Limited Role of Rhamnolipids on Cadmium Resistance for an Endogenous-Secretion Bacterium
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
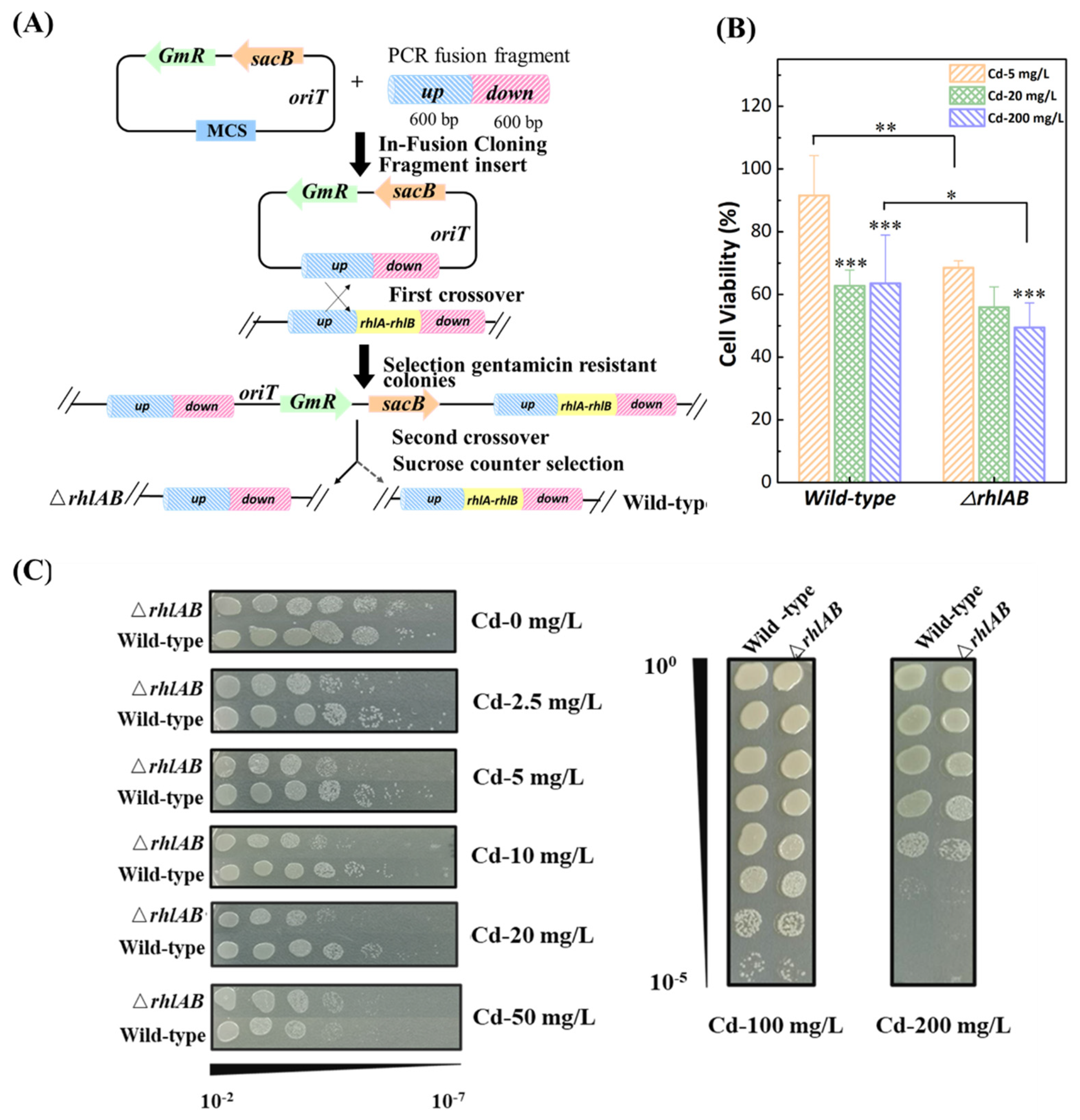
2.2. Plasmid Construction and Gene Deletion Mutant
2.3. Detection Analysis of Rhamnolipids
2.4. Characteristics Analysis of Rhamnolipids
2.5. Distribution of Cd on Bacterial Media
2.6. Bacterial Surface Properties Analysis
2.7. Bacterial Viability Assays
2.8. Bacterial Biochemical Characteristic
2.9. Statistical Analysis
3. Results and Discussion
3.1. Effect of Cd2+ on Cells Viability
3.2. Distribution of Cd2+ in Bacteria
3.3. Effect of Cd2+ on Bacterial Characteristics
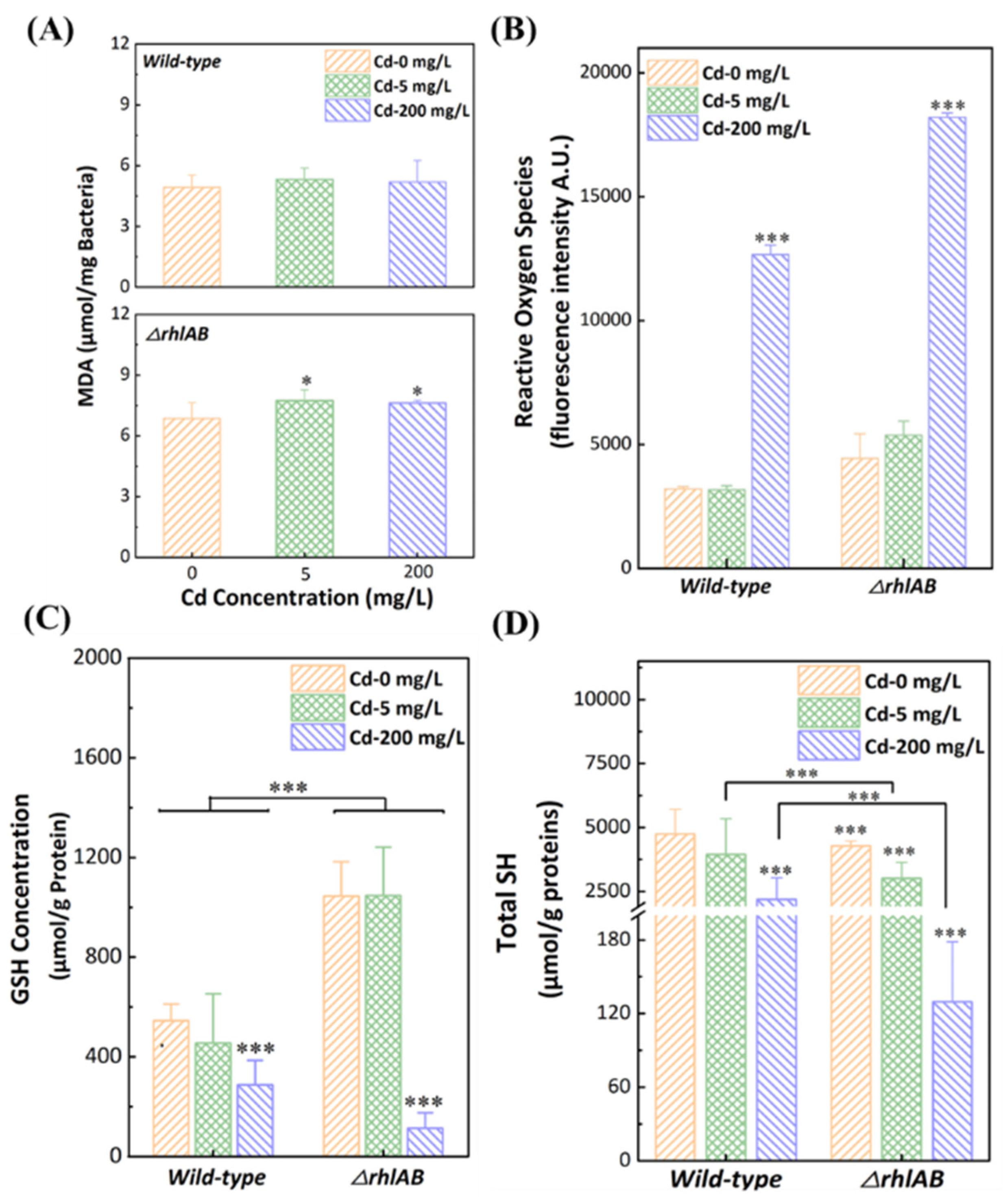
3.4. Response of Antioxidant Systems to Cd2+ Stress
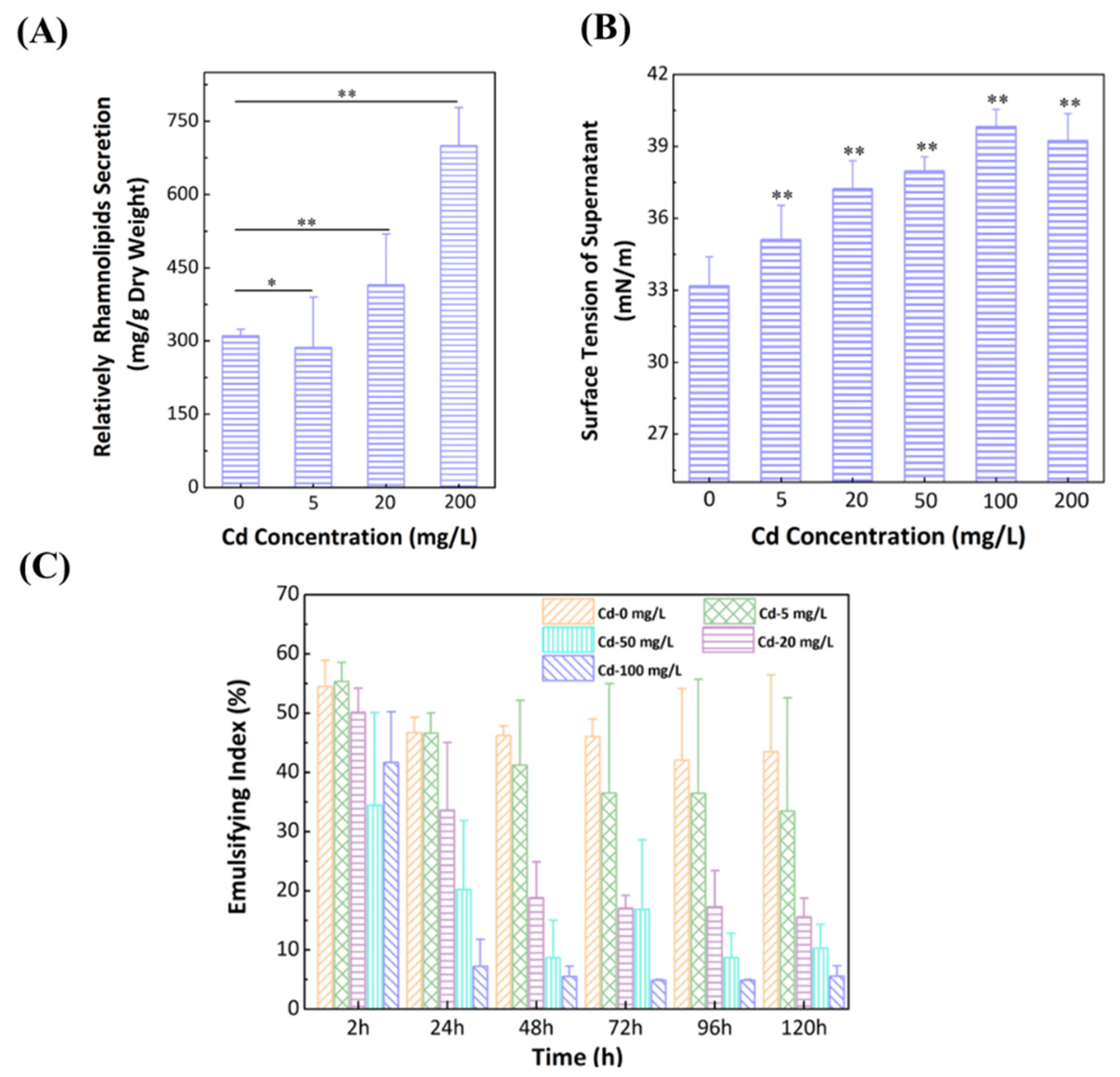
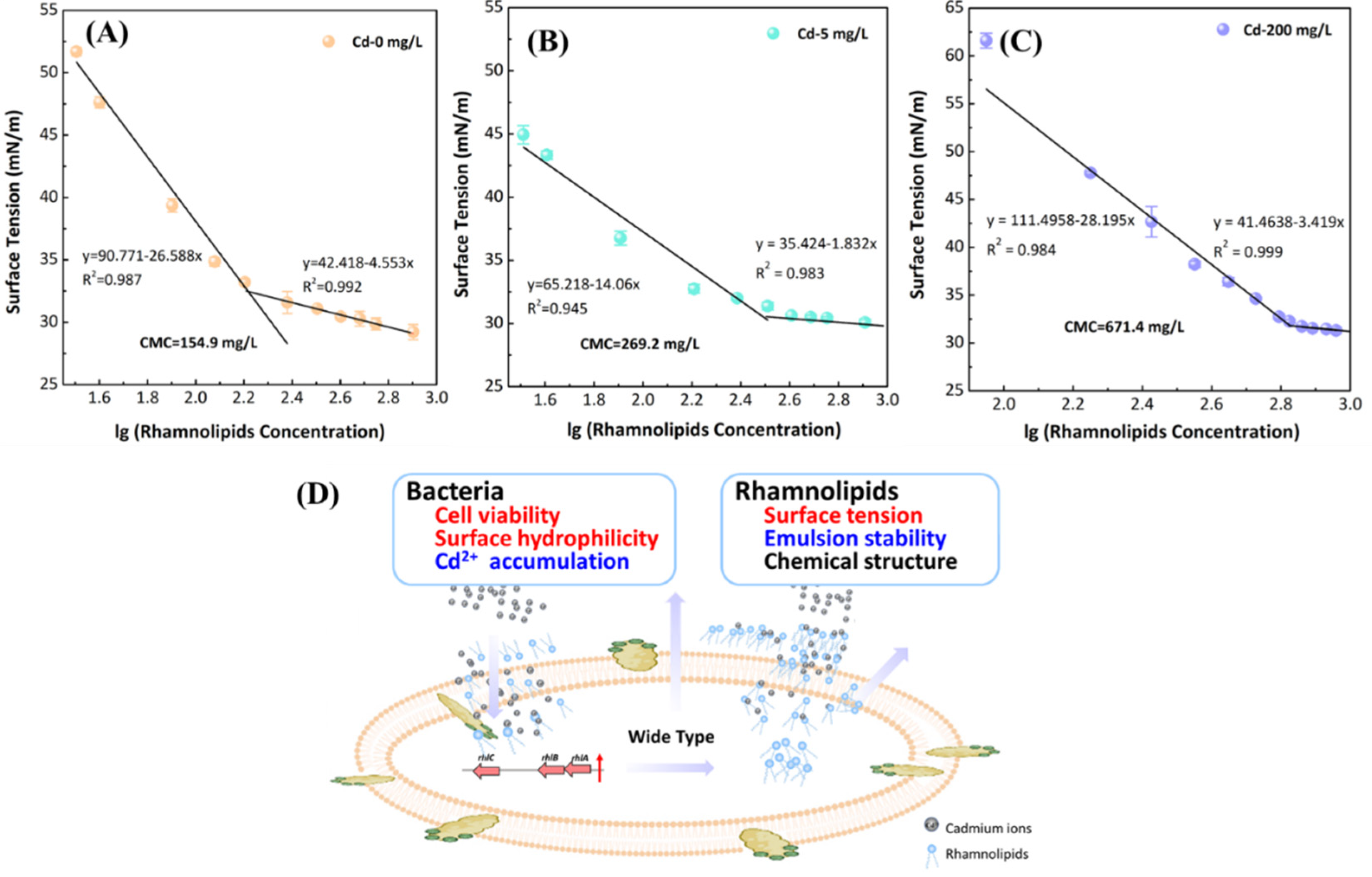
3.5. Effect of Cd2+ on Secretion of Rhamnolipids
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Módenes, A.N.; de Abreu Pietrobelli, J.M.T.; dos Santos, G.H.F.; Borba, C.E.; da Silva Sá Ravagnani, M.A.; Espinoza-Quiñones, F.R. Multi-component mathematical model based on mass transfer coefficients for prediction of the Zn and Cd ions biosorption data by E. densa in a continuous system. J. Environ. Chem. Eng. 2018, 6, 5141–5149. [Google Scholar] [CrossRef]
- Hamid, Y.; Tang, L.; Sohail, M.I.; Cao, X.; Hussain, B.; Aziz, M.Z.; Usman, M.; He, Z.-l.; Yang, X. An explanation of soil amendments to reduce cadmium phytoavailability and transfer to food chain. Sci. Total Environ. 2019, 660, 80–96. [Google Scholar] [CrossRef] [PubMed]
- da Silva Oliveira, A.; Bocio, A.; Trevilato, T.M.B.; Takayanagui, A.M.M.; Domingo, J.L.; Segura-Muñoz, S.I. Heavy metals in untreated/treated urban effluent and sludge from a biological wastewater treatment plant. Environ. Sci. Pollut. Res. Int. 2007, 14, 483. [Google Scholar] [CrossRef] [PubMed]
- Tu, C.; Wei, J.; Guan, F.; Liu, Y.; Sun, Y.H.; Luo, Y.M. Biochar and bacteria inoculated biochar enhanced Cd and Cu immobilization and enzymatic activity in a polluted soil. Environ. Int. 2020, 137, 105576. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.-R.; Schoenung, J.M. Human health and ecological toxicity potentials due to heavy metal content in waste electronic devices with flat panel displays. J. Hazard. Mater. 