Prenatal Exposure to Traffic-Related Air Pollution and the DNA Methylation in Cord Blood Cells: MOCEH Study
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Assessment of Air-Pollutant Concentration
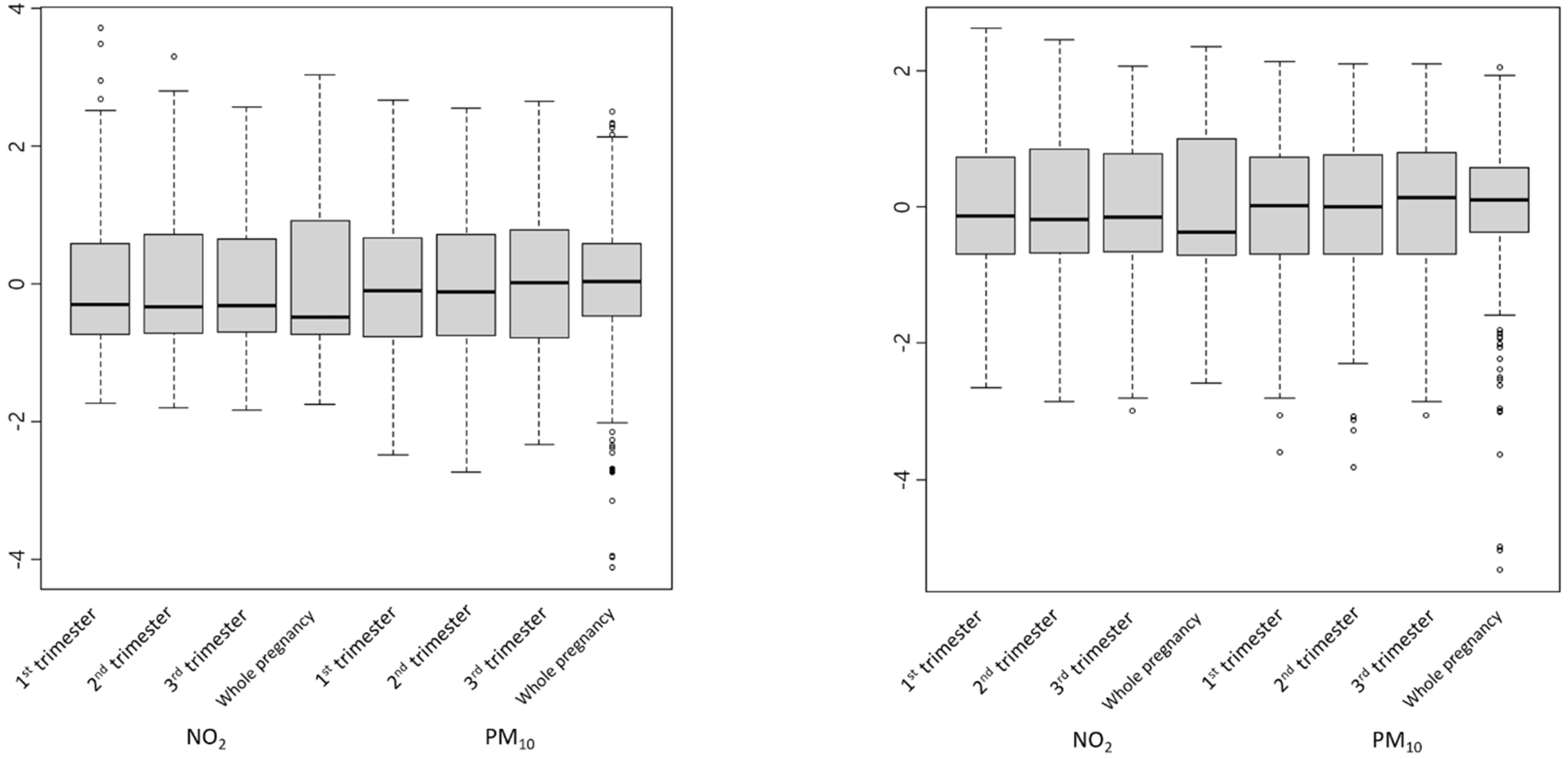
2.3. Data Extraction, Cleaning, and Imputation
2.4. Genotyping and Assessment of the Methylation Levels
2.5. Epigenome-Wide Association Analysis (EWAS)
2.6. Reverse Causation Identification
3. Results
3.1. General Characteristics and Exposure to Ambient Air Pollutants
3.2. Multipollutant Analysis
3.2.1. Whole Pregnancy
3.2.2. The First Trimester of Pregnancy
3.2.3. The Second Trimester of Pregnancy
3.2.4. The Third Trimester of Pregnancy
3.3. Models with a Single Pollutant
3.4. Sensitivity Analysis
3.5. MRC-IEU EWAS Catalog
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cai, Y.; Hansell, A.L.; Granell, R.; Blangiardo, M.; Zottoli, M.; Fecht, D.; Gulliver, J.; Henderson, A.J.; Elliott, P. Prenatal, Early-Life, and Childhood Exposure to Air Pollution and Lung Function: The ALSPAC Cohort. Am. J. Respir. Crit. Care Med. 2020, 202, 112–123. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Hong, Y.C.; Park, H.; Kim, Y.; Ha, M.; Ha, E. Combined effects of multiple prenatal exposure to pollutants on birth weight: The Mothers and Children’s Environmental Health (MOCEH) study. Environ. Res. 2020, 181, 108832. [Google Scholar] [CrossRef] [PubMed]
- Rider, C.F.; Carlsten, C. Air pollution and DNA methylation: Effects of exposure in humans. Clin. Epigenetics 2019, 11, 131. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jirtle, R.L.; Skinner, M.K. Environmental epigenomics and disease susceptibility. Nat. Rev. Genet. 2007, 8, 253–262. [Google Scholar] [CrossRef]
- Gruzieva, O.; Xu, C.J.; Breton, C.V.; Annesi-Maesano, I.; Antó, J.M.; Auffray, C.; Ballereau, S.; Bellander, T.; Bousquet, J.; Bustamante, M.; et al. Epigenome-Wide Meta-Analysis of Methylation in Children Related to Prenatal NO2 Air Pollution Exposure. Environ. Health Perspect. 2017, 125, 104–110. [Google Scholar] [CrossRef] [Green Version]
- Gruzieva, O.; Xu, C.J.; Yousefi, P.; Relton, C.; Merid, S.K.; Breton, C.V.; Gao, L.; Volk, H.E.; Feinberg, J.I.; Ladd-Acosta, C.; et al. Prenatal Particulate Air Pollution and DNA Methylation in Newborns: An Epigenome-Wide Meta-Analysis. Environ. Health Perspect. 2019, 127, 57012. [Google Scholar] [CrossRef]
- Cai, J.; Zhao, Y.; Liu, P.; Xia, B.; Zhu, Q.; Wang, X.; Song, Q.; Kan, H.; Zhang, Y. Exposure to particulate air pollution during early pregnancy is associated with placental DNA methylation. Sci. Total Environ. 2017, 607–608, 1103–1108. [Google Scholar] [CrossRef]
