AhlX, an N-acylhomoserine Lactonase with Unique Properties
Abstract
:1. Introduction
2. Results
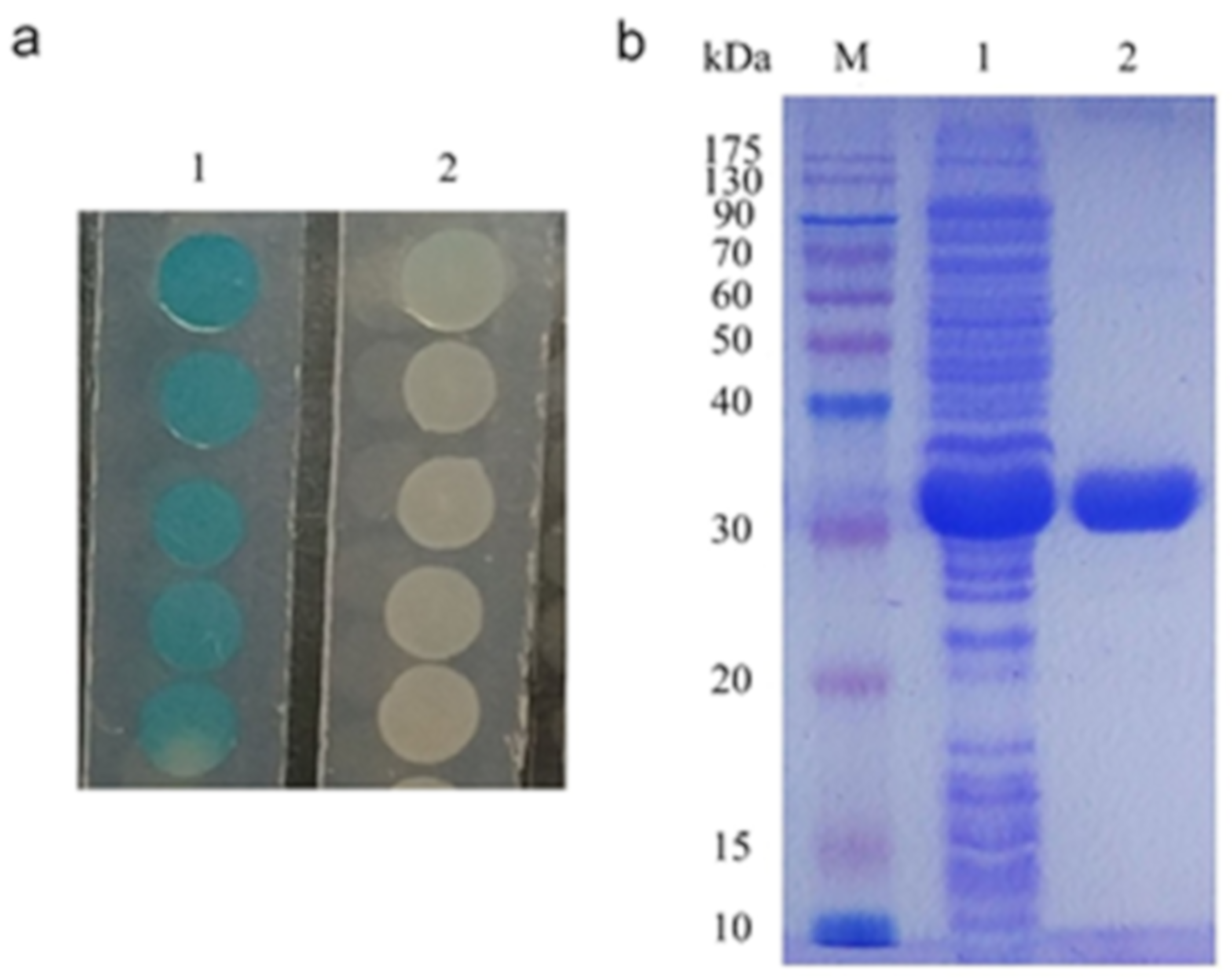
2.1. Cloning of the Lactonase-Encoding Gene AhlX
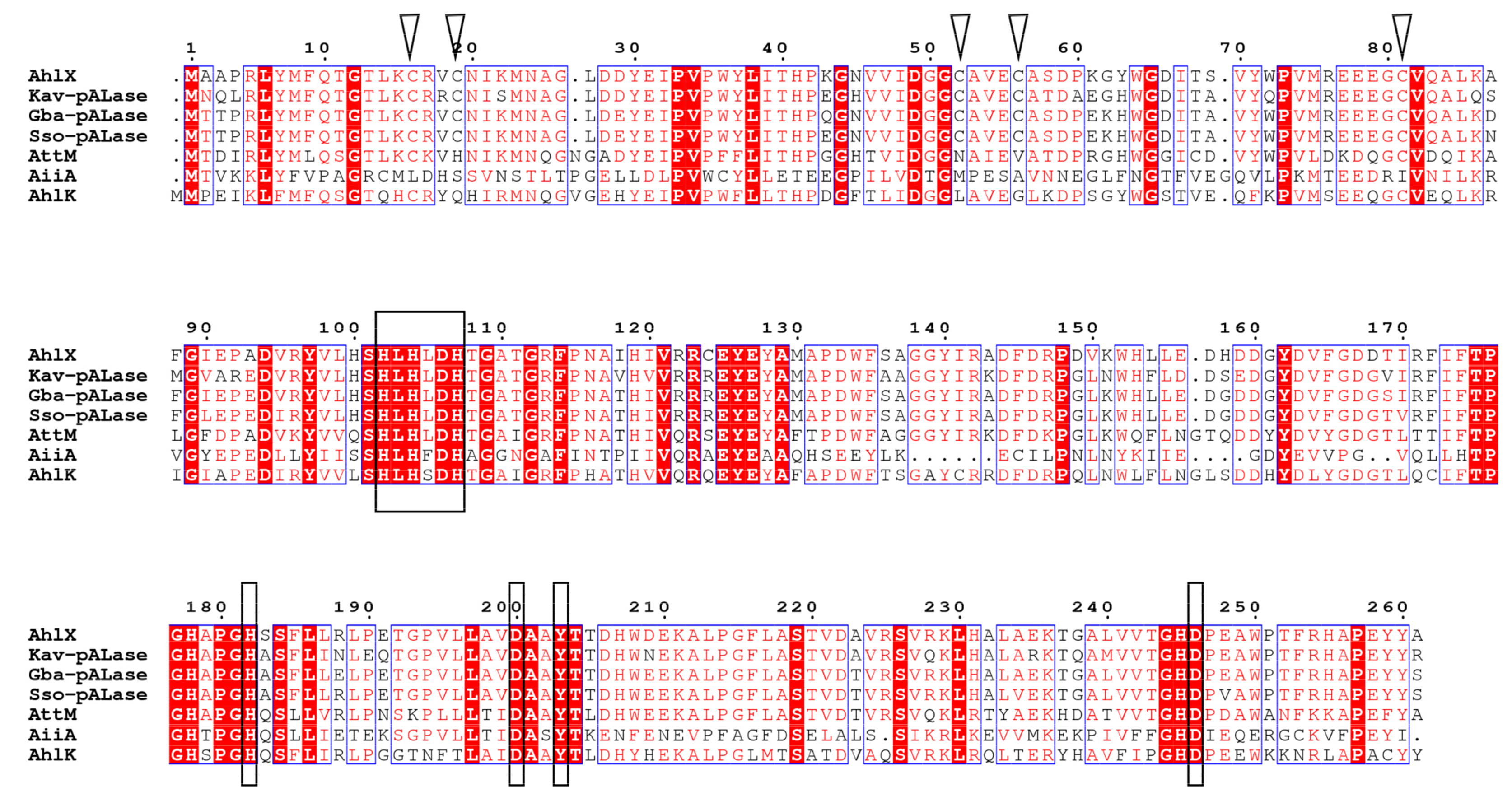
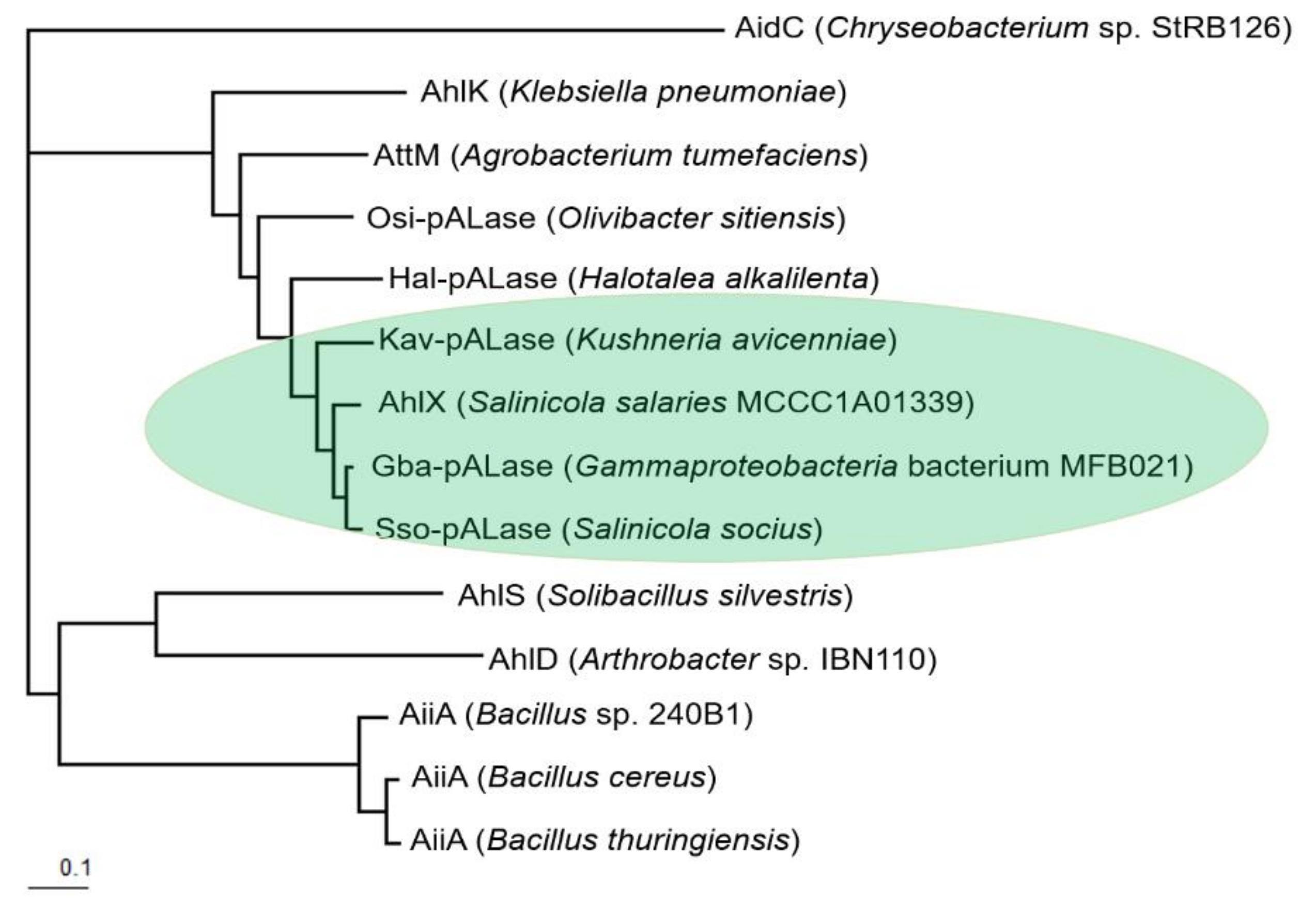
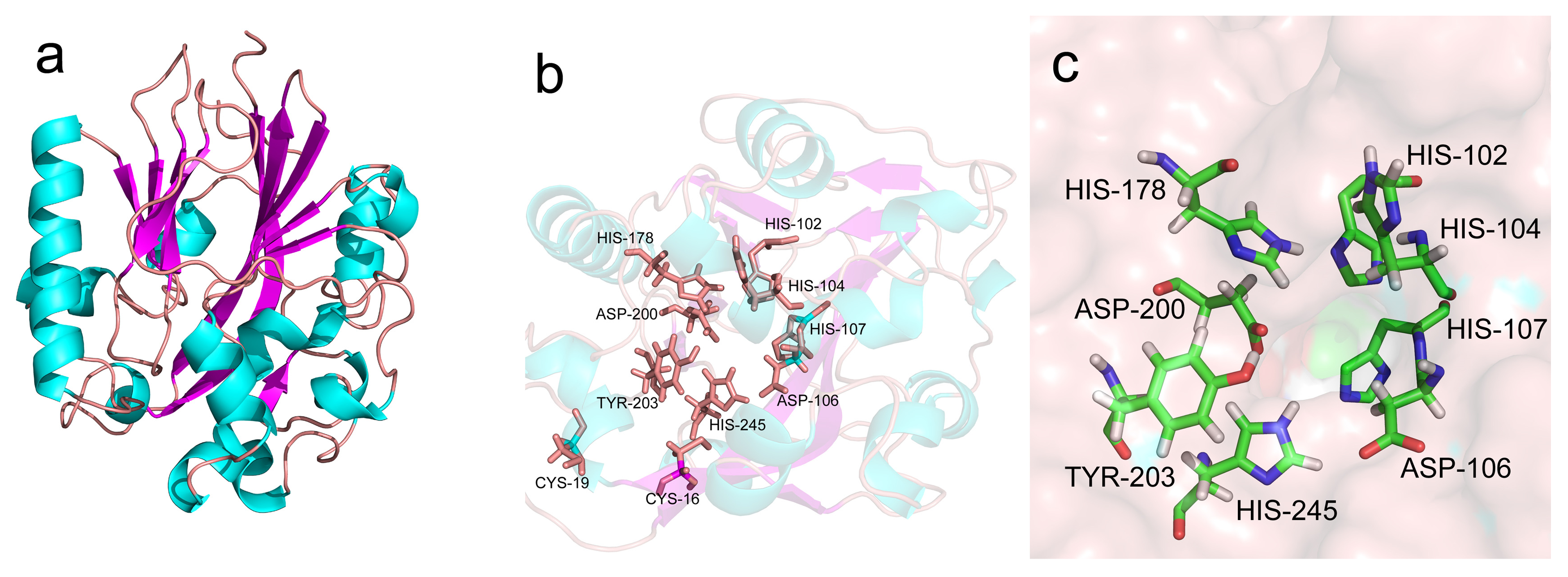
2.2. Bioinformatic Analysis of AhlX
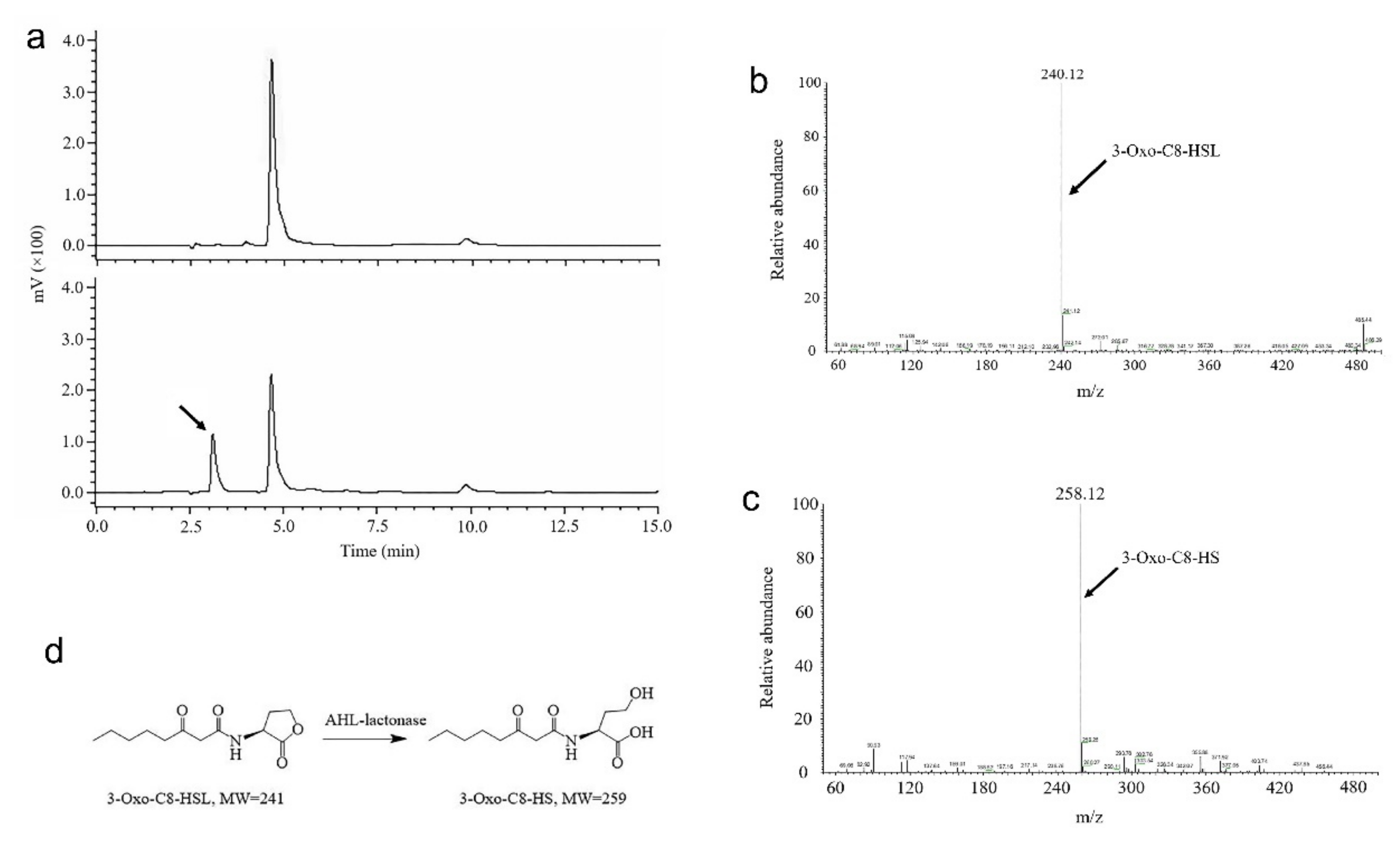
2.3. Characterization of 3OC8-HSL Degradation by AhlX
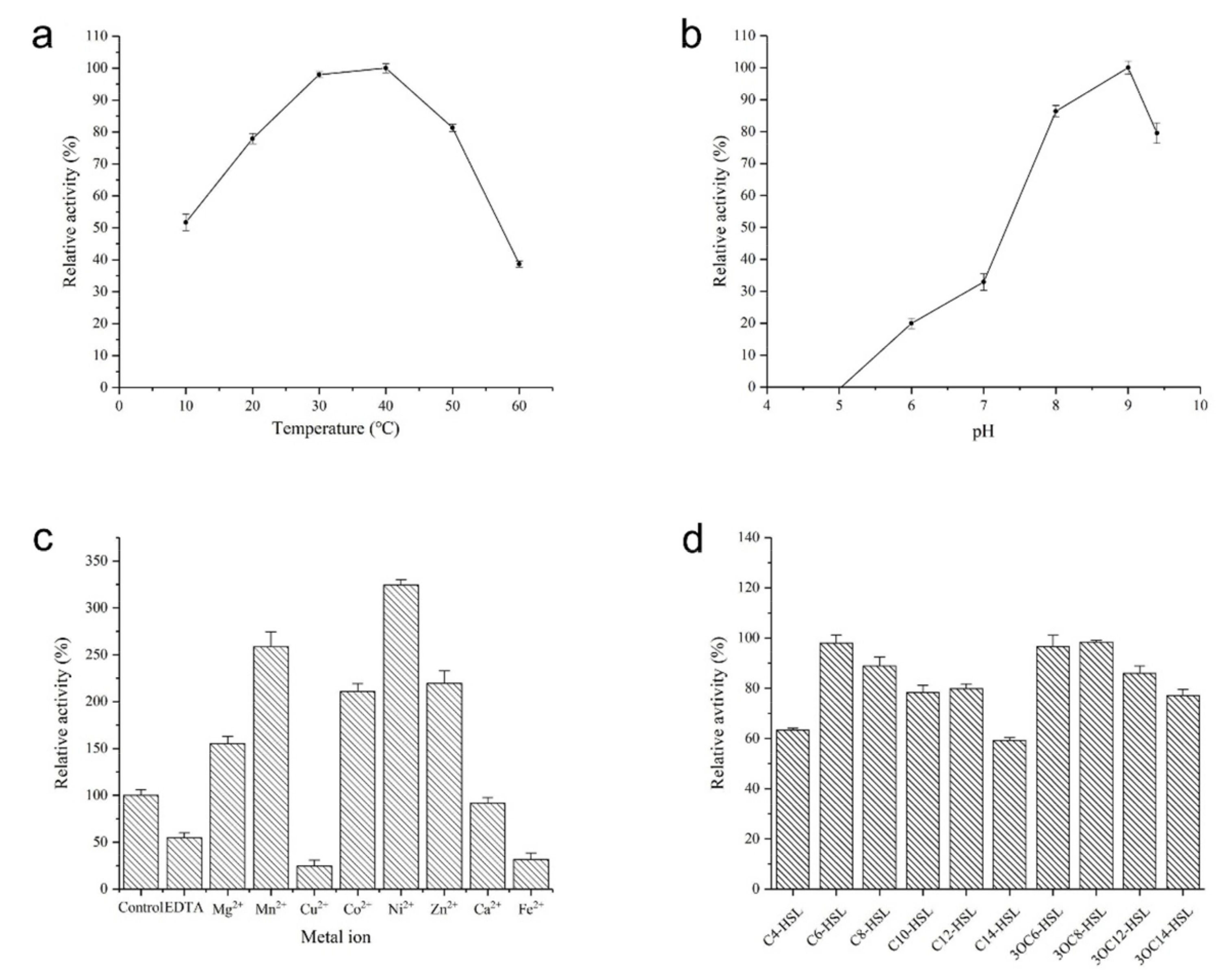
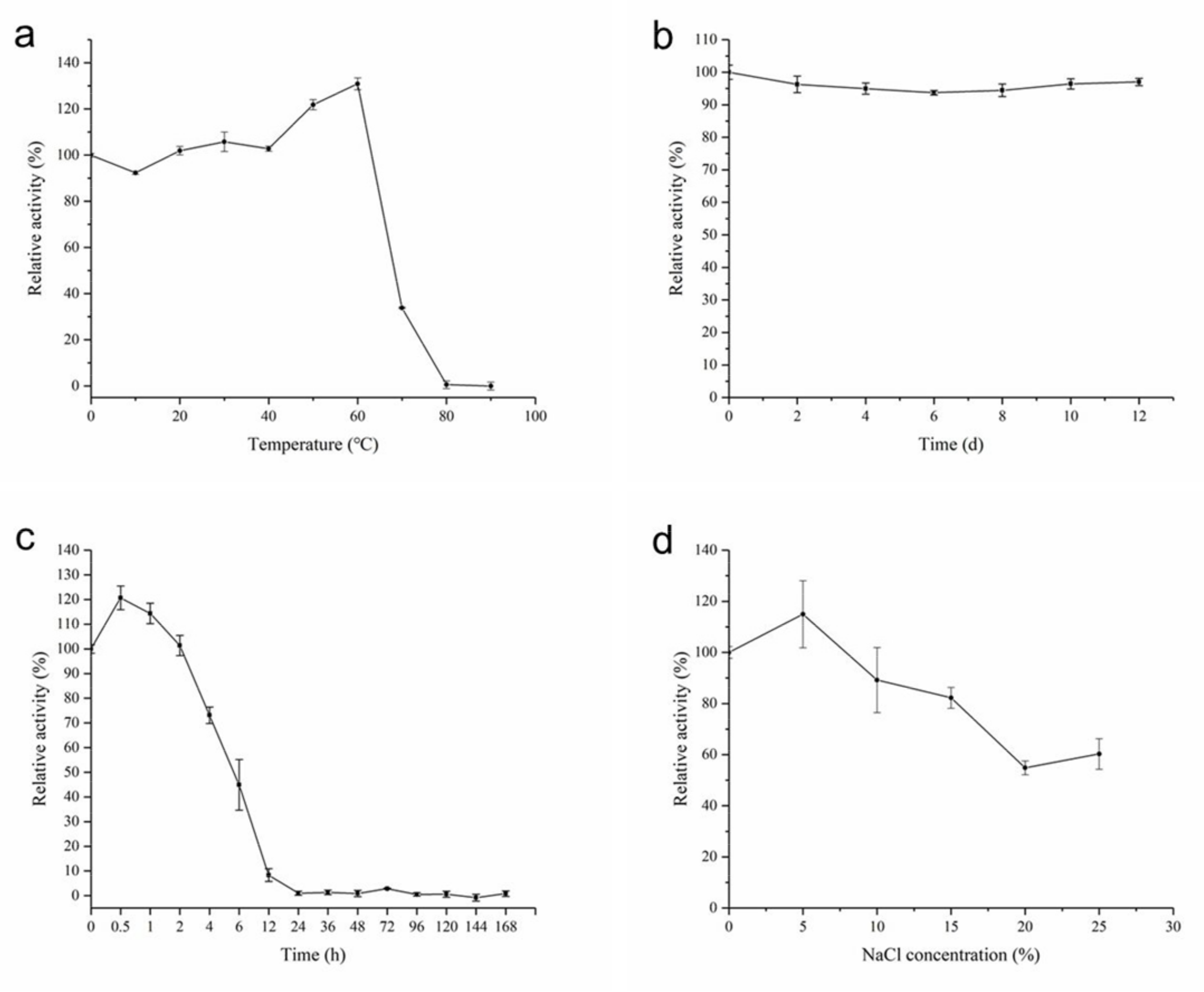
2.4. Biochemical Characterization of AhlX
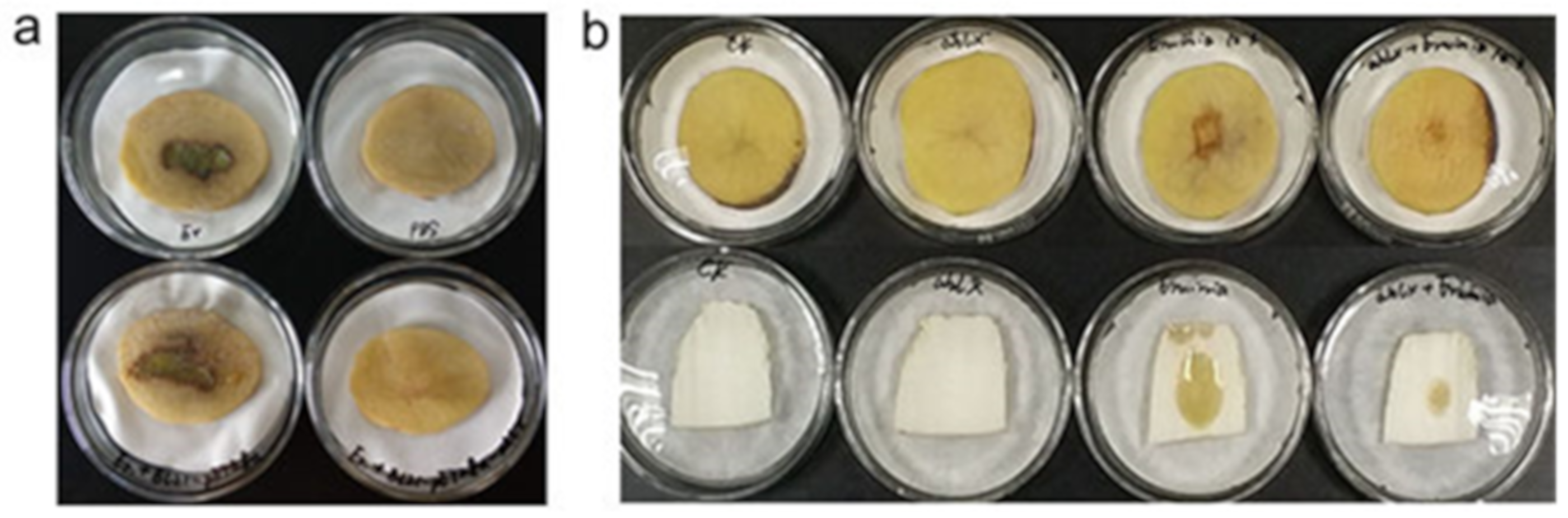
2.5. Quenching the E. carotovora Infection by AhlX
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Chemicals
4.2. Genome Sequencing and Bioinformatic Analysis
4.3. Expression and Purification of AhlX
4.4. Standard Activity Assay of AhlX
4.5. Determination of the Mechanism of AHL Degradation by AhlX
4.6. Biochemical Characterization of AhlX
4.7. Quenching the E. carotovora Infection by AhlX
4.8. Nucleotide Sequence Accession Number
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bassler, B.L.; Greenberg, E.P.; Stevens, A.M. Cross-species induction of luminescence in the quorum-sensing bacterium Vibrio harveyi. J. Bacteriol. 1997, 179, 4043–4045. [Google Scholar] [CrossRef]
