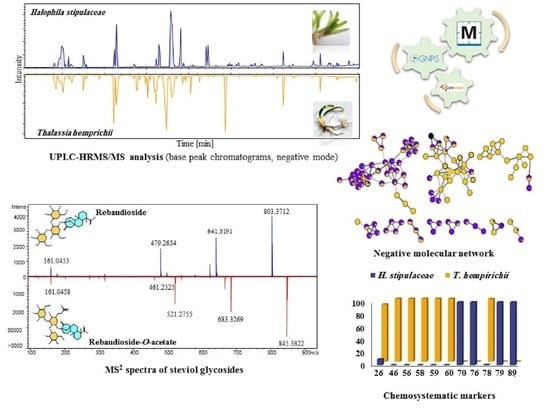
Molecular Networking Leveraging the Secondary Metabolomes Space of Halophila stipulaceae (Forsk.) Aschers. and Thalassia hemprichii (Ehrenb. ex Solms) Asch. in Tandem with Their Chemosystematics and Antidiabetic Potentials
Abstract
1. Introduction
2. Results and Discussion
2.1. Phytochemical Analysis
2.1.1. Acid Hydrolysis
2.1.2. High-Performance Liquid Chromatography (HPLC) and Ultra-High Performance Liquid Chromatography-High-Resolution Mass Spectrometry (UPLC-HRMS/MS) Analyses
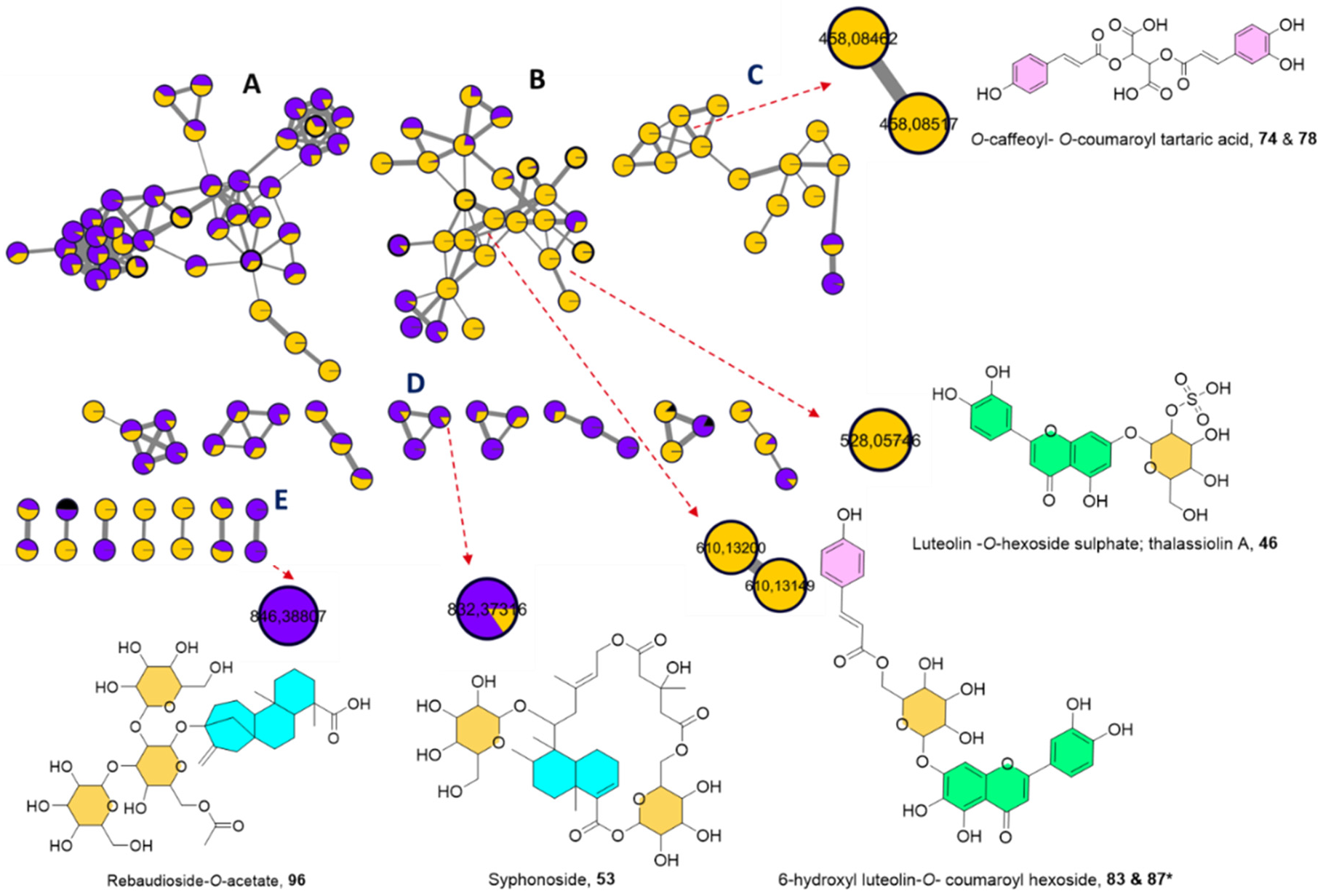
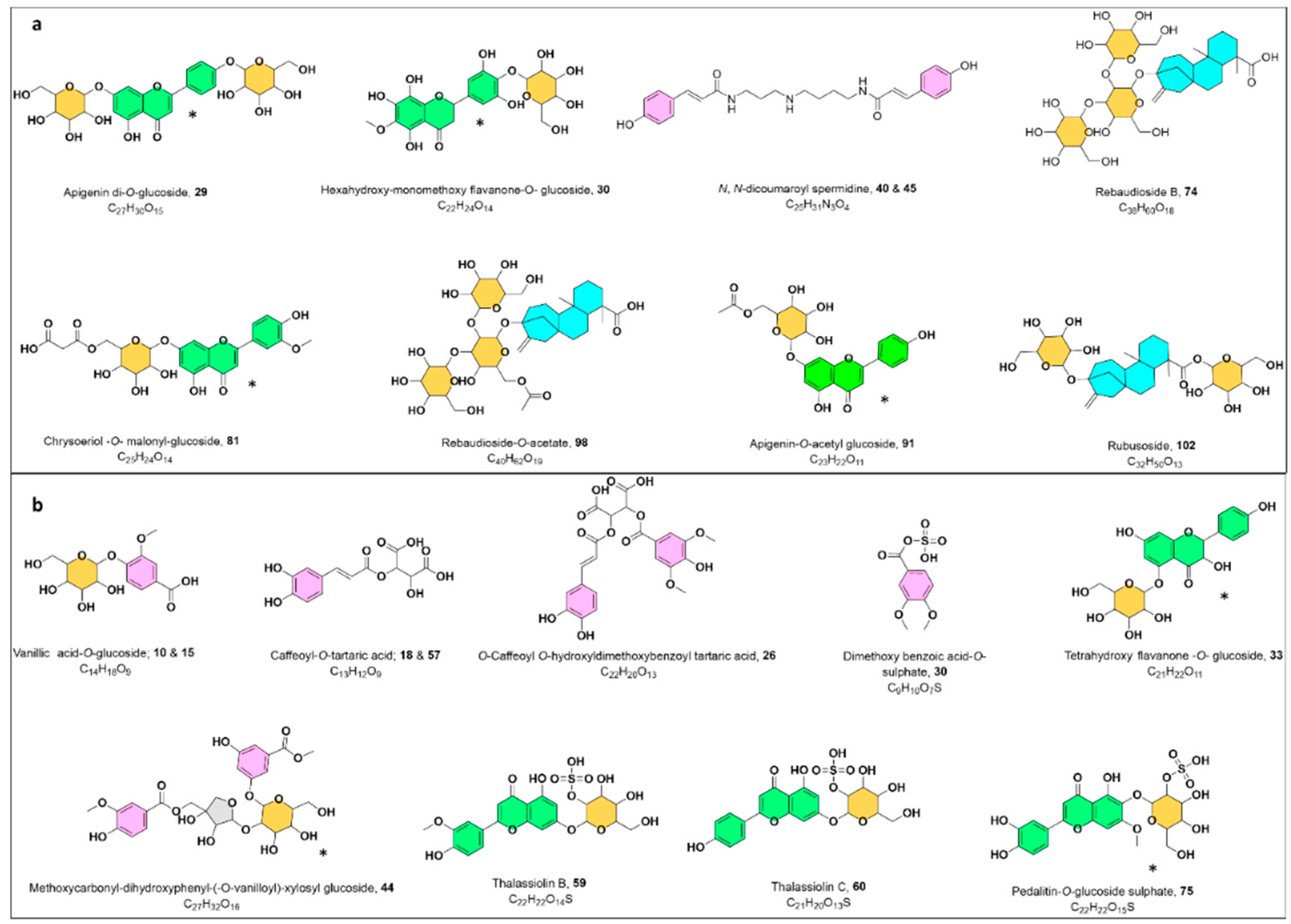
2.1.3. UPLC-HRMS/MS Metabolite Annotation Aided with Molecular Networking
Phenolic Acids
Benzoic Acids
Cinnamic Acids and Their Derivatives
Flavonoids
Sulphated Flavonoids
Acylated Flavonoids
Flavonoid Aglycones
Diterpenoids
Steviol Glycosides
Macrocyclic Glycoterpenoids
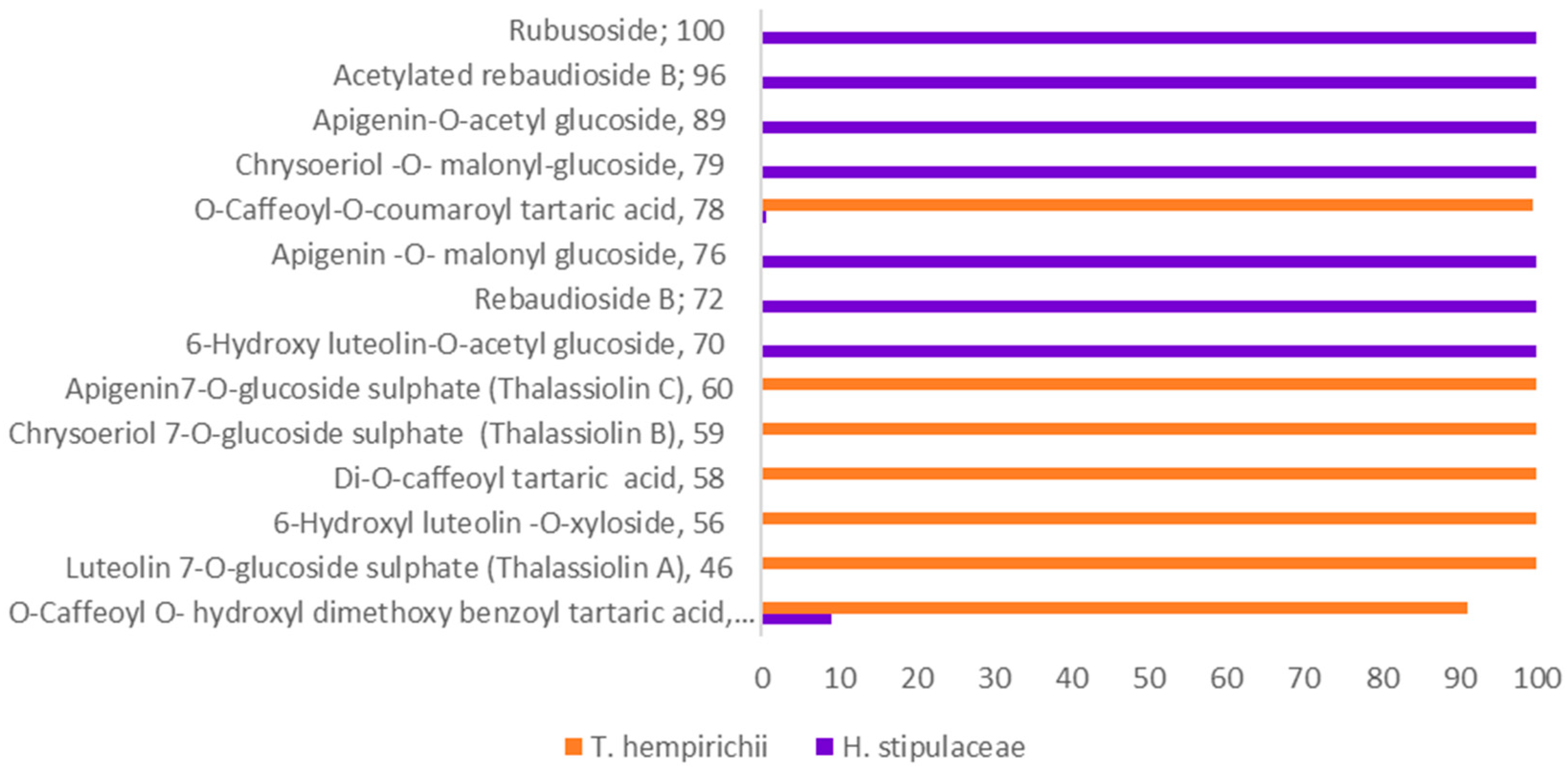
2.2. Chemosystematic Significance
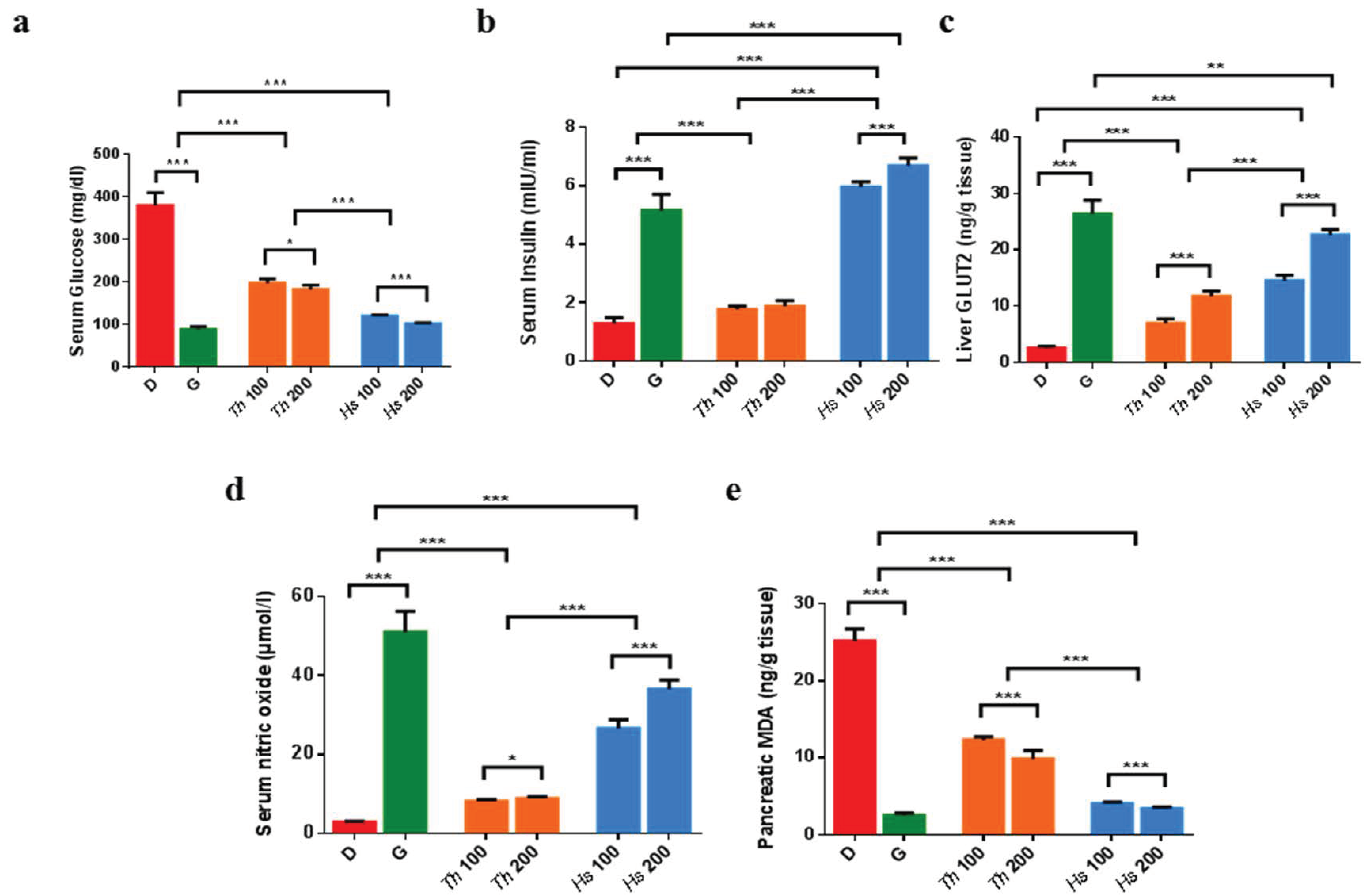
2.3. In Vitro and In Vivo Antidiabetic Assays
3. Materials and Methods
3.1. Plant Material
3.2. Chemicals and Reagents
3.3. Preparation of the Crude Extracts
3.4. Phytochemical Analysis
3.4.1. Acid Hydrolysis
3.4.2. Sample Preparation, HPLC Profiling and MS Analyses
Sample Preparation
HPLC Profiling
UPLC–HRMS-MS Analysis
Data Analysis
Molecular Networking and Compounds Dereplication
3.5. Biological Assays
3.5.1. In Vitro Anti-Diabetic Assays (Enzymes Inhibitory Assays)
3.5.2. In Vivo Antidiabetic Study
Experimental Animals
Induction of Diabetes Mellitus
Experimental Design
Preparation of Serum and Tissue Samples
Biochemical Analyses
3.6. Statistical Analyses
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Ethics Approval
References
- El Shaffai, A. Field Guide to Seagrasses of the Red Sea; Total Foundation, Courbevoie, France, International Union for the Conservation of Nature: Gland, Switzerland, 2011. [Google Scholar]
- Den Hartog, C.; Kuo, J. Taxonomy and biogeography of seagrasses. In Seagrasses: Biology, Ecologyand Conservation; Springer: Berlin, Germany, 2007; pp. 1–23. [Google Scholar]
- Les, D.H.; Tippery, N.P. In time and with water... the systematics of alismatid monocotyledons. Early Events Monocot Evol. 2013, 83, 118–164. [Google Scholar]
- Christenhusz, M.J.; Byng, J.W. The number of known plants species in the world and its annual increase. Phytotaxa 2016, 261, 201–217. [Google Scholar] [CrossRef]
