Prospective Monitoring of Circulating Epithelial Tumor Cells (CETC) Reveals Changes in Gene Expression during Adjuvant Radiotherapy of Breast Cancer Patients
Abstract
:1. Introduction
2. Materials and Methods
2.1. Inclusion Criteria
2.2. Study Procedures
2.3. Radiotherapy
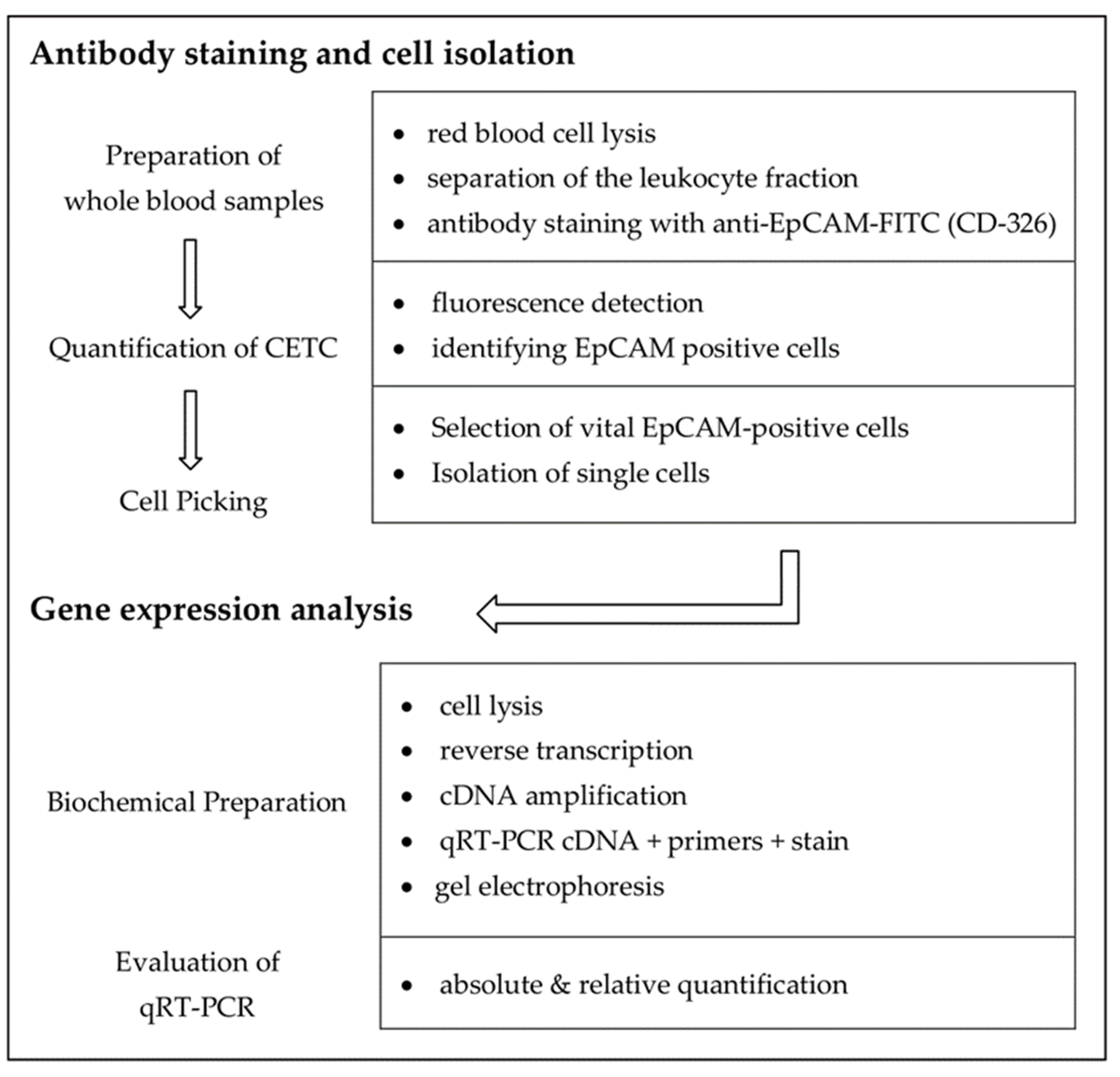
2.4. Sample Processing and Analysis
- quantitative immunofluorescence detection of viable CETC and
- quantitative gene expression analysis of single CETC by qRT-PCR to analyze for differential expression of selected genes.
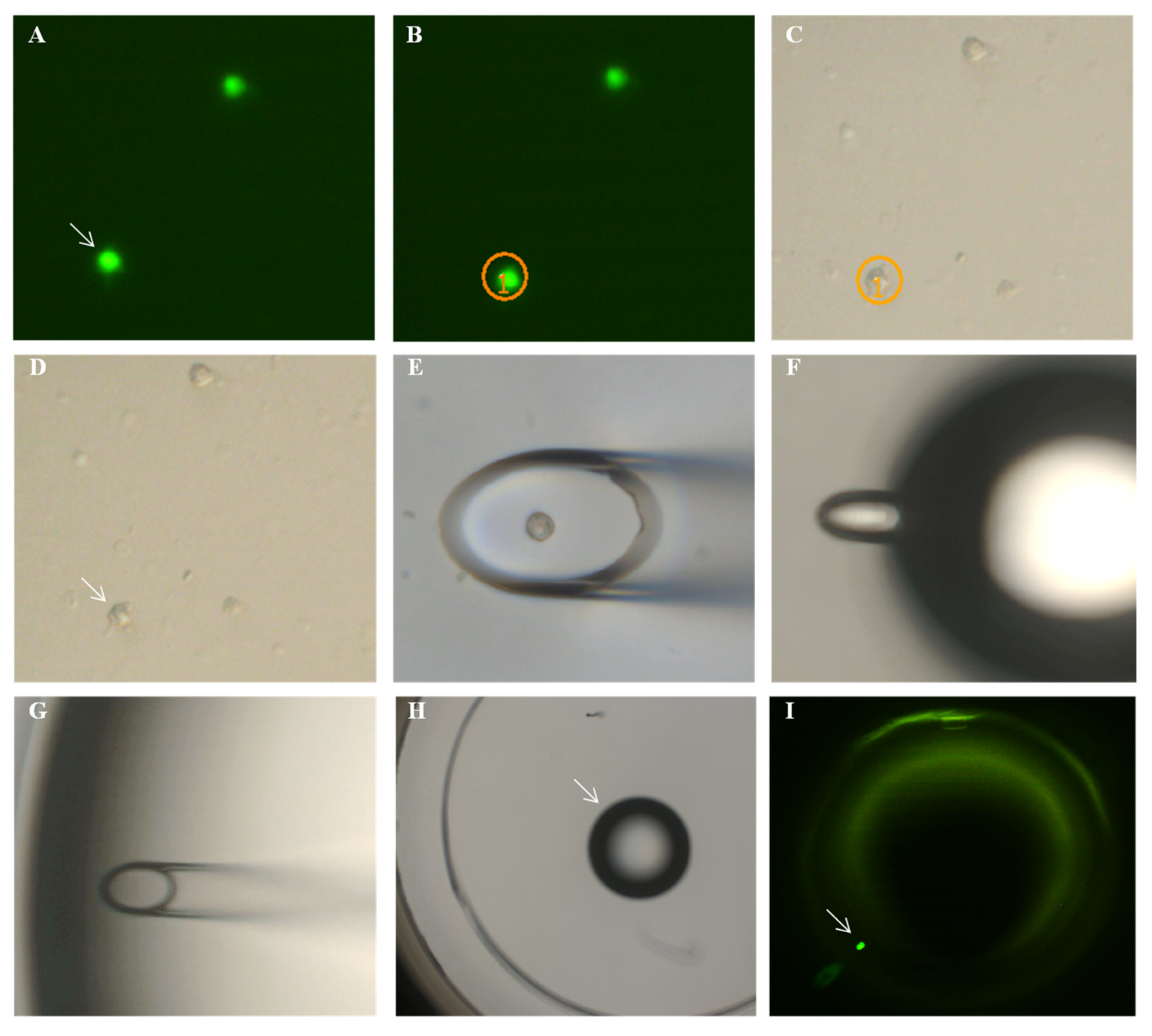
2.5. Immunofluorescence Assay and Quantification
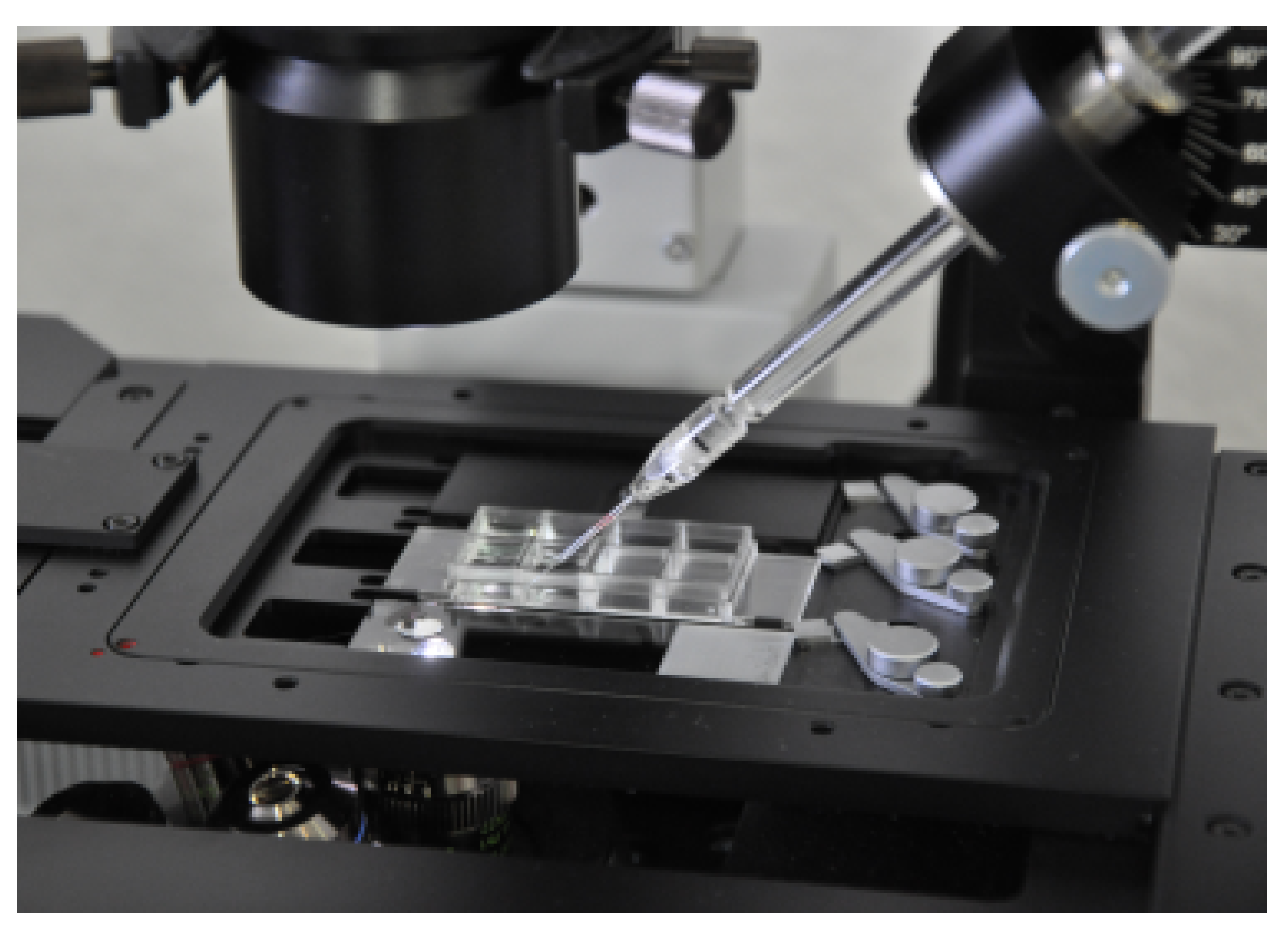
2.6. Single Cell Isolation and Quantitative Real-Time PCR
2.7. Statistical Analysis
3. Results
3.1. Changes in CETC Number
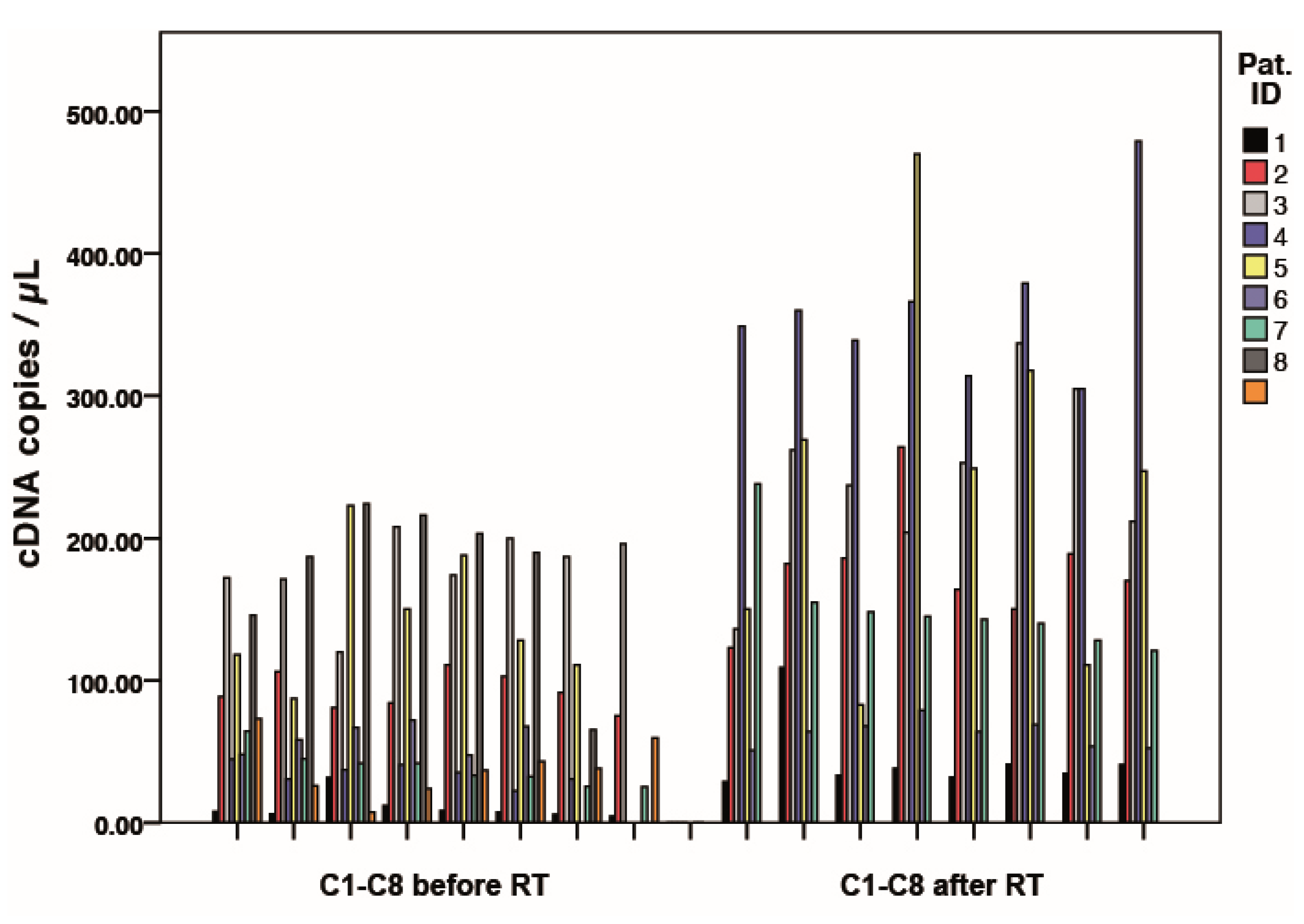
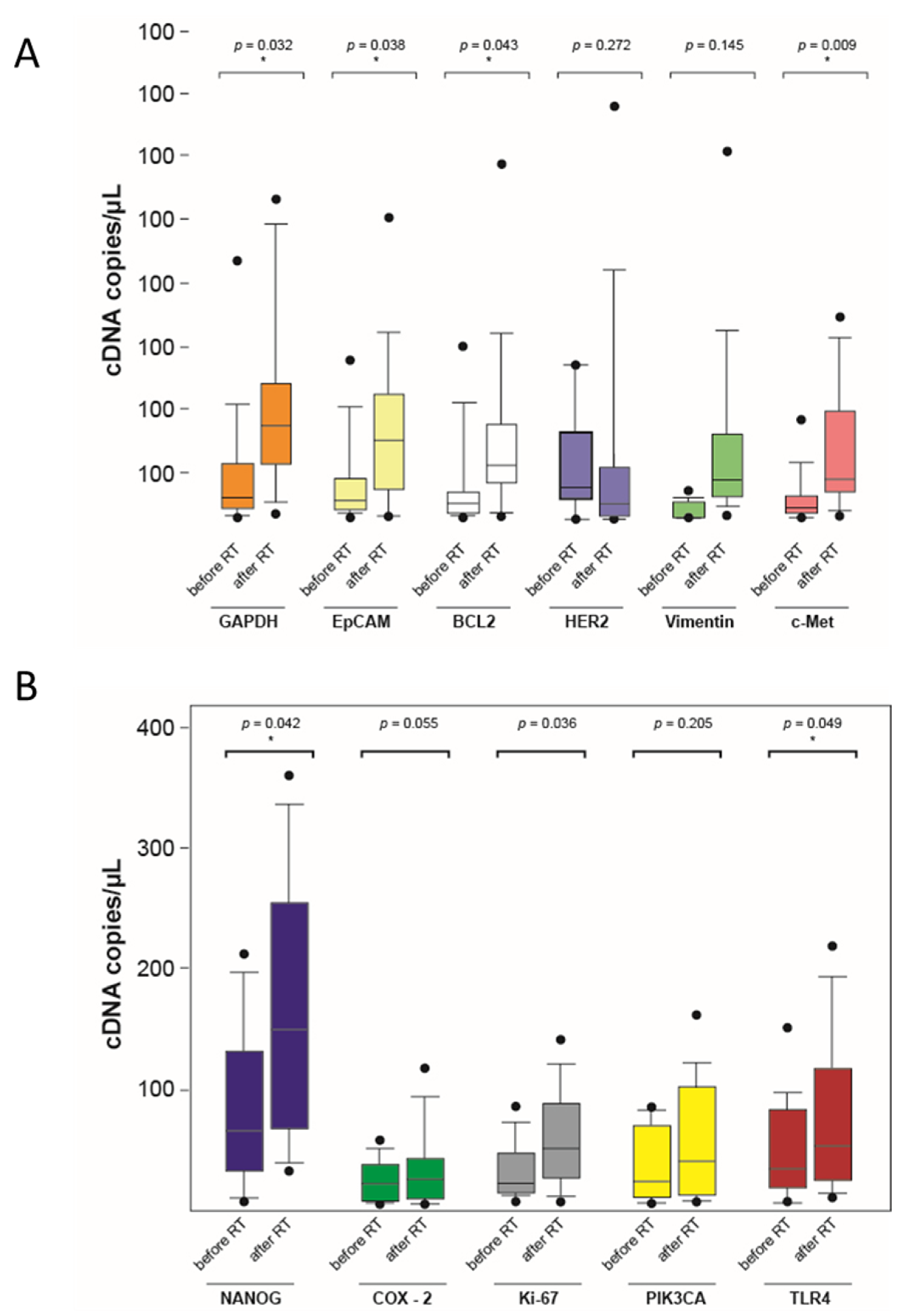
3.2. Gene Expression Profiles of Single Circulating Epithelial Cells
4. Discussion
4.1. Comparison of Methods for Detection of CETC
4.2. Expression Pattern of EpCAM
4.3. Quantitative Polymerase Chain Reaction
4.4. Quantitative Measurement of the CETC
4.5. Gene Expression Analysis
4.6. Influence of Endocrine Therapy
4.7. Clinical Follow-Up
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Glossary
| cDNA | Complementary deoxyribonucleic acid |
| CETC | Circulating epithelial tumor cells |
| EpCAM | Epithelial cell adhesion molecule |
| FITC | Fluoroisothiocyanate |
| GAPDH | Glyceraldehyde 3-phosphate dehydrogenase |
| mRNA | Messenger ribonucleic acid |
| qRT-PCR | Quantitative real-time PCR |
| RT | Radiotherapy |
References
- Early Breast Cancer Trialists’ Collaborative, G.; Darby, S.; McGale, P.; Correa, C.; Taylor, C.; Arriagada, R.; Clarke, M.; Cutter, D.; Davies, C.; Ewertz, M.; et al. Effect of radiotherapy after breast-conserving surgery on 10-year recurrence and 15-year breast cancer death: Meta-analysis of individual patient data for 10,801 women in 17 randomised trials. Lancet 2011, 378, 1707–1716. [Google Scholar]
- Taubert, H.; Blumke, K.; Bilkenroth, U.; Meye, A.; Kutz, A.; Bartel, F.; Lautenschlager, C.; Ulbrich, E.J.; Nass, N.; Holzhausen, H.J.; et al. Detection of disseminated tumor cells in peripheral blood of patients with breast cancer: Correlation to nodal status and occurrence of metastases. Gynecol. Oncol. 2004, 92, 256–261. [Google Scholar] [CrossRef]
- Fabisiewicz, A.; Szostakowska-Rodzos, M.; Zaczek, A.J.; Grzybowska, E.A. Circulating tumor cells in early and advanced breast cancer; biology and prognostic value. Int. J. Mol. Sci. 2020, 21, 1671. [Google Scholar] [CrossRef] [Green Version]
- Nicolini, A.; Ferrari, P.; Duffy, M.J. Prognostic and predictive biomarkers in breast cancer: Past, present and future. Semin. Cancer Biol. 2018, 52, 56–73. [Google Scholar] [CrossRef]
- Pantel, K.; Muller, V.; Auer, M.; Nusser, N.; Harbeck, N.; Braun, S. Detection and clinical implications of early systemic tumor cell dissemination in breast cancer. Clin. Cancer Res. 2003, 9, 6326–6334. [Google Scholar]
- Goodman, C.R.; Seagle, B.L.; Friedl, T.W.P.; Rack, B.; Lato, K.; Fink, V.; Cristofanilli, M.; Donnelly, E.D.; Janni, W.; Shahabi, S.; et al. Association of circulating tumor cell status with benefit of radiotherapy and survival in early-stage breast cancer. JAMA Oncol. 2018, 4, e180163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reinhardt, F.; Franken, A.; Meier-Stiegen, F.; Driemel, C.; Stoecklein, N.H.; Fischer, J.C.; Niederacher, D.; Ruckhaeberle, E.; Fehm, T.; Neubauer, H. Diagnostic leukapheresis enables reliable transcriptomic profiling of single circulating tumor cells to characterize inter-cellular heterogeneity in terms of endocrine resistance. Cancers 2019, 11, 903. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liang, D.H.; Hall, C.; Lucci, A. Circulating tumor cells in breast cancer. Recent Results Cancer Res. 2020, 215, 127–145. [Google Scholar]
- Janni, W.J.; Rack, B.; Terstappen, L.W.; Pierga, J.Y.; Taran, F.A.; Fehm, T.; Hall, C.; de Groot, M.R.; Bidard, F.C.; Friedl, T.W.; et al. Pooled analysis of the prognostic relevance of circulating tumor cells in primary breast cancer. Clin. Cancer Res. 2016, 22, 2583–2593. [Google Scholar] [CrossRef] [Green Version]