2010, 177, 251–259. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Liu, H.; Liu, F.; Ju, X.; Dinis, F.; Yu, E.; Yu, Z. Source identification and superposition effect of heavy metals (HMs) in agricultural soils at a high geological background area of Karst: A case study in a typical watershed. Int. J. Environ. Res. Public Health 2022, 19, 11374. [Google Scholar] [CrossRef]
- Liu, X.; Hu, Q.; Yang, J.; Huang, S.; Wei, T.; Chen, W.; He, Y.; Wang, D.; Liu, Z.; Wang, K.; et al. Selective cadmium regulation mediated by a cooperative binding mechanism in CadR. Proc. Natl. Acad. Sci. USA 2019, 116, 20398–20403. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Y.; Ma, J.; Chen, F.; Yu, R.; Hu, G.; Zhang, S. Remediation of Soil polluted with Cd in a postmining area using thiourea-modified biochar. Int. J. Environ. Res. Public Health 2020, 17, 7654. [Google Scholar] [CrossRef]
- Mishra, S.; Lin, Z.; Pang, S.; Zhang, Y.; Bhatt, P.; Chen, S. Biosurfactant is a powerful tool for the bioremediation of heavy metals from contaminated soils. J. Hazard. Mater. 2021, 418, 126253. [Google Scholar] [CrossRef]
- Agnello, A.C.; Bagard, M.; van Hullebusch, E.D.; Esposito, G.; Huguenot, D. Comparative bioremediation of heavy metals and petroleum hydrocarbons co-contaminated soil by natural attenuation, phytoremediation, bioaugmentation and bioaugmentation-assisted phytoremediation. Sci. Total Environ. 2016, 563–564, 693–703. [Google Scholar] [CrossRef]
- Yang, K.; Zhu, L.; Zhao, Y.; Wei, Z.; Chen, X.; Yao, C.; Meng, Q.; Zhao, R. A novel method for removing heavy metals from composting system: The combination of functional bacteria and adsorbent materials. Bioresour. Technol. 2019, 293, 122095. [Google Scholar] [CrossRef] [PubMed]
- Asad, S.A.; Farooq, M.; Afzal, A.; West, H. Integrated phytobial heavy metal remediation strategies for a sustainable clean environment—A review. Chemosphere 2019, 217, 925–941. [Google Scholar] [CrossRef] [PubMed]
- Arora, P.K. Bacilli-mediated degradation of xenobiotic compounds and heavy metals. Front. Bioeng. Biotechnol. 2020, 8, 570307. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S.; Lin, Z.; Pang, S.; Zhang, W.; Bhatt, P.; Chen, S. Recent advanced technologies for the characterization of xenobiotic-degrading microorganisms and microbial communities. Front. Bioeng. Biotechnol. 2021, 9, 632059. [Google Scholar] [CrossRef]