- Mabberley, D. Mabberley’s Plant-Book, 3rd ed.; Cambridge University Press: Cambridge, UK, 2008. [Google Scholar]
- Boulos, L. Flora of Egypt Checklist; Revised Annotated Edition; Al-Hadara Publishing: Cairo, Egypt, 2009; pp. 198–201. [Google Scholar]
- Boulos, L. Flora of Egypt; Al-Hadara Publishing: Cairo, Egypt, 2005; Volume 4. [Google Scholar]
- El-Hady, H.; Hamed, E.; Shehata, A.N. Molecular identification, antimicrobial and antioxidant activities of the tropical seagrass Halophila stipulacea grown in El-Bardawil lake, Egypt. Aust. J. Basic Appl. Sci. 2012, 6, 474–481. [Google Scholar]
- Gavagnin, M.; Carbone, M.; Amodeo, P.; Mollo, E.; Vitale, R.M.; Roussis, V.; Cimino, G. Structure and absolute stereochemistry of syphonoside, a unique macrocyclic glycoterpenoid from marine organisms. J. Org. Chem. 2007, 72, 5625–5630. [Google Scholar] [CrossRef] [PubMed]
- Meng, Y.; Krzysiak, A.J.; Durako, M.J.; Kunzelman, J.I.; Wright, J.L. Flavones and flavone glycosides from Halophila johnsonii. Phytochemistry 2008, 69, 2603–2608. [Google Scholar] [CrossRef] [PubMed]
- Hawas, U.W. A new 8-hydroxy flavone O-xyloside sulfate and antibacterial activity from the Egyptian seagrass Thalassia hemprichii. Chem. Nat. Compd. 2014, 50, 629–632. [Google Scholar] [CrossRef]
- Jafriati, J.; Hatta, M.; Yuniar, N.; Junita, A.R.; Dwiyanti, R.; Sabir, M.; Primaguna, M.R. Thalassia hemprichii Seagrass Extract as antimicrobial and antioxidant potential on human: A mini review of the benefits of seagrass. J. Biol. Sci. 2019, 19, 363–371. [Google Scholar] [CrossRef]
- Kannan, R.R.R.; Arumugam, R.; Anantharaman, P. Antibacterial potential of three seagrasses against human pathogens. Asian Pac. J. Trop. Med. 2010, 3, 890–893. [Google Scholar] [CrossRef]
- Kannan Rengasamy, R.R.; Rajasekaran, A.; Micheline, G.-D.; Perumal, A. Antioxidant activity of seagrasses of the Mandapam coast, India. Pharm. Biol. 2012, 50, 182–187. [Google Scholar] [CrossRef]
- El-Ghffar, E.A.A.; Hegazi, N.M.; Saad, H.H.; Soliman, M.M.; El-Raey, M.A.; Shehata, S.M.; Barakat, A.; Yasri, A.; Sobeh, M. HPLC-ESI-MS/MS analysis of beet (Beta vulgaris) leaves and its beneficial properties in type 1 diabetic rats. Biomed. Pharmacother. 2019, 120, 109541. [Google Scholar] [CrossRef] [PubMed]
- Ezzat, S.M.; Bishbishy, M.H.E.; Habtemariam, S.; Salehi, B.; Sharifi-Rad, M.; Martins, N.; Sharifi-Rad, J. Looking at marine-derived bioactive molecules as upcoming anti-diabetic agents: A special emphasis on PTP1B inhibitors. Molecules 2018, 23, 3334. [Google Scholar] [CrossRef]