- Frick, M.A.; Feigenberg, S.J.; Jean-Baptiste, S.R.; Aguarin, L.A.; Mendes, A.; Chinniah, C.; Swisher-McClure, S.; Berman, A.; Levin, W.; Cengel, K.A.; et al. Circulating tumor cells are associated with recurrent disease in patients with early-stage non-small cell lung cancer treated with stereotactic body radiotherapy. Clin. Cancer Res. 2020, 26, 2372–2380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pachmann, K.; Camara, O.; Kavallaris, A.; Krauspe, S.; Malarski, N.; Gajda, M.; Kroll, T.; Jorke, C.; Hammer, U.; Altendorf-Hofmann, A.; et al. Monitoring the response of circulating epithelial tumor cells to adjuvant chemotherapy in breast cancer allows detection of patients at risk of early relapse. J. Clin. Oncol. 2008, 26, 1208–1215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bidard, F.C.; Peeters, D.J.; Fehm, T.; Nole, F.; Gisbert-Criado, R.; Mavroudis, D.; Grisanti, S.; Generali, D.; Garcia-Saenz, J.A.; Stebbing, J.; et al. Clinical validity of circulating tumour cells in patients with metastatic breast cancer: A pooled analysis of individual patient data. Lancet Oncol. 2014, 15, 406–414. [Google Scholar] [CrossRef]
- Cristofanilli, M.; Hayes, D.F.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Reuben, J.M.; Doyle, G.V.; Matera, J.; Allard, W.J.; Miller, M.C.; et al. Circulating tumor cells: A novel prognostic factor for newly diagnosed metastatic breast cancer. J. Clin. Oncol. 2005, 23, 1420–1430. [Google Scholar] [CrossRef]
- Cherdyntseva, N.V.; Litviakov, N.V.; Denisov, E.V.; Gervas, P.A.; Cherdyntsev, E.S. Circulating tumor cells in breast cancer: Functional heterogeneity, pathogenetic and clinical aspects. Exp. Oncol. 2017, 39, 2–11. [Google Scholar] [CrossRef]
- Baccelli, I.; Schneeweiss, A.; Riethdorf, S.; Stenzinger, A.; Schillert, A.; Vogel, V.; Klein, C.; Saini, M.; Bäuerle, T.; Wallwiener, M.; et al. Identification of a population of blood circulating tumor cells from breast cancer patients that initiates metastasis in a xenograft assay. Nat. Biotechnol. 2013, 31, 539–544. [Google Scholar] [CrossRef]
- Hall, C.; Valad, L.; Lucci, A. Circulating tumor cells in breast cancer patients. Crit. Rev. Oncog. 2016, 21, 125–139. [Google Scholar] [CrossRef] [PubMed]
- Sorg, S.; Pachmann, K.; Brede-Hekimian, K.; Freesmeyer, M.; Winkens, T. Determining tissue origin of circulating epithelial cells (cec) in patients with differentiated thyroid cancer by real-time pcr using thyroid mrna probes. Cancer Lett. 2015, 356, 491–495. [Google Scholar] [CrossRef]
- Lapin, M.; Tjensvoll, K.; Oltedal, S.; Javle, M.; Smaaland, R.; Gilje, B.; Nordgard, O. Single-cell mrna profiling reveals transcriptional heterogeneity among pancreatic circulating tumour cells. BMC Cancer 2017, 17, 390. [Google Scholar] [CrossRef]
- Wenz, F.; Budach, W. Personalized radiotherapy for invasive breast cancer in 2017 national s3 guidelines and degro and ago recommendations. Strahlenther. Onkol. 2017, 193, 601–603. [Google Scholar] [CrossRef] [Green Version]
- Pachmann, K.; Clement, J.H.; Schneider, C.P.; Willen, B.; Camara, O.; Pachmann, U.; Hoffken, K. Standardized quantification of circulating peripheral tumor cells from lung and breast cancer. Clin. Chem. Lab. Med. 2005, 43, 617–627. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W.; Hageleit, M. Validities of mrna quantification using recombinant rna and recombinant DNA external calibration curves in real-time rt-pcr. Biotechnol. Lett. 2001, 23, 275–282. [Google Scholar] [CrossRef]
- Ståhlberg, A.; Kubista, M. The workflow of single-cell expression profiling using quantitative real-time pcr. Expert Rev. Mol. Diagn. 2014, 14, 323–331. [Google Scholar] [CrossRef]
- Mäurer, M.A. Analysen Zirkulierender Epithelialer Tumorzellen im Periphervenösen blut bei Patientinnen mit Primär nicht Metastasiertem Mammakarzinom unter Adjuvanter Radiotherapie. Ph.D. Thesis, Friedrich-Schiller-University Jena, Jena, Germany, 2 December 2014. [Google Scholar]
