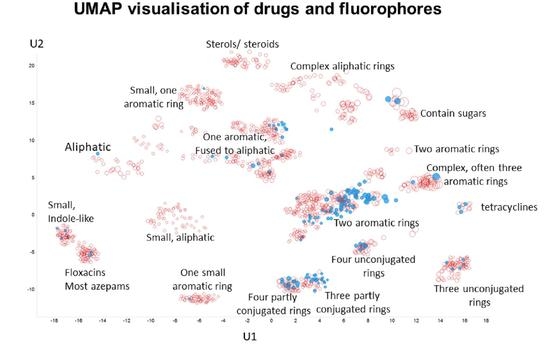
Structural Similarities between Some Common Fluorophores Used in Biology, Marketed Drugs, Endogenous Metabolites, and Natural Products
Abstract
1. Introduction
2. Results
3. Discussion
| Dye | Transporter | Comments | Reference |
|---|---|---|---|
| Amiloride | OCT2 (SLC22A2) | A drug. Rhodamine 123 and 6G also served as substrates. | [134] |
| 4′,6-diamidino-2-phenylindol (DAPI) | OCT1 (SLC22A1) | Potently inhibited by desipramine and also by various organophosphate pesticides. | [135,136] |
| DiBAC(4)3 | Na+/HCO3− NBCe1-A SLC4A4 | Competes with 4,4′-Diisothiocyanatostilbene-2,2′-disulfonic acid | [137] |
| 5-carboxyfluorescein | OAT3 (SLC22A8 | Very high Vmax | [94] |
| 6-carboxyfluorescein | OAT1 (SLC22A6) | [94,138] | |
| 2′,7′-dichlorofluorescein | OATP1B1 (SLCO1B1) | Good substrate | [139] |
| 4-(4-(Dimethylamino) styryl)-N-methylpyridinium (ASP+) | Dopamine transporter (SLC6A3) | [140,141] | |
| Noradrenaline transporter (SLC6A2) | [140,142,143] | ||
| Serotonin transporter SLC6A4 | [140,144] | ||
| Various monoamine transporters | [145] | ||
| OCT1/OCT2 (SLC22A1/2); | Seen as a model substrate | [146] | |
| Various OCT transporters | [147] | ||
| Other, unknown (non-OCT1/2) transporters with low affinity | [148] | ||
| Ethidium | OCT1/2/3 (SLC22A1/2/3) | Substrate | [149] |
| FFN511 | VMAT2 (SLC18A2) | ‘False fluorescent neurotransmitter’ (i.e., surrogate substrates) concept | [79] |
| FFN54/246 | SLC6A4, SLC18 | ‘False’ fluorescent substrates for serotonin and VMAT transporters. Potent inhibition by imipramine and citalopram | [80] |
| FFN270 | SLC6A4, SLC18 | Another example of a fluorescent false neurotransmitter | [150] |
| Fluorescein | SLCO1B1/3B1 | Effective substrate; analysis of inhibitors | [92,93] |
| OAT6 (SLC22A20) | [94] | ||
| SLC16A1, SLC16A4 | [91] | ||
| Many OATPs (SLCO family) expressed in insect cells | [90] | ||
| Lucifer yellow | Sodium-dependent anion transporters | Inhibited by probenecid | [151,152] |
| Rhodamine 123 | OCT1/OCT2 (SLC22A1/2) | Potent substrate | [153] |
| Stilbazolium dyes | Norepinephrine transporter (SLC6) | Dyes related to ASP+ | [154] |
| Zombie Violet, Live/Dead Green, Cascade Blue, Alexa Fluor 405 | OATP (SLCO) 1B1/1B3 and 2B1 | All shown to be direct substrates, and uptake inhibited by known transporter inhibitors | [155] |
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| PCA | Principal Components Analysis |
| QED | Quantitative Estimate of Drug-likeness |
| QSAR | Quantitative Structure–Activity Relationship |
| SLC | solute carrier |
| TS | Tanimoto similarity |
| UNPD | Universal Natural Products Database |
References
- Chalfie, M.; Kain, S. Green Fluorescent Protein: Properties, Applications, and Protocols; Wiley-Liss: New York, NY, USA, 1998. [Google Scholar]
- Hemmilä, I.A. Applications of Fluorescence in Immunoassays; Wiley: New York, NY, USA, 1991. [Google Scholar]
- Waggoner, A.S. Fluorescent probes for cytometry. In Flow Cytometry and Sorting, 2nd ed.; Melamed, M.R., Lindmo, T., Mendelsohn, M.L., Eds.; Wiley-Liss Inc.: New York, NY, USA, 1990; pp. 209–225. [Google Scholar]
- Kyrychenko, A. Using fluorescence for studies of biological membranes: A review. Methods Appl. Fluoresc. 2015, 3, 042003. [Google Scholar] [PubMed]
- Lagorio, M.G.; Cordon, G.B.; Iriel, A. Reviewing the relevance of fluorescence in biological systems. Photochem. Photobiol. Sci. 2015, 14, 1538–1559. [Google Scholar] [PubMed]
- Specht, E.A.; Braselmann, E.; Palmer, A.E. A critical and comparative review of fluorescent tools for live-cell imaging. Annu. Rev. Physiol. 2017, 79, 93–117. [Google Scholar] [PubMed]
- Sahl, S.J.; Hell, S.W.; Jakobs, S. Fluorescence nanoscopy in cell biology. Nat. Rev. Mol. Cell Biol. 2017, 18, 685–701. [Google Scholar] [PubMed]
- Mavrakis, M.; Pourquie, O.; Lecuit, T. Lighting up developmental mechanisms: How fluorescence imaging heralded a new era. Development 2010, 137, 373–387. [Google Scholar] [PubMed]
- Johnson, I. Fluorescent probes for living cells. Histochem. J. 1998, 30, 123–140. [Google Scholar] [PubMed]
- Kolanowski, J.L.; Liu, F.; New, E.J. Fluorescent probes for the simultaneous detection of multiple analytes in biology. Chem. Soc. Rev. 2018, 47, 195–208. [Google Scholar]
- Zhang, J.; Campbell, R.E.; Ting, A.Y.; Tsien, R.Y. Creating new fluorescent probes for cell biology. Nat. Rev. Mol. Cell Biol. 2002, 3, 906–918. [Google Scholar]
- Jiang, X.; Wang, L.; Carroll, S.L.; Chen, J.; Wang, M.C.; Wang, J. Challenges and opportunities for small-molecule fluorescent probes in redox biology applications. Antioxid. Redox Signal. 2018, 29, 518–540. [Google Scholar]
- Winterbourn, C.C. The challenges of using fluorescent probes to detect and quantify specific reactive oxygen species in living cells. Biochim. Biophys. Acta 2014, 1840, 730–738. [Google Scholar]
- Davey, H.M.; Kell, D.B. Flow cytometry and cell sorting of heterogeneous microbial populations: The importance of single-cell analysis. Microbiol. Rev. 1996, 60, 641–696. [Google Scholar] [PubMed]
- Shapiro, H.M. Practical Flow Cytometry, 3rd ed.; John Wiley: New York, NY, USA, 2003. [Google Scholar]
- Lavis, L.D.; Raines, R.T. Bright ideas for chemical biology. ACS Chem. Biol. 2008, 3, 142–155. [Google Scholar] [PubMed]
- Kell, D.B. A Protet-Based, Protonic Charge Transfer Model of Energy Coupling in Oxidative and Photosynthetic Phosphorylation. OSF Preprint. 2020. Available online: http://osf.io/2xsz8 (accessed on 23 November 2020).
