Induction of Phlorotannins and Gene Expression in the Brown Macroalga Fucus vesiculosus in Response to the Herbivore Littorina littorea
Abstract
1. Introduction
2. Results
2.1. Phenol Contents
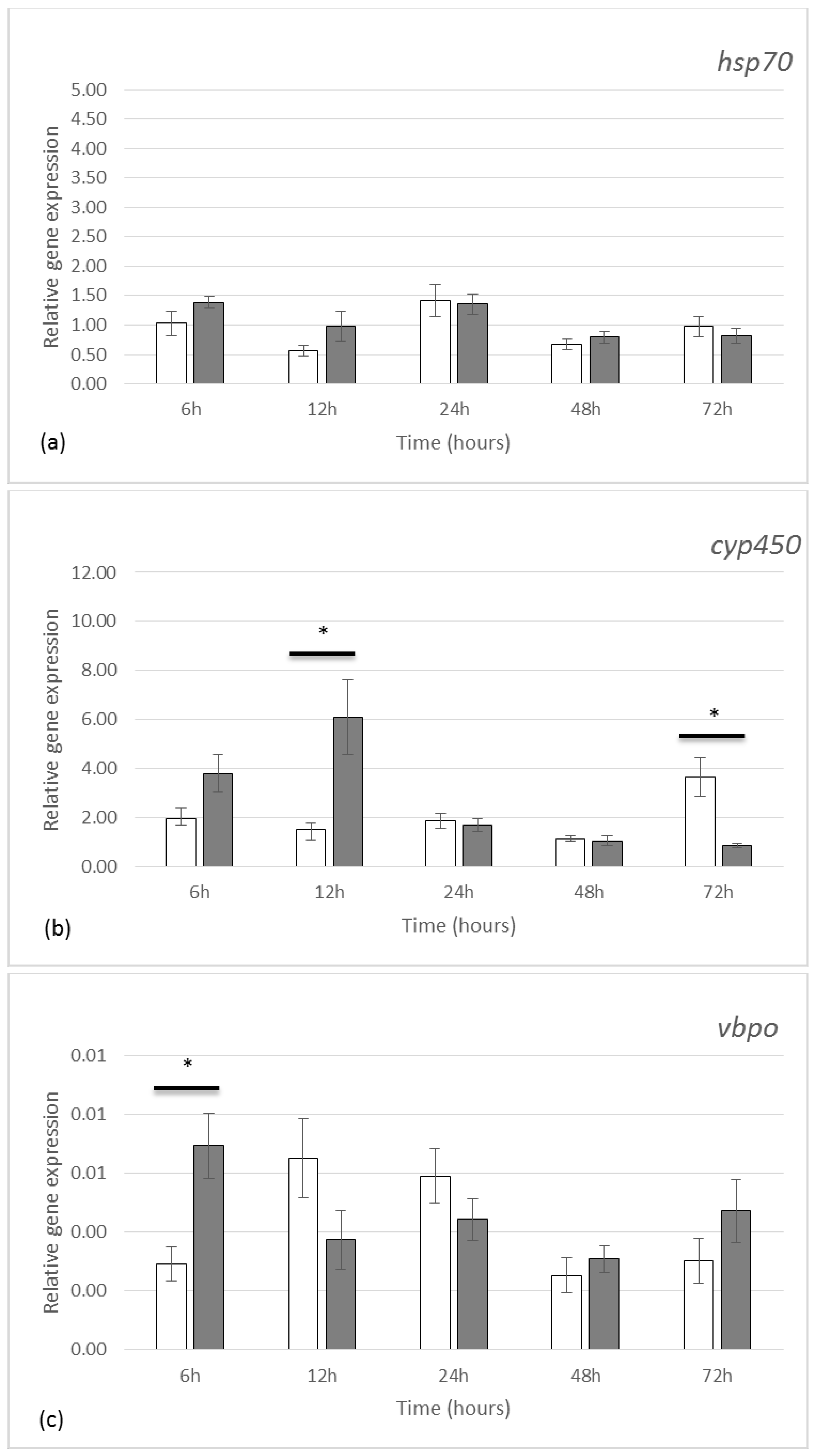
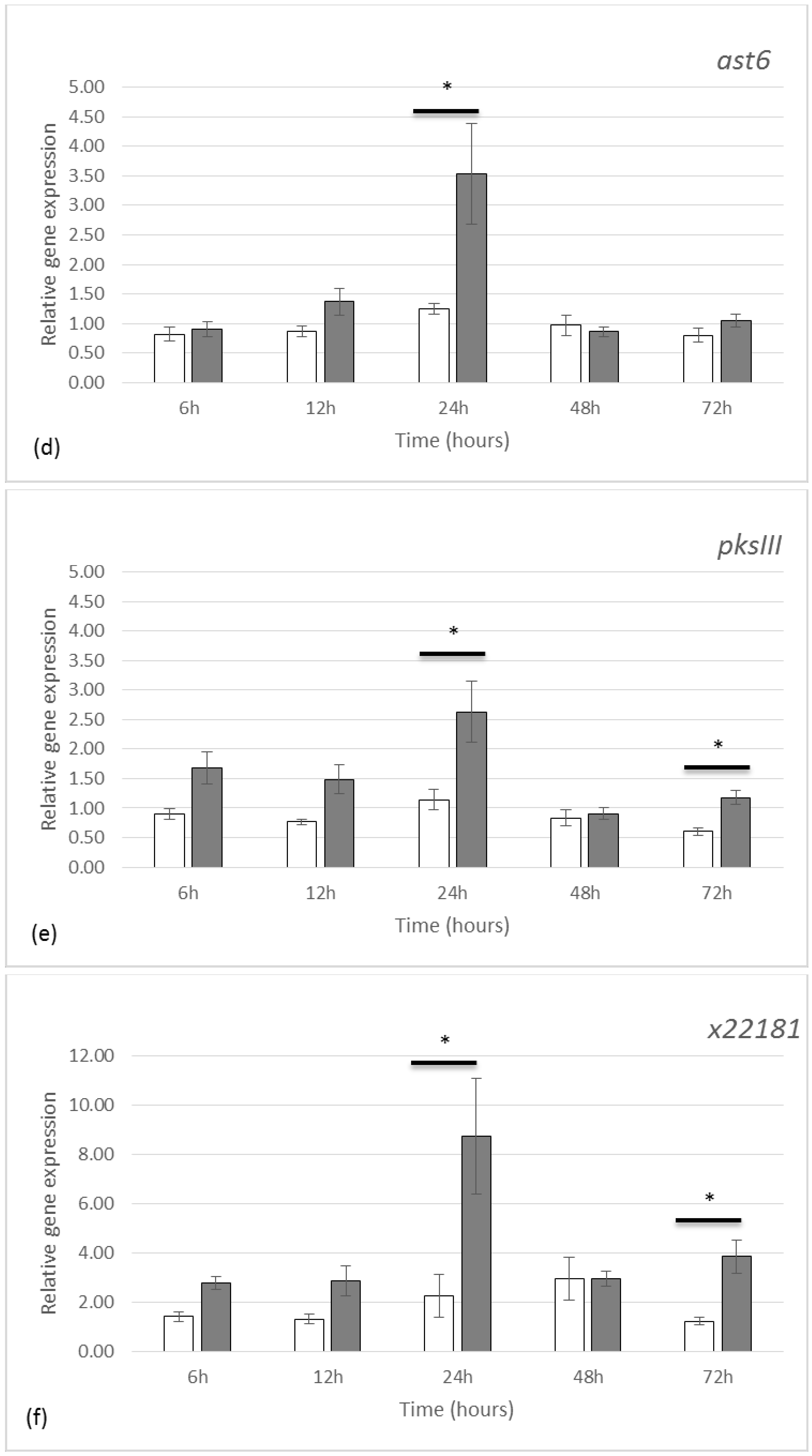
2.2. Gene Expression by qRT-PCR
3. Discussion
3.1. Quantification of Soluble Phenolic Compounds
3.2. Gene Expression
4. Materials and Methods
4.1. Biological Materials and Experimental Design
4.2. RNA Extraction
4.3. Quantitative Real-Time PCR (qRT-PCR)
4.4. Gene Expression Study
4.5. CYP450 Phylogenetic Analysis
4.6. Phlorotannin Extraction
4.7. Quantification of Soluble Phlorotannins
4.8. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Ragan, M.A.; Glombitza, K.W. Phlorotannins, brown algal polyphenols. Prog. Phycol. Res. 1986, 4, 129–241. [Google Scholar]
- Meslet-Cladiere, L.; Delage, L.; Leroux, C.J.-J.; Goulitquer, S.; Leblanc, C.; Creis, E.; Gall, E.A.; Stiger-Pouvreau, V.; Czjzek, M.; Potin, P. Structure/Function analysis of a type III polyketide synthase in the brown alga Ectocarpus siliculosus reveals a biochemical pathway in phlorotannin monomer biosynthesis. Plant Cell 2013, 25, 3089–3103. [Google Scholar] [CrossRef] [PubMed]
- Creis, E.; Ar Gall, E.; Potin, P. Ubiquitous phlorotannins prospects and perspectives. In Blue Biotechnology; Wiley-VCH Verlag GmbH & Co. KGaA: Weinheim, Germany, 2018; pp. 67–116. [Google Scholar]
- Koivikko, R. Brown Algal Phlorotannins Improving and Applying Chemical Methods. Ph.D. Thesis, University of Turku, Turku, Finland, 2008. [Google Scholar]
- Schoenwaelder, M.E.A.; Clayton, M.N. Secretion of phenolic substances into the zygote wall and cell plate in embryos of Hormosira and Acrocarpia (Fucales, Phaeophyceae). J. Phycol. 1998, 34, 969–980. [Google Scholar] [CrossRef]
- Arnold, T.M.; Targett, N.M. To grow and defend: Lack of tradeoffs for brown algal phlorotannins. Oikos 2003, 100, 406–408. [Google Scholar] [CrossRef]
- Lemesheva, V.; Birkemeyer, C.; Garbary, D.; Tarakhovskaya, E. Vanadium-dependent haloperoxidase activity and phlorotannin incorporation into the cell wall during early embryogenesis of Fucus vesiculosus (Phaeophyceae). Eur. J. Phycol. 2020, 55, 275–284. [Google Scholar] [CrossRef]
