1. Introduction
Inflammatory bowel diseases (IBD) including ulcerative colitis (UC) and Crohn’s disease are characterised by chronic and recurrent inflammation in the gastrointestinal tract (GIT) [
1,
2,
3]. The aetiology of IBD involves a complex interaction between host genetics, lifestyle/environmental factors and the gut microbiome. The IBD patient loses immunotolerance towards enteric bacteria, resulting in uncontrolled inflammation and epithelial damage [
3,
4,
5]. The composition of the gut microbiota strongly influences the outcome of such an immune interaction, with some microbes promoting immune regulation, while other pathogenic organisms drive inflammation at stressful times in susceptible hosts [
6].
Conventional medicines used for the management and treatment of IBDs include aminosalicylates [
7], corticosteroids [
8] and immunosuppressants [
7,
8]. These medicines have variable remission responses and many create significant side effects with long-term usage [
7,
9]. Hence, there is a desire to identify natural bioactives that have anti-microbial and anti-inflammatory activities that could be used to manage IBD without negatively impacting on other aspects of health. Such bioactives would be particularly useful and desirable for patients in remission or with milder forms of disease [
10]. To effectively influence the environment of the large intestine, bioactive supplements must be resistant to digestion in the upper gastrointestinal tract. A number of bioactives, such as laminarin and chitosan, have such properties. Laminarin is a seaweed-derived polysaccharide composed of β-(1,3)-linked glucans with β-(1,6)-linked side chains of varying length and distribution [
11]. Laminarin is classified as a dietary fibre as the presence of glycosidic bonds make it resistant to hydrolytic enzymes and it is non-digestible in in-vitro studies [
12]. The anti-inflammatory potential of laminarin in the intestine of weaned pigs has been clearly identified, with reductions in the expression of pro-inflammatory cytokines [
13,
14,
15,
16] and increased mucin expression in the ileum and colon [
17,
18]. Laminarin also favourably influences microbial populations in the post-weaned pig, as evidenced by a reduction in the abundance of attaching and effacing
Escherichia coli (AEEC) strains and
Enterobacteriaceae in the large intestine [
13,
14,
16,
19]. Chitosan is another marine-derived polysaccharide that has biological characteristics that make it an interesting candidate. Chitosan is formed by the partial deacetylation of chitin, which is found in the exoskeletons of crustaceans. It comprises copolymers of glucosamine and N-acetylglucosamine [
20]. It is resistant to hydrolysis by digestive enzymes [
21]. In vitro, chitosan has had inhibitory effects against eight species of pathogenic bacteria [
22]. Recently, chitosan reduced the degree of body weight loss, disease activity index scores, histological injury and increased the expression of tight junction proteins in dextran sodium sulphate (DSS)-challenged mice [
23]. Oral administration of oligosaccharides of chitosan (chito-oligosaccharide (COS)) to mice in two DSS challenge studies also reduced the clinical signs, reduced mucosal injury, reduced NF-κB activation, supressed TNF-α and IL6 levels [
24], prevented colonic shortening and inhibited myeloperoxidase, NF-κB, COX-2, and iNOS activation [
25]. However, it is unclear if these beneficial effects observed in mice will be observed in other animal models of intestinal inflammation.
Therapeutic candidates for the treatment of IBD are often tested using animal models of intestinal inflammation including pig models as the GIT of pigs is both morphologically and physiologically similar to that of humans [
26,
27,
28]. Dextran sodium sulphate, a chemical colitogen which induces epithelial damage, is commonly used to model colitis [
29]. Whilst more frequently used in rodents, the number of studies using the DSS model in pigs is increasing. These models often examine the effects of a supplement on the clinical signs (weight loss and incidence of diarrhoea), histological features, the gene expression and protein levels of cytokines, intestinal permeability, myeloperoxidase activity and immunoglobulin concentrations [
2,
10,
26,
30,
31]. Human colitis and the DSS model share several characteristics including superficial ulceration, mucosal damage, leukocyte infiltration, the production of cytokines and other inflammatory mediators [
6]. Thus, the objective of this study was to investigate the effects of the prior consumption of laminarin or chitosan on post- DSS challenge faecal scores, daily gains, histological features, the expression of genes involved in inflammation and the diversity and composition of the colonic microbiota. Considering the previously demonstrated growth enhancing, anti-microbial and anti-inflammatory effects of laminarin and chitosan, it was hypothesised that laminarin or chitosan will remediate colitis-type epithelial damage via the divergent mechanisms including the modification of the enteric microbiota and inflammatory transcriptome in the colon, collectively improving the pathology characteristics of colitis.
