O.; Cohen, D.J.; Desai, A.J.; Ginther, C.; Atefi, M.; Chen, I.; Fowst, C.; et al. PD 0332991, a selective cyclin D kinase 4/6 inhibitor, preferentially inhibits proliferation of luminal estrogen receptor-positive human breast cancer cell lines in vitro. Breast Cancer Res. 2009, 11, R77. [Google Scholar] [CrossRef] [Green Version]
- Subik, K.; Lee, J.-F.; Baxter, L.; Strzepek, T.; Costello, D.; Crowley, P.; Xing, L.; Hung, M.-C.; Bonfiglio, T.; Hicks, D.G.; et al. The Expression Patterns of ER, PR, HER2, CK5/6, EGFR, Ki-67 and AR by Immunohistochemical analysis in breast cancer cell lines. Breast Cancer Basic Clin. Res. 2010, 4, 35–41. [Google Scholar] [CrossRef]
- Dai, X.; Cheng, H.; Bai, Z.; Li, J. Breast Cancer Cell Line Classification and Its Relevance with Breast Tumor Subtyping. J. Cancer 2017, 8, 3131–3141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Neve, R.M.; Chin, K.; Fridlyand, J.; Yeh, J.; Baehner, F.L.; Fevr, T.; Clark, L.; Bayani, N.; Coppe, J.-P.; Tong, F.; et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 2006, 10, 515–527. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hongisto, V.; Jernström, S.; Fey, V.; Mpindi, J.P.; Kleivi Sahlberg, K.; Kallioniemi, O.; Perälä, M. High-throughput 3D screening reveals differences in drug sensitivities between culture models of JIMT1 breast cancer cells. PLoS ONE 2013, 8, e77232. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Araki, K.; Miyoshi, Y. Mechanism of resistance to endocrine therapy in breast cancer: The important role of PI3K/Akt/mTOR in estrogen receptor-positive, HER2-negative breast cancer. Breast Cancer 2017, 25, 392–401. [Google Scholar] [CrossRef] [PubMed]
- Dustin, D.; Gu, G.; Fuqua, S.A.W. ESR1 mutations in breast cancer. Cancer 2019, 125, 3714–3728. [Google Scholar] [CrossRef] [PubMed]
- Rozeboom, B.; Dey, N.; De, P. ER+ metastatic breast cancer: Past, present, and a prescription for an apoptosis-targeted future. Am. J. Cancer Res. 2019, 9, 2821–2831. [Google Scholar]
- Keck, K.J.; Breheny, P.; Braun, T.A.; Darbro, B.; Li, G.; Dillon, J.S.; Bellizzi, A.M.; O’Dorisio, T.M.; Howe, J.R. Changes in gene expression in small bowel neuroendocrine tumors associated with progression to metastases. Surgery 2017, 163, 232–239. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Cao, J.; Lin, W.; Chen, H.; Xiong, X.; Ao, H.; Yu, M.; Lin, J.; Cui, Q. The Roles of Cyclin-Dependent Kinases in Cell-Cycle Progression and Therapeutic Strategies in Human Breast Cancer. Int. J. Mol. Sci. 2020, 21, 1960. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Gene | Official Full Name | Correlation Coefficient | p-Value |
|---|---|---|---|
| MDM2 | MDM2 proto-oncogene | 0.797 | 0.001 |
| KRAS | KRAS proto-oncogene, GTPase | 0.791 | 0.001 |
| KIT | KIT proto-oncogene, receptor tyrosine kinase | 0.725 | 0.005 |
| NFKBIA | NFKB inhibitor alpha | 0.725 | 0.005 |
| MYB | MYB proto-oncogene, transcription factor | 0.709 | 0.007 |
| AKT1 | AKT serine/threonine kinase 1 | 0.681 | 0.010 |
| WWOX | WW domain containing oxidoreductase | 0.681 | 0.010 |
| XRCC1 | X-ray repair cross complementing 1 | 0.681 | 0.010 |
| BCL2 | BCL2 apoptosis regulator | 0.670 | 0.012 |
| TP73 | tumor protein p73 | 0.665 | 0.013 |
| APC | APC regulator of WNT signaling pathway | 0.654 | 0.015 |
| ESR1 | estrogen receptor 1 | 0.654 | 0.015 |
| BRCA2 | BRCA2 DNA repair associated | 0.648 | 0.017 |
| BRCA1 | BRCA1 DNA repair associated | 0.643 | 0.018 |
| RARA | retinoic acid receptor alpha | 0.637 | 0.019 |
| MYCN | MYCN proto-oncogene, bHLH transcription factor | 0.610 | 0.027 |
| CDK4 | cyclin dependent kinase 4 | 0.599 | 0.031 |
| NF1 | neurofibromin 1 | 0.593 | 0.033 |
| CDH1 | cadherin 1 | 0.566 | 0.044 |
| Clinicopathological Parameter | |
|---|---|
| Age, median (range) | 55 (27–78) |
| ≤60 years | 107 (64.8%) |
| >60 years | 58 (35.2%) |
| Histology | |
| DCIS | 7 (4.2%) |
| IDC | 146 (88.6%) |