- Tarakhovskaya, E.; Lemesheva, V.; Bilova, T.; Birkemeyer, C. Early embryogenesis of brown alga Fucus vesiculosus L. is characterized by significant changes in carbon and energy metabolism. Molecules 2017, 22, 1509. [Google Scholar] [CrossRef] [PubMed]
- Tarakhovskaya, E.R. Mechanisms of bioadhesion of macrophytic algae. Russ. J. Plant Physiol. 2013, 61, 19–25. [Google Scholar] [CrossRef]
- Potin, P.; Bouarab, K.; Salaun, J.-P.; Pohnert, G.; Kloareg, B. Biotic interactions of marine algae. Curr. Opin. Plant Biol. 2002, 5, 308–317. [Google Scholar] [CrossRef]
- Lopes, G.; Sousa, C.; Silva, L.R.; Pinto, E.; Andrade, P.B.; Bernardo, J.; Mouga, T.; Valentão, P. Can phlorotannins purified extracts constitute a novel pharmacological alternative for microbial infections with associated inflammatory conditions? PLoS ONE 2012, 7, e31145. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Jónsdóttir, R.; Liu, H.; Gu, L.; Kristinsson, H.G.; Raghavan, S.; Ólafsdóttir, G. Antioxidant capacities of phlorotannins extracted from the brown algae Fucus vesiculosus. J. Agric. Food Chem. 2012, 60, 5874–5883. [Google Scholar] [CrossRef]
- Gomez, I.; Huovinen, P. Induction of phlorotannins during UV exposure mitigates inhibition of photosynthesis and DNA damage in the kelp Lessonia nigrescens. Photochem. Photobiol. 2010, 86, 1056–1063. [Google Scholar] [CrossRef] [PubMed]
- Feeny, P. Plant apparency and chemical defense. In Biochemical Interaction Between Plants and Insects; Wallace, J.W., Mansell, R.L., Eds.; Springer US: Boston, MA, USA, 1976; pp. 1–40. ISBN 978-1-4684-2646-5. [Google Scholar]
- Stern, J.; Hagerman, A.; Steinberg, P. Phlorotannin-protein interactions. J. Chem. 1996, 22. [Google Scholar] [CrossRef] [PubMed]
- Rhoades, D.F.; Cates, R.G. Toward a general theory of plant antiherbivore chemistry. In Biochemical Interaction between Plants and Insects; Wallace, J.W., Mansell, R.L., Eds.; Springer US: Boston, MA, USA, 1976; pp. 168–213. ISBN 978-1-4684-2646-5. [Google Scholar]
- Coleman, R.A.; Ramchunder, S.J.; Moody, A.J.; Foggo, A. An enzyme in snail saliva induces herbivore-resistance in a marine alga. Funct. Ecol. 2007, 21, 101–106. [Google Scholar] [CrossRef]
- Lüder, U.H.; Clayton, M.N. Induction of phlorotannins in the brown macroalga Ecklonia radiata (Laminariales, Phaeophyta) in response to simulated herbivory--the first microscopic study. Planta 2004, 218, 928–937. [Google Scholar] [CrossRef]