3. Discussion
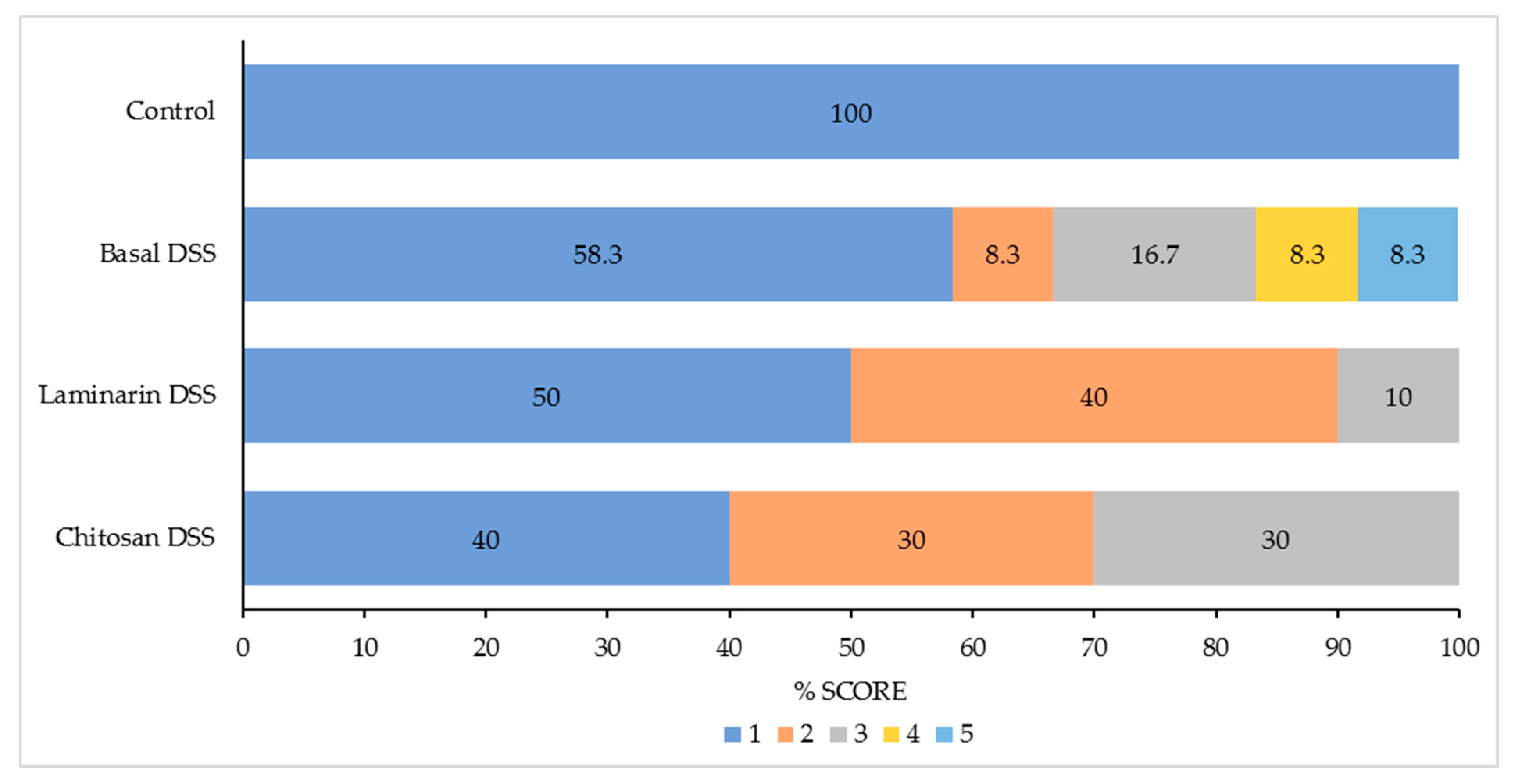
The potential of laminarin or chitosan to protect the colon from the dysbiosis, excessive inflammation and the epithelial injury that typically occurs in an acute DSS challenge model of colitis were investigated. As expected, the acute DSS challenge increased the pathology scores, as well as the markers of inflammation and tissue remodelling in the colon, while it negatively impacted growth performance, faecal scores and colonic microbiota. The supplementation of neither 200 ppm laminarin nor 300 ppm chitosan (MW = 1.526 kDa) was successful in counteracting the effects of the acute DSS challenge in terms of growth performance and inflammation, although none of the pigs in these supplemented groups had a pathology score greater than 3. The pigs supplemented with laminarin had increased molar proportions of acetate and a reduced relative abundance of members of the genus
Escherichia/Shigella, a genus which is typically increased in IBD patients [
32] and associated with severe disease [
33]. These changes suggest that laminarin has potential to control pathogen proliferation in an acute experimental pig model of colitis. However, a study of longer duration post-DSS challenge is warranted to ascertain evidence of the resolution of inflammation and tissue remodelling in the colon of the laminarin-supplemented pigs.
