Anti-SARS-CoV-2 Activity of Rhamnan Sulfate from Monostroma nitidum
Abstract
:1. Introduction
2. Results and Discussion
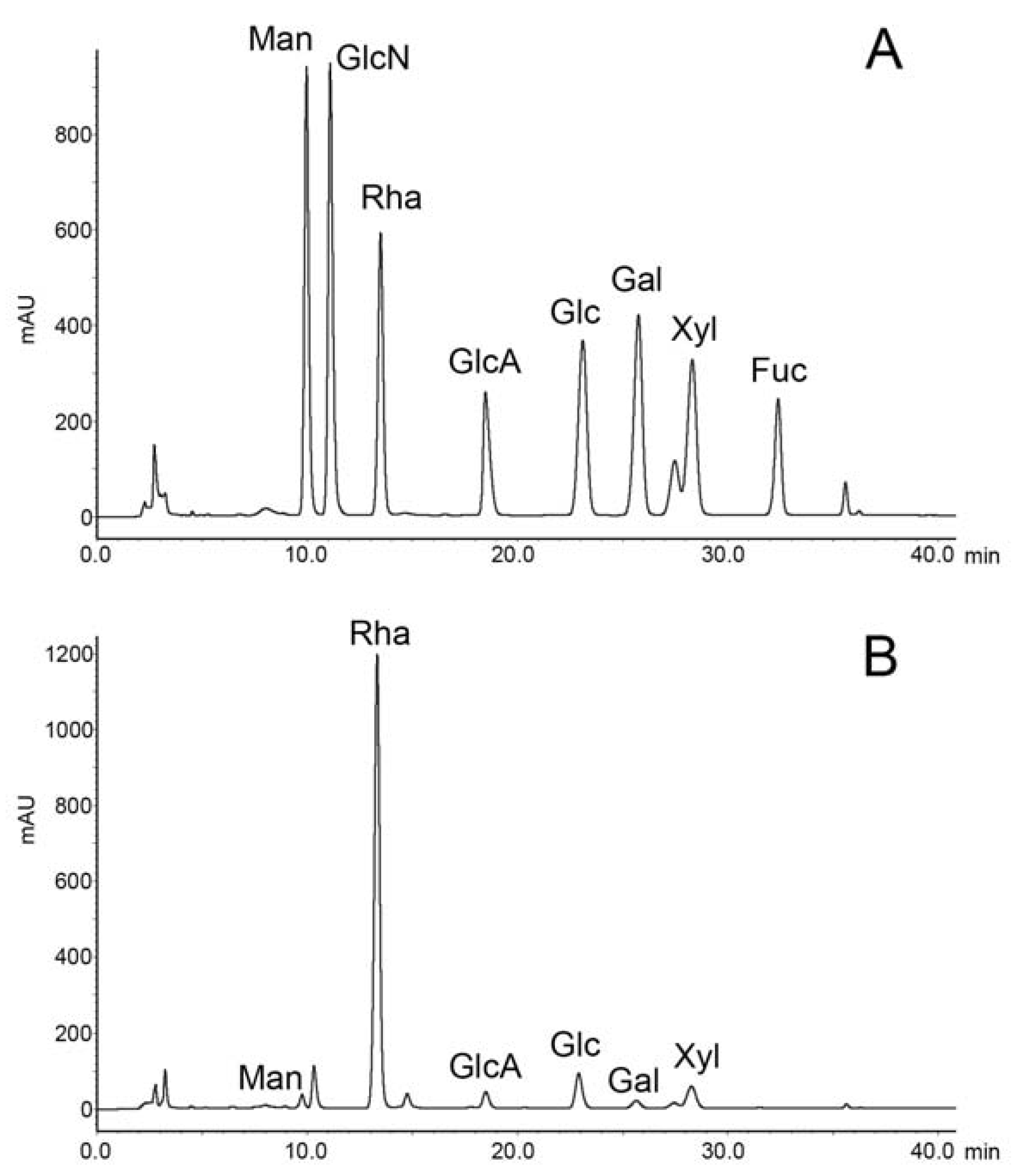
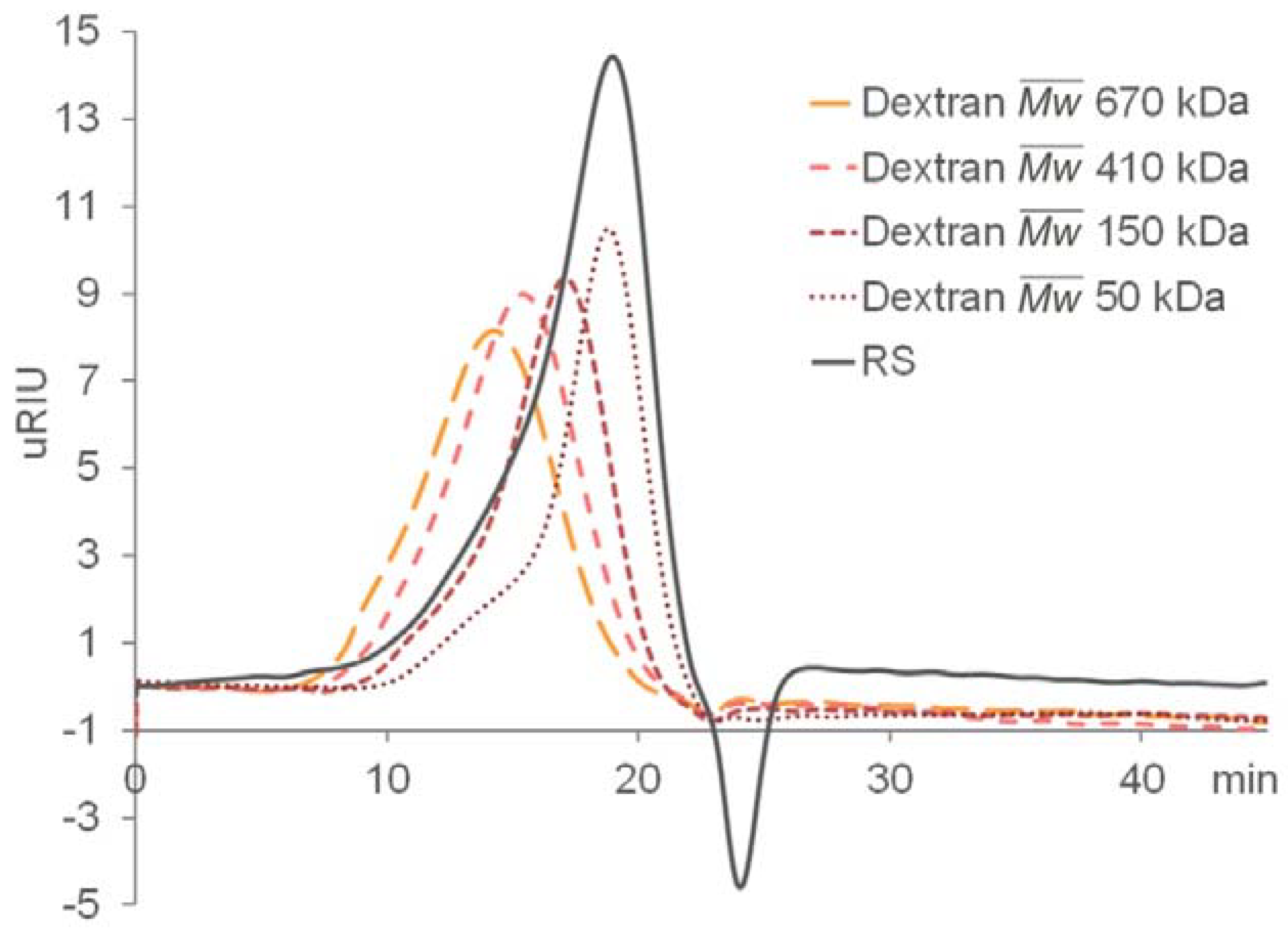
2.1. Preparation and Compositional Analysis of RS
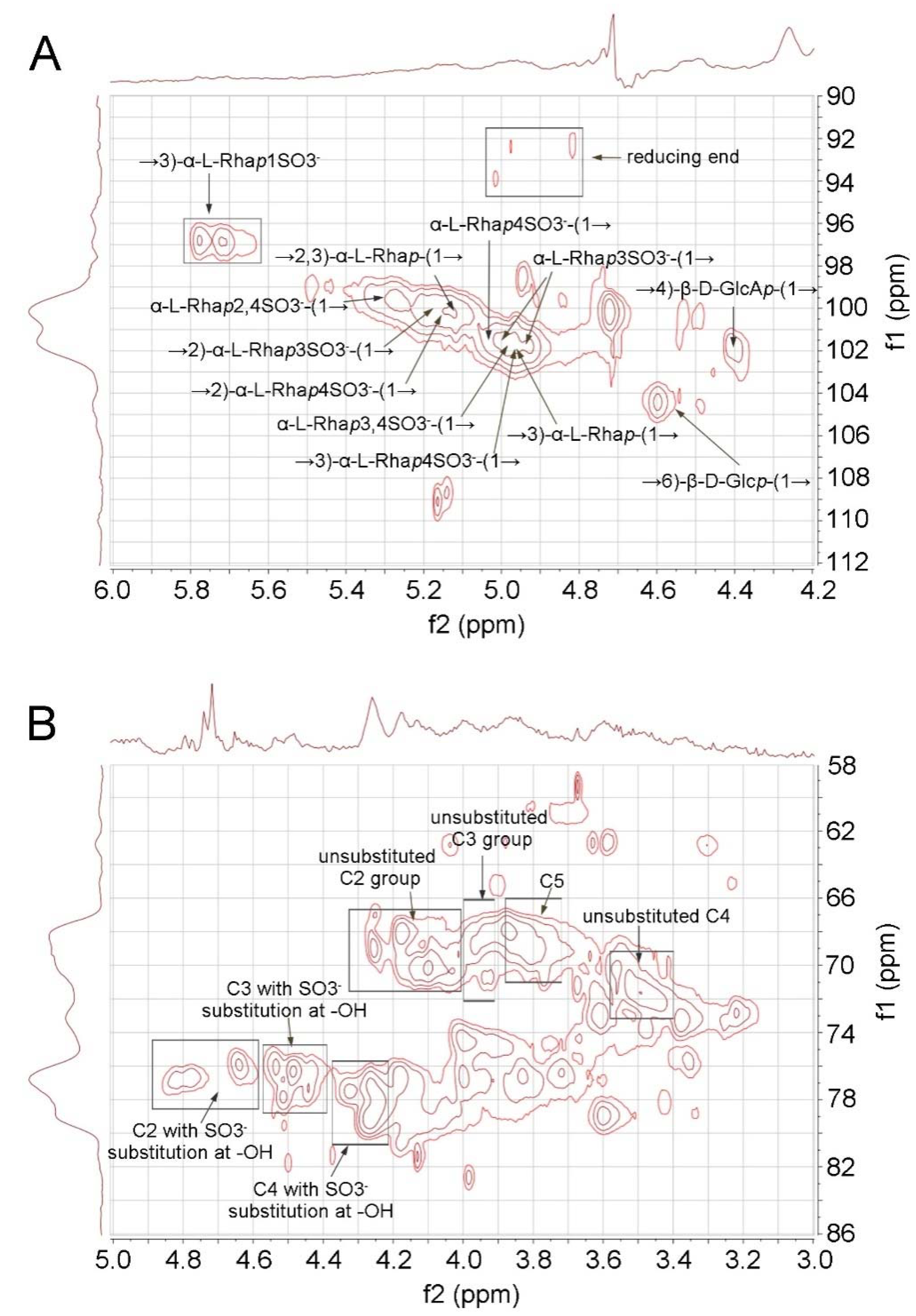
2.2. 2D-NMR Analysis of RS
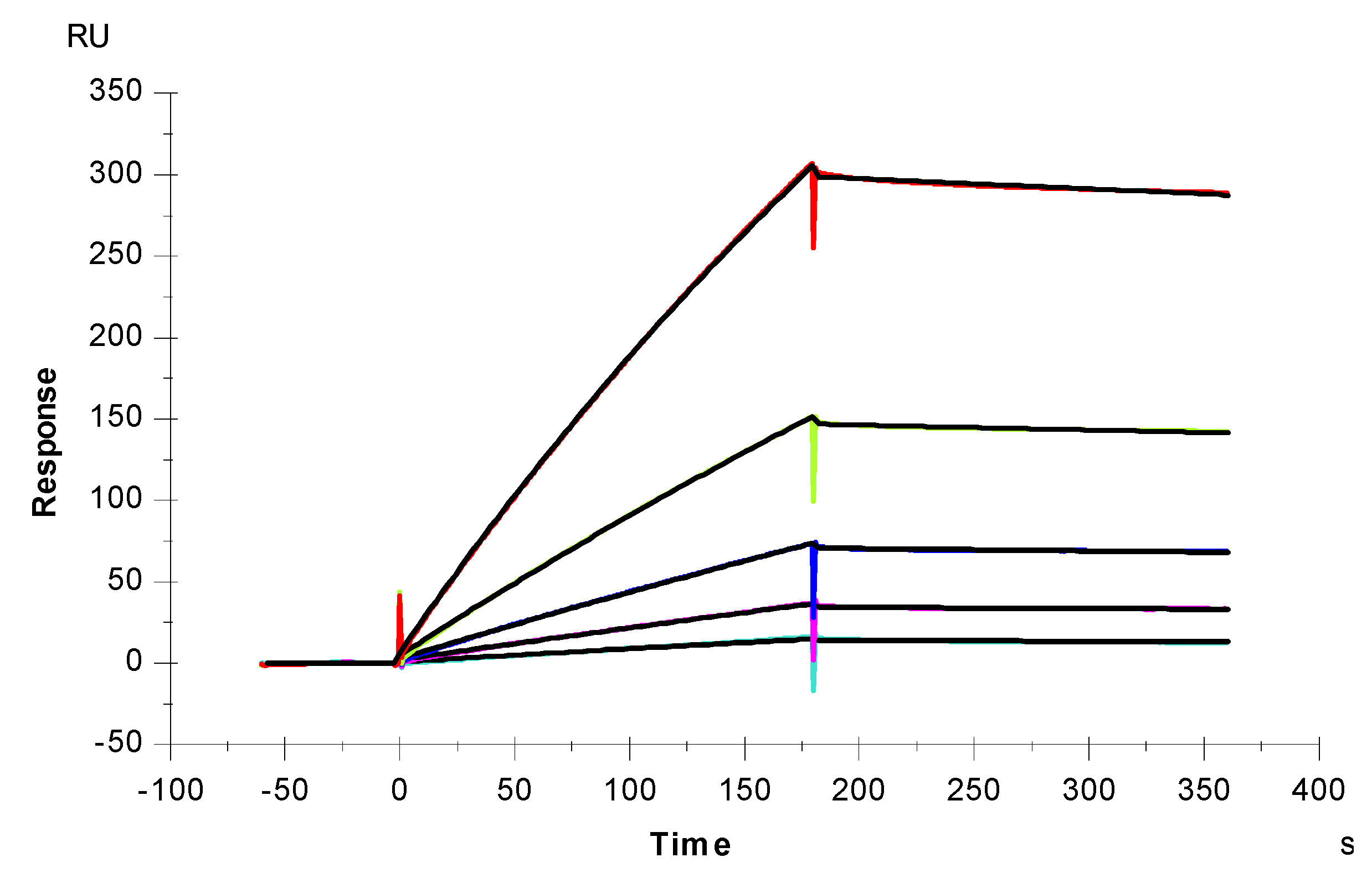
2.3. Interaction between SARS-CoV-2 S-Protein RBD and Heparin
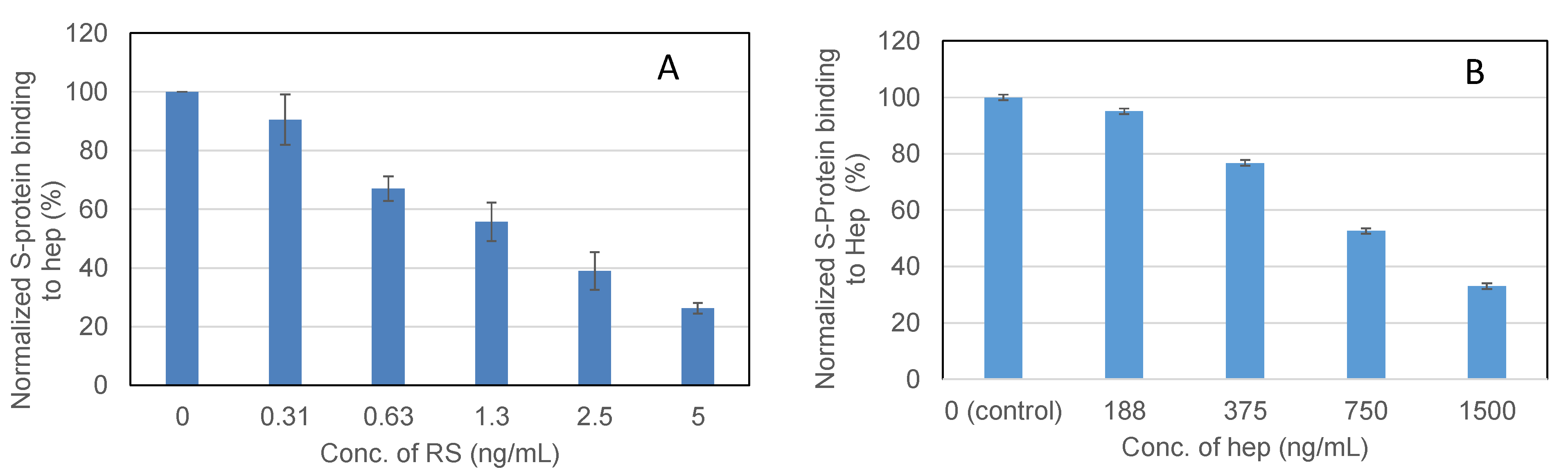
2.4. IC50 Measurement on the Inhibition of S-Protein Binding to Heparin by RS Using Solution Competition SPR
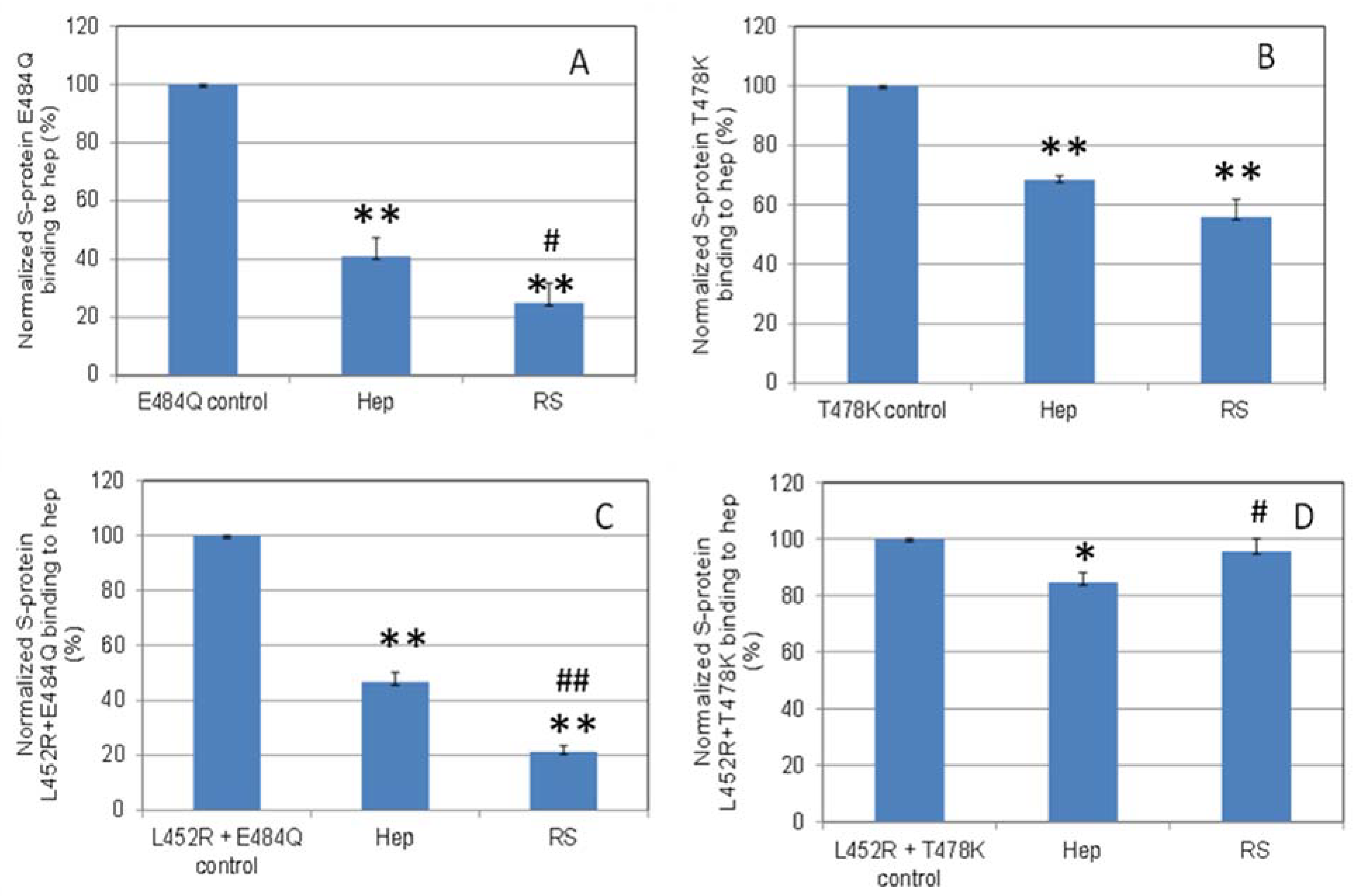
2.5. Inhibition of S-Protein RBD Variants Binding to Heparin by RS Using Solution Competition SPR
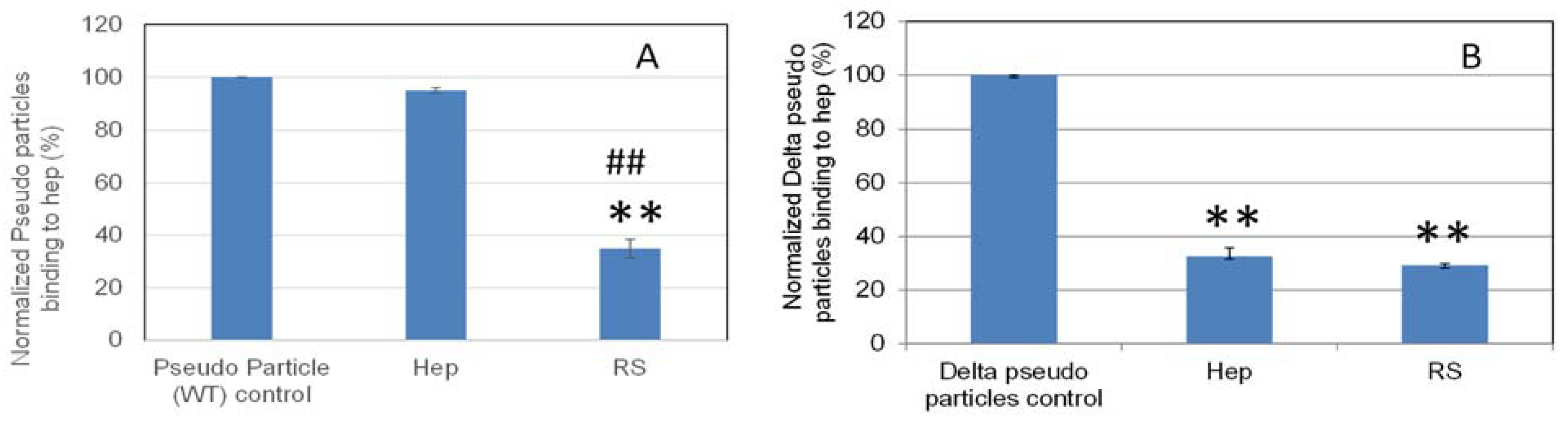
2.6. SPR Solution Competition Study on the Inhibition by RS of Heparin Interaction with Pseudovirus Particles
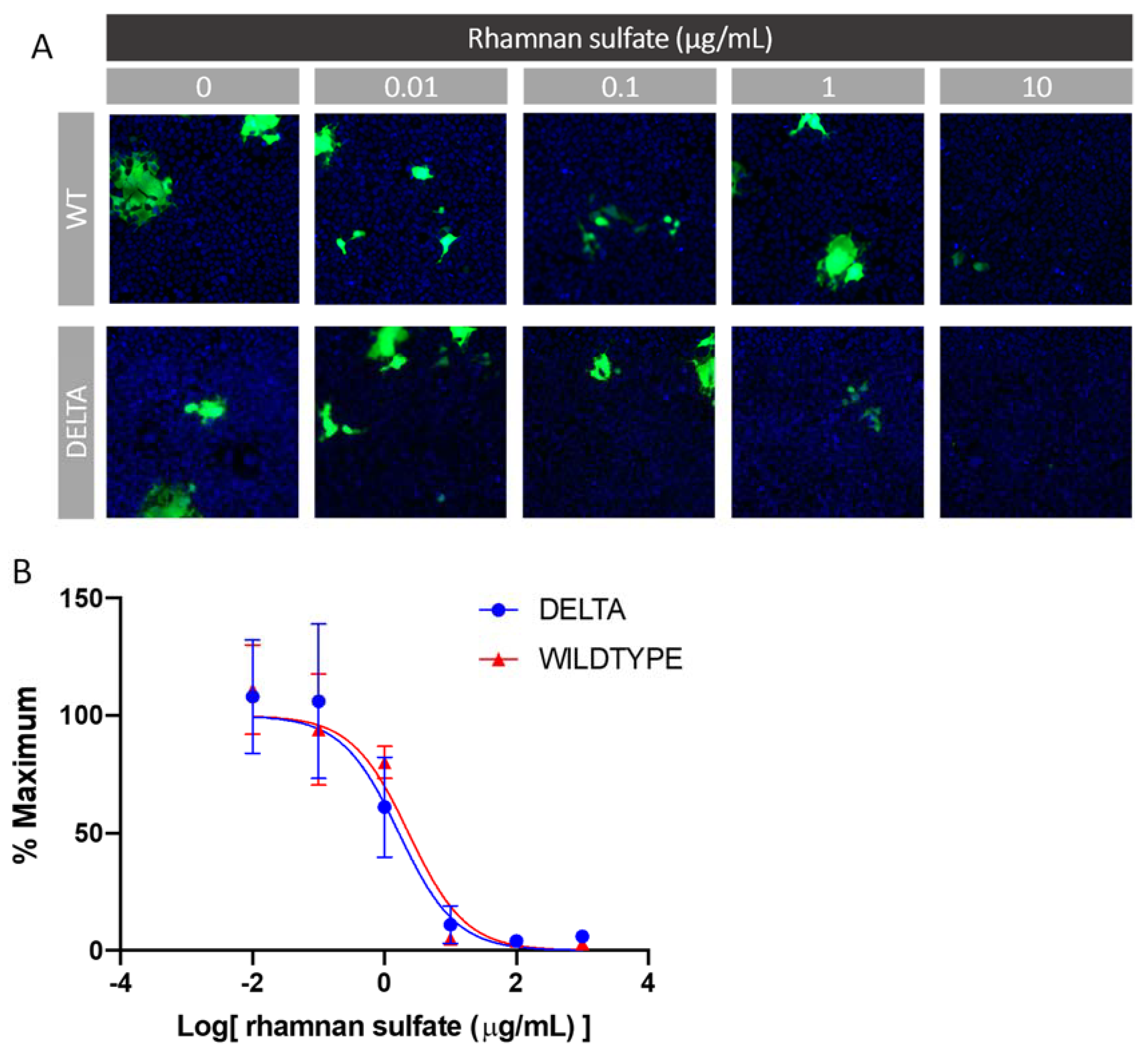
2.7. In Vitro SARS-CoV-2 Pseudotyped Virus Neutralization
3. Materials and Methods
3.1. Biological Materials and Reagents
3.2. Preparation of RS
3.3. Compositional Analysis of RS
3.4. 2D NMR Analysis
3.5. Measurement of Interaction between Heparin and S-Proteins RBD Using SPR
3.6. SPR Solution Competition Study of RS
3.7. In Vitro SARS-CoV-2 Pseudotyped Virus Neutralization Assay
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Peng, R.; Wu, L.-A.; Wang, Q.; Qi, J.; Gao, G.F. Cell entry of SARS-CoV-2. Trends Biochem. Sci. 2021, 117, 11727–11734. [Google Scholar] [CrossRef]