- Alarcon-Aguilara, F.J.; Roman-Ramos, R.; Perez-Gutierrez, S.; Aguilar-Contreras, A.; Contreras-Weber, C.C.; Flores-Saenz, J.L. Study of the anti-hyperglycemic effect of plants used as antidiabetics. J. Ethnopharmacol. 1998, 61, 101–110. [Google Scholar] [CrossRef]
- Kannan, R.R.R.; Arumugam, R.; Iyapparaj, P.; Thangaradjou, T.; Anantharaman, P. In vitro antibacterial, cytotoxicity and haemolytic activities and phytochemical analysis of seagrasses from the Gulf of Mannar, South India. Food Chem. 2013, 136, 1484–1489. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Carver, J.J.; Phelan, V.V.; Sanchez, L.M.; Garg, N.; Peng, Y.; Nguyen, D.D.; Watrous, J.; Kapono, C.A.; Luzzatto-Knaan, T.; et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 2016, 34, 828–837. [Google Scholar] [CrossRef] [PubMed]
- Böcker, S.; Dührkop, K. Fragmentation trees reloaded. J. Cheminform. 2016, 8, 5. [Google Scholar] [CrossRef] [PubMed]
- Dührkop, K.; Shen, H.; Meusel, M.; Rousu, J.; Böcker, S. Searching molecular structure databases with tandem mass spectra using CSI: FingerID. Proc. Natl. Acad. Sci. USA 2015, 112, 12580–12585. [Google Scholar] [CrossRef] [PubMed]
- Zapata, O.; McMillan, C. Phenolic acids in seagrasses. Aquat. Bot. 1979, 7, 307–317. [Google Scholar] [CrossRef]
- Qi, S.; Huang, L.; He, F.; Zhang, S.; Dong, J. Phytochemical and chemotaxonomic investigation of seagrass Thalassia hemprichii (Ehrenb.) Aschers (Hydrocharitaceae). Biochem. Syst. Ecol. 2012, 43, 128–131. [Google Scholar] [CrossRef]
- Ishii, T.; Okino, T.; Suzuki, M.; Machiguchi, Y. Tichocarpols A and B, Two novel phenylpropanoids with feeding-deterrent activity from the red alga Tichocarpus crinitus. J. Nat. Prod. 2004, 67, 1764–1766. [Google Scholar] [CrossRef]
- Regalado, E.L.; Menendez, R.; Valdés, O.; Morales, R.A.; Laguna, A.; Thomas, O.P.; Hernandez, Y.; Nogueiras, C.; Kijjoa, A. Phytochemical analysis and antioxidant capacity of BM-21, a bioactive extract rich in polyphenolic metabolites from the sea grass Thalassia testudinum. Nat. Prod. Commun. 2012, 7, 1934578X1200700117. [Google Scholar] [CrossRef]
- Bel Mabrouk, S.; Reis, M.; Sousa, M.L.; Ribeiro, T.; Almeida, J.R.; Pereira, S.; Antunes, J.; Rosa, F.; Vasconcelos, V.; Achour, L. The marine seagrass Halophila stipulacea as a source of bioactive metabolites against obesity and biofouling. Mar. Drugs 2020, 18, 88. [Google Scholar] [CrossRef] [PubMed]