- Yates, J.C.; Peckol, P. Effects of nutrient availability and herbivory on polyphenolics in the seaweed Fucus vesiculosus. Ecology 1993, 74, 1757–1766. [Google Scholar] [CrossRef]
- Deal, M.S.; Hay, M.E.; Wilson, D.; Fenical, W. Galactolipids rather than phlorotannins as herbivore deterrents in the brown seaweed Fucus vesiculosus. Oecologia 2003, 136, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Karban, R.; Agrawal, A.A.; Thaler, J.S.; Adler, L.S. Induced plant responses and information content about risk of herbivory. Trends Ecol. Evol. 1999, 14, 443–447. [Google Scholar] [CrossRef]
- Stamp, N. Out of the quagmire of plant defense hypotheses. Q. Rev. Biol. 2003, 78, 23–55. [Google Scholar] [CrossRef] [PubMed]
- Flöthe, C.R.; Molis, M.; John, U. Induced resistance to periwinkle grazing in the brown seaweed Fucus vesiculosus (Phaeophyceae): Molecular insights and seaweed-mediated effects on herbivore interactions. J. Phycol. 2014, 50, 564–576. [Google Scholar] [CrossRef] [PubMed]
- Flöthe, C.R.; John, U.; Molis, M. Comparing the relative importance of water-borne cues and direct grazing for the induction of defenses in the brown seaweed Fucus vesiculosus. PLoS ONE 2014, 9, e109247. [Google Scholar] [CrossRef]
- Flöthe, C.R.; Molis, M.; Kruse, I.; Weinberger, F.; John, U. Herbivore-induced defence response in the brown seaweed Fucus vesiculosus (Phaeophyceae): Temporal pattern and gene expression. Eur. J. Phycol. 2014, 49, 356–369. [Google Scholar] [CrossRef]
- Hammerstrom, K.; Dethier, M.; Duggins, D. Rapid phlorotannin induction and relaxation in five Washington kelps. Mar. Ecol. Prog. Ser. 1998, 165, 293–305. [Google Scholar] [CrossRef]
- Pearson, G.; Hoarau, G.; Lago-Leston, A.; Coyer, J.; Kube, M.; Reinhardt, R.; Henckel, K.; Serrão, E.; Corre, E.; Olsen, J. An expressed sequence tag analysis of the intertidal brown seaweeds Fucus serratus(L.) and F. vesiculosus (L.) (Heterokontophyta, Phaeophyceae) in response to abiotic stressors. Mar. Biotechnol. 2010, 12, 195–213. [Google Scholar] [CrossRef]