- Ghaith, E.-S.I.; Rizvi, S.; Namasivayam, C.; Rahman, P. Removal of Cd++ from contaminated water using bio-surfactant modified ground grass as a bio-sorbent. In Proceedings of the 2019 Advances in Science and Engineering Technology International Conferences (ASET), Dubai, United Arab Emirates, 26 March–10 April 2019; IEEE: Piscataway, NJ, USA, 2019; pp. 1–7. [Google Scholar]
- Qin, H.; Hu, T.; Zhai, Y.; Lu, N.; Aliyeva, J. The improved methods of heavy metals removal by biosorbents: A review. Environ. Pollut. 2020, 258, 113777. [Google Scholar] [CrossRef]
- Wen, J.; McLaughlin, M.J.; Stacey, S.P.; Kirby, J.K. Aseptic hydroponics to assess rhamnolipid-Cd and rhamnolipid-Zn bioavailability for sunflower (Helianthus annuus): A phytoextraction mechanism study. Environ. Sci. Pollut. Res. 2016, 23, 21327–21335. [Google Scholar] [CrossRef]
- da Rocha Junior, R.B.; Meira, H.M.; Almeida, D.G.; Rufino, R.D.; Luna, J.M.; Santos, V.A.; Sarubbo, L.A. Application of a low-cost biosurfactant in heavy metal remediation processes. Biodegradation 2019, 30, 215–233. [Google Scholar] [CrossRef]
- Yang, Z.; Zhang, Z.; Chai, L.; Wang, Y.; Liu, Y.; Xiao, R. Bioleaching remediation of heavy metal-contaminated soils using Burkholderia sp. Z-90. J. Hazard. Mater. 2016, 301, 145–152. [Google Scholar] [CrossRef]
- Liu, G.; Zhong, H.; Yang, X.; Liu, Y.; Shao, B.; Liu, Z. Advances in applications of rhamnolipids biosurfactant in environmental remediation: A review. Biotechnol. Bioeng. 2018, 115, 796–814. [Google Scholar] [CrossRef]
- Sachdev, D.P.; Cameotra, S.S. Biosurfactants in agriculture. Appl. Microbiol. Biotechnol. 2013, 97, 1005–1016. [Google Scholar] [CrossRef] [Green Version]
- Bondarenko, O.; Rahman, P.K.S.M.; Rahman, T.J.; Kahru, A.; Ivask, A. Effects of rhamnolipids from Pseudomonas aeruginosa DS10-129 on iuminescent bacteria: Toxicity and modulation of cadmium bioavailability. Microb. Ecol. 2010, 59, 588–600. [Google Scholar] [CrossRef] [Green Version]