- Lang, J.; Faure, D. Functions and regulation of quorum-sensing in Agrobacterium tumefaciens. Front. Plant Sci. 2014, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Duerkop, B.A.; Varga, J.; Chandler, J.R.; Peterson, S.B.; Herman, J.P.; Churchill, M.E.; Parsek, M.R.; Nierman, W.C.; Greenberg, E.P. Quorum-sensing control of antibiotic synthesis in Burkholderia thailandensis. J. Bacteriol. 2009, 191, 3909–3918. [Google Scholar] [CrossRef]
- Allison, D.G.; Ruiz, B.; SanJose, C.; Jaspe, A.; Gilbert, P. Extracellular products as mediators of the formation and detachment of Pseudomonas fluorescens biofilms. FEMS Microbiol. Lett. 1998, 167, 179–184. [Google Scholar] [CrossRef] [PubMed]
- De Kievit, T.R.; Iglewski, B.H. Bacterial quorum sensing in pathogenic relationships. Infect. Immun. 2000, 68, 4839–4849. [Google Scholar] [CrossRef] [PubMed]
- Waters, C.M.; Bassler, B.L. Quorum sensing: cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 2005, 21, 319–346. [Google Scholar] [CrossRef] [PubMed]
- Parsek, M.R.; Greenberg, E.P. Acyl-homoserine lactone quorum sensing in gram-negative bacteria: a signaling mechanism involved in associations with higher organisms. Proc. Natl. Acad. Sci. USA 2000, 97, 8789–8793. [Google Scholar] [CrossRef] [PubMed]
- Andersson, R.A.; Eriksson, A.R.; Heikinheimo, R.; Mae, A.; Pirhonen, M.; Koiv, V.; Hyytiainen, H.; Tuikkala, A.; Palva, E.T. Quorum sensing in the plant pathogen Erwinia carotovora subspcarotovora: the role of expR(Ecc). Mol. Plant-Microbe Interact. 2000, 13, 384–393. [Google Scholar]