- Creis, E.; Delage, L.; Charton, S.; Goulitquer, S.; Leblanc, C.; Potin, P.; Ar Gall, E. Constitutive or inducible protective mechanisms against UV-B radiation in the brown alga Fucus vesiculosus? A study of gene expression and phlorotannin content responses. PLoS ONE 2015, 10, e0128003. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Einig, E.; Almeida-Trapp, M.; Albert, M.; Fliegmann, J.; Mithöfer, A.; Kalbacher, H.; Felix, G. The systemin receptor SYR1 enhances resistance of tomato against herbivorous insects. Nat. Plants 2018, 4, 152–156. [Google Scholar] [CrossRef]
- Erb, M.; Reymond, P. Molecular Interactions between Plants and Insect Herbivores. Annu. Rev. Plant Biol. 2019, 70, 527–557. [Google Scholar] [CrossRef]
- Thomas, F.; Cosse, A.; Le Panse, S.; Kloareg, B.; Potin, P.; Leblanc, C. Kelps feature systemic defense responses: Insights into the evolution of innate immunity in multicellular eukaryotes. New Phytol. 2014, 204, 567–576. [Google Scholar] [CrossRef] [PubMed]
- Arnold, T.M.; Targett, N.M. Evidence for metabolic turnover of polyphenolics in tropical brown algae. J. Chem. Ecol. 2000, 26, 1393–1410. [Google Scholar] [CrossRef]
- Birkemeyer, C.; Lemesheva, V.; Billig, S.; Tarakhovskaya, E. Composition of intracellular and cell wall-bound phlorotannin fractions in fucoid algae indicates specific functions of these metabolites dependent on the chemical structure. Metabolites 2020, 10, 369. [Google Scholar] [CrossRef]
- Zhang, H.; Li, W.; Li, J.; Fu, W.; Yao, J.; Duan, D. Characterization and expression analysis of hsp70 gene from Ulva prolifera J. Agardh (Chlorophycophyta, Chlorophyceae). Mar. Genom. 2012, 5, 53–58. [Google Scholar] [CrossRef] [PubMed]
- Henkel, S.K.; Hofmann, G.E. Differing patterns of hsp70 gene expression in invasive and native kelp species: Evidence for acclimation-induced variation. In Nineteenth International Seaweed Symposium, Proceedings of the 19th International Seaweed Symposium, Kobe, Japan, 26–31 March 2007; Borowitzka, M.A., Critchley, A.T., Kraan, S., Peters, A., Sjøtun, K., Notoya, M., Eds.; Springer: Dordrecht, The Netherlands, 2009; pp. 465–474. ISBN 978-1-4020-9619-8. [Google Scholar]
- Smolina, I.; Kollias, S.; Jueterbock, A.; Coyer, J.A.; Hoarau, G. Variation in thermal stress response in two populations of the brown seaweed, Fucus distichus, from the Arctic and subarctic intertidal. R. Soc. Open Sci. 2016, 3. [Google Scholar] [CrossRef]
- Leblanc, C.; Schaal, G.; Cosse, A.; Destombe, C.; Valero, M.; Riera, P.; Potin, P. Trophic and biotic interactions in Laminaria digitata beds: Which factors could influence the persistence of marine kelp forests in northern brittany? Cah. Biol. Mar. 2011, 52, 415–427. [Google Scholar]