- Salcedo-Sora, J.E.; Jindal, S.; O’Hagan, S.; Kell, D.B. A palette of fluorophores that are differentially accumulated by wild-type and mutant strains of Escherichia coli: Surrogate ligands for bacterial membrane transporters. bioRxiv 2020. [Google Scholar] [CrossRef]
- Karasev, M.M.; Stepanenko, O.V.; Rumyantsev, K.A.; Turoverov, K.K.; Verkhusha, V.V. Near-infrared fluorescent proteins and their applications. Biochemistry (Moscow) 2019, 84, S32–S50. [Google Scholar]
- Liu, W.F.; Deng, M.Y.; Yang, C.M.; Liu, F.; Guan, X.M.; Du, Y.C.; Wang, L.; Chu, J. Genetically encoded single circularly permuted fluorescent protein-based intensity indicators. J. Phys. D 2020, 53, 113001. [Google Scholar]
- Pedelacq, J.D.; Cabantous, S. Development and applications of superfolder and split fluorescent protein detection systems in biology. Int. J. Mol. Sci. 2019, 20, 3479. [Google Scholar]
- Shcherbakova, D.M.; Stepanenko, O.V.; Turoverov, K.K.; Verkhusha, V.V. Near-infrared fluorescent proteins: Multiplexing and optogenetics across scales. Trends Biotechnol. 2018, 36, 1230–1243. [Google Scholar]
- Jindal, S.; Yang, L.; Day, P.J.; Kell, D.B. Involvement of multiple influx and efflux transporters in the accumulation of cationic fluorescent dyes by Escherichia coli. BMC Microbiol. 2019, 19, 195. [Google Scholar]
- Kell, D.B. Control of metabolite efflux in microbial cell factories: Current advances and future prospects. In Fermentation Microbiology and Biotechnology, 4th ed.; El-Mansi, E.M.T., Nielsen, J., Mousdale, D., Allman, T., Carlson, R., Eds.; CRC Press: Boca Raton, FL, USA, 2019; pp. 117–138. [Google Scholar]
- Szakács, G.; Váradi, A.; Özvegy-Laczka, C.; Sarkadi, B. The role of ABC transporters in drug absorption, distribution, metabolism, excretion and toxicity (ADME-Tox). Drug Discov. Today 2008, 13, 379–393. [Google Scholar]
- Schinkel, A.H.; Jonker, J.W. Mammalian drug efflux transporters of the ATP binding cassette (ABC) family: An overview. Adv. Drug Deliv. Rev. 2012, 64, 138–153. [Google Scholar]
- Tarling, E.J.; de Aguiar Vallim, T.Q.; Edwards, P.A. Role of ABC transporters in lipid transport and human disease. Trends Endocrinol. Metab. 2013, 24, 342–350. [Google Scholar] [PubMed]
- Andreoletti, P.; Raas, Q.; Gondcaille, C.; Cherkaoui-Malki, M.; Trompier, D.; Savary, S. Predictive structure and topology of peroxisomal ATP-binding cassette (ABC) transporters. Int. J. Mol. Sci. 2017, 18, 1593. [Google Scholar]
- Sharom, F.J. ABC multidrug transporters: Structure, function and role in chemoresistance. Pharmacogenomics 2008, 9, 105–127. [Google Scholar]
- Pahnke, J.; Fröhlich, C.; Paarmann, K.; Krohn, M.; Bogdanovic, N.; Årsland, D.; Winblad, B. Cerebral ABC transporter-common mechanisms may modulate neurodegenerative diseases and depression in elderly subjects. Arch. Med. Res. 2014, 45, 738–743. [Google Scholar] [PubMed]
- Abuznait, A.H.; Kaddoumi, A. Role of ABC transporters in the pathogenesis of alzheimer’s disease. ACS Chem. Neurosci. 2012, 3, 820–831. [Google Scholar] [PubMed]
- Lam, F.C.; Liu, R.; Lu, P.; Shapiro, A.B.; Renoir, J.M.; Sharom, F.J.; Reiner, P.B. Beta-amyloid efflux mediated by P-glycoprotein. J. Neurochem. 2001, 76, 1121–1128. [Google Scholar] [PubMed]
- Molnar, J.; Ocsovszki, I.; Pusztai, R. Amyloid-beta interactions with ABC transporters and resistance modifiers. Anticancer Res. 2018, 38, 3407–3410. [Google Scholar]
- Wijnholds, J.; Evers, R.; van Leusden, M.R.; Mol, C.A.A.M.; Zaman, G.J.R.; Mayer, U.; Beijnen, J.H.; van der Valk, M.; Krimpenfort, P.; Borst, P. Increased sensitivity to anticancer drugs and decreased inflammatory response in mice lacking the multidrug resistance-associated protein. Nat. Med. 1997, 3, 1275–1279. [Google Scholar]
- Prates, R.A.; Kato, I.T.; Ribeiro, M.S.; Tegos, G.P.; Hamblin, M.R. Influence of multidrug efflux systems on methylene blue-mediated photodynamic inactivation of Candida albicans. J. Antimicrob. Chemother. 2011, 66, 1525–1532. [Google Scholar] [PubMed]
- Forster, S.; Thumser, A.E.; Hood, S.R.; Plant, N. Characterization of rhodamine-123 as a tracer dye for use in in vitro drug transport assays. PLoS ONE 2012, 7, e33253. [Google Scholar]
- Ivnitski-Steele, I.; Holmes, A.R.; Lamping, E.; Monk, B.C.; Cannon, R.D.; Sklar, L.A. Identification of nile red as a fluorescent substrate of the Candida albicans ATP-binding cassette transporters cdr1p and cdr2p and the major facilitator superfamily transporter mdr1p. Anal. Biochem. 2009, 394, 87–91. [Google Scholar] [PubMed]
- Strouse, J.J.; Ivnitski-Steele, I.; Waller, A.; Young, S.M.; Perez, D.; Evangelisti, A.M.; Ursu, O.; Bologa, C.G.; Carter, M.B.; Salas, V.M.; et al. Fluorescent substrates for flow cytometric evaluation of efflux inhibition in ABCB1, ABCC1, and ABCG2 transporters. Anal. Biochem. 2013, 437, 77–87. [Google Scholar] [PubMed]
- Kell, D.B. Control of Metabolite Efflux in Microbial Cell Factories: Current Advances and Future Prospects. Available online: https://osf.io/xg9jh/download?format=pdf (accessed on 8 November 2020).