- Juwarkar, A.A.; Dubey, K.V.; Nair, A.; Singh, S.K. Bioremediation of multi-metal contaminated soil using biosurfactant—A novel approach. Indian J. Microbiol. 2008, 48, 142–146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, F.; Yuan, M.; Lei, L.; Li, C.; Xu, X. Enhanced production of mono-rhamnolipid in Pseudomonas aeruginosa and application potential in agriculture and petroleum industry. Bioresour. Technol. 2021, 323, 124605. [Google Scholar] [CrossRef] [PubMed]
- Zhao, F.; Han, S.; Zhang, Y. Comparative studies on the structural composition, surface/interface activity and application potential of rhamnolipids produced by Pseudomonas aeruginosa using hydrophobic or hydrophilic substrates. Bioresour. Technol. 2020, 295, 122269. [Google Scholar] [CrossRef] [PubMed]
- Eslami, P.; Hajfarajollah, H.; Bazsefidpar, S. Recent advancements in the production of rhamnolipid biosurfactants by Pseudomonas aeruginosa. RSC Adv. 2020, 10, 34014–34032. [Google Scholar] [CrossRef]
- Chakraborty, J.; Das, S. Characterization and cadmium-resistant gene expression of biofilm-forming marine bacterium Pseudomonas aeruginosa JP-11. Environ. Sci. Pollut. Res. 2014, 21, 14188–14201. [Google Scholar] [CrossRef]
- Brocklehurst, K.; Megit, S.; Morby, A.P. Characterisation of CadR from Pseudomonas aeruginosa: A Cd (II)-responsive MerR homologue. Biochem. Biophys. Res. Commun. 2003, 308, 234–239. [Google Scholar] [CrossRef]
- Abbas, S.; Rafatullah, M.; Hossain, K.; Ismail, N.; Tajarudin, H.; Khalil, H.A. A review on mechanism and future perspectives of cadmium-resistant bacteria. Int. J. Environ. Sci. Technol. 2018, 15, 243–262. [Google Scholar] [CrossRef]
- Singh, R.; Bishnoi, N.R.; Kirrolia, A. Evaluation of Pseudomonas aeruginosa an innovative bioremediation tool in multi metals ions from simulated system using multi response methodology. Bioresour. Technol. 2013, 138, 222–234. [Google Scholar] [CrossRef]
- Jain, R.M.; Mody, K.; Mishra, A.; Jha, B. Isolation and structural characterization of biosurfactant produced by an alkaliphilic bacterium Cronobacter sakazakii isolated from oil contaminated wastewater. Carbohydr. Polym. 2012, 87, 2320–2326. [Google Scholar] [CrossRef]