- Xu, J.; Wang, X.Y.; Guo, W.Z. The cytochrome P450 superfamily: Key players in plant development and defense. J. Integr. Agric. 2015, 14, 1673–1686. [Google Scholar] [CrossRef]
- Pandian, B.A.; Sathishraj, R.; Djanaguiraman, M.; Prasad, P.V.V.; Jugulam, M. Role of cytochrome P450 enzymes in plant stress response. Antioxidants 2020, 9, 454. [Google Scholar] [CrossRef] [PubMed]
- Toporkova, Y.Y.; Fatykhova, V.S.; Gogolev, Y.V.; Khairutdinov, B.I.; Mukhtarova, L.S.; Grechkin, A.N. Epoxyalcohol synthase of Ectocarpus siliculosus. First CYP74-related enzyme of oxylipin biosynthesis in brown algae. BBA Mol. Cell Biol. Lipids 2016, 1862, 167–175. [Google Scholar] [CrossRef]
- Teng, L.; Fan, X.; Nelson, D.R.; Han, W.; Zhang, X.; Xu, D.; Renault, H.; Markov, G.V.; Ye, N. Diversity and evolution of cytochromes P450 in stramenopiles. Planta 2019, 249, 647–661. [Google Scholar] [CrossRef] [PubMed]
- Bitton, R.; Ben-Yehuda, M.; Davidovich, M.; Balazs, Y.; Potin, P.; Delage, L.; Colin, C.; Bianco-Peled, H. Structure of algal-born phenolic polymeric adhesives. Macromol. Biosci. 2006, 6, 737–746. [Google Scholar] [CrossRef]
- Salgado, L.T.; Cinelli, L.P.; Viana, N.B.; Tomazetto de Carvalho, R.; de Souza Mourão, P.A.; Teixeira, V.L.; Farina, M.; Filho, G.M.A. A vanadium bromoperoxidase catalyzes the formation of high-molecular-weight complexes between brown algal phenolic substances and alginates. J. Phycol. 2009, 45, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Vreeland, V.; Waite, J.H.; Epstein, L. Minireview-polyphenols and oxidases in substratum adhesion by marine algae and mussels. J. Phycol. 1998, 34, 1–8. [Google Scholar] [CrossRef]
- Ye, N.; Zhang, X.; Miao, M.; Fan, X.; Zheng, Y.; Xu, D.; Wang, J.; Zhou, L.; Wang, D.; Gao, Y.; et al. Saccharina genomes provide novel insight into kelp biology. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Van Der Horst, M.A.; Hartog, A.F.; El Morabet, R.; Marais, A.; Kircz, M.; Wever, R. Enzymatic sulfation of phenolic hydroxy groups of various plant metabolites by an arylsulfotransferase. Eur. J. Org. Chem. 2015, 2015, 534–541. [Google Scholar] [CrossRef]