- Kalra, R.S.; Kandimalla, R. Engaging the spikes: Heparan sulfate facilitates SARS-CoV-2 spike protein binding to ACE2 and potentiates viral infection. Signal Transduct. Target. Ther. 2021, 6, 1–2. [Google Scholar] [CrossRef]
- Zhang, Q.; Chen, C.Z.; Swaroop, M.; Xu, M.; Wang, L.; Lee, J.; Wang, A.Q.; Pradhan, M.; Hagen, N.; Chen, L. Heparan sulfate assists SARS-CoV-2 in cell entry and can be targeted by approved drugs in vitro. Cell Discov. 2020, 6, 1–14. [Google Scholar] [CrossRef]
- Liu, L.; Chopra, P.; Li, X.; Bouwman, K.M.; Tompkins, S.M.; Wolfert, M.A.; de Vries, R.P.; Boons, G.-J. Heparan sulfate proteoglycans as attachment factor for SARS-CoV-2. ACS Cent. Sci. 2021, 7, 1009–1018. [Google Scholar] [CrossRef]
- Clausen, T.M.; Sandoval, D.R.; Spliid, C.B.; Pihl, J.; Perrett, H.R.; Painter, C.D.; Narayanan, A.; Majowicz, S.A.; Kwong, E.M.; McVicar, R.N. SARS-CoV-2 infection depends on cellular heparan sulfate and ACE2. Cell 2020, 183, 1043–1057. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Jin, W.; Sood, A.; Montgomery, D.W.; Grant, O.C.; Fuster, M.M.; Fu, L.; Dordick, J.S.; Woods, R.J.; Zhang, F.; et al. Characterization of heparin and severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) spike glycoprotein binding interactions. Antivir. Res. 2020, 181, 104873. [Google Scholar] [CrossRef] [PubMed]
- Synytsya, A.; Čopíková, J.; Kim, W.J.; Park, Y.I. Cell wall polysaccharides of marine algae. In Hb25_Springer Handbook of Marine Biotechnology; Springer: Singapore, 2015; pp. 543–590. [Google Scholar]