- Zidorn, C. Secondary metabolites of seagrasses (Alismatales and Potamogetonales; Alismatidae): Chemical diversity, bioactivity, and ecological function. Phytochemistry 2016, 124, 5–28. [Google Scholar] [CrossRef] [PubMed]
- Nuissier, G.; Rezzonico, B.; Grignon-Dubois, M. Chicoric acid from Syringodium filiforme. Food Chem. 2010, 120, 783–788. [Google Scholar] [CrossRef]
- Grignon-Dubois, M.; Rezzonico, B. The economic potential of beach-cast seagrass—Cymodocea nodosa: A promising renewable source of chicoric acid. Bot. Mar. 2013, 56, 303–311. [Google Scholar] [CrossRef]
- Farid, M.; Marzouk, M.; Hussein, S.; Elkhateeb, A.; Abdel-Hameed, E. Comparative study of Posidonia oceanica L.: LC/ESI/MS analysis, cytotoxic activity and chemosystematic significance. J. Mater. Environ. Sci. 2018, 9, 1676–1682. [Google Scholar]
- Subhashini, P.; Dilipan, E.; Thangaradjou, T.; Papenbrock, J. Bioactive natural products from marine angiosperms: Abundance and functions. Nat. Prod. Bioprospect. 2013, 3, 129–136. [Google Scholar] [CrossRef]
- Rowley, D.C.; Hansen, M.S.; Rhodes, D.; Sotriffer, C.A.; Ni, H.; McCammon, J.A.; Bushman, F.D.; Fenical, W. Thalassiolins A–C: New marine-derived inhibitors of HIV cDNA integrase. Bioorg. Med. Chem. 2002, 10, 3619–3625. [Google Scholar] [CrossRef]
- Hawas, U.W.; Abou El-Kassem, L.T. Thalassiolin D: A new flavone O-glucoside Sulphate from the seagrass Thalassia hemprichii. Nat. Prod. Res. 2017, 31, 2369–2374. [Google Scholar] [CrossRef] [PubMed]
- Bitam, F.; Ciavatta, M.L.; Carbone, M.; Manzo, E.; Mollo, E.; Gavagnin, M. Chemical analysis of flavonoid constituents of the seagrass Halophila stipulacea: First finding of malonylated derivatives in marine phanerogams. Biochem. Syst. Ecol. 2010, 38, 686–690. [Google Scholar] [CrossRef]
- Abbas Momtazi-Borojeni, A.; Esmaeili, S.-A.; Abdollahi, E.; Sahebkar, A. A review on the pharmacology and toxicology of steviol glycosides extracted from Stevia rebaudiana. Curr. Pharm. Des. 2017, 23, 1616–1622. [Google Scholar] [CrossRef]
- Pól, J.; Hohnová, B.; Hyötyläinen, T. Characterisation of Stevia rebaudiana by comprehensive two-dimensional liquid chromatography time-of-flight mass spectrometry. J. Chromatogr. A 2007, 1150, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Molina-Calle, M.; de Medina, V.S.; de la Torre, M.D.; Priego-Capote, F.; de Castro, M.L. Development and application of a quantitative method based on LC–QqQ MS/MS for determination of steviol glycosides in Stevia leaves. Talanta 2016, 154, 263–269. [Google Scholar] [CrossRef]