- Szabó, E.; Türk, D.; Telbisz, Á.; Kucsma, N.; Horváth, T.; Szakács, G.; Homolya, L.; Sarkadi, B.; Várady, G. A new fluorescent dye accumulation assay for parallel measurements of the ABCG2, ABCB1 and ABCC1 multidrug transporter functions. PLoS ONE 2018, 13, e0190629. [Google Scholar]
- Gökirmak, T.; Shipp, L.E.; Campanale, J.P.; Nicklisch, S.C.T.; Hamdoun, A. Transport in technicolor: Mapping ATP-binding cassette transporters in sea urchin embryos. Mol. Reprod Dev. 2014, 81, 778–793. [Google Scholar]
- Fardel, O.; Le Vee, M.; Jouan, E.; Denizot, C.; Parmentier, Y. Nature and uses of fluorescent dyes for drug transporter studies. Expert Opin. Drug Metab. Toxicol. 2015, 11, 1233–1251. [Google Scholar]
- Fredriksson, R.; Nordström, K.J.; Stephansson, O.; Hägglund, M.G.; Schiöth, H.B. The solute carrier (SLC) complement of the human genome: Phylogenetic classification reveals four major families. FEBS Lett. 2008, 582, 3811–3816. [Google Scholar]
- Darbani, B.; Kell, D.B.; Borodina, I. Energetic evolution of cellular transportomes. BMC Genom. 2018, 19, 418. [Google Scholar]
- O’Hagan, S.; Swainston, N.; Handl, J.; Kell, D.B. A ‘rule of 0.5’ for the metabolite-likeness of approved pharmaceutical drugs. Metabolomics 2015, 11, 323–339. [Google Scholar]
- O’Hagan, S.; Kell, D.B. Understanding the foundations of the structural similarities between marketed drugs and endogenous human metabolites. Front. Pharmacol. 2015, 6, 105. [Google Scholar]
- O’Hagan, S.; Kell, D.B. Metmaxstruct: A Tversky-similarity-based strategy for analysing the (sub)structural similarities of drugs and endogenous metabolites. Front. Pharmacol. 2016, 7, 266. [Google Scholar]
- O’Hagan, S.; Kell, D.B. Analysis of drug-endogenous human metabolite similarities in terms of their maximum common substructures. J. Cheminform. 2017, 9, 18. [Google Scholar] [PubMed]
- Nigam, S.K. What do drug transporters really do? Nat. Rev. Drug Discov. 2015, 14, 29–44. [Google Scholar] [PubMed]
- Girardi, E.; César-Razquin, A.; Lindinger, S.; Papakostas, K.; Lindinger, S.; Konecka, J.; Hemmerich, J.; Kickinger, S.; Kartnig, F.; Gürtl, B.; et al. A widespread role for SLC transmembrane transporters in resistance to cytotoxic drugs. Nat. Chem. Biol. 2020, 16, 469–478. [Google Scholar] [PubMed]
- Pizzagalli, M.D.; Bensimon, A.; Superti-Furga, G. A guide to plasma membrane solute carrier proteins. FEBS J. 2020. [Google Scholar] [CrossRef]
- Superti-Furga, G.; Lackner, D.; Wiedmer, T.; Ingles-Prieto, A.; Barbosa, B.; Girardi, E.; Goldman, U.; Gürtl, B.; Klavins, K.; Klimek, C. The RESOLUTE consortium: Unlocking SLC transporters for drug discovery. Nat. Rev. Drug Discov. 2020, 19, 429–430. [Google Scholar]
- Kell, D.B. Hitchhiking into the cell. Nat. Chem. Biol. 2020, 16, 367–368. [Google Scholar]
- Gründemann, D.; Harlfinger, S.; Golz, S.; Geerts, A.; Lazar, A.; Berkels, R.; Jung, N.; Rubbert, A.; Schömig, E. Discovery of the ergothioneine transporter. Proc. Natl. Acad. Sci. USA 2005, 102, 5256–5261. [Google Scholar]
- O’Hagan, S.; Kell, D.B. Consensus rank orderings of molecular fingerprints illustrate the ‘most genuine’ similarities between marketed drugs and small endogenous human metabolites, but highlight exogenous natural products as the most important ‘natural’ drug transporter substrates. ADMET DMPK 2017, 5, 85–125. [Google Scholar]
- Gautier, J.; Munnier, E.; Souce, M.; Chourpa, I.; Douziech Eyrolles, L. Analysis of doxorubicin distribution in mcf-7 cells treated with drug-loaded nanoparticles by combination of two fluorescence-based techniques, confocal spectral imaging and capillary electrophoresis. Anal. Bioanal. Chem. 2015, 407, 3425–3435. [Google Scholar]
- Khader, H.; Solodushko, V.; Al-Mehdi, A.B.; Audia, J.; Fouty, B. Overlap of doxycycline fluorescence with that of the redox-sensitive intracellular reporter roGFP. J. Fluoresc. 2014, 24, 305–311. [Google Scholar]
- Motlagh, N.S.H.; Parvin, P.; Ghasemi, F.; Atyabi, F. Fluorescence properties of several chemotherapy drugs: Doxorubicin, paclitaxel and bleomycin. Biomed. Opt. Express 2016, 7, 2400–2406. [Google Scholar] [PubMed]
- Baldini, G.; Doglia, S.; Dolci, S.; Sassi, G. Fluorescence-determined preferential binding of quinacrine to DNA. Biophys. J. 1981, 36, 465–477. [Google Scholar] [PubMed]
- Nguyen, M.; Marcellus, R.C.; Roulston, A.; Watson, M.; Serfass, L.; Murthy Madiraju, S.R.; Goulet, D.; Viallet, J.; Belec, L.; Billot, X.; et al. Small molecule obatoclax (GX15-070) antagonizes MCL-1 and overcomes MCL-1-mediated resistance to apoptosis. Proc. Natl. Acad. Sci. USA 2007, 104, 19512–19517. [Google Scholar] [PubMed]
- Stamelos, V.A.; Fisher, N.; Bamrah, H.; Voisey, C.; Price, J.C.; Farrell, W.E.; Redman, C.W.; Richardson, A. The bh3 mimetic obatoclax accumulates in lysosomes and causes their alkalinization. PLoS ONE 2016, 11, e0150696. [Google Scholar]