- Yamashiro, Y.; Nakamura, M.; Yogi, T.; Teruya, T.; Konishi, T.; Uechi, S.; Tako, M. Anticoagulant activity of rhamnan sulfate isolated from commercially cultured Monostroma nitidum. Int. J. Biomed. Mater. Res. 2017, 5, 37. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.-J.; Mao, W.-J.; Fang, F.; Li, H.-Y.; Sun, H.-H.; Chen, Y.; Qi, X.-H. Chemical characteristics and anticoagulant activities of a sulfated polysaccharide and its fragments from Monostroma latissimum. Carbohydr. Polym. 2008, 71, 428–434. [Google Scholar] [CrossRef]
- Mao, W.-J.; Fang, F.; Li, H.-Y.; Qi, X.-H.; Sun, H.-H.; Chen, Y.; Guo, S.-D. Heparinoid-active two sulfated polysaccharides isolated from marine green algae Monostroma nitidum. Carbohydr. Polym. 2008, 74, 834–839. [Google Scholar] [CrossRef]
- Li, H.; Mao, W.; Hou, Y.; Gao, Y.; Qi, X.; Zhao, C.; Chen, Y.; Chen, Y.; Li, N.; Wang, C. Preparation, structure and anticoagulant activity of a low molecular weight fraction produced by mild acid hydrolysis of sulfated rhamnan from Monostroma latissimum. Bioresour. Technol. 2012, 114, 414–418. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wang, S.; Cao, S.; He, X.; Qin, L.; He, M.; Yang, Y.; Hao, J.; Mao, W. Structural characteristics and anticoagulant property in vitro and in vivo of a seaweed sulfated Rhamnan. Mar. Drugs 2018, 16, 243. [Google Scholar] [CrossRef] [Green Version]