Dysbiosis is commonly reported in studies of IBD and is characterised by an increase in pathogenic bacteria concomitant with a reduction in beneficial bacterial species, although it is not clear whether it is a cause or consequence of inflammation [
34]. The microbial profile of the DSS-challenged pigs in this study corresponded with a broad microbial pattern identified in IBD patients, characterised by reduced bacterial diversity, the reduced abundance of bacterial taxa in the phyla Firmicutes and Bacteroidetes and increased Gammaproteobacteria (as reviewed by Zuo and Ng [
35]). In this study, basal-fed-DSS-challenged pigs had reduced alpha diversity, characterised by the reduced abundance of taxa within the phyla Firmicutes (
Roseburia, Phascolarctobacterium and Oribacterium) and Bacteroidetes (
Prevotella and
Rikenellaceae) and the increased abundance of genera within the phylum Proteobacteria (
Escherichia/Shigella and Campylobacter) in colonic digesta. Interestingly, many of the reduced bacterial taxa are associated with VFA production. Previously,
Roseburia and
Phascolarctobacterium were reduced in IBD patients [
32,
36]. The concentrations of total VFAs were unaffected by the DSS challenge, however, there was a reduction in the molar proportions of acetate and increased propionate, valerate and branched-chain fatty acids (BCFA). Intestinal concentrations of BCFA may be used as an indicator of the extent of the microbial amino acid (AA) metabolism [
37], as they are produced exclusively through the fermentation of branched-chain AA [
38]. Thus, the increase in BCFA suggests that the DSS-challenged pigs had a greater amount of undigested protein reaching the colon. In the laminarin-supplemented pigs, the molar proportions of acetate were increased, suggesting there was more substrate available for bacterial fermentation. The VFAs are essential for the growth and proliferation of colonocytes [
39], and while the majority of acetate produced is taken up by the liver and used as an energy source, it may also be used as an energy source for the colonocytes [
40]. Thus, this increase in acetate may suggest an improvement in colonic health in laminarin-supplemented pigs. During dysbiosis, facultative anaerobic Proteobacteria including the family
Enterobacteriaceae expanded at the expense of oxygen-sensitive butyrate producers, disrupting the anaerobic intestinal environment [
41,
42]. This is consistent with the reduced abundance of
Lachnospiraceae and
Roseburia and the increase in
Escherichia/Shigella in this study. In agreement with the results of this study, a DSS challenge in wild type mice also led to an increase in
Enterobacteriaceae and
Mucispirillum and a reduction in
Rikenellaceae [
42].
E. coli were previously increased in the mucosa of ulcerative colitis patients and associated with disease activity [
43,
44]. The
E. coli strains isolated from IBD patients were frequently adherent and invasive in nature and exhibited more pathogenic potential [
45,
46]. While many of the
E. coli present in the human intestine were commensals, during an IBD flare-up these strains may also proliferate and contribute to disease development [
47]. Laminarin supplementation caused a reduction in the genus
Escherichia/Shigella. Laminarin has previously been shown to reduce
Enterobacteriaceae in newly-weaned and DSS-challenged pigs [
48,
49] and to reduce
E. coli levels in weaned pigs [
13,
50], indicating that laminarin possesses anti-microbial activities against members of the
Enterobacteriaceae family. As members of the phylum Proteobacteria—particularly
E. coli—have pro-inflammatory properties and are increased in IBD patients (reviewed by [
51]), the ability of laminarin to reduce them suggests that laminarin may have potential to help maintain a healthy microbial profile in IBD patients.