- Pautke, C.; Vogt, S.; Kreutzer, K.; Haczek, C.; Wexel, G.; Kolk, A.; Imhoff, A.B.; Zitzelsberger, H.; Milz, S.; Tischer, T. Characterization of eight different tetracyclines: Advances in fluorescence bone labeling. J. Anat. 2010, 217, 76–82. [Google Scholar]
- Burke, T.G.; Malak, H.; Gryczynski, I.; Mi, Z.; Lakowicz, J.R. Fluorescence detection of the anticancer drug topotecan in plasma and whole blood by two-photon excitation. Anal. Biochem. 1996, 242, 266–270. [Google Scholar] [PubMed]
- Yonezawa, A.; Inui, K. Novel riboflavin transporter family rfvt/slc52: Identification, nomenclature, functional characterization and genetic diseases of rfvt/slc52. Mol. Aspects Med. 2013, 34, 693–701. [Google Scholar]
- Zhang, S.; Sakuma, M.; Deora, G.S.; Levy, C.W.; Klausing, A.; Breda, C.; Read, K.D.; Edlin, C.D.; Ross, B.P.; Wright Muelas, M.; et al. A brain-permeable inhibitor of the neurodegenerative disease target kynurenine 3-monooxygenase prevents accumulation of neurotoxic metabolites. Commun. Biol. 2019, 2, 271. [Google Scholar]
- Liu, Q.; Liu, Y.; Guo, M.; Luo, X.; Yao, S. A simple and sensitive method of nonaqueous capillary electrophoresis with laser-induced native fluorescence detection for the analysis of chelerythrine and sanguinarine in chinese herbal medicines. Talanta 2006, 70, 202–207. [Google Scholar]
- Duval, R.; Duplais, C. Fluorescent natural products as probes and tracers in biology. Nat. Prod. Rep. 2017, 34, 161–193. [Google Scholar]
- Taniguchi, M.; Lindsey, J.S. Database of absorption and fluorescence spectra of >300 common compounds for use in photochemcad. Photochem. Photobiol. 2018, 94, 290–327. [Google Scholar] [PubMed]
- Li, X.Z.; Plésiat, P.; Nikaido, H. The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin. Microbiol. Rev. 2015, 28, 337–418. [Google Scholar] [PubMed]
- Holmes, A.H.; Moore, L.S.P.; Sundsfjord, A.; Steinbakk, M.; Regmi, S.; Karkey, A.; Guerin, P.J.; Piddock, L.J.V. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 2016, 387, 176–187. [Google Scholar]
- Salcedo-Sora, J.E.; Kell, D.B. A quantitative survey of bacterial persistence in the presence of antibiotics: Towards antipersister antimicrobial discovery. Antibiotics 2020, 9, 508. [Google Scholar]
- Munita, J.M.; Arias, C.A. Mechanisms of antibiotic resistance. Microbiol. Spectrum. 2016, 4, VMBF-0016-2015. [Google Scholar]
- Saleem, M.; Nazir, M.; Ali, M.S.; Hussain, H.; Lee, Y.S.; Riaz, N.; Jabbar, A. Antimicrobial natural products: An update on future antibiotic drug candidates. Nat. Prod. Rep. 2010, 27, 238–254. [Google Scholar]
- Roemer, T.; Davies, J.; Giaever, G.; Nislow, C. Bugs, drugs and chemical genomics. Nat. Chem. Biol. 2012, 8, 46–56. [Google Scholar]
- Gyawali, R.; Ibrahim, S.A. Natural products as antimicrobial agents. Food Control 2014, 46, 412–429. [Google Scholar]
- Harvey, A.L.; Edrada-Ebel, R.; Quinn, R.J. The re-emergence of natural products for drug discovery in the genomics era. Nat. Rev. Drug Discov. 2015, 14, 111–129. [Google Scholar]
- Moloney, M.G. Natural products as a source for novel antibiotics. Trends Pharmacol. Sci. 2016, 37, 689–701. [Google Scholar]
- Wright, G.D. Opportunities for natural products in 21st century antibiotic discovery. Nat. Prod. Rep. 2017, 34, 694–701. [Google Scholar] [PubMed]
- Gubernator, N.G.; Zhang, H.; Staal, R.G.; Mosharov, E.V.; Pereira, D.B.; Yue, M.; Balsanek, V.; Vadola, P.A.; Mukherjee, B.; Edwards, R.H.; et al. Fluorescent false neurotransmitters visualize dopamine release from individual presynaptic terminals. Science 2009, 324, 1441–1444. [Google Scholar] [PubMed]
- Henke, A.; Kovalyova, Y.; Dunn, M.; Dreier, D.; Gubernator, N.G.; Dincheva, I.; Hwu, C.; Sebej, P.; Ansorge, M.S.; Sulzer, D.; et al. Toward serotonin fluorescent false neurotransmitters: Development of fluorescent dual serotonin and vesicular monoamine transporter substrates for visualizing serotonin neurons. ACS Chem. Neurosci. 2018, 9, 925–934. [Google Scholar] [PubMed]
- O’Hagan, S.; Kell, D.B. Structural similarities between some common fluorophores used in biology and marketed drugs, endogenous metabolites, and natural products. bioRxiv 2019, 2019, 834325. [Google Scholar]
- van der Maaten, L.; Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Thiele, I.; Heinken, A.; Fleming, R.M. A systems biology approach to studying the role of microbes in human health. Curr. Opin. Biotechnol. 2013, 24, 4–12. [Google Scholar]
- O’Hagan, S.; Kell, D.B. Analysing and navigating natural products space for generating small, diverse, but representative chemical libraries. Biotechnol. J. 2018, 13, 1700503. [Google Scholar]