- Pachmann, K.; Pizon, M.; Zimon, D.; Stein, E.L. Mc13-0080 somatic mutations of the egfr, kras and braf genes: Heterogeneity in circulating epithalial tumor cells (cetc) as determined using the cobas® z 480 analyzer. Eur. J. Cancer 2013, 49, S34. [Google Scholar] [CrossRef]
- Alix-Panabieres, C.; Pantel, K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 2016, 6, 479–491. [Google Scholar] [CrossRef] [Green Version]
- Koonce, N.A.; Juratli, M.A.; Cai, C.; Sarimollaoglu, M.; Menyaev, Y.A.; Dent, J.; Quick, C.M.; Dings, R.P.M.; Nedosekin, D.; Zharov, V.; et al. Real-time monitoring of circulating tumor cell (ctc) release after nanodrug or tumor radiotherapy using in vivo flow cytometry. Biochem. Biophys. Res. Commun. 2017, 492, 507–512. [Google Scholar] [CrossRef]
- Wang, C.; Mu, Z.; Chervoneva, I.; Austin, L.; Ye, Z.; Rossi, G.; Palazzo, J.P.; Sun, C.; Abu-Khalaf, M.; Myers, R.E.; et al. Longitudinally collected ctcs and ctc-clusters and clinical outcomes of metastatic breast cancer. Breast Cancer Res. Treat. 2017, 161, 83–94. [Google Scholar] [CrossRef] [PubMed]
- Tewes, M.; Aktas, B.; Welt, A.; Mueller, S.; Hauch, S.; Kimmig, R.; Kasimir-Bauer, S. Molecular profiling and predictive value of circulating tumor cells in patients with metastatic breast cancer: An option for monitoring response to breast cancer related therapies. Breast Cancer Res. Treat. 2009, 115, 581–590. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pizon, M.; Zimon, D.; Carl, S.; Pachmann, U.; Pachmann, K.; Camara, O. Heterogeneity of circulating epithelial tumour cells from individual patients with respect to expression profiles and clonal growth (sphere formation) in breast cancer. Ecancermedicalscience 2013, 7, 343. [Google Scholar] [PubMed]
- Li, Y.; Wu, S.; Bai, F. Molecular characterization of circulating tumor cells-from bench to bedside. Semin Cell Dev. Biol. 2018, 75, 88–97. [Google Scholar] [CrossRef] [PubMed]
- Cristofanilli, M.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Matera, J.; Miller, M.C.; Reuben, J.M.; Doyle, G.V.; Allard, W.J.; Terstappen, L.W.; et al. Circulating tumor cells, disease progression, and survival in metastatic breast cancer. N. Engl. J. Med. 2004, 351, 781–791. [Google Scholar] [CrossRef] [Green Version]
- Muller, V.; Riethdorf, S.; Rack, B.; Janni, W.; Fasching, P.A.; Solomayer, E.; Aktas, B.; Kasimir-Bauer, S.; Pantel, K.; Fehm, T.; et al. Prognostic impact of circulating tumor cells assessed with the cellsearch system and adnatest breast in metastatic breast cancer patients: The detect study. Breast Cancer Res. 2012, 14, R118. [Google Scholar] [CrossRef] [PubMed]
- Riethdorf, S.; Fritsche, H.; Muller, V.; Rau, T.; Schindlbeck, C.; Rack, B.; Janni, W.; Coith, C.; Beck, K.; Janicke, F.; et al. Detection of circulating tumor cells in peripheral blood of patients with metastatic breast cancer: A validation study of the cellsearch system. Clin. Cancer Res. 2007, 13, 920–928. [Google Scholar] [CrossRef] [Green Version]
- Nole, F.; Munzone, E.; Zorzino, L.; Minchella, I.; Salvatici, M.; Botteri, E.; Medici, M.; Verri, E.; Adamoli, L.; Rotmensz, N.; et al. Variation of circulating tumor cell levels during treatment of metastatic breast cancer: Prognostic and therapeutic implications. Ann. Oncol. 2008, 19, 891–897. [Google Scholar] [CrossRef]
- Deng, G.; Herrler, M.; Burgess, D.; Manna, E.; Krag, D.; Burke, J.F. Enrichment with anti-cytokeratin alone or combined with anti-epcam antibodies significantly increases the sensitivity for circulating tumor cell detection in metastatic breast cancer patients. Breast Cancer Res. 2008, 10, R69. [Google Scholar] [CrossRef] [Green Version]
- Coumans, F.A.W.; Doggen, C.J.M.; Attard, G.; de Bono, J.S.; Terstappen, L. All circulating epcam+ck+cd45- objects predict overall survival in castration-resistant prostate cancer. Ann. Oncol. 2010, 21, 1851–1857. [Google Scholar] [CrossRef] [PubMed]
- Fehm, T.; Muller, V.; Alix-Panabieres, C.; Pantel, K. Micrometastatic spread in breast cancer: Detection, molecular characterization and clinical relevance. Breast Cancer Res. 2008, 10 (Suppl. 1), S1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pachmann, U.; Hekimian, K.; Pachmann, K. Comparing sequential steps for detection of circulating tumor cells: More specific or just less sensitive? Oncol. Rep. 2011, 37, 3219–3226. [Google Scholar]