- Dong, Y.H.; Xu, J.L.; Li, X.Z.; Zhang, L.H. AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora. Proc. Natl. Acad. Sci. USA 2000, 97, 3526–3531. [Google Scholar] [CrossRef]
- Dong, Y.H.; Wang, L.H.; Xu, J.L.; Zhang, H.B.; Zhang, X.F.; Zhang, L.H. Quenching quorum-sensing-dependent bacterial infection by an N-acyl homoserine lactonase. Nature 2001, 411, 813–817. [Google Scholar] [CrossRef]
- Fetzner, S. Quorum quenching enzymes. J. Bacteriol. 2015, 201, 2–14. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.H.; Zhang, L.H. Quorum sensing and quorum-quenching enzymes. J. Microbiol. 2005, 43, 101–109. [Google Scholar] [PubMed]
- Cai, X.; Yu, M.; Shan, H.; Tian, X.; Zheng, Y.; Xue, C.; Zhang, X. Characterization of a Novel N-Acylhomoserine Lactonase RmmL from Ruegeria mobilis YJ3. Mar. Drugs 2018, 16, 370. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Lin, Y.; Yi, S.; Liu, P.; Shen, J.; Shao, Z.; Liu, Z. QsdH, a novel AHL lactonase in the RND-type inner membrane of marine Pseudoalteromonas byunsanensis strain 1A01261. PLoS ONE 2012, 7, e46587. [Google Scholar] [CrossRef] [PubMed]