- Rosa, M.J.; Hsu, H.L.; Just, A.C.; Brennan, K.J.; Bloomquist, T.; Kloog, I.; Pantic, I.; Mercado García, A.; Wilson, A.; Coull, B.A.; et al. Association between prenatal particulate air pollution exposure and telomere length in cord blood: Effect modification by fetal sex. Environ. Res. 2019, 172, 495–501. [Google Scholar] [CrossRef]
- Lee, M.K.; Xu, C.J.; Carnes, M.U.; Nichols, C.E.; Ward, J.M.; Kwon, S.O.; Kim, S.Y.; Kim, W.J.; London, S.J. Genome-wide DNA methylation and long-term ambient air pollution exposure in Korean adults. Clin. Epigenetics 2019, 11, 37. [Google Scholar] [CrossRef] [Green Version]
- Kim, B.M.; Ha, M.; Park, H.S.; Lee, B.E.; Kim, Y.J.; Hong, Y.C.; Kim, Y.; Chang, N.; Roh, Y.M.; Kim, B.N.; et al. The Mothers and Children’s Environmental Health (MOCEH) study. Eur. J. Epidemiol. 2009, 24, 573–583. [Google Scholar] [CrossRef] [Green Version]
- Kim, E.; Park, H.; Hong, Y.C.; Ha, M.; Kim, Y.; Kim, B.N.; Kim, Y.; Roh, Y.M.; Lee, B.E.; Ryu, J.M.; et al. Prenatal exposure to PM10 and NO2 and children’s neurodevelopment from birth to 24 months of age: Mothers and Children’s Environmental Health (MOCEH) study. Sci. Total Environ. 2014, 481, 439–445. [Google Scholar] [CrossRef] [PubMed]
- Henderson, S.B.; Beckerman, B.; Jerrett, M.; Brauer, M. Application of land use regression to estimate long-term concentrations of traffic-related nitroen oxides and fine particulate matter. Environ. Sci. Technol. 2007, 41, 2422–2428. [Google Scholar] [CrossRef] [PubMed]
- Stekhovan, D.J.; Bühlmann, P. MissForest-non-parametric missing value imputation for mixed-type data. Bioinformatics 2012, 28, 112–119. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heiss, J.A.; Just, A.C. Identifying mislabeled and contaminated DNA methylation microarray data: An extended quality control toolset with examples from GEO. Clin. Epigenetics 2018, 10, 73. [Google Scholar] [CrossRef] [Green Version]
- Heiss, J.A.; Just, A.C. Improved filtering of DNA methylation microarray data by detection p values and its impact on downstream analyses. Clin. Epigenetics 2019, 11, 15. [Google Scholar] [CrossRef] [Green Version]
- Aryee, M.J.; Jaffe, A.E.; Corrada-Bravo, H.; Ladd-Acosta, C.; Feinberg, A.P.; Hansen, K.D.; Irizarry, R.A. Minfi: A flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 2014, 30, 1363–1369. [Google Scholar] [CrossRef] [Green Version]
- Park, J.; Kim, J.; Kim, E.; Kim, W.J.; Won, S. Prenatal lead exposure and cord blood DNA methylation in the Korean Exposome Study. Environ. Res. 2021, 195, 110767. [Google Scholar] [CrossRef]
- Gervin, K.; Salas, L.A.; Bakulski, K.M.; van Zelm, M.C.; Koestler, D.C.; Wiencke, J.K.; Duijts, L.; Moll, H.A.; Kelsey, K.T.; Kobor, M.S.; et al. Systemic evaluation and validation of reference and library selection methods for deconvolution of cord blood DNA methylation data. Clin. Epigenetics 2019, 11, 125. [Google Scholar] [CrossRef] [Green Version]