- Gaforio, J.J.; Serrano, M.J.; Sanchez-Rovira, P.; Sirvent, A.; Delgado-Rodriguez, M.; Campos, M.; de la Torre, N.; Algarra, I.; Duenas, R.; Lozano, A. Detection of breast cancer cells in the peripheral blood is positively correlated with estrogen-receptor status and predicts for poor prognosis. Int. J. Cancer 2003, 107, 984–990. [Google Scholar] [CrossRef]
- Lankiewicz, S.; Rivero, B.G.; Bocher, O. Quantitative real-time rt-pcr of disseminated tumor cells in combination with immunomagnetic cell enrichment. Mol. Biotechnol. 2006, 34, 15–27. [Google Scholar] [CrossRef]
- Tandon, A.K.; Clark, G.M.; Chamness, G.C.; McGuire, W.L. Association of the 323/a3 surface glycoprotein with tumor characteristics and behavior in human breast cancer. Cancer Res. 1990, 50, 3317–3321. [Google Scholar]
- Balzar, M.; Winter, M.J.; de Boer, C.J.; Litvinov, S.V. The biology of the 17-1a antigen (ep-cam). J. Mol. Med. (Berl) 1999, 77, 699–712. [Google Scholar] [CrossRef] [PubMed]
- Osta, W.A.; Chen, Y.; Mikhitarian, K.; Mitas, M.; Salem, M.; Hannun, Y.A.; Cole, D.J.; Gillanders, W.E. Epcam is overexpressed in breast cancer and is a potential target for breast cancer gene therapy. Cancer Res. 2004, 64, 5818–5824. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moldenhauer, G.; Momburg, F.; Moller, P.; Schwartz, R.; Hammerling, G.J. Epithelium-specific surface glycoprotein of mr 34,000 is a widely distributed human carcinoma marker. Br. J. Cancer 1987, 56, 714–721. [Google Scholar] [CrossRef] [Green Version]
- Keilholz, U.; Willhauck, M.; Scheibenbogen, C.; deVries, T.J.; Burchill, S. Polymerase chain reaction detection of circulating tumour cells. Melanoma Res. 1997, 7, S133–S141. [Google Scholar] [CrossRef]
- Kvastad, L.; Werne Solnestam, B.; Johansson, E.; Nygren, A.O.; Laddach, N.; Sahlen, P.; Vickovic, S.; Bendigtsen, S.C.; Aaserud, M.; Floer, L.; et al. Single cell analysis of cancer cells using an improved rt-mlpa method has potential for cancer diagnosis and monitoring. Sci. Rep. 2015, 5, 16519. [Google Scholar] [CrossRef]
- Daskalaki, A.; Agelaki, S.; Perraki, M.; Apostolaki, S.; Xenidis, N.; Stathopoulos, E.; Kontopodis, E.; Hatzidaki, D.; Mavroudis, D.; Georgoulias, V. Detection of cytokeratin-19 mrna-positive cells in the peripheral blood and bone marrow of patients with operable breast cancer. Br. J. Cancer 2009, 101, 589–597. [Google Scholar] [CrossRef]
- Rajeevan, M.S.; Vernon, S.D.; Taysavang, N.; Unger, E.R. Validation of array-based gene expression profiles by real-time (kinetic) rt-pcr. J. Mol. Diagn. 2001, 3, 26–31. [Google Scholar] [CrossRef] [Green Version]
- Fehm, T.; Sagalowsky, A.; Clifford, E.; Beitsch, P.; Saboorian, H.; Euhus, D.; Meng, S.; Morrison, L.; Tucker, T.; Lane, N.; et al. Cytogenetic evidence that circulating epithelial cells in patients with carcinoma are malignant. Clin. Cancer Res. 2002, 8, 2073–2084. [Google Scholar]
- Klein, C.A. Parallel progression of primary tumours and metastases. Nat. Rev. Cancer 2009, 9, 302–312. [Google Scholar] [CrossRef] [PubMed]
- Klopp, A.H.; Spaeth, E.L.; Dembinski, J.L.; Woodward, W.A.; Munshi, A.; Meyn, R.E.; Cox, J.D.; Andreeff, M.; Marini, F.C. Tumor irradiation increases the recruitment of circulating mesenchymal stem cells into the tumor microenvironment. Cancer Res. 2007, 67, 11687–11695. [Google Scholar] [CrossRef] [Green Version]
- Martin, O.A.; Anderson, R.L.; Narayan, K.; MacManus, M.P. Does the mobilization of circulating tumour cells during cancer therapy cause metastasis? Nat. Rev. Clin. Oncol. 2017, 14, 32–44. [Google Scholar] [CrossRef] [PubMed]
- Sherry, A.D.; von Eyben, R.; Newman, N.B.; Gutkin, P.; Mayer, I.; Horst, K.; Chakravarthy, A.B.; Rafat, M. Systemic inflammation after radiation predicts locoregional recurrence, progression, and mortality in stage ii-iii triple-negative breast cancer. Int. J. Radiat. Oncol. Biol. Phys. 2019, 108, 268–276. [Google Scholar] [CrossRef]