- Ritter, A.; Cabioch, L.; Brillet-Guéguen, L.; Corre, E.; Cosse, A.; Dartevelle, L.; Duruflé, H.; Fasshauer, C.; Goulitquer, S.; Thomas, F.; et al. Herbivore-induced chemical and molecular responses of the kelps Laminaria digitata and Lessonia spicata. PLoS ONE 2017, 12, 1–17. [Google Scholar] [CrossRef]
- Targett, N.M.; Arnold, T.M. Minireview-predicting the effects of brown algal phlorotannins on marine herbivores in tropical and temperate oceans. J. Phycol. 1998, 34, 195–205. [Google Scholar] [CrossRef]
- Van Alstyne, K. Herbivore grazing increases polyphenolic defenses in the intertidal brown alga Fucus distichus. Ecology 1988, 69, 655–663. [Google Scholar] [CrossRef]
- Long, J.D.; Hamilton, R.S.; Mitchell, J.L. Asymmetric competition via induced resistance: Specialist herbivores indirectly suppress generalist preference and populations. Ecology 2007, 88, 1232–1240. [Google Scholar] [CrossRef]
- Yun, H.Y.; Rohde, S.; Linnane, K.; Wahl, M.; Molis, M. Seaweed-mediated indirect interaction between two species of meso-herbivores. Mar. Ecol. Prog. Ser. 2010, 408, 47–53. [Google Scholar] [CrossRef]
- Hammann, M.; Wang, G.; Rickert, E.; Boo, S.M.; Weinberger, F. Invasion success of the seaweed Gracilaria vermiculophylla correlates with low palatibility. Mar. Ecol. Prog. Ser. 2013, 486, 93–103. [Google Scholar] [CrossRef][Green Version]
- Apt, K.E.; Grossman, A.R. Characterization and transcript analysis of the major phycobiliprotein subunit genes from Aglaothamnion neglectum (Rhodophyta). Plant Mol. Biol. 1993, 21, 27–38. [Google Scholar] [CrossRef]
- Apt, K.E.; Clendennen, S.K.; Powers, D.A.; Grossman, A.R. The gene family encoding the fucoxanthin chlorophyll proteins from the brown alga Macrocystis pyrifera. Mol. Gen. Genet. 1995, 246, 455–464. [Google Scholar] [CrossRef]
- Pearson, G.; Lago-Leston, A.; Valente, M.; Serrão, E. Simple and rapid RNA extraction from freeze-dried tissue of brown algae and seagrasses. Eur. J. Phycol. 2006, 41, 97–104. [Google Scholar] [CrossRef]
- Fang, G.; Hammar, S.; Grumet, R. A quick and inexpensive method for removing polysaccharides from plant genomic DNA. Biotechniques 1992, 13, 52–56. [Google Scholar]
- Kinnby, A.; Jonsson, P.R.; Ortega-Martinez, O.; Töpel, M.; Pavia, H.; Pereyra, R.T.; Johannesson, K. Combining an ecological experiment and a genome scan show idiosyncratic responses to salinity stress in local populations of a seaweed. Front. Mar. Sci. 2020, 7, 1–12. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL): An online tool for phylogenetic tree display and annotation. Bioinformatics 2007, 23, 127–128. [Google Scholar] [CrossRef] [PubMed]