- Li, N.; Liu, X.; He, X.; Wang, S.; Cao, S.; Xia, Z.; Xian, H.; Qin, L.; Mao, W. Structure and anticoagulant property of a sulfated polysaccharide isolated from the green seaweed Monostroma angicava. Carbohydr. Polym. 2017, 159, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Cao, S.; He, X.; Qin, L.; He, M.; Yang, Y.; Liu, Z.; Mao, W. Anticoagulant and antithrombotic properties in vitro and in vivo of a novel sulfated polysaccharide from marine green alga Monostroma nitidum. Mar. Drugs 2019, 17, 247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, X.; Du, P.; Liu, X.; Cao, S.; Qin, L.; He, M.; He, X.; Mao, W. Anticoagulant properties of a green algal Rhamnan-type sulfated polysaccharide and its low-molecular-weight fragments prepared by mild acid degradation. Mar. Drugs 2018, 16, 445. [Google Scholar] [CrossRef] [Green Version]
- Suzuki, K.; Terasawa, M. Biological activities of Rhamnan sulfate extract from the green algae Monostroma nitidum (Hitoegusa). Mar. Drugs 2020, 18, 228. [Google Scholar] [CrossRef]
- Hoang, M.H.; Kim, J.-Y.; Lee, J.H.; You, S.G.; Lee, S.-J. Antioxidative, hypolipidemic, and anti-inflammatory activities of sulfated polysaccharides from Monostroma nitidum. Food Sci. Biotechnol. 2015, 24, 199–205. [Google Scholar] [CrossRef]
- Liu, X.; Hao, J.; He, X.; Wang, S.; Cao, S.; Qin, L.; Mao, W. A rhamnan-type sulfated polysaccharide with novel structure from Monostroma angicava Kjellm (Chlorophyta) and its bioactivity. Carbohydr. Polym. 2017, 173, 732–748. [Google Scholar] [CrossRef] [PubMed]