- Neilson, J.W.; Zhang, L.; Veres-Schalnat, T.A.; Chandler, K.B.; Neilson, C.H.; Crispin, J.D.; Pemberton, J.E.; Maier, R.M. Cadmium effects on transcriptional expression of rhlB/rhlC genes and congener distribution of monorhamnolipid and dirhamnolipid in Pseudomonas aeruginosa IGB83. Appl. Microbiol. Biotechnol. 2010, 88, 953–963. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Wu, Q.; Hua, Y.; Chen, J.; Zhang, H.; Wang, H. Potential applications of biosurfactant rhamnolipids in agriculture and biomedicine. Appl. Microbiol. Biotechnol. 2017, 101, 8309–8319. [Google Scholar] [CrossRef] [PubMed]
- Hmelo, L.R.; Borlee, B.R.; Almblad, H.; Love, M.E.; Randall, T.E.; Tseng, B.S.; Lin, C.; Irie, Y.; Storek, K.M.; Yang, J.J. Precision-engineering the Pseudomonas aeruginosa genome with two-step allelic exchange. Nat. Protoc. 2015, 10, 1820–1841. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, F.; Shi, R.; Ma, F.; Han, S.; Zhang, Y. Oxygen effects on rhamnolipids production by Pseudomonas aeruginosa. Microb. Cell Factories 2018, 17, 39. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Setoodeh, P.; Jahanmiri, A.; Eslamloueyan, R.; Niazi, A.; Ayatollahi, S.S.; Aram, F.; Mahmoodi, M.; Hortamani, A. Statistical screening of medium components for recombinant production of Pseudomonas aeruginosa ATCC 9027 rhamnolipids by nonpathogenic cell factory Pseudomonas putida KT2440. Mol. Biotechnol. 2014, 56, 175–191. [Google Scholar] [CrossRef]
- Somani, B.L.; Khanade, J.; Sinha, R. A Modified Anthrone Sulfuric-Acid Method for the Determination of Fructose in the Presence of Certain Proteins. Anal. Biochem. 1987, 167, 327–330. [Google Scholar] [CrossRef]
- Benincasa, M.; Abalos, A.; Oliveira, I.; Manresa, A. Chemical structure, surface properties and biological activities of the biosurfactant produced by Pseudomonas aeruginosa LBI from soapstock. Antonie Van Leeuwenhoek 2004, 85, 1–8. [Google Scholar] [CrossRef]
- Subbiahdoss, G.; Reimhult, E. Biofilm formation at oil-water interfaces is not a simple function of bacterial hydrophobicity. Colloids Surf. B Biointerfaces 2020, 194, 111163. [Google Scholar] [CrossRef]
- Matthews, L.A.; Simmons, L.A. The Bacillus subtilis PriA Winged Helix Domain Is Critical for Surviving DNA Damage. J. Bacteriol. 2022, 204, e00539-21. [Google Scholar] [CrossRef]