- Nakai, T. Ordines, Familiae, Tribi, Genera, Sectiones, Species, Varietates, Formae et Combinationes Novae a Prof. Nakai-Takenosin adhuc ut novis edita: Appendix: Quaestiones Characterium Naturalium Plantarum vel Extractus ex Praelectionibus pro Aluminis Botanicis Universitatis Imperialis Tokyoensis per Annos 1926–1941, 1943; Alexander Doweld.
- Kimura, Y. Système et phylogénie des monocotylédones. Notul. Syst. 1956, 15, 137–159. [Google Scholar]
- Les, D.H.; Cleland, M.A.; Waycott, M. Phylogenetic studies in Alismatidae, II: Evolution of marine angiosperms (seagrasses) and hydrophily. Syst. Bot. 1997, 22, 443–463. [Google Scholar] [CrossRef]
- den Hartog, C.d. Structure, function, and classification in seagrass communities. In Seagrass Ecosystems a Scientific Perspective; McRoy, C.P., Helfferich, C., Eds.; Marcel Dekker, Inc.: New York, NY, USA, 1973; pp. 89–121. [Google Scholar]
- Qi, S.-H.; Zhang, S.; Qian, P.-Y.; Wang, B.-G. Antifeedant, antibacterial, and antilarval compounds from the South China Sea seagrass Enhalus acoroides. Bot. Mar. 2008, 51, 441–447. [Google Scholar] [CrossRef]
- Erhard, D.; Pohnert, G.; Gross, E.M. Chemical defense in Elodea nuttallii reduces feeding and growth of aquatic herbivorous Lepidoptera. J. Chem. Ecol. 2007, 33, 1646–1661. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.-h.; Tian, C.-r.; Gao, C.-y.; Wang, X.-y.; Yang, X.; Chen, Y.-x.; Liu, Z.-z. Phenolic profile, antioxidant and enzyme inhibitory activities of Ottelia acuminata, an endemic plant from southwestern China. Ind. Crops Prod. 2019, 138, 111423. [Google Scholar] [CrossRef]
- Spínola, V.; Castilho, P.C. Evaluation of Asteraceae herbal extracts in the management of diabetes and obesity. Contribution of caffeoylquinic acids on the inhibition of digestive enzymes activity and formation of advanced glycation end-products (in vitro). Phytochemistry 2017, 143, 29–35. [Google Scholar] [CrossRef]
- Vani, M.; Vasavi, T.; Uma, M.D.P. Evaluation of in vitro antidiabetic activity of methanolic extract of seagrass Halophila beccarii. Evaluation 2018, 11, 150–153. [Google Scholar]
- Widiyanto, A.; Anwar, E.; Nurhayati, T. In vitro assay of alpha-glucosidase inhibitor activities of three seagrasses from Banten Bay, Indonesia. Pharmacogn. J. 2018, 10, 907–910. [Google Scholar] [CrossRef]