| ILC | 7 (4.2%) |
| Others | 5 (3.0%) |
| UICC T factor | |
| Tis | 7 (4.2%) |
| T1 | 70 (42.4%) |
| T2 | 74 (44.9%) |
| T3 | 8 (4.9%) |
| T4 | 6 (3.6%) |
| Node status | |
| Negative | 85 (51.5%) |
| Positive | 80 (48.5%) |
| UICC pathological stage | |
| 0 | 7 (4.2%) |
| I | 47 (18.2%) |
| II | 78 (57.3%) |
| III | 32 (22.9%) |
| IV | 1 (0.6%) |
| ER status | |
| Positive | 125 (75.8%) |
| Negative | 40 (24.2%) |
| PgR status | |
| Positive | 113 (68.5%) |
| Negative | 52 (31.5%) |
| HER2 status | |
| Positive | 38 (23.0%) |
| Negative | 118 (71.5%) |
| Unknown | 9 (5.5%) |
| Triple-negative | |
| Yes | 18 (10.9%) |
| No | 146 (88.5%) |
| Unknown | 1 (0.6%) |
| Adjuvant therapy | |
| Endocrine therapy alone | 57 (34.5%) |
| Chemotherapy alone | 30 (18.2%) |
| Endocrine and chemotherapy | 62 (37.6%) |
| None | 16 (9.7%) |
| Clinicopathological Parameter | High SYT13 Group (n = 42) | Others (n = 123) | p-Value * |
|---|---|---|---|
| Age | 0.965 | ||
| ≤60 year | 28 | 79 | |
| >60 year | 15 | 43 | |
| Histology | 0.985 | ||
| DCIS | 2 | 5 | |
| IDC | 38 | 108 | |
| ILC | 3 | 4 | |
| Others | 0 | 5 | |
| UICC T factor | 0.162 | ||
| Tis/T1 | 24 | 53 | |
| T2/T3/T4 | 19 | 69 | |
| Node status | 0.084 | ||
| Negative | 27 | 58 | |
| Positive | 16 | 64 | |
| UICC pathological stage | 0.095 | ||
| 0/I/II | 38 | 94 | |
| III/IV | 5 | 28 | |
| ER status | 0.018 * | ||
| Positive | 38 | 87 | |
| Negative | 5 | 35 | |
| PgR status | 0.009 * | ||
| Positive | 36 | 77 | |
| Negative | 7 | 45 | |
| HER2 status | 0.020 * | ||
| Positive | 4 | 34 | |
| Negative | 33 | 85 | |
| Triple-negative | 0.337 | ||
| Yes | 3 | 15 | |
| No | 39 | 107 | |
| Adjuvant therapy | 0.124 | ||
| Endocrine therapy alone | 17 | 40 | |
| Chemotherapy alone | 3 | 27 | |
| Endocrine and chemotherapy | 18 | 44 | |
| None | 5 | 11 |
| Clinicopathological Parameter | |
|---|---|
| Age, median (range) | 58 (29–90) |
| ≤60 years | 391 (57.4%) |
| >60 years | 290 (42.6%) |
| Sex | |
| Male | 9 (1.3%) |
| Female | 672 (98.7%) |
| Histology | |
| IDC | 494 (72.5%) |
| ILC | 113 (16.6%) |
| Others | 74 (10.9%) |
| UICC T factor | |
| T1 | 188 (27.6%) |
| T2 | 382 (56.1%) |
| T3 | 88 (12.9%) |
| T4 | 22 (3.2%) |
| Unknown | 1 (0.2%) |
| Node status | |
| Negative | 323 (47.4%) |
| Positive | 354 (52.0%) |
| Unknown | 4 (0.6%) |
| UICC pathological stage | |
| I | 124 (18.2%) |
| II | 390 (57.3%) |
| III | 156 (22.9%) |
| IV | 6 (0.9%) |
| Unknown | 5 (0.7%) |
| ER status | |
| Positive | 532 (78.1%) |
| Negative | 149 (21.9%) |
| PgR status | |
| Positive | 456 (67.0%) |
| Negative | 225 (33.0%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ichikawa, T.; Shibata, M.; Inaishi, T.; Soeda, I.; Kanda, M.; Hayashi, M.; Takano, Y.; Takeuchi, D.; Tsunoda, N.; Kodera, Y.; et al. Synaptotagmin 13 Is Highly Expressed in Estrogen Receptor-Positive Breast Cancer. Curr. Oncol. 2021, 28, 4080-4092. https://doi.org/10.3390/curroncol28050346
Ichikawa T, Shibata M, Inaishi T, Soeda I, Kanda M, Hayashi M, Takano Y, Takeuchi D, Tsunoda N, Kodera Y, et al. Synaptotagmin 13 Is Highly Expressed in Estrogen Receptor-Positive Breast Cancer. Current Oncology. 2021; 28(5):4080-4092. https://doi.org/10.3390/curroncol28050346
Chicago/Turabian StyleIchikawa, Takahiro, Masahiro Shibata, Takahiro Inaishi, Ikumi Soeda, Mitsuro Kanda, Masamichi Hayashi, Yuko Takano, Dai Takeuchi, Nobuyuki Tsunoda, Yasuhiro Kodera, and et al. 2021. "Synaptotagmin 13 Is Highly Expressed in Estrogen Receptor-Positive Breast Cancer" Current Oncology 28, no. 5: 4080-4092. https://doi.org/10.3390/curroncol28050346
APA StyleIchikawa, T., Shibata, M., Inaishi, T., Soeda, I., Kanda, M., Hayashi, M., Takano, Y., Takeuchi, D., Tsunoda, N., Kodera, Y., & Kikumori, T. (2021). Synaptotagmin 13 Is Highly Expressed in Estrogen Receptor-Positive Breast Cancer. Current Oncology, 28(5), 4080-4092. https://doi.org/10.3390/curroncol28050346