- Meng, S.; Tripathy, D.; Frenkel, E.P.; Shete, S.; Naftalis, E.Z.; Huth, J.F.; Beitsch, P.D.; Leitch, M.; Hoover, S.; Euhus, D.; et al. Circulating tumor cells in patients with breast cancer dormancy. Clin. Cancer Res. 2004, 10, 8152–8162. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Y.; Meucci, S.; Sheng, L.; Keilholz, U. Meta-analysis of the mutational status of circulation tumor cells and paired primary tumor tissues from colorectal cancer patients. Oncotarget. 2017, 8, 77928–77941. [Google Scholar] [CrossRef] [Green Version]
- Bolke, E.; Orth, K.; Gerber, P.A.; Lammering, G.; Mota, R.; Peiper, M.; Matuschek, C.; Budach, W.; Rusnak, E.; Shaikh, S.; et al. Gene expression of circulating tumour cells and its correlation with tumour stage in breast cancer patients. Eur. J. Med. Res. 2009, 14, 359–363. [Google Scholar] [CrossRef] [Green Version]
- Slamon, D.J.; Godolphin, W.; Jones, L.A.; Holt, J.A.; Wong, S.G.; Keith, D.E.; Levin, W.J.; Stuart, S.G.; Udove, J.; Ullrich, A.; et al. Studies of the her-2/neu proto-oncogene in human breast and ovarian cancer. Science 1989, 244, 707–712. [Google Scholar] [CrossRef] [PubMed]
- Takayama, H.; LaRochelle, W.J.; Sharp, R.; Otsuka, T.; Kriebel, P.; Anver, M.; Aaronson, S.A.; Merlino, G. Diverse tumorigenesis associated with aberrant development in mice overexpressing hepatocyte growth factor/scatter factor. Proc. Natl. Acad. Sci. USA 1997, 94, 701–706. [Google Scholar] [CrossRef] [Green Version]
- Laurenzi, I.J. An analytical solution of the stochastic master equation for reversible bimolecular reaction kinetics. J. Chem. Phys. 2000, 113, 3315–3322. [Google Scholar] [CrossRef]
- Simmons, D.L.; Botting, R.M.; Hla, T. Cyclooxygenase isozymes: The biology of prostaglandin synthesis and inhibition. Pharmacol. Rev. 2004, 56, 387–437. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [Green Version]
- Li, S.Y.; Rong, M.; Grieu, F.; Iacopetta, B. Pik3ca mutations in breast cancer are associated with poor outcome. Breast Cancer Res. Treat. 2006, 96, 91–95. [Google Scholar] [CrossRef]
- Ivaska, J.; Pallari, H.M.; Nevo, J.; Eriksson, J.E. Novel functions of vimentin in cell adhesion, migration, and signaling. Exp. Cell Res. 2007, 313, 2050–2062. [Google Scholar] [CrossRef]
- Maetzel, D.; Denzel, S.; Mack, B.; Canis, M.; Went, P.; Benk, M.; Kieu, C.; Papior, P.; Baeuerle, P.A.; Munz, M.; et al. Nuclear signalling by tumour-associated antigen epcam. Nat. Cell Biol. 2009, 11, 162–171. [Google Scholar] [CrossRef]
- Yang, H.; Zhou, H.; Feng, P.; Zhou, X.; Wen, H.; Xie, X.; Shen, H.; Zhu, X. Reduced expression of toll-like receptor 4 inhibits human breast cancer cells proliferation and inflammatory cytokines secretion. J. Exp. Clin. Cancer Res. 2010, 29, 92. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hou, Y.Y.; Cao, W.W.; Li, L.; Li, S.P.; Liu, T.; Wan, H.Y.; Liu, M.; Li, X.; Tang, H. Microrna-519d targets mki67 and suppresses cell growth in the hepatocellular carcinoma cell line qgy-7703. Cancer Lett. 2011, 307, 182–190. [Google Scholar] [CrossRef] [PubMed]
- Muraro, E.; Furlan, C.; Avanzo, M.; Martorelli, D.; Comaro, E.; Rizzo, A.; Fae, D.A.; Berretta, M.; Militello, L.; Del Conte, A.; et al. Local high-dose radiotherapy induces systemic immunomodulating effects of potential therapeutic relevance in oligometastatic breast cancer. Front Immunol. 2017, 8, 1476. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, G.; Li, Y.; He, Y.; Zeng, B.; Yi, C.; Wang, C.; Zhang, X.; Zhao, W.; Yu, D. Upregulation of circular rna circatrnl1 to sensitize oral squamous cell carcinoma to irradiation. Mol. Ther. Nucleic Acids 2020, 19, 961–973. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.M.; Yang, X.L.; Jiang, F.; Pan, Y.C.; Zhang, L. Matrine involves in the progression of gastric cancer through inhibiting mir-93-5p and upregulating the expression of target gene ahnak. J. cell. Biochem. 2020, 121, 2467–2477. [Google Scholar] [CrossRef]
- Jabbari, N.; Nawaz, M.; Rezaie, J. Bystander effects of ionizing radiation: Conditioned media from x-ray irradiated mcf-7 cells increases the angiogenic ability of endothelial cells. Cell commun. Signal. 2019, 17, 165. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.Y.; Jeong, E.K.; Ju, M.K.; Jeon, H.M.; Kim, M.Y.; Kim, C.H.; Park, H.G.; Han, S.I.; Kang, H.S. Induction of metastasis, cancer stem cell phenotype, and oncogenic metabolism in cancer cells by ionizing radiation. Mol. cancer 2017, 16, 10. [Google Scholar] [CrossRef] [Green Version]