- De la Haba, R.R.; Sanchez-Porro, C.; Marquez, M.C.; Ventosa, A. Taxonomic study of the genus Salinicola: transfer of Halomonas salaria and Chromohalobacter salarius to the genus Salinicola as Salinicola salarius comb. nov. and Salinicola halophilus nom. nov., respectively. Int. J. Syst. Evol. Microbiol. 2010, 60, 963–971. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.B.; Wang, L.H.; Zhang, L.H. Genetic control of quorum-sensing signal turnover in Agrobacterium tumefaciens. Proc. Natl. Acad. Sci. USA 2002, 99, 4638–4643. [Google Scholar] [CrossRef] [PubMed]
- Joseph, T.C.; Baby, A.; Reghunathan, D.; Varghese, A.M.; Murugadas, V.; Lalitha, K.V. Draft Genome Sequence of the Halophilic and Highly Halotolerant Gammaproteobacteria Strain MFB021. Genome Announc. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Anan’ina, L.N.; Plotnikova, E.G.; Gavrish, E.; Demakov, V.A.; Evtushenko, L.I. Salinicola socius gen. nov., sp. nov., a moderately halophilic bacterium from a naphthalene-utilizing microbial association. Mikrobiologiia 2007, 76, 369–376. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Porro, C.; De la Haba, R.R.; Soto-Ramirez, N.; Marquez, M.C.; Montalvo-Rodriguez, R.; Ventosa, A. Description of Kushneria aurantia gen. nov., sp. nov., a novel member of the family Halomonadaceae, and a proposal for reclassification of Halomonas marisflavi as Kushneria marisflavi comb. nov., of Halomonas indalinina as Kushneria indalinina comb. nov. and of Halomonas avicenniae as Kushneria avicenniae comb. nov. Int. J. Syst. Evol. Microbiol. 2009, 59, 397–405. [Google Scholar] [PubMed]