- Terasawa, M.; Hayashi, K.; Lee, J.-B.; Nishiura, K.; Matsuda, K.; Hayashi, T.; Kawahara, T. Anti-Influenza a virus activity of rhamnan sulfate from green algae Monostroma nitidum in mice with normal and compromised immunity. Mar. Drugs 2020, 18, 254. [Google Scholar] [CrossRef]
- Wang, S.; Wang, W.; Hao, C.; Yunjia, Y.; Qin, L.; He, M.; Mao, W. Antiviral activity against enterovirus 71 of sulfated rhamnan isolated from the green alga Monostroma latissimum. Carbohydr. Polym. 2018, 200, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-B.; Koizumi, S.; Hayashi, K.; Hayashi, T. Structure of rhamnan sulfate from the green alga Monostroma nitidum and its anti-herpetic effect. Carbohydr. Polym. 2010, 81, 572–577. [Google Scholar] [CrossRef]
- Wang, S.; Wang, W.; Hou, L.; Qin, L.; He, M.; Li, W.; Mao, W. A sulfated glucuronorhamnan from the green seaweed Monostroma nitidum: Characteristics of its structure and antiviral activity. Carbohydr. Polym. 2020, 227, 115280. [Google Scholar] [CrossRef] [PubMed]
- Tree, J.A.; Turnbull, J.E.; Buttigieg, K.R.; Elmore, M.J.; Coombes, N.; Hogwood, J.; Mycroft-West, C.J.; Lima, M.A.; Skidmore, M.A.; Karlsson, R. Unfractionated heparin inhibits live wild type SARS-CoV-2 cell infectivity at therapeutically relevant concentrations. Br. J. Pharmacol. 2021, 178, 626–635. [Google Scholar] [CrossRef] [PubMed]