- Jarosz-Biej, M.; Smolarczyk, R.; Cichon, T.; Kulach, N. Tumor microenvironment as a "game changer" in cancer radiotherapy. Int. J. Mol. Sci. 2019, 20, 3212. [Google Scholar] [CrossRef] [Green Version]
- Eke, I.; Makinde, A.Y.; Aryankalayil, M.J.; Sandfort, V.; Palayoor, S.T.; Rath, B.H.; Liotta, L.; Pierobon, M.; Petricoin, E.F.; Brown, M.F.; et al. Exploiting radiation-induced signaling to increase the susceptibility of resistant cancer cells to targeted drugs: Akt and mtor inhibitors as an example. Mol. Cancer Ther. 2018, 17, 355–367. [Google Scholar] [CrossRef] [Green Version]
- Leeman, J.E.; Schoenfeld, J.D. Radiation therapy and immune modulation. Hematol. Oncol. Clin. N. Am. 2019, 33, 233–248. [Google Scholar] [CrossRef]
- Lewin, N.L.; Luetragoon, T.; Andersson, B.A.; Oliva, D.; Nilsson, M.; Strandeus, M.; Lofgren, S.; Rutqvist, L.E.; Lewin, F. The influence of single nucleotide polymorphisms and adjuvant radiotherapy on systemic inflammatory proteins, chemokines and cytokines of patients with breast cancer. Anticancer Res. 2019, 39, 1287–1292. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, D.; Yusuf, N. Expression of toll-like receptors on breast tumors: Taking a toll on tumor microenvironment. Int. J. Breast Cancer 2012, 2012, 716564. [Google Scholar] [CrossRef]
- Kim, Y.B.; Kim, G.E.; Cho, N.H.; Pyo, H.R.; Shim, S.J.; Chang, S.K.; Park, H.C.; Suh, C.O.; Park, T.K.; Kim, B.S. Overexpression of cyclooxygenase-2 is associated with a poor prognosis in patients with squamous cell carcinoma of the uterine cervix treated with radiation and concurrent chemotherapy. Cancer 2002, 95, 531–539. [Google Scholar] [CrossRef] [PubMed]
- Mansoori, M.; Madjd, Z.; Janani, L.; Rasti, A. Circulating cancer stem cell markers in breast carcinomas: A systematic review protocol. Syst. Rev. 2017, 6, 262. [Google Scholar] [CrossRef] [Green Version]
- Wang, F.; Li, Y.C.; Liu, L.P.; Zhang, H.M.; Tong, S. Circulating tumor cells and tumor stem cells detection in the peripheral blood mononuclear cells of breast cancer. J. Clin. Lab. Anal. 2016, 30, 616–622. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ni, J.; Cozzi, P.; Hao, J.L.; Beretov, J.; Chang, L.; Duan, W.; Shigdar, S.; Delprado, W.; Graham, P.; Bucci, J.; et al. Epithelial cell adhesion molecule (epcam) is associated with prostate cancer metastasis and chemo/radioresistance via the pi3k/akt/mtor signaling pathway. Int. J. Biochem. Cell B 2013, 45, 2736–2748. [Google Scholar] [CrossRef]
- Deutsch, T.M.; Riethdorf, S.; Nees, J.; Hartkopf, A.D.; Schonfisch, B.; Domschke, C.; Sprick, M.R.; Schutz, F.; Brucker, S.Y.; Stefanovic, S.; et al. Impact of apoptotic circulating tumor cells (actc) in metastatic breast cancer. Breast Cancer Res. Treat. 2016, 160, 277–290. [Google Scholar] [CrossRef]
- Luzhna, L.; Golubov, A.; Ilnytskyy, S.; Chekhun, V.F.; Kovalchuk, O. Molecular mechanisms of radiation resistance in doxorubicin-resistant breast adenocarcinoma cells. Int. J. Oncol. 2013, 42, 1692–1708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Badve, S.; Nakshatri, H. Breast-cancer stem cells-beyond semantics. Lancet Oncol. 2012, 13, e43–e48. [Google Scholar] [CrossRef]
- Olivares-Urbano, M.A.; Grinan-Lison, C.; Rios-Arrabal, S.; Artacho-Cordon, F.; Torralbo, A.I.; Lopez-Ruiz, E.; Marchal, J.A.; Nunez, M.I. Radiation and stemness phenotype may influence individual breast cancer outcomes: The crucial role of mmps and microenvironment. Cancers 2019, 11, 1781. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saga, R.; Matsuya, Y.; Takahashi, R.; Hasegawa, K.; Date, H.; Hosokawa, Y. Analysis of the high-dose-range radioresistance of prostate cancer cells, including cancer stem cells, based on a stochastic model. J. Radiat. Res. 2019, 60, 298–307. [Google Scholar] [CrossRef]