- Wang, W.Z.; Morohoshi, T.; Someya, N.; Ikeda, T. AidC, a novel N-acylhomoserine lactonase from the potato root-associated cytophaga-flavobacteria-bacteroides (CFB) group bacterium Chryseobacterium sp. strain StRB126. Appl. Environ. Microbiol. 2012, 78, 7985–7992. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Lee, S.J.; Oh, T.K.; Oh, J.W.; Koo, B.T.; Yum, D.Y.; Lee, J.K. AhlD, an N-acylhomoserine lactonase in Arthrobacter sp., and predicted homologues in other bacteria. Microbiology 2003, 149, 1541–1550. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.H.; Choi, W.C.; Kang, H.O.; Lee, J.S.; Kang, B.S.; Kim, K.J.; Derewenda, Z.S.; Oh, T.K.; Lee, C.H.; Lee, J.K. The molecular structure and catalytic mechanism of a quorum-quenching N-acyl-L-homoserine lactone hydrolase. Proc. Natl. Acad. Sci. USA 2005, 102, 17606–17611. [Google Scholar] [CrossRef] [PubMed]
- Momb, J.; Wang, C.; Liu, D.; Thomas, P.W.; Petsko, G.A.; Guo, H.; Ringe, D.; Fast, W. Mechanism of the quorum-quenching lactonase (AiiA) from Bacillus thuringiensis. 2. Substrate modeling and active site mutations. Biochemistry 2008, 47, 7715–7725. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef] [PubMed]