- Park, J.; Kwon, S.O.; Kim, S.-H.; Kim, S.J.; Koh, E.J.; Won, S.; Kim, W.J.; Hwang, S.Y. Methylation quantitative trait loci analysis in Korean exposome study. Mol. Cell. Toxicol. 2020, 16, 175–183. [Google Scholar] [CrossRef]
- Chang, C.C.; Chow, C.C.; Tellier, L.C.; Vattikuti, S.; Purcell, S.M.; Lee, J.J. Second-generation PLINK: Rising to the challenge of larger and richer datasets. Gigascience 2015, 4, 7. [Google Scholar] [CrossRef]
- Delaneau, O.; Marchini, J.; Zagury, J.F. A linear complexity phasing method for thousands of genomes. Nat. Methods 2011, 9, 179–181. [Google Scholar] [CrossRef] [PubMed]
- Howie, B.; Fuchsberger, C.; Stephens, M.; Marchini, J.; Abecasis, G.R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 2012, 44, 955–959. [Google Scholar] [CrossRef] [PubMed]
- Howie, B.; Marchini, J.; Stephens, M. Genotype imputation with thousands of genomes. G3 Genes Genomes Genet. 2011, 1, 457–470. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Howie, B.N.; Donnelly, P.; Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009, 5, e1000529. [Google Scholar] [CrossRef] [Green Version]
- Zeileis, C.K.A. Applied Econometrics with R, 1st ed.; Springer: New York, NY, USA, 2008. [Google Scholar]
- Mori, M.; Saito, K.; Ohta, Y. ARHGAP22 localizes at endosomes and regulates actin cytoskeleton. PLoS ONE 2014, 9, e100271. [Google Scholar] [CrossRef] [Green Version]
- Park, K.M.; Kang, E.; Jeon, Y.J.; Kim, N.; Kim, N.S.; Yoo, H.S.; Yeom, Y.I.; Kim, S.J. Identification of Novel Regulators of Apoptosis Using a High-Throughput Cell-based Screen. Mol. Cells 2007, 23, 170–174. [Google Scholar]
- Grundberg, E.; Meduri, E.; Sandling, J.K.; Hedman, A.K.; Keildson, S.; Buil, A.; Busche, S.; Yuan, W.; Nisbet, J.; Sekowska, M.; et al. Global analysis of DNA methylation variation in adipose tissue from twins reveals links to disease-associated variants in distal regulatory elements. Am. J. Hum. Genet. 2013, 93, 876–890. [Google Scholar] [CrossRef] [Green Version]
- Lakhani, S.A.; Masud, A.; Kuida, K.; Porter, G.A., Jr.; Booth, C.J.; Mehal, W.Z.; Inayat, I.; Flavell, R.A. Caspases 3 and 7: Key mediators of mitochondrial events of apoptosis. Science 2006, 311, 847–851. [Google Scholar] [CrossRef] [Green Version]
- Chiu, Y.M.; Hsu, H.L.; Wilson, A.; Coull, B.A.; Pendo, M.P.; Baccarelli, A.; Kloog, I.; Schwartz, J.; Wright, R.O.; Taveras, E.M.; et al. Prenatal particulate air pollution exposure and body composition in urban preschool children: Examining sensitive windows and sex-specific associations. Environ. Res. 2017, 158, 798–805. [Google Scholar] [CrossRef]