- Gu, J.Y.; Gui, Y.S.; Chen, L.R.; Yuan, G.; Lu, H.Z.; Xu, X.J. Use of natural products as chemical library for drug discovery and network pharmacology. PLoS ONE 2013, 8, e62839. [Google Scholar]
- Eisen, M.B.; Spellman, P.T.; Brown, P.O.; Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 1998, 95, 14863–14868. [Google Scholar]
- Thiele, I.; Swainston, N.; Fleming, R.M.T.; Hoppe, A.; Sahoo, S.; Aurich, M.K.; Haraldsdottír, H.; Mo, M.L.; Rolfsson, O.; Stobbe, M.D.; et al. A community-driven global reconstruction of human metabolism. Nat. Biotechnol. 2013, 31, 419–425. [Google Scholar]
- Bakkar, M.M.; Hardaker, L.; March, P.; Morgan, P.B.; Maldonado-Codina, C.; Dobson, C.B. The cellular basis for biocide-induced fluorescein hyperfluorescence in mammalian cell culture. PLoS ONE 2014, 9, e84427. [Google Scholar]
- Khan, T.F.; Price, B.L.; Morgan, P.B.; Maldonado-Codina, C.; Dobson, C.B. Cellular fluorescein hyperfluorescence is dynamin-dependent and increased by tetronic 1107 treatment. Int. J. Biochem. Cell Biol. 2018, 101, 54–63. [Google Scholar] [PubMed]
- Patik, I.; Kovacsics, D.; Német, O.; Gera, M.; Várady, G.; Stieger, B.; Hagenbuch, B.; Szakács, G.; Özvegy-Laczka, C. Functional expression of the 11 human organic anion transporting polypeptides in insect cells reveals that sodium fluorescein is a general OATP substrate. Biochem. Pharmacol. 2015, 98, 649–658. [Google Scholar] [PubMed]
- Sun, Y.C.; Liou, H.M.; Yeh, P.T.; Chen, W.L.; Hu, F.R. Monocarboxylate transporters mediate fluorescein uptake in corneal epithelial cells. Investig. Ophthalmol. Vis. Sci. 2017, 58, 3716–3722. [Google Scholar]
- De Bruyn, T.; Fattah, S.; Stieger, B.; Augustijns, P.; Annaert, P. Sodium fluorescein is a probe substrate for hepatic drug transport mediated by OATP1B1 and OATP1B3. J. Pharm. Sci. 2011, 100, 5018–5030. [Google Scholar]
- De Bruyn, T.; van Westen, G.J.; Ijzerman, A.P.; Stieger, B.; de Witte, P.; Augustijns, P.F.; Annaert, P.P. Structure-based identification of OATP1B1/3 inhibitors. Mol. Pharmacol. 2013, 83, 1257–1267. [Google Scholar]
- Truong, D.M.; Kaler, G.; Khandelwal, A.; Swaan, P.W.; Nigam, S.K. Multi-level analysis of organic anion transporters 1, 3, and 6 reveals major differences in structural determinants of antiviral discrimination. J. Biol. Chem. 2008, 283, 8654–8663. [Google Scholar]
- Patil, S.A.; Moss, A.C. Balsalazide disodium for the treatment of ulcerative colitis. Expert Rev. Gastroenterol. Hepatol. 2008, 2, 177–184. [Google Scholar]
- Egesel, T.; Ünsal, I.; Dikmen, G.; Bayraktar, Y. The assessment of pancreatic exocrine function by bentiromide test in patients with chronic portal vein thrombosis. Pancreas 2002, 25, 355–359. [Google Scholar]
- Singal, A. Butenafine and superficial mycoses: Current status. Expert Opin. Drug Metab. Toxicol 2008, 4, 999–1005. [Google Scholar]
- Blair, H.A. Tolvaptan: A review in autosomal dominant polycystic kidney disease. Drugs 2019, 79, 303–313. [Google Scholar] [PubMed]
- Brown, N.; Jacoby, E. On scaffolds and hopping in medicinal chemistry. Mini Rev. Med. Chem. 2006, 6, 1217–1229. [Google Scholar] [PubMed]
- Geppert, H.; Bajorath, J. Advances in 2d fingerprint similarity searching. Expert Opin. Drug Discov. 2010, 5, 529–542. [Google Scholar] [PubMed]
- Lamberth, C. Agrochemical lead optimization by scaffold hopping. Pest Manag. Sci. 2018, 74, 282–292. [Google Scholar] [PubMed]
- Mauser, H.; Guba, W. Recent developments in de novo design and scaffold hopping. Curr. Opin. Drug Discov. Dev. 2008, 11, 365–374. [Google Scholar]
- Sun, H.; Tawa, G.; Wallqvist, A. Classification of scaffold-hopping approaches. Drug Discov. Today 2012, 17, 310–324. [Google Scholar]
- Zhao, H. Scaffold selection and scaffold hopping in lead generation: A medicinal chemistry perspective. Drug Disc. Today 2007, 12, 149–155. [Google Scholar]
- Das, A.M. Clinical utility of nitisinone for the treatment of hereditary tyrosinemia type-1 (ht-1). Appl. Clin. Genet 2017, 10, 43–48. [Google Scholar]
- Lock, E.; Ranganath, L.R.; Timmis, O. The role of nitisinone in tyrosine pathway disorders. Curr. Rheumatol. Rep. 2014, 16, 457. [Google Scholar]
- Ranganath, L.R.; Khedr, M.; Milan, A.M.; Davison, A.S.; Hughes, A.T.; Usher, J.L.; Taylor, S.; Loftus, N.; Daroszewska, A.; West, E.; et al. Nitisinone arrests ochronosis and decreases rate of progression of alkaptonuria: Evaluation of the effect of nitisinone in the United Kingdom national alkaptonuria centre. Mol. Genet Metab. 2018, 125, 127–134. [Google Scholar]
- Bickerton, G.R.; Paolini, G.V.; Besnard, J.; Muresan, S.; Hopkins, A.L. Quantifying the chemical beauty of drugs. Nat. Chem. 2012, 4, 90–98. [Google Scholar] [PubMed]