- Van Alstyne, K. Comparison of three methods for quantifying brown algal polyphenolic compounds. J. Chem. Ecol. 1995, 21, 45–58. [Google Scholar] [CrossRef] [PubMed]
- Lenth, R.; Singmann, H.; Love, J.; Buerkner, P.; Herve, M. Emmeans: Estimated Marginal Means, Aka Least-Squares Means (Version 1.4.5) [Computer Software]. 2020. Available online: https://cran.r-project.org/package=emmeans (accessed on 20 September 2020).
- Koivikko, R.; Loponen, J.; Honkanen, T.; Jormalainen, V. Contents of soluble, cell-wall-bound and exuded phlorotannins in the brown alga Fucus vesiculosus, with implications on their ecological functions. J. Chem. Ecol. 2005, 31, 195–212. [Google Scholar] [CrossRef]
- Thomas, N.V.; Kim, S.-K. Potential pharmacological applications of polyphenolic derivatives from marine brown algae. Environ. Toxicol. Pharmacol. 2011, 32, 325–335. [Google Scholar] [CrossRef]
- Joe, M.-J.; Kim, S.-N.; Choi, H.-Y.; Shin, W.-S.; Park, G.-M.; Kang, D.-W.; Kim, Y.K. The inhibitory effects of eckol and dieckol from Ecklonia stolonifera on the expression of matrix metalloproteinase-1 in human dermal fibroblasts. Biol. Pharm. Bull. 2006, 29, 1735–1739. [Google Scholar] [CrossRef]
- Yoon, N.Y.; Kim, H.R.; Chung, H.Y.; Choi, J.S. Anti-hyperlipidemic effect of an edible brown algae, Ecklonia stolonifera, and its constituents on poloxamer 407-induced hyperlipidemic and cholesterol-fed rats. Arch. Pharm. Res. 2008, 31, 1564–1571. [Google Scholar] [CrossRef]
- Moon, H.E.; Islam, N.; Ahn, B.R.; Chowdhury, S.S.; Sohn, H.S.; Jung, H.A.; Choi, J.S. Protein tyrosine phosphatase 1B and α-glucosidase inhibitory phlorotannins from edible brown algae, Ecklonia stolonifera and Eisenia bicyclis. Biosci. Biotechnol. Biochem. 2011, 75, 1472–1480. [Google Scholar] [CrossRef]
- Lee, J.H.; Hye-Young, O.; Park, J. S. Preventive effect of Ecklonia stolonifera on the frequency of benzo(a)pyrene-induced chromosomal aberration. J. Food Sci. Nutr. 1996, 1, 64–68. [Google Scholar]

| Gene | Forward | Reverse | Amplicon Length (bp) | Tm °C | Accession Number | References |
|---|---|---|---|---|---|---|
| ef1.alpha | TGCGTACAATCGCATTCG | CGAAACATGAAGGACAGTTGC | 198 | 58 | GH706096 | EST (Pearson et al. 2010) |