- Bougas, N.; Rancière, F.; Beydon, N.; Viola, M.; Perrot, X.; Gabet, S.; Lezmi, G.; Amat, F.; De Blic, J.; Just, J.; et al. Traffic-related Air Pollution, Lung Function, and Host Vulnerability. New Insights from the PARIS Birth Cohort. Ann. Am. Thorac. Soc. 2018, 15, 599–607. [Google Scholar] [CrossRef]
- Chen, Y.; Hodgson, S.; Gulliver, J.; Granell, R.; Henderson, A.J.; Cai, Y.; Hansell, A.L. Trimester effects of source-specific PM(10) on birth weight outcomes in the Avon Longitudinal Study of Parents and Children (ALSPAC). Environ. Health 2021, 20, 4. [Google Scholar] [CrossRef] [PubMed]
- Morales, E.; Garcia-Esteban, R.; de la Cruz, O.A.; Basterrechea, M.; Lertxundi, A.; de Dicastillo, M.D.; Zabaleta, C.; Sunyer, J. Intrauterine and early postnatal exposure to outdoor air pollution and lung function at preschool age. Thorax 2015, 70, 64–73. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghazi, T.; Naidoo, P.; Naidoo, R.N.; Chuturgoon, A.A. Prenatal air pollution exposure and placental DNA methylation changes:Implications on fetal development and future disease susceptibility. Cells 2021, 10, 3025. [Google Scholar] [CrossRef] [PubMed]
- Gross, A.M.; Jaeger, P.A.; Kreisberg, J.F.; Licon, K.; Jepsen, K.L.; Khosroheidari, M.; Morsey, B.M.; Swindells, S.; Shen, H.; Ng, C.T.; et al. Methylome-wide Analysis of Chronic HIV Infection Reveals Five-Year Increase in Biological Age and Epigenetic Targeting of HLA. Mol. Cell 2016, 62, 157–168. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Merid, S.K.; Bustamante, M.; Standl, M.; Sunyer, J.; Heinrich, J.; Lemonnier, N.; Aguilar, D.; Antó, J.M.; Bousquet, J.; Santa-Marina, L.; et al. Integration of gene expression and DNA methylation identifies epigenetically controlled modules related to PM(2.5) exposure. Environ. Int. 2021, 146, 106248. [Google Scholar] [CrossRef]
| Air Pollutants (Unit) | Pregnancy | |||
|---|---|---|---|---|
| Whole Period Mean (min–max) | 1st Trimester Mean (min–max) | 2nd Trimester Mean (min–max) | 3rd Trimester Mean (min–max) | |
| NO2 (ppm) | 0.0260 (0.0113–0.0559) | 0.0249 (0.0098–0.0628) | 0.0261 (0.0098–0.0609) | 0.0264 (0.0091–0.0557) |
| PM10 (ug/m3) | 53.49 (24.96–71.93) | 52.30 (21.47–86.38) | 52.02 (23.09–81.23) | 53.71 (26.03–83.06) |
| Variables | Summary |
|---|---|
| Covariates, N (%) | |
| Child Sex | |
| Male | 180 (50.28%) |
| Female | 178 (49.72%) |
| Maternal smoking, current smokers | 7 (1.96%) |
| Estimated leukocyte composition of the cord blood samples | |
| CD4 T cells (%) | 20.70 (0.449) |
| CD8 T cells (%) | 4.46 (0.173) |
| Natural killer cells (%) | 3.31 (0.141) |
| B cells (%) | 5.18 (0.130) |
| Granulocytes (%) | 55.86 (0.651) |
| Monocytes (%) | 9.34 (0.184) |