A number of genes with known relevance to IBD were upregulated in the basal DSS group compared to the untreated controls. These included the cytokines
IL13 and
IL23 and the matrix metalloproteinase
MMP1. IL13 is a crucial effector cytokine acting on epithelial cell function and initiating apoptosis in UC [
52]. Previously, IL13 gene expression was elevated in the colonic mucosa of UC patients [
53]. Similarly, serum IL23 levels are increased in UC patients and are proportional to disease severity [
54]. IL23 is most highly expressed within the GIT, and it is thought to be involved in the development of a strong immune response against environmental pathogens [
55]. MMP1 is upregulated in ulcerated and inflamed areas of the colonic mucosa and its expression levels correlate with disease severity in UC patients [
56]. MMPs degrade the extracellular matrix and basement membrane, facilitating cell migration and tissue remodelling [
57]. Supplementation with laminarin or chitosan did not reduce the expression levels of the pro-inflammatory genes compared with the basal DSS group. This was surprising as both laminarin and chitosan reduced the expression of a range of pro-inflammatory cytokines in newly weaned pigs—a stressful time characterised by impaired GIT health [
13,
14,
58]. However, the ability of these bioactives to reduce inflammatory responses may vary depending on the degree of challenge to the intestine (as reviewed by Sweeney and O’Doherty [
59]). The bioactivity of chitosan varies depending on its degree of deacetylation (DDA) and molecular weight (MW) [
60]. A recent study in which chitosan improved the outcome of a DSS challenge in mice [
23] used chitosan with a greater DDA (>95% vs. 90.2%) and MW (217kDa vs. 1.526kDa) compared with the chitosan used in the present study. These authors also observed a greater response with 250 vs. 150 ppm chitosan, suggesting that a greater response may have been observed with higher inclusion levels of laminarin and chitosan. The current study was an acute DSS challenge and it is possible that the anti-inflammatory properties may have become evident in a longer-term study once the harsh DSS challenge was removed. As laminarin possesses anti-microbial activities against members of the
Enterobacteriaceae family which have pro-inflammatory activity, it would be of interest to observe the inflammatory and tissue-remodelling profile of these tissues in a longer-term study.
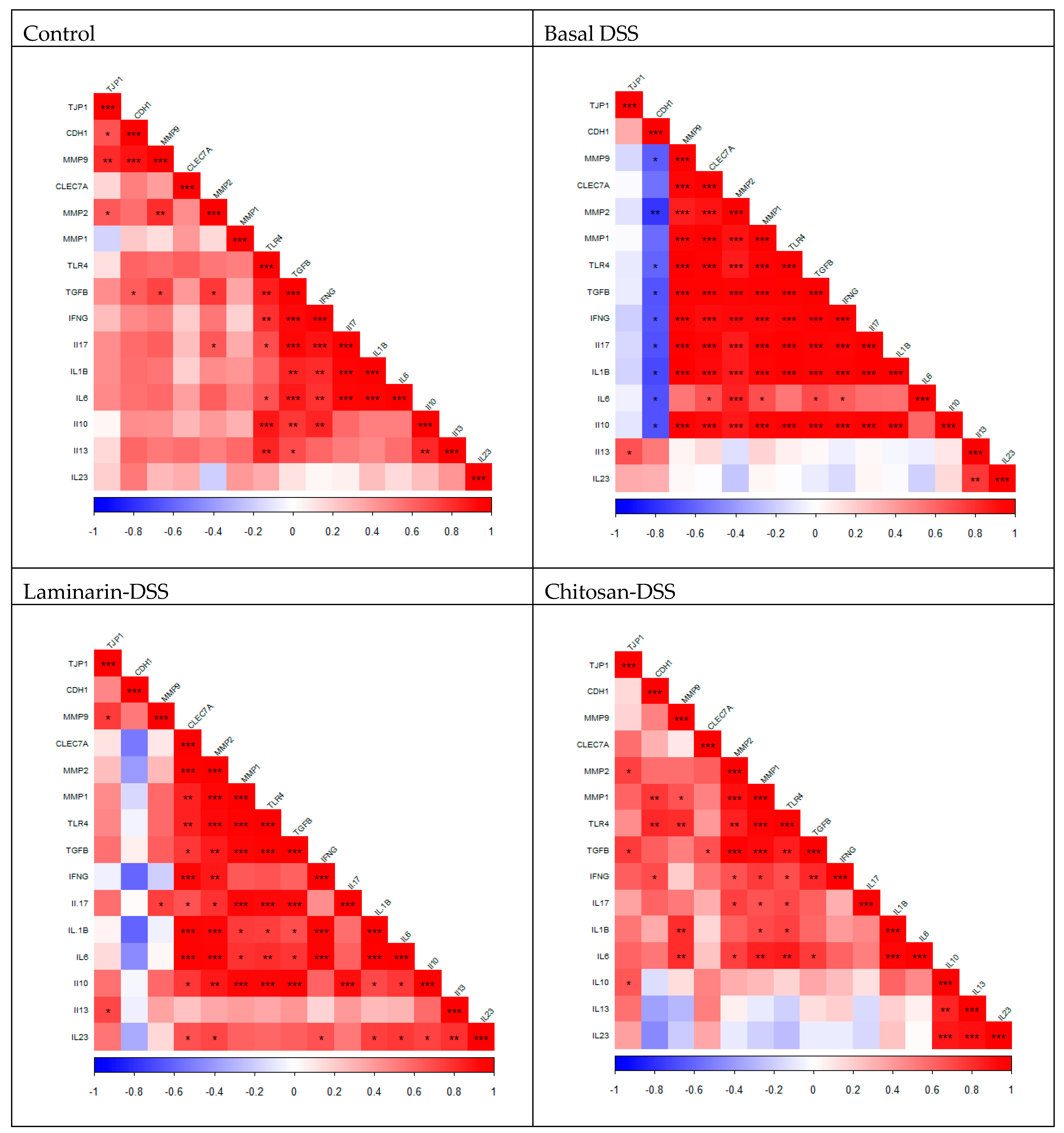
While the differences in gene expression levels between the DSS-challenged groups were not significant, it is interesting to note that there were alterations in the pattern of co-expressed genes across the treatment groups suggesting that both laminarin and chitosan were having an influence on the transcriptome. The gene
CDH1 encodes a crucial cell–cell adhesion molecule called E-cadherin that holds epithelial cells together. The expression of