- Shultz, M.D. Two decades under the influence of the rule of five and the changing properties of approved oral drugs. J. Med. Chem. 2019, 62, 1701–1714. [Google Scholar] [PubMed]
- Lovering, F.; Bikker, J.; Humblet, C. Escape from flatland: Increasing saturation as an approach to improving clinical success. J. Med. Chem. 2009, 52, 6752–6756. [Google Scholar] [PubMed]
- Meyers, J.; Carter, M.; Mok, N.Y.; Brown, N. On the origins of three-dimensionality in drug-like molecules. Future Med. Chem. 2016, 8, 1753–1767. [Google Scholar]
- Campbell, P.S.; Jamieson, C.; Simpson, I.; Watson, A.J.B. Practical synthesis of pharmaceutically relevant molecules enriched in sp(3) character. Chem. Commun. (Camb.) 2017, 54, 46–49. [Google Scholar]
- Blakemore, D.C.; Castro, L.; Churcher, I.; Rees, D.C.; Thomas, A.W.; Wilson, D.M.; Wood, A. Organic synthesis provides opportunities to transform drug discovery. Nat. Chem. 2018, 10, 383–394. [Google Scholar]
- Boström, J.; Brown, D.G.; Young, R.J.; Keserü, G.M. Expanding the medicinal chemistry synthetic toolbox. Nat. Rev. Drug Discov. 2018, 17, 709–727. [Google Scholar]
- Yuan, L.; Lin, W.; Zheng, K.; He, L.; Huang, W. Far-red to near infrared analyte-responsive fluorescent probes based on organic fluorophore platforms for fluorescence imaging. Chem. Soc. Rev. 2013, 42, 622–661. [Google Scholar]
- McInnes, L.; Healy, J.; Melville, J. UMAP: Uniform manifold approximation and projection for dimension reduction. arXiv 2018, arXiv:1802.03426. [Google Scholar]
- McInnes, L.; Healy, J.; Saul, N.; Großberger, L. Umap: Uniform manifold approximation and projection. J. Open Source Softw. 2018. [Google Scholar] [CrossRef]
- Koehn, F.E.; Carter, G.T. The evolving role of natural products in drug discovery. Nat. Rev. Drug Discov. 2005, 4, 206–220. [Google Scholar] [PubMed]
- Dobson, P.D.; Patel, Y.; Kell, D.B. “Metabolite-likeness” as a criterion in the design and selection of pharmaceutical drug libraries. Drug Disc. Today 2009, 14, 31–40. [Google Scholar]
- Doak, B.C.; Over, B.; Giordanetto, F.; Kihlberg, J. Oral druggable space beyond the rule of 5: Insights from drugs and clinical candidates. Chem. Biol. 2014, 21, 1115–1142. [Google Scholar] [PubMed]
- Quinn, R.J.; Carroll, A.R.; Pham, N.B.; Baron, P.; Palframan, M.E.; Suraweera, L.; Pierens, G.K.; Muresan, S. Developing a drug-like natural product library. J. Nat. Prod. 2008, 71, 464–468. [Google Scholar]
- Krämer, S.D.; Aschmann, H.E.; Hatibovic, M.; Hermann, K.F.; Neuhaus, C.S.; Brunner, C.; Belli, S. When barriers ignore the “rule-of-five”. Adv. Drug Deliv. Rev. 2016, 101, 62–74. [Google Scholar]
- Guimarães, C.R.W.; Mathiowetz, A.M.; Shalaeva, M.; Goetz, G.; Liras, S. Use of 3d properties to characterize beyond rule-of-5 property space for passive permeation. J. Chem. Inf. Model. 2012, 52, 882–890. [Google Scholar]
- Leeson, P.D. Molecular inflation, attrition and the rule of five. Adv. Drug Deliv. Rev. 2016, 101, 22–33. [Google Scholar]
- DeGoey, D.A.; Chen, H.J.; Cox, P.B.; Wendt, M.D. Beyond the rule of 5: Lessons learned from abbvie’s drugs and compound collection. J. Med. Chem. 2018, 61, 2636–2651. [Google Scholar]
- Chen, Y.; Garcia de Lomana, M.; Friedrich, N.O.; Kirchmair, J. Characterization of the chemical space of known and readily obtainable natural products. J. Chem. Inf. Model. 2018, 58, 1518–1532. [Google Scholar]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J. Nat. Prod. 2020, 83, 770–803. [Google Scholar]
- Ivnitski-Steele, I.; Larson, R.S.; Lovato, D.M.; Khawaja, H.M.; Winter, S.S.; Oprea, T.I.; Sklar, L.A.; Edwards, B.S. High-throughput flow cytometry to detect selective inhibitors of ABCB1, ABCC1, and ABCG2 transporters. Assay Drug Dev. Technol. 2008, 6, 263–276. [Google Scholar] [PubMed]
- Tegos, G.P.; Evangelisti, A.M.; Strouse, J.J.; Ursu, O.; Bologa, C.; Sklar, L.A. A high throughput flow cytometric assay platform targeting transporter inhibition. Drug Disc. Today Technol. 2014, 12, e95–e103. [Google Scholar]
- Windt, T.; Tóth, S.; Patik, I.; Sessler, J.; Kucsma, N.; Szepesi, A.; Zdrazil, B.; Özvegy-Laczka, C.; Szakács, G. Identification of anticancer OATP2B1 substrates by an in vitro triple-fluorescence-based cytotoxicity screen. Arch. Toxicol. 2019, 93, 953–964. [Google Scholar] [PubMed]
- Bednarczyk, D.; Mash, E.A.; Aavula, B.R.; Wright, S.H. Nbd-tma: A novel fluorescent substrate of the peritubular organic cation transporter of renal proximal tubules. Pflügers Arch. 2000, 440, 184–192. [Google Scholar]
- Stone, M.R.L.; Butler, M.S.; Phetsang, W.; Cooper, M.A.; Blaskovich, M.A.T. Fluorescent antibiotics: New research tools to fight antibiotic resistance. Trends Biotechnol. 2018, 36, 523–536. [Google Scholar]