- Qi, X.S.; Pajonk, F.; McCloskey, S.; Low, D.A.; Kupelian, P.; Steinberg, M.; Sheng, K. Radioresistance of the breast tumor is highly correlated to its level of cancer stem cell and its clinical implication for breast irradiation. Radiother. Oncol. 2017, 124, 455–461. [Google Scholar] [CrossRef]
- Schott, D.S.; Pizon, M.; Pachmann, U.; Pachmann, K.; Schobert, R.; Wittig, A.; Mäurer, M. Influence of adjuvant radiotherapy on circulating epithelial tumor cells and circulating cancer stem cells in primary non-metastatic breast cancer. Transl. Oncol. 2021, 14, 101009. [Google Scholar] [CrossRef]
- Liu, T.; Xu, H.; Huang, M.; Ma, W.; Saxena, D.; Lustig, R.A.; Alonso-Basanta, M.; Zhang, Z.; O’Rourke, D.M.; Zhang, L.; et al. Circulating glioma cells exhibit stem cell-like properties. Cancer Res. 2018, 78, 6632–6642. [Google Scholar] [CrossRef] [Green Version]
- Dutsch-Wicherek, M. Rcas1, mt, and vimentin as potential markers of tumor microenvironment remodeling. Am. J. Reprod. Immunol. 2010, 63, 181–188. [Google Scholar] [CrossRef]
- Pachmann, K.; Schuster, S. The value of monitoring the behavior of circulating tumor cells at the end of endocrine therapy in breast cancer patients. Cancers 2018, 10, 407. [Google Scholar] [CrossRef] [Green Version]
- Patriarca, C.; Macchi, R.M.; Marschner, A.K.; Mellstedt, H. Epithelial cell adhesion molecule expression (cd326) in cancer: A short review. Cancer Treat. Rev. 2012, 38, 68–75. [Google Scholar] [CrossRef] [PubMed]

| Gene Name | Sequence (5′–3′) Sense (s)—Antisense (a) | Product Length Base Pairs (bp) |
|---|---|---|
| GAPDH | s: GAC AGT CAG CCG CAT CTT CT a: GCG CCC AAT ACG ACC AAA TC | 104 bp |
| EpCAM | s: GGG AAA TAG CAA ATG GAC ACA a: CGA TGG AGT CCA AGT TCT GG | 219 bp |
| NANOG | s: GGA TCC AGC TTG TCC CCA AA a: TGC ACC AGG TCT GAG TGT TC | 674 bp |
| Bcl-2 | s: TTT GTG GAA CTG TAC GGC CC a: CCG GCC AAC AAC ATG GAA AG | 519 bp |
| TLR4 | s: GGT CAG ACG GTG ATA GCG AG a: ATT AGG AAC CAC CAC GC | 179 bp |
| COX-2 | s: GAT GAT TGC CCG ACT CCC TT a: TGA AAA GGC GCA GTT TAC GC | 274 bp |
| PIK3CA | s: CCC AGG TGG AAT GAA TGG CT a: CCA AAA GCA GGC CAA ACC TC | 925 bp |
| HER2 | s: AGG TAA CCC TGG CCC CTT T a: TTC AGC GGG TCT CCA TTG TC | 539 bp |
| Vimentin | s: TCC GCA CAT TCG AGC AAA GA a: ATT CAA GTC TCA GCG GGC TC | 161 bp |
| Ki-67 | s: CCT CAG CAC CTG CTT GTT TG a: TCC CTG AGC AAC ACT GTC TTT | 466 bp |
| c-Met | s: GGT CTT CAA GTA GCC AAA GCG a: TTC TTG CAG CCA AGT TGT | 73 bp |
| Pat. ID | Age [Years] | Date of First Diagnosis | Initial Tumor Stage and Tumor Characteristics | Receptor Status of Tumor Cells | Endocrine Therapy during RT | Locoregional Recurrence after 5 Years | Distant Metastasis after 5 Years |
|---|---|---|---|---|---|---|---|
| 1 | 60 | 05/2012 | pT1c pN0(0/1sn) cM0 L0 V0 R0 G2 | ER: 80% PR: 100% HER2: 0 | Anastrozol | - | - |
| 2 | 73 | 02/2012 | pT2 pN0(0/2sn) cM0 L0 V0 R0 G2 | ER: 85% PR: 85% HER2: 1+ | Letrozol | - | - |
| 3 | 68 | 03/2012 | pT1c pN0(0/1sn) cM0 L0 V0 R0 G2 | ER: 80% PR: 0% HER2: 2+ | Letrozol | - | - |
| 4 | 52 | 03/2012 | pT1c (m) pN0(0/1sn) cM0 L0 V0 R0 G2 | ER: 80% PR: 70% HER2: 1+ | Tamoxifen | - | - |
| 5 | 46 | 06/2012 | pT1b(m) pN0(0/1sn) cM0 L0 V0 R0 G1 | ER: 90% PR: 90% HER2: 2+ | Tamoxifen | Local relapse 78 months after RT | - |
| 6 | 55 | 04/2012 | pT1b pN0(0/2 sn) cM0 L0 V0 R0 G2 | ER: 90%, PR: 80% HER2: 1+ | Letrozol | - | - |
| 7 | 59 | 10/2011 | pT1a+Tis pN0(0/1sn) cM0 L0 V0 R0 G1 | ER: 100% PR: 100% HER2: 1+ | Letrozol | - | - |
| 8 | 51 | 11/2011 | pT1c pN0(0/3 sn) cM0 L0 V0 R0 G3 | ER: 0% PR: 0% HER2: 0 | - | - | - |
| 9 | 41 | 11/2011 | pT2 pN0(0/1 sn) cM0 L0 V0 R0 G3 | ER: 45% PR: 70% HER2: 2+ | Tamoxifen | - | - |
| Pat. ID | before Radiotherapy | after Radiotherapy | Quotient |
|---|---|---|---|
| Number of Viable CETC/mL Whole Blood | Number of Viable CETC/mL Whole Blood | ||
| 1 | 3170 | 8640 | 2.72 |
| 2 | 2200 | 5860 | 2.66 |
| 3 | 2440 | 6100 | 2.50 |
| 4 | 18070 | 17340 | 0.96 |
| 5 | 1460 | 1220 | 0.83 |
| 6 | 2690 | 5150 | 1.91 |
| 7 | 4200 | 11480 | 2.73 |
| 8 | 2930 | 4150 | 1.42 |
| 9 | 11720 | 18120 | 1.55 |
| Cell Number | GAPDH | EpCAM | NANOG | BCL-2 | ||||
|---|---|---|---|---|---|---|---|---|