- Palzkill, T. Metallo-beta-lactamase structure and function. Ann. NY Acad. Sci. 2013, 1277, 91–104. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wang, J.; Feng, T.; Du, R.; Tian, X.; Wang, Y.; Zhang, X. Heterologous Expression of the Marine-Derived Quorum Quenching Enzyme MomL Can Expand the Antibacterial Spectrum of Bacillus brevis. Mar. Drugs 2019, 17, 128. [Google Scholar] [CrossRef]
- Jones, S.; Yu, B.; Bainton, N.J.; Birdsall, M.; Bycroft, B.W.; Chhabra, S.R.; Cox, A.J.R.; Golby, P.; Reeves, P.J.; Stephens, S.; et al. The lux autoinducer regulates the production of exoenzyme virulence determinants in Erwinia carotovora and Pseudomonas aeruginosa. EMBO J. 1993, 12, 2477–2482. [Google Scholar] [CrossRef]
- Yin, J.; Chen, J.C.; Wu, Q.; Chen, G.Q. Halophiles, coming stars for industrial biotechnology. Biotechnol. Adv. 2015, 33, 1433–1442. [Google Scholar] [CrossRef]
- Fujimoto, K.; Morita, T. Aerobic removal of technetium by a marine Halomonas strain. Appl. Environ. Microbiol. 2006, 72, 7922–7924. [Google Scholar] [CrossRef]
- Tahrioui, A.; Schwab, M.; Quesada, E.; Llamas, I. Quorum sensing in some representative species of halomonadaceae. Life 2013, 3, 260–275. [Google Scholar] [CrossRef]