- Jiang, M.; Li, H.; Johnson, A.; Karasawa, T.; Zhang, Y.; Meier, W.B.; Taghizadeh, F.; Kachelmeier, A.; Steyger, P.S. Inflammation up-regulates cochlear expression of TRPV1 to potentiate drug-induced hearing loss. Sci. Adv. 2019, 5, eaaw1836. [Google Scholar]
- Ugwu, M.C.; Pelis, R.; Esimone, C.O.; Agu, R.U. Fluorescent organic cations for human OCT2 transporters screening: Uptake in CHO cells stably expressing hoct2. ADMET DMPK 2017, 5, 135–145. [Google Scholar]
- Yasujima, T.; Ohta, K.; Inoue, K.; Yuasa, H. Characterization of human OCT1-mediated transport of dapi as a fluorescent probe substrate. J. Pharm. Sci. 2011, 100, 4006–4012. [Google Scholar]
- Chedik, L.; Bruyere, A.; Fardel, O. Interactions of organophosphorus pesticides with solute carrier (SLC) drug transporters. Xenobiotica 2018, 49, 363–374. [Google Scholar] [CrossRef]
- Liu, X.; Williams, J.B.; Sumpter, B.R.; Bevensee, M.O. Inhibition of the Na/bicarbonate cotransporter NBCE1-A by DIBAC oxonol dyes relative to niflumic acid and a stilbene. J. Membr. Biol. 2007, 215, 195–204. [Google Scholar]
- Zou, L.; Stecula, A.; Gupta, A.; Prasad, B.; Chien, H.C.; Yee, S.W.; Wang, L.; Unadkat, J.D.; Stahl, S.H.; Fenner, K.S.; et al. Molecular mechanisms for species differences in organic anion transporter 1, oat1: Implications for renal drug toxicity. Mol. Pharmacol. 2018, 94, 689–699. [Google Scholar] [PubMed]
- Izumi, S.; Nozaki, Y.; Komori, T.; Takenaka, O.; Maeda, K.; Kusuhara, H.; Sugiyama, Y. Investigation of fluorescein derivatives as substrates of organic anion transporting polypeptide (OATP) 1B1 to develop sensitive fluorescence-based OATP1B1 inhibition assays. Mol. Pharm. 2016, 13, 438–448. [Google Scholar] [PubMed]
- Schwartz, J.W.; Blakely, R.D.; DeFelice, L.J. Binding and transport in norepinephrine transporters. Real-time, spatially resolved analysis in single cells using a fluorescent substrate. J. Biol. Chem. 2003, 278, 9768–9777. [Google Scholar]
- Inyushin, M.U.; Arencibia-Albite, F.; de la Cruz, A.; Vazquez-Torres, R.; Colon, K.; Sanabria, P.; Jimenez-Rivera, C.A. New method to visualize neurons with dat in slices of rat vta using fluorescent substrate for DAT, ASP+. J. Neurosci. Neuroeng 2013, 2, 98–103. [Google Scholar] [PubMed]
- Schwartz, J.W.; Novarino, G.; Piston, D.W.; DeFelice, L.J. Substrate binding stoichiometry and kinetics of the norepinephrine transporter. J. Biol. Chem. 2005, 280, 19177–19184. [Google Scholar]
- Haunsø, A.; Buchanan, D. Pharmacological characterization of a fluorescent uptake assay for the noradrenaline transporter. J. Biomol. Screen 2007, 12, 378–384. [Google Scholar]
- Oz, M.; Libby, T.; Kivell, B.; Jaligam, V.; Ramamoorthy, S.; Shippenberg, T.S. Real-time, spatially resolved analysis of serotonin transporter activity and regulation using the fluorescent substrate, ASP+. J. Neurochem. 2010, 114, 1019–1029. [Google Scholar]
- Zwartsen, A.; Verboven, A.H.A.; van Kleef, R.; Wijnolts, F.M.J.; Westerink, R.H.S.; Hondebrink, L. Measuring inhibition of monoamine reuptake transporters by new psychoactive substances (NPS) in real-time using a high-throughput, fluorescence-based assay. Toxicol In Vitro 2017, 45, 60–71. [Google Scholar]
- Kido, Y.; Matsson, P.; Giacomini, K.M. Profiling of a prescription drug library for potential renal drug-drug interactions mediated by the organic cation transporter 2. J. Med. Chem. 2011, 54, 4548–4558. [Google Scholar]
- Salomon, J.J.; Endter, S.; Tachon, G.; Falson, F.; Buckley, S.T.; Ehrhardt, C. Transport of the fluorescent organic cation 4-(4-(dimethylamino)styryl)-n-methylpyridinium iodide (ASP+) in human respiratory epithelial cells. Eur. J. Pharm. Biopharm. 2012, 81, 351–359. [Google Scholar]
- Rytting, E.; Bryan, J.; Southard, M.; Audus, K.L. Low-affinity uptake of the fluorescent organic cation 4-(4-(dimethylamino)styryl)-n-methylpyridinium iodide (4-di-1-ASP) in BeWo cells. Biochem. Pharmacol. 2007, 73, 891–900. [Google Scholar] [PubMed]
- Lee, W.K.; Reichold, M.; Edemir, B.; Ciarimboli, G.; Warth, R.; Koepsell, H.; Thévenod, F. Organic cation transporters OCT1, 2, and 3 mediate high-affinity transport of the mutagenic vital dye ethidium in the kidney proximal tubule. Am. J. Physiol. Renal. Physiol. 2009, 296, F1504–F1513. [Google Scholar] [PubMed]
- Dunn, M.; Henke, A.; Clark, S.; Kovalyova, Y.; Kempadoo, K.A.; Karpowicz, R.J., Jr.; Kandel, E.R.; Sulzer, D.; Sames, D. Designing a norepinephrine optical tracer for imaging individual noradrenergic synapses and their activity in vivo. Nat. Commun. 2018, 9, 2838. [Google Scholar] [PubMed]
- Masereeuw, R.; Moons, M.M.; Toomey, B.H.; Russel, F.G.M.; Miller, D.S. Active lucifer yellow secretion in renal proximal tubule: Evidence for organic anion transport system crossover. J. Pharmacol. Exp. Ther. 1999, 289, 1104–1111. [Google Scholar]