- Partridge, L.J.; Urwin, L.; Nicklin, M.J.; James, D.C.; Green, L.R.; Monk, P.N. ACE2-Independent Interaction of SARS-CoV-2 spike protein with human epithelial cells is inhibited by unfractionated heparin. Cells 2021, 10, 1419. [Google Scholar] [CrossRef] [PubMed]
- Tandon, R.; Sharp, J.S.; Zhang, F.; Pomin, V.H.; Ashpole, N.M.; Mitra, D.; McCandless, M.G.; Jin, W.; Liu, H.; Sharma, P. Effective inhibition of SARS-CoV-2 entry by heparin and enoxaparin derivatives. J. Virol. 2020, 95, e01987-20. [Google Scholar] [CrossRef]
- Kwon, P.S.; Oh, H.; Kwon, S.-J.; Jin, W.; Zhang, F.; Fraser, K.; Hong, J.J.; Linhardt, R.J.; Dordick, J.S. Sulfated polysaccharides effectively inhibit SARS-CoV-2 in vitro. Cell Discov. 2020, 6, 1–4. [Google Scholar] [CrossRef]
- Hans, N.; Malik, A.; Naik, S. Antiviral activity of sulfated polysaccharides from marine algae and its application in combating COVID-19: Mini review. Bioresour. Technol. Rep. 2021, 13, 100623. [Google Scholar] [CrossRef]
- Jin, W.; Zhang, W.; Mitra, D.; McCandless, M.G.; Sharma, P.; Tandon, R.; Zhang, F.; Linhardt, R.J. The structure-activity relationship of the interactions of SARS-CoV-2 spike glycoproteins with glucuronomannan and sulfated galactofucan from Saccharina japonica. Int. J. Biol. Macromol. 2020, 163, 1649–1658. [Google Scholar] [CrossRef]
- Yim, S.-K.; Kim, K.; Kim, I.; Chun, S.; Oh, T.; Kim, J.-U.; Kim, J.; Jung, W.; Moon, H.; Ku, B. Inhibition of SARS-CoV-2 virus entry by the crude polysaccharides of seaweeds and abalone viscera in vitro. Mar. Drugs 2021, 19, 219. [Google Scholar] [CrossRef]