- Dolezalova, E.; Lukes, P. Membrane damage and active but nonculturable state in liquid cultures of Escherichia coli treated with an atmospheric pressure plasma jet. Bioelectrochemistry 2015, 103, 7–14. [Google Scholar] [CrossRef]
- Tan, H.; Champion, J.T.; Artiola, J.F.; Brusseau, M.L.; Miller, R.M. Complexation of Cadmium by a Rhamnolipid Biosurfactant. Env. Sci. Technol. 1994, 28, 2402–2406. [Google Scholar] [CrossRef]
- Mohanty, S.; Mukherji, S. Surfactant aided biodegradation of NAPLs by Burkholderia multivorans: Comparison between Triton X-100 and rhamnolipid JBR-515. Colloids Surf. B Biointerfaces 2013, 102, 644–652. [Google Scholar] [CrossRef] [PubMed]
- He, T.-F.; Zhang, Z.-H.; Zeng, X.-A.; Wang, L.-H.; Brennan, C.S. Determination of membrane disruption and genomic DNA binding of cinnamaldehyde to Escherichia coli by use of microbiological and spectroscopic techniques. J. Photochem. Photobiol. B Biol. 2018, 178, 623–630. [Google Scholar] [CrossRef] [PubMed]
- Yun, O.; Liu, Z.-W.; Zeng, X.-A.; Han, Z. Salmonella typhimurium resistance on pulsed electric fields associated with membrane fluidity and gene regulation. Innov. Food Sci. Emerg. Technol. 2016, 36, 252–259. [Google Scholar] [CrossRef]
- Hamida, R.S.; Ali, M.A.; Goda, D.A.; Khalil, M.I.; Al-Zaban, M.I. Novel biogenic silver nanoparticle-induced reactive oxygen species inhibit the biofilm formation and virulence activities of methicillin-resistant Staphylococcus aureus (MRSA) Strain. Front. Bioeng. Biotechnol. 2020, 8, 433. [Google Scholar] [CrossRef]
- Simão, F.; Matté, A.; Matté, C.; Soares, F.M.S.; Wyse, A.T.S.; Netto, C.A.; Salbego, C.G. Resveratrol prevents oxidative stress and inhibition of Na+K+-ATPase activity induced by transient global cerebral ischemia in rats. J. Nutr. Biochem. 2011, 22, 921–928. [Google Scholar] [CrossRef]
- Wang, X.; Fan, L.; Cheng, L.; Sun, Y.; Wang, X.; Zhong, X.; Shi, Q.; Gong, F.; Yang, Y.; Ma, Y.; et al. Biodegradable nickel disulfide nanozymes with GSH-depleting function for high-efficiency photothermal-catalytic antibacterial therapy. iScience 2020, 23, 101281. [Google Scholar] [CrossRef] [PubMed]
- Benhalima, L.; Amri, S.; Bensouilah, M.; Ouzrout, R. Heavy metal resistance and metallothionein induction in bacteria isolated from Seybouse river, Algeria. Appl. Ecol. Env. Res. 2020, 18, 1721–1737. [Google Scholar] [CrossRef]