- Kawser Hossain, M.; Abdal Dayem, A.; Han, J.; Yin, Y.; Kim, K.; Kumar Saha, S.; Yang, G.-M.; Choi, H.Y.; Cho, S.-G. Molecular mechanisms of the anti-obesity and anti-diabetic properties of flavonoids. Int. J. Mol. Sci. 2016, 17, 569. [Google Scholar] [CrossRef]
- Adisakwattana, S. Cinnamic acid and its derivatives: Mechanisms for prevention and management of diabetes and its complications. Nutrients 2017, 9, 163. [Google Scholar] [CrossRef] [PubMed]
- Myint, K.Z.; Chen, J.-m.; Zhou, Z.-y.; Xia, Y.-m.; Lin, J.; Zhang, J. Structural dependence of antidiabetic effect of steviol glycosides and their metabolites on streptozotocin-induced diabetic mice. J. Sci. Food Agric. 2020, 100, 3841–3849. [Google Scholar] [CrossRef] [PubMed]
- Villanueva-Penacarrillo, M.L.; Puente, J.; Redondo, A.; Clemente, F.; Valverde, I. Effect of GLP-1 treatment on GLUT2 and GLUT4 expression in type 1 and type 2 rat diabetic models. Endocrine 2001, 15, 241–248. [Google Scholar] [CrossRef]
- Anker, C.C.B.; Rafiq, S.; Jeppesen, P.B. Effect of steviol glycosides on human health with emphasis on type 2 diabetic biomarkers: A systematic review and meta-analysis of randomized controlled trials. Nutrients 2019, 11, 1965. [Google Scholar] [CrossRef]
- Rathinam, A.; Pari, L. Myrtenal ameliorates hyperglycemia by enhancing GLUT2 through Akt in the skeletal muscle and liver of diabetic rats. Chem.-Biol. Interact. 2016, 256, 161–166. [Google Scholar] [CrossRef] [PubMed]
- Yonamine, C.Y.; Pinheiro-Machado, E.; Michalani, M.L.; Freitas, H.S.; Okamoto, M.M.; Correa-Giannella, M.L.; Machado, U.F. Resveratrol improves glycemic control in insulin-treated diabetic rats: Participation of the hepatic territory. Nutr. Metab. 2016, 13, 44. [Google Scholar] [CrossRef] [PubMed]
- Marghani, B.; Ateya, A.; Saleh, R.; Eltaysh, R. Assessing of antidiabetic and ameliorative effect of lupin seed aqueous extract on hyperglycemia, hyperlipidemia and effect on pdx1, Nkx6.1, Insulin-1, GLUT-2 and glucokinase genes expression in streptozotocin-induced diabetic rats. J. Food Nutr. Res. 2019, 7, 333–341. [Google Scholar] [CrossRef]
- Chidambaram, K.; John, C.; Das, S.; Venkatesan, K.; Siddalingam, R. Modulation of glucose transporter proteins by polyphenolic extract of Ichnocarpus frutescens (L.) W. T. Aiton in experimental type 2 diabetic rats. Indian J. Exp. Biol. 2020, 58, 172–180. [Google Scholar]