| Nucleated red blood cells (%) | 3.08 (0.241) |
| NO2 | PM10 | |||||||
|---|---|---|---|---|---|---|---|---|
| Whole Pregnancy | 1st Trimester of Pregnancy | 2nd Trimester of Pregnancy | 3rd Trimester of Pregnancy | Whole Pregnancy | 1st Trimester of Pregnancy | 2nd Trimester of Pregnancy | 3rd Trimester of Pregnancy | |
| Before transformation | ||||||||
| Skewness | 0.756 | 0.944 | 0.869 | 0.828 | −0.682 | 0.331 | 0.221 | 0.021 |
| Kurtosis | 2.386 | 3.478 | 2.864 | 2.640 | 4.949 | 2.477 | 2.660 | 2.194 |
| After transformation | ||||||||
| Skewness | 0.315 | 0.161 | 0.280 | 0.210 | −1.585 | −0.230 | −0.357 | −0.420 |
| Kurtosis | 2.195 | 2.624 | 2.361 | 2.470 | 8.461 | 2.800 | 3.232 | 2.498 |
| CpG ID | p-Value (All Pollutants) | Log-FC (NO2) | p-Value (NO2) | Log-FC (PM10) | p-Value (PM10) | Gene Annotation | CpG Island Annotation |
|---|---|---|---|---|---|---|---|
| All infants (N = 358) | |||||||
| cg14547404 | 1.78 × 10−11 | −0.0177 | 0.237 | 0.231 | ARHGAP22 TSS1500/body | chr10: 49863620–49864601 N-shore | |
| cg06517429 | 2.37 × 10−8 | 0.0664 | 5.05 × 10−8 | −0.0195 | 0.472 | CASP7 5′ UTR/1st exon | chr10: 115439007–115440196 island |
| cg08906307 | 1.81 × 10−7 | 0.0641 | 5.81 × 10−8 | −0.0440 | 0.0959 | - | chr1: 221064889–221065600 island |
| Male infants (N = 180) | |||||||
| cg19390934 | 2.60 × 10−7 | −0.0577 | 3.86 × 10−8 | 0.0699 | 0.0124 | MTERFD2 body SNED1 body | - |
| Female infants (N = 178) | |||||||
| cg14547404 | 4.36 × 10−10 | −0.0112 | 0.619 | ARHGAP22 TSS1500/body | chr10: 49863620–49864601 N-shore | ||
| cg00670246 | 4.44 × 10−10 | −0.0315 | 0.309 | 0.405 | GRHL1 body | - | |
| cg14561322 | 1.47 × 10−7 | −0.0482 | 1.81 × 10−3 | 0.184 | - | - | |
| cg02737288 | 2.75 × 10−7 | −0.0504 | 3.90 × 10−3 | SGPP1 body | Unknown | ||
| CpG ID | p-Value (All Pollutants) | Log-FC (NO2) | p-Value (NO2) | Log-FC (PM10) | p-Value (PM10) | Gene Annotation | CpG Island Annotation |
|---|---|---|---|---|---|---|---|
| All infants (N = 358) | |||||||
| cg19190403 | 8.38 × 10−8 | −0.0478 | 1.23 × 10−8 | 0.0354 | 4.76 × 10−3 | - | - |
| cg06517429 | 9.37 × 10−8 | 0.0641 | 2.65 × 10−8 | −0.0343 | 0.0456 | CASP7 5′ UTR/1st exon | chr10: 115439007–115440196 island |
| Male infants (N = 180) | |||||||
| cg27535616 | 2.59 × 10−7 | 0.0974 | 3.99 × 10−8 | −0.0660 | 0.0123 | Unknown | Unknown |
| CpG ID | p-Value (All Pollutants) | Log-FC (NO2) | p-Value (NO2) | Log-FC (PM10) | p-Value (PM10) | Gene Annotation | CpG Island Annotation |
|---|---|---|---|---|---|---|---|
| All infants (N = 358) | |||||||
| cg04129282 | 4.37 × 10−9 | 0.0217 | 9.61 × 10−4 | 0.0334 | 2.12 × 10−3 | WDR93 TSS1500 PEX11A body | chr15: 90233530–90234083 island |