- Morokutti-Kurz, M.; Fröba, M.; Graf, P.; Große, M.; Grassauer, A.; Auth, J.; Schubert, U.; Prieschl-Grassauer, E. Iota-carrageenan neutralizes SARS-CoV-2 and inhibits viral replication in vitro. PLoS ONE 2021, 16, e0237480. [Google Scholar] [CrossRef]
- Dubois, M.; Gilles, K.A.; Hamilton, J.K.; Rebers, P.A.; Smith, F. Colorimetric method for determination of sugars and related substances. Anal. Chem. 1956, 28, 350–356. [Google Scholar] [CrossRef]
- Song, Y.; Wang, Q.; He, Y.; Ren, D.; Kow, F.; Li, J.; Liu, S.; Cong, H. The positive effects of fucoidans extracted from the brown seaweed Saccharina japonica on protection against CCl4-induced liver injury. J. Appl. Phycol. 2017, 29, 2077–2087. [Google Scholar] [CrossRef]
- Yan, L.; Song, Y.; Xia, K.; He, P.; Zhang, F.; Chen, S.; Pouliot, R.; Weiss, D.J.; Tandon, R.; Bates, J.T. Heparan sulfates from bat and human lung and their binding to the spike protein of SARS-CoV-2 virus. Carbohydr. Polym. 2021, 260, 117797. [Google Scholar] [CrossRef] [PubMed]
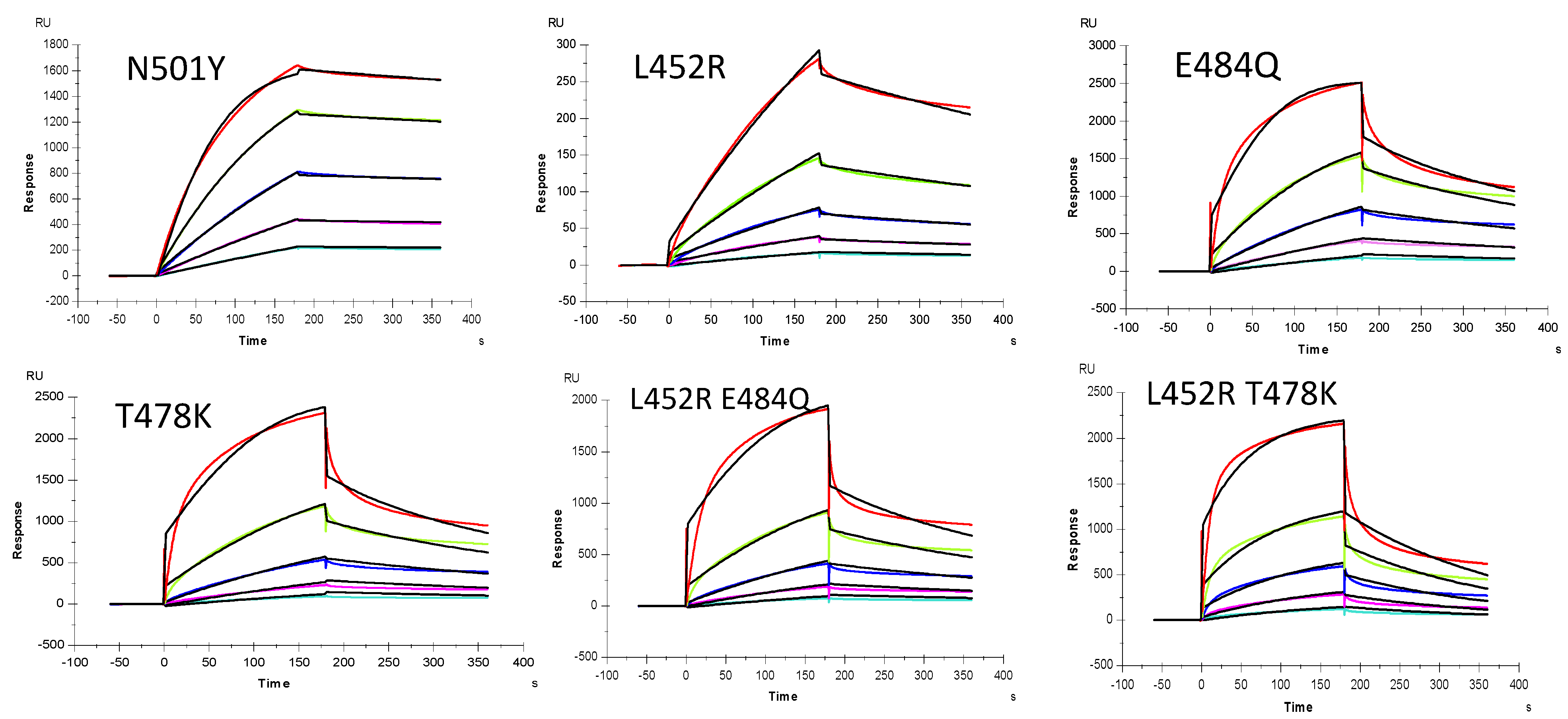
| Interaction | ka (1/MS) | kd (1/S) | KD (M) |
|---|---|---|---|
| SARS-CoV-2 S-protein RBD wt | 2144 (± 19) | 2.2 × 10−4 (±2 × 10−6) | 1.1 × 10−7 |
| SARS-CoV-2 S-protein RBD N501Y | 2.2 × 104 (±66) | 3.1 × 10−4 (±2.7 × 10−6) | 1.4 × 10−8 |
| SARS-CoV-2 S-protein RBD E484Q | 3.7 × 104 (±640) | 5.0 × 10−3 (±7.9 × 10−5) | 1.3 × 10−7 |
| SARS-CoV-2 S-protein RBD L452R | 1036 (±11) | 1.3 × 10−3 (±4.1 × 10−6) | 1.2 × 10−6 |