- Tatsch, E.; Bochi, G.V.; Piva, S.J.; De Carvalho, J.A.; Kober, H.; Torbitz, V.D.; Duarte, T.; Signor, C.; Coelho, A.C.; Duarte, M.M.; et al. Association between DNA strand breakage and oxidative, inflammatory and endothelial biomarkers in type 2 diabetes. Mutat. Res. 2012, 732, 16–20. [Google Scholar] [CrossRef]
- Kim, F.; Pham, M.; Maloney, E.; Rizzo, N.O.; Morton, G.J.; Wisse, B.E.; Kirk, E.A.; Chait, A.; Schwartz, M.W. Vascular inflammation, insulin resistance, and reduced nitric oxide production precede the onset of peripheral insulin resistance. Arterioscler. Thromb. Vasc. Biol. 2008, 28, 1982–1988. [Google Scholar] [CrossRef] [PubMed]
- Righi, N.; Boumerfeg, S.; Fernandes, P.A.; Deghima, A.; Baali, F.; Coelho, E.; Cardoso, S.M.; Coimbra, M.A.; Baghiani, A. Thymus algeriensis Bioss & Reut: Relationship of phenolic compounds composition with in vitro/in vivo antioxidant and antibacterial activity. Food Res. Int. 2020, 136, 109500. [Google Scholar]
- Eid, S.; Sas, K.M.; Abcouwer, S.F.; Feldman, E.L.; Gardner, T.W.; Pennathur, S.; Fort, P.E. New insights into the mechanisms of diabetic complications: Role of lipids and lipid metabolism. Diabetologia 2019, 62, 1539–1549. [Google Scholar] [CrossRef]
- Mabry, T.J.; Markham, K.; Thomas, M. The ultraviolet spectra of flavones and flavonols. In The Systematic Identification of Flavonoids; Springer: Berlin, Germany, 1970; pp. 41–164. [Google Scholar]
- Hegazi, N.M.; Radwan, R.A.; Ali, S.M.; Saad, H.H. Molecular networking aided metabolomic profiling of beet leaves using three extraction solvents and in relation to its anti-obesity effects. J. Adv. Res. 2020, 24, 545–555. [Google Scholar] [CrossRef] [PubMed]
- Garg, N.; Kapono, C.A.; Lim, Y.W.; Koyama, N.; Vermeij, M.J.A.; Conrad, D.; Rohwer, F.; Dorrestein, P.C. Mass spectral similarity for untargeted metabolomics data analysis of complex mixtures. Int. J. Mass Spectrom. 2015, 377, 719–727. [Google Scholar] [CrossRef] [PubMed]
- Olmo-García, L.; Wendt, K.; Kessler, N.; Bajoub, A.; Fernández-Gutiérrez, A.; Baessmann, C.; Carrasco-Pancorbo, A. Exploring the capability of LC-MS and GC-MS multi-class methods to discriminate virgin olive oils from different geographical indications and to identify potential origin markers. Eur. J. Lipid Sci. Technol. 2019, 121, 1800336. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Zani, C.L.; Carroll, A.R. Database for rapid dereplication of known natural products using data from MS and fast NMR experiments. J. Nat. Prod. 2017, 80, 1758–1766. [Google Scholar] [CrossRef] [PubMed]