- Llamas, I.; Quesada, E.; Martinez-Canovas, M.J.; Gronquist, M.; Eberhard, A.; Gonzalez, J.E. Quorum sensing in halophilic bacteria: detection of N-acyl-homoserine lactones in the exopolysaccharide-producing species of Halomonas. Extremophiles 2005, 9, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Fomenko, D.E.; Gladyshev, V.N. Identity and functions of CxxC-derived motifs. Biochemistry 2003, 42, 11214–11225. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.H.; Weng, L.X.; Dong, Y.H.; Zhang, L.H. Specificity and enzyme kinetics of the quorum-quenching N-Acyl homoserine lactone lactonase (AHL-lactonase). J. Biol. Chem. 2004, 279, 13645–13651. [Google Scholar] [CrossRef] [PubMed]
- Seo, M.; Lee, B.S.; Pyun, Y.; Park, H. Isolation and Characterization of N-Acylhomoserine Lactonase from the Thermophilic Bacterium, Geobacillus caldoxylosilyticus YS-8. Biosci. Biotechnol. Biochem. 2011, 75, 1789–1795. [Google Scholar] [CrossRef] [PubMed]
- Stevens, D.A.; Hamilton, J.R.; Johnson, N.; Kim, K.K.; Lee, J.S. Halomonas, a newly recognized human pathogen causing infections and contamination in a dialysis center: Three new species. Medicine 2009, 88, 244–249. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.Z.; Morohoshi, T.; Ikenoya, M.; Someya, N.; Ikeda, T. AiiM, a novel class of N-acylhomoserine lactonase from the leaf-associated bacterium Microbacterium testaceum. Appl. Environ. Microbiol. 2010, 76, 2524–2530. [Google Scholar] [CrossRef]