- Lee, S.W.; Glickmann, E.; Cooksey, D.A. Chromosomal locus for cadmium resistance in Pseudomonas putida consisting of a cadmium-transporting ATPase and a MerR family response regulator. Appl. Env. Microb. 2001, 67, 1437–1444. [Google Scholar] [CrossRef] [Green Version]
- Hassan, M.E.T.; van der Lelie, D.; Springael, D.; Romling, U.; Ahmed, N.; Mergeay, M. Identification of a gene cluster, czr, involved in cadmium and zinc resistance in Pseudomonas aeruginosa. Gene 1999, 238, 417–425. [Google Scholar] [CrossRef]
- Rondeau, P.; Sers, S.; Jhurry, D.; Cadet, F. Sugar interaction with metals in aqueous solution: Indirect determination from infrared and direct determination from nuclear magnetic resonance spectroscopy. Appl. Spectrosc. 2003, 57, 466–472. [Google Scholar] [CrossRef]
- Tang, S.; Yin, H.; Zhou, S.; Chen, S.; Peng, H.; Liu, Z.; Dang, Z. Simultaneous Cr(VI) removal and 2,2′,4,4′-tetrabromodiphenyl ether (BDE-47) biodegradation by Pseudomonas aeruginosa in liquid medium. Chemosphere 2016, 150, 24–32. [Google Scholar] [CrossRef]
- Zeng, Z.T.; Liu, Y.; Zhong, H.; Xiao, R.; Zeng, G.M.; Liu, Z.F.; Cheng, M.; Lai, C.; Zhang, C.; Liu, G.S.; et al. Mechanisms for rhamnolipids-mediated biodegradation of hydrophobic organic compounds. Sci. Total Env. 2018, 634, 1–11. [Google Scholar] [CrossRef]
- Nitschke, M.; Costa, S.G.V.A.O.; Contiero, J. Rhamnolipid surfactants: An update on the general aspects of these remarkable biomolecules. Biotechnol. Prog. 2005, 21, 1593–1600. [Google Scholar] [CrossRef]
- Helvacı, Ş.Ş.; Peker, S.; Özdemir, G. Effect of electrolytes on the surface behavior of rhamnolipids R1 and R2. Colloids Surf. B Biointerfaces 2004, 35, 225–233. [Google Scholar] [CrossRef]
- Kłosowska-Chomiczewska, I.E.; Mędrzycka, K.; Hallmann, E.; Karpenko, E.; Pokynbroda, T.; Macierzanka, A.; Jungnickel, C. Rhamnolipid CMC prediction. J. Colloid Interface Sci. 2017, 488, 10–19. [Google Scholar] [CrossRef]
- Costa, S.G.V.A.O.; Nitschke, M.; Lépine, F.; Déziel, E.; Contiero, J. Structure, properties and applications of rhamnolipids produced by Pseudomonas aeruginosa L2-1 from cassava wastewater. Process Biochem. 2010, 45, 1511–1516. [Google Scholar] [CrossRef]