| tua | GTCACACCGATGTAGAGGA | GGCTTCCAGACAATTACCC | 96 | 58 | GH702736 | EST (Pearson et al. 2010) |
| pks | TTGCACGTATGTCTCTGTTGC | GCGCGAATAACCTGATGG | 135 | 60 | GH706741 | EST (Pearson et al. 2010) |
| vbpo | CCAAGGCGTCGAGTCATATC | GCACTTACTGCAATCCAATGTAC | 129 | 59 | CUST_1779_PI408257168 | Microarray accession number GSE47975 (Flöthe et al 2014 Eur J.Phycology) |
| hsp70 | AGATCGAGGAGATTGACTAGATGG | CGACTTGCATCACACATATCG | 161 | 60 | GH704979 | EST (Pearson et al. 2010) |
| ast6 | GACCCTTCCCTGATCTTCC | CCAGATGCGGTCATTTCAC | 83 | 59 | JACAZD010018410.1 (19554-20266) | Fucus vesiculosus genome |
| x22181 | TGGTCGAGACGGAGGAAG | TGCACTTCAAGCTATTACTCTTGC | 132 | 60 | CUST_44291_PI408257168 | Microarray accession number GSE47975 (Flöthe et al 2014 Eur J.Phycology) |
| cyp450 | TAACGACATGGCTCAAATCAC | ACACAACAAACACCCACAC | 84 | 59 | CUST_44854_PI408257168 | Microarray accession number GSE47975 (Flöthe et al 2014 Eur J.Phycology) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Emeline, C.B.; Ludovic, D.; Laurent, V.; Catherine, L.; Kruse, I.; Erwan, A.G.; Florian, W.; Philippe, P. Induction of Phlorotannins and Gene Expression in the Brown Macroalga Fucus vesiculosus in Response to the Herbivore Littorina littorea. Mar. Drugs 2021, 19, 185. https://doi.org/10.3390/md19040185
Emeline CB, Ludovic D, Laurent V, Catherine L, Kruse I, Erwan AG, Florian W, Philippe P. Induction of Phlorotannins and Gene Expression in the Brown Macroalga Fucus vesiculosus in Response to the Herbivore Littorina littorea. Marine Drugs. 2021; 19(4):185. https://doi.org/10.3390/md19040185
Chicago/Turabian StyleEmeline, Creis Bendelac, Delage Ludovic, Vallet Laurent, Leblanc Catherine, Inken Kruse, Ar Gall Erwan, Weinberger Florian, and Potin Philippe. 2021. "Induction of Phlorotannins and Gene Expression in the Brown Macroalga Fucus vesiculosus in Response to the Herbivore Littorina littorea" Marine Drugs 19, no. 4: 185. https://doi.org/10.3390/md19040185
APA StyleEmeline, C. B., Ludovic, D., Laurent, V., Catherine, L., Kruse, I., Erwan, A. G., Florian, W., & Philippe, P. (2021). Induction of Phlorotannins and Gene Expression in the Brown Macroalga Fucus vesiculosus in Response to the Herbivore Littorina littorea. Marine Drugs, 19(4), 185. https://doi.org/10.3390/md19040185