| before RT | after RT | before RT | after RT | before RT | after RT | before RT | after RT | |
| 1 | 10.20 | 29.40 | 13.00 | 32.60 | 88.60 | 123.00 | 9.40 | 33.90 |
| 2 | 15.50 | 53.30 | 9.13 | 69.50 | 106.00 | 182.00 | 7.43 | 28.40 |
| 3 | 9.94 | 553.00 | 10.60 | 29.70 | 80.80 | 186.00 | 7.81 | 20.7 |
| 4 | 146.00 | 16.20 | 10.70 | 17.50 | 84.10 | 264.00 | 7.76 | 33.70 |
| 5 | 47.60 | 25.60 | 14.90 | 28.50 | 111.00 | 164.00 | 11.40 | 29.50 |
| 6 | 14.40 | 298.00 | 10.60 | 52.20 | 103.00 | 150.00 | 6.27 | 199.00 |
| 7 | 15.30 | 404.00 | 16.00 | 104.00 | 91.60 | 189.00 | 15.30 | 233.00 |
| 8 | 12.90 | 200.00 | 15.20 | 45.30 | 75.20 | 170.00 | 8.22 | 72.40 |
| Cell Number | COX-2 | PIK3CA | HER2 | Vimentin | ||||
| before RT | after RT | before RT | after RT | before RT | after RT | before RT | after RT | |
| 1 | 5.09 | 4.41 | 1.31 | 6.71 | 2.28 | 1.87 | 4.98 | 17.55 |
| 2 | 2.25 | 5.56 | 2.86 | 21.80 | 1.55 | 2.72 | 3.84 | 21.50 |
| 3 | 1.75 | 4.48 | 3.28 | 7.64 | 1.69 | 1.73 | 3.41 | 7.22 |
| 4 | 2.41 | 6.69 | 1.51 | 13.50 | 2.35 | 1.69 | 3.98 | 109.00 |
| 5 | 4.00 | 18.60 | 11.00 | 43.00 | 1.78 | 137.00 | 7.52 | 34.70 |
| 6 | 3.00 | 407.00 | 8.97 | 147.00 | 2.66 | 71.60 | 3.85 | 33.50 |
| 7 | 3.40 | 508.00 | 2.09 | 107.00 | 2.40 | 134.00 | 7.24 | 56.30 |
| 8 | 1.41 | 16.20 | 1.25 | 127.00 | 2.36 | 9.14 | 3.40 | 134.00 |
| Cell Number | c-Met | Ki-67 | TLR 4 | |||||
| before RT | after RT | before RT | after RT | before RT | after RT | |||
| 1 | 4.93 | 6.44 | 15.80 | 27.40 | 3.03 | 5.55 | ||
| 2 | 3.24 | 6.42 | 14.10 | 68.80 | 29.80 | 7.27 | ||
| 3 | 3.41 | 7.32 | 10.23 | 43.90 | 1.70 | 25.40 | ||
| 4 | 3.93 | 4.94 | 16.80 | 30.00 | 4.01 | 24.90 | ||
| 5 | 5.52 | 13.10 | 27.10 | 67.40 | 3.60 | 85.60 | ||
| 6 | 3.85 | 32.20 | 23.90 | 104.00 | 2.30 | 724.00 | ||
| 7 | 7.42 | 53.30 | 18.10 | 131.00 | 3.44 | 864.00 | ||
| 8 | 3.70 | 19.10 | 17.40 | 87.00 | 1.34 | 93.20 | ||
| Pat. ID | CETC Number | GAPDH | EpCAM | NANOG | BCL-2 | COX-2 |
|---|---|---|---|---|---|---|
| 1 | 2.72 1 | 3.97 | 8.12 | 4.26 | 8.79 | 18.67 |
| 2 | 2.66 | 5.81 | 3.79 | 1.93 | 8.84 | 41.65 |
| 3 | 2.50 | 3.59 | 2.31 | 1.36 | 3.00 | 4.69 |
| 4 | 0.96 | 1.85 | 0.67 | 10.45 | 0.87 | 0.51 |
| 5 | 0.83 | 1.12 | 1.56 | 1.65 | 1.63 | 1.58 |
| 6 | 1.91 | 1.30 | 1.70 | 1.04 | 1.63 | 2.69 |
| 7 | 2.73 | 4.70 | 6.01 | 3.93 | 4.41 | 4.90 |
| Average | 2.04 | 3.19 | 3.45 | 03.52 | 4.17 | 10.67 |
| Pat. ID | PIK3CA | HER2 | Vimentin | c-Met | Ki-67 | TLR 4 |
| 1 | 5.74 | 10.46 | 6.59 | 8.03 | 3.96 | 9.31 |
| 2 | 14.68 | 21.07 | 10.83 | 3.97 | 4.19 | 37.18 |
| 3 | 6.47 | 4.65 | 1.27 | 2.25 | 1.14 | 3.27 |
| 4 | 0.25 | 0.16 | 0.50 | 0.84 | 2.82 | 0.23 |
| 5 | 1.17 | 0.98 | 2.26 | 1.95 | 1.24 | 1.82 |
| 6 | 1.78 | 2.05 | 1.99 | 4.90 | 1.05 | 1.33 |
| 7 | 2.40 | 15.16 | 1.57 | 7.47 | 4.28 | 5.16 |
| Average | 4.64 | 7.79 | 3.57 | 4.20 | 2.67 | 8.33 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mäurer, M.; Pachmann, K.; Wendt, T.; Schott, D.; Wittig, A. Prospective Monitoring of Circulating Epithelial Tumor Cells (CETC) Reveals Changes in Gene Expression during Adjuvant Radiotherapy of Breast Cancer Patients. Curr. Oncol. 2021, 28, 3507-3524. https://doi.org/10.3390/curroncol28050302
Mäurer M, Pachmann K, Wendt T, Schott D, Wittig A. Prospective Monitoring of Circulating Epithelial Tumor Cells (CETC) Reveals Changes in Gene Expression during Adjuvant Radiotherapy of Breast Cancer Patients. Current Oncology. 2021; 28(5):3507-3524. https://doi.org/10.3390/curroncol28050302
Chicago/Turabian StyleMäurer, Matthias, Katharina Pachmann, Thomas Wendt, Dorothea Schott, and Andrea Wittig. 2021. "Prospective Monitoring of Circulating Epithelial Tumor Cells (CETC) Reveals Changes in Gene Expression during Adjuvant Radiotherapy of Breast Cancer Patients" Current Oncology 28, no. 5: 3507-3524. https://doi.org/10.3390/curroncol28050302
APA StyleMäurer, M., Pachmann, K., Wendt, T., Schott, D., & Wittig, A. (2021). Prospective Monitoring of Circulating Epithelial Tumor Cells (CETC) Reveals Changes in Gene Expression during Adjuvant Radiotherapy of Breast Cancer Patients. Current Oncology, 28(5), 3507-3524. https://doi.org/10.3390/curroncol28050302