- Patel, K.; Patel, M. Improving bioremediation process of petroleum wastewater using biosurfactants producing Stenotrophomonas sp. S1VKR-26 and assessment of phytotoxicity. Bioresour. Technol. 2020, 315, 123861. [Google Scholar] [CrossRef]
- Datta, P.; Tiwari, P.; Pandey, L.M. Isolation and characterization of biosurfactant producing and oil degrading Bacillus subtilis MG495086 from formation water of Assam oil reservoir and its suitability for enhanced oil recovery. Bioresour. Technol. 2018, 270, 439–448. [Google Scholar] [CrossRef]
- Zhu, L.; Qi, H.-y.; Kong, Y.; Yu, Y.-w.; Xu, X.-y. Component analysis of extracellular polymeric substances (EPS) during aerobic sludge granulation using FTIR and 3D-EEM technologies. Bioresour. Technol. 2012, 124, 455–459. [Google Scholar] [CrossRef]
- Bazire, A.; Dheilly, A.; Diab, F.; Morin, D.; Jebbar, M.; Haras, D.; Dufour, A. Osmotic stress and phosphate limitation alter production of cell-to-cell signal molecules and rhamnolipid biosurfactant by Pseudomonas aeruginosa. Fems. Microbiol. Lett. 2005, 253, 125–131. [Google Scholar] [CrossRef] [PubMed] [Green Version]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xing, S.; Yan, Z.; Song, C.; Tian, H.; Wang, S. Limited Role of Rhamnolipids on Cadmium Resistance for an Endogenous-Secretion Bacterium. Int. J. Environ. Res. Public Health 2022, 19, 12555. https://doi.org/10.3390/ijerph191912555
Xing S, Yan Z, Song C, Tian H, Wang S. Limited Role of Rhamnolipids on Cadmium Resistance for an Endogenous-Secretion Bacterium. International Journal of Environmental Research and Public Health. 2022; 19(19):12555. https://doi.org/10.3390/ijerph191912555
Chicago/Turabian StyleXing, Sufang, Zhen Yan, Chao Song, Huifang Tian, and Shuguang Wang. 2022. "Limited Role of Rhamnolipids on Cadmium Resistance for an Endogenous-Secretion Bacterium" International Journal of Environmental Research and Public Health 19, no. 19: 12555. https://doi.org/10.3390/ijerph191912555
APA StyleXing, S., Yan, Z., Song, C., Tian, H., & Wang, S. (2022). Limited Role of Rhamnolipids on Cadmium Resistance for an Endogenous-Secretion Bacterium. International Journal of Environmental Research and Public Health, 19(19), 12555. https://doi.org/10.3390/ijerph191912555