| cg06772824 | 4.84 × 10−9 | 0.0278 | 4.29 × 10−6 | 0.0144 | 0.146 | FAM176A TSS200/5′ UTR | chr2: 75787717–75788312 island |
| cg03233931 | 2.85 × 10−8 | 8.09 × 10−3 | 0.166 | LRRC20 TSS200/5′ UTR/ body | chr10: 72141560–72142637 island | ||
| cg06517429 | 3.47 × 10−8 | 0.0622 | 6.83 × 10−8 | −0.0119 | 0.523 | CASP7 5′ UTR/1st exon | chr10: 115439007–115440196 island |
| cg16274061 | 4.50 × 10−8 | 0.0252 | 1.30 × 10−3 | 0.0355 | 6.14 × 10−3 | SAP30L TSS1500 | chr5: 153825417–153826526 N-shore |
| cg23560755 | 5.06 × 10−8 | 0.0135 | 0.123 | SORBS3 body | chr8: 22422534–22423702 N-shelf | ||
| cg00894435 | 5.60 × 10−8 | 0.0337 | 4.76 × 10−4 | 0.0380 | 0.0169 | SV2B TSS1500 | chr15: 91642908–91643702 island |
| Female infants (N = 178) | |||||||
| cg14262371 | 3.54 × 10−8 | 0.0150 | 2.97 × 10−4 | MOV10 5ʹ UTR/1st exon | chr1: 113217475–113218097 N-shore | ||
| CpG ID | p-Value (All Pollutants) | Log-FC (NO2) | p-Value (NO2) | Log-FC (PM10) | p-Value (PM10) | Gene Annotation | CpG Island Annotation |
|---|---|---|---|---|---|---|---|
| All infants (N = 358) | |||||||
| cg06517429 | 2.92 × 10−7 | 0.0616 | 5.46 × 10−8 | −0.0399 | 0.0206 | CASP7 5ʹ UTR/1st exon | chr10: 115439007–115440196 island |
| Female infants (N = 178) | |||||||
| cg20654468 | 2.80 × 10−7 | −0.0398 | 6.87 × 10−3 | 0.123 | 4.58 × 10−8 | LPXN body | chr11: 58345673–58347321 N-shelf |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Park, J.; Kim, W.J.; Kim, J.; Jeong, C.-Y.; Park, H.; Hong, Y.-C.; Ha, M.; Kim, Y.; Won, S.; Ha, E. Prenatal Exposure to Traffic-Related Air Pollution and the DNA Methylation in Cord Blood Cells: MOCEH Study. Int. J. Environ. Res. Public Health 2022, 19, 3292. https://doi.org/10.3390/ijerph19063292
Park J, Kim WJ, Kim J, Jeong C-Y, Park H, Hong Y-C, Ha M, Kim Y, Won S, Ha E. Prenatal Exposure to Traffic-Related Air Pollution and the DNA Methylation in Cord Blood Cells: MOCEH Study. International Journal of Environmental Research and Public Health. 2022; 19(6):3292. https://doi.org/10.3390/ijerph19063292
Chicago/Turabian StylePark, Jaehyun, Woo Jin Kim, Jeeyoung Kim, Chan-Yeong Jeong, Hyesook Park, Yun-Chul Hong, Mina Ha, Yangho Kim, Sungho Won, and Eunhee Ha. 2022. "Prenatal Exposure to Traffic-Related Air Pollution and the DNA Methylation in Cord Blood Cells: MOCEH Study" International Journal of Environmental Research and Public Health 19, no. 6: 3292. https://doi.org/10.3390/ijerph19063292
APA StylePark, J., Kim, W. J., Kim, J., Jeong, C.-Y., Park, H., Hong, Y.-C., Ha, M., Kim, Y., Won, S., & Ha, E. (2022). Prenatal Exposure to Traffic-Related Air Pollution and the DNA Methylation in Cord Blood Cells: MOCEH Study. International Journal of Environmental Research and Public Health, 19(6), 3292. https://doi.org/10.3390/ijerph19063292