- Xavier, R.; Patrice, G. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, 320–324. [Google Scholar]

| Purification Step | Total Protein (mg) | Specific Activity (Units/mg) | Total Activity (Units) | Enzyme Activity Recovery (%) |
|---|---|---|---|---|
| Lysate supernatant | 507.06 | 108.89 | 5.52 × 104 | 100 |
| Heat treatment | 247.35 | 223.76 | 5.53 × 104 | 100.24 |
| Heat treatment + Spray drying | 203.36 | 228.25 | 4.64 × 104 | 84.06 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, P.; Chen, Y.; Shao, Z.; Chen, J.; Wu, J.; Guo, Q.; Shi, J.; Wang, H.; Chu, X. AhlX, an N-acylhomoserine Lactonase with Unique Properties. Mar. Drugs 2019, 17, 387. https://doi.org/10.3390/md17070387
Liu P, Chen Y, Shao Z, Chen J, Wu J, Guo Q, Shi J, Wang H, Chu X. AhlX, an N-acylhomoserine Lactonase with Unique Properties. Marine Drugs. 2019; 17(7):387. https://doi.org/10.3390/md17070387
Chicago/Turabian StyleLiu, Pengfu, Yan Chen, Zongze Shao, Jianwei Chen, Jiequn Wu, Qian Guo, Jiping Shi, Hong Wang, and Xiaohe Chu. 2019. "AhlX, an N-acylhomoserine Lactonase with Unique Properties" Marine Drugs 17, no. 7: 387. https://doi.org/10.3390/md17070387
APA StyleLiu, P., Chen, Y., Shao, Z., Chen, J., Wu, J., Guo, Q., Shi, J., Wang, H., & Chu, X. (2019). AhlX, an N-acylhomoserine Lactonase with Unique Properties. Marine Drugs, 17(7), 387. https://doi.org/10.3390/md17070387