| SARS-CoV-2 S-protein RBD T478K | 3.2 × 104 (±1.1 × 103) | 7.9 × 10−3(±2.2 × 10−4) | 2.5 × 10−7 |
| SARS-CoV-2 S-protein RBD L452R+E484Q | 2.0 × 104 (±800) | 5.4 × 10−3 (±1.6 × 10−4) | 2.7 × 10−7 |
| SARS-CoV-2 S-protein RBD L452R+T478K | 1.2 × 104 (±89) | 4.8 × 10−3 (±2.7 × 10−5) | 4.1 × 10−7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Song, Y.; He, P.; Rodrigues, A.L.; Datta, P.; Tandon, R.; Bates, J.T.; Bierdeman, M.A.; Chen, C.; Dordick, J.; Zhang, F.; et al. Anti-SARS-CoV-2 Activity of Rhamnan Sulfate from Monostroma nitidum. Mar. Drugs 2021, 19, 685. https://doi.org/10.3390/md19120685
Song Y, He P, Rodrigues AL, Datta P, Tandon R, Bates JT, Bierdeman MA, Chen C, Dordick J, Zhang F, et al. Anti-SARS-CoV-2 Activity of Rhamnan Sulfate from Monostroma nitidum. Marine Drugs. 2021; 19(12):685. https://doi.org/10.3390/md19120685
Chicago/Turabian StyleSong, Yuefan, Peng He, Andre L. Rodrigues, Payel Datta, Ritesh Tandon, John T. Bates, Michael A. Bierdeman, Chen Chen, Jonathan Dordick, Fuming Zhang, and et al. 2021. "Anti-SARS-CoV-2 Activity of Rhamnan Sulfate from Monostroma nitidum" Marine Drugs 19, no. 12: 685. https://doi.org/10.3390/md19120685
APA StyleSong, Y., He, P., Rodrigues, A. L., Datta, P., Tandon, R., Bates, J. T., Bierdeman, M. A., Chen, C., Dordick, J., Zhang, F., & Linhardt, R. J. (2021). Anti-SARS-CoV-2 Activity of Rhamnan Sulfate from Monostroma nitidum. Marine Drugs, 19(12), 685. https://doi.org/10.3390/md19120685