- de Gier, R.P.E.; Feitz, W.F.J.; Masereeuw, R.; Wouterse, A.C.; Smits, D.; Russel, F.G.M. Anionic and cationic drug secretion in the isolated perfused rat kidney after neonatal surgical induction of ureteric obstruction. BJU Int. 2003, 92, 452–458. [Google Scholar]
- Jouan, E.; Le Vee, M.; Denizot, C.; Da Violante, G.; Fardel, O. The mitochondrial fluorescent dye rhodamine 123 is a high-affinity substrate for organic cation transporters (OCTs) 1 and 2. Fundam Clin. Pharmacol. 2014, 28, 65–77. [Google Scholar]
- Wilson, J.N.; Brown, A.S.; Babinchak, W.M.; Ridge, C.D.; Walls, J.D. Fluorescent stilbazolium dyes as probes of the norepinephrine transporter: Structural insights into substrate binding. Org. Biomol. Chem. 2012, 10, 8710–8719. [Google Scholar]
- Patik, I.; Székely, V.; Német, O.; Szepesi, Á.; Kucsma, N.; Várady, G.; Szakács, G.; Bakos, É.; Özvegy-Laczka, C. Identification of novel cell-impermeant fluorescent substrates for testing the function and drug interaction of organic anion-transporting polypeptides, OATP1B1/1B3 and 2B1. Sci. Rep. 2018, 8, 2630. [Google Scholar]
- Lajiness, M.S.; Maggiora, G.M.; Shanmugasundaram, V. Assessment of the consistency of medicinal chemists in reviewing sets of compounds. J. Med. Chem. 2004, 47, 4891–4896. [Google Scholar]
- Samanta, S.; O’Hagan, S.; Swainston, N.; Roberts, T.J.; Kell, D.B. Vae-sim: A novel molecular similarity measure based on a variational autoencoder. Molecules 2020, 25, 3446. [Google Scholar]
- Kell, D.B. Finding novel pharmaceuticals in the systems biology era using multiple effective drug targets, phenotypic screening, and knowledge of transporters: Where drug discovery went wrong and how to fix it. FEBS J. 2013, 280, 5957–5980. [Google Scholar] [PubMed]
- Kell, D.B.; Wright Muelas, M.; O’Hagan, S.; Day, P.J. The role of drug transporters in phenotypic screening. Drug Target Rev. 2018, 4, 16–19. [Google Scholar]
- Prior, M.; Chiruta, C.; Currais, A.; Goldberg, J.; Ramsey, J.; Dargusch, R.; Maher, P.A.; Schubert, D. Back to the future with phenotypic screening. ACS Chem. Neurosci. 2014, 5, 503–513. [Google Scholar] [PubMed]
- Swinney, D.C. Opportunities for phenotypic screening in drug discovery. Drug Disc. World 2014, 15, 33–42. [Google Scholar]
- Yamaguchi, Y.; Matsubara, Y.; Ochi, T.; Wakamiya, T.; Yoshida, Z. How the π conjugation length affects the fluorescence emission efficiency. J. Am. Chem. Soc. 2008, 130, 13867–13869. [Google Scholar]
- Lavis, L.D.; Raines, R.T. Bright building blocks for chemical biology. ACS Chem. Biol. 2014, 9, 855–866. [Google Scholar]
- Zambianchi, M.; Di Maria, F.; Cazzato, A.; Gigli, G.; Piacenza, M.; Della Sala, F.; Barbarella, G. Microwave-assisted synthesis of thiophene fluorophores, labeling and multilabeling of monoclonal antibodies, and long lasting staining of fixed cells. J. Am. Chem. Soc. 2009, 131, 10892–10900. [Google Scholar]
- Di Maria, F.; Palamà, I.E.; Baroncini, M.; Barbieri, A.; Bongini, A.; Bizzarri, R.; Gigli, G.; Barbarella, G. Live cell cytoplasm staining and selective labeling of intracellular proteins by non-toxic cell-permeant thiophene fluorophores. Org. Biomol. Chem. 2014, 12, 1603–1610. [Google Scholar]
- O’Hagan, S.; Kell, D.B. The KNIME workflow environment and its applications in genetic programming and machine learning. Genet. Progr. Evol. Mach. 2015, 16, 387–391. [Google Scholar]
- O’Hagan, S.; Kell, D.B. The apparent permeabilities of caco-2 cells to marketed drugs: Magnitude, and independence from both biophysical properties and endogenite similarities. PeerJ 2015, 3, e1405. [Google Scholar]
- O’Hagan, S.; Kell, D.B. Generation of a small library of natural products designed to cover chemical space inexpensively. Pharm. Front. 2019, 1, e190005. [Google Scholar] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
O’Hagan, S.; Kell, D.B. Structural Similarities between Some Common Fluorophores Used in Biology, Marketed Drugs, Endogenous Metabolites, and Natural Products. Mar. Drugs 2020, 18, 582. https://doi.org/10.3390/md18110582
O’Hagan S, Kell DB. Structural Similarities between Some Common Fluorophores Used in Biology, Marketed Drugs, Endogenous Metabolites, and Natural Products. Marine Drugs. 2020; 18(11):582. https://doi.org/10.3390/md18110582
Chicago/Turabian StyleO’Hagan, Steve, and Douglas B. Kell. 2020. "Structural Similarities between Some Common Fluorophores Used in Biology, Marketed Drugs, Endogenous Metabolites, and Natural Products" Marine Drugs 18, no. 11: 582. https://doi.org/10.3390/md18110582
APA StyleO’Hagan, S., & Kell, D. B. (2020). Structural Similarities between Some Common Fluorophores Used in Biology, Marketed Drugs, Endogenous Metabolites, and Natural Products. Marine Drugs, 18(11), 582. https://doi.org/10.3390/md18110582