- Miller, G.L. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 1959, 31, 426–428. [Google Scholar] [CrossRef]
- Sancheti, S.; Seo, S. Chaenomeles sinensis: A potent α-and β-glucosidase inhibitor. Am. J. Pharmacol. Toxicol. 2009, 4, 8–11. [Google Scholar] [CrossRef]
- Conforti, F.; Perri, V.; Menichini, F.; Marrelli, M.; Uzunov, D.; Statti, G.A.; Menichini, F. Wild Mediterranean dietary plants as inhibitors of pancreatic lipase. Phytother. Res. 2012, 26, 600–604. [Google Scholar] [CrossRef]
- Chiocchio, I.; Mandrone, M.; Sanna, C.; Maxia, A.; Tacchini, M.; Poli, F. Screening of a hundred plant extracts as tyrosinase and elastase inhibitors, two enzymatic targets of cosmetic interest. Ind. Crops Prod. 2018, 122, 498–505. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hegazi, N.M.; Saad, H.H.; Marzouk, M.M.; Abdel Rahman, M.F.; El Bishbishy, M.H.; Zayed, A.; Ulber, R.; Ezzat, S.M. Molecular Networking Leveraging the Secondary Metabolomes Space of Halophila stipulaceae (Forsk.) Aschers. and Thalassia hemprichii (Ehrenb. ex Solms) Asch. in Tandem with Their Chemosystematics and Antidiabetic Potentials. Mar. Drugs 2021, 19, 279. https://doi.org/10.3390/md19050279
Hegazi NM, Saad HH, Marzouk MM, Abdel Rahman MF, El Bishbishy MH, Zayed A, Ulber R, Ezzat SM. Molecular Networking Leveraging the Secondary Metabolomes Space of Halophila stipulaceae (Forsk.) Aschers. and Thalassia hemprichii (Ehrenb. ex Solms) Asch. in Tandem with Their Chemosystematics and Antidiabetic Potentials. Marine Drugs. 2021; 19(5):279. https://doi.org/10.3390/md19050279
Chicago/Turabian StyleHegazi, Nesrine M., Hamada H. Saad, Mona M. Marzouk, Mohamed F. Abdel Rahman, Mahitab H. El Bishbishy, Ahmed Zayed, Roland Ulber, and Shahira M. Ezzat. 2021. "Molecular Networking Leveraging the Secondary Metabolomes Space of Halophila stipulaceae (Forsk.) Aschers. and Thalassia hemprichii (Ehrenb. ex Solms) Asch. in Tandem with Their Chemosystematics and Antidiabetic Potentials" Marine Drugs 19, no. 5: 279. https://doi.org/10.3390/md19050279
APA StyleHegazi, N. M., Saad, H. H., Marzouk, M. M., Abdel Rahman, M. F., El Bishbishy, M. H., Zayed, A., Ulber, R., & Ezzat, S. M. (2021). Molecular Networking Leveraging the Secondary Metabolomes Space of Halophila stipulaceae (Forsk.) Aschers. and Thalassia hemprichii (Ehrenb. ex Solms) Asch. in Tandem with Their Chemosystematics and Antidiabetic Potentials. Marine Drugs, 19(5), 279. https://doi.org/10.3390/md19050279