CDH1 was negatively correlated with most cytokines (
IL6,
IFNG,
IL10,
IL17, and
IL1B) and MMPs (
MMP2 and
MMP9) in the basal DSS group. This is in line with other studies, where the expression of E-cadherin was downregulated in biopsy specimens from IBD patients [
61,
62,
63] and is considered a feature of the pathophysiological characteristics of Crohn’s disease, another form of IBD [
64]. In the basal DSS group,
MMP9 was positively correlated with several cytokines (
IFNG, IL10, IL17 and
IL1B) and MMPs (
MMP1 and
MMP2). In contrast to the laminarin and chitosan DSS groups, both numbers of correlations (2 and 4, respectively) and strength of the correlations between MMP9 and other genes were reduced. MMP9 has been implicated in the pathogenesis of colitis, with MMP-9
−/− mice challenged with DSS having reduced inflammation and mucosal injury, whilst in Caco2-BBE cells, MMP-9 inhibited cell attachment and wound healing [
65]. Thus, despite the absence of differences in gene expression levels, the co-expression patterns in the laminarin DSS and chitosan DSS groups differed from the basal DSS group and this may have contributed to the improved distribution of pathology scores within these groups. The signature of expression may be more important in this regard than the effects of individual genes, with the changes to the relationships between these key genes driven by regulatory elements not explored in this study.
No single model of colitis sufficiently represents the complexity of human IBDs. However, the DSS-induced colitis model is frequently used as it is a simple, inexpensive and reproducible model [
29], which can be used to study the effects of a dietary supplement on various parameters of colonic health which are affected by IBD. Although the DSS-challenge replicated several of the characteristics observed in human IBD in the current study and several others, there has been a lot of variation between the porcine DSS experiments conducted to date [
2,
10,
26,
30,
31]. Differences include the ages of the pigs used, the method of DSS administration, the dosage used and the frequency of dosing, the duration of recovery prior to slaughter and the differences in the administration of the compound being tested. The age of the pig used is an important consideration. Many studies have used very young pigs; however, the immune system of neonates is functionally different from that of adults, with the young pig being highly dependent on maternally derived passive immunity [
66]. The age at which the pig is weaned and the time between weaning and challenge is also an important consideration as weaning itself is a challenge to the GIT leading to disturbances in gastrointestinal physiology, immunology and microbiology [
67]. Thus, these factors should be considered when interpreting the findings of these porcine model